Effect of the Aerobic Denitrifying Bacterium Pseudomonas furukawaii ZS1 on Microbiota Compositions in Grass Carp Culture Water
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strain and Experimental Design
2.2. Sample Collection and Assessment of Water Quality
2.3. Microbial Composition Analysis via High-throughput Sequencing
2.4. Data Analysis
3. Results
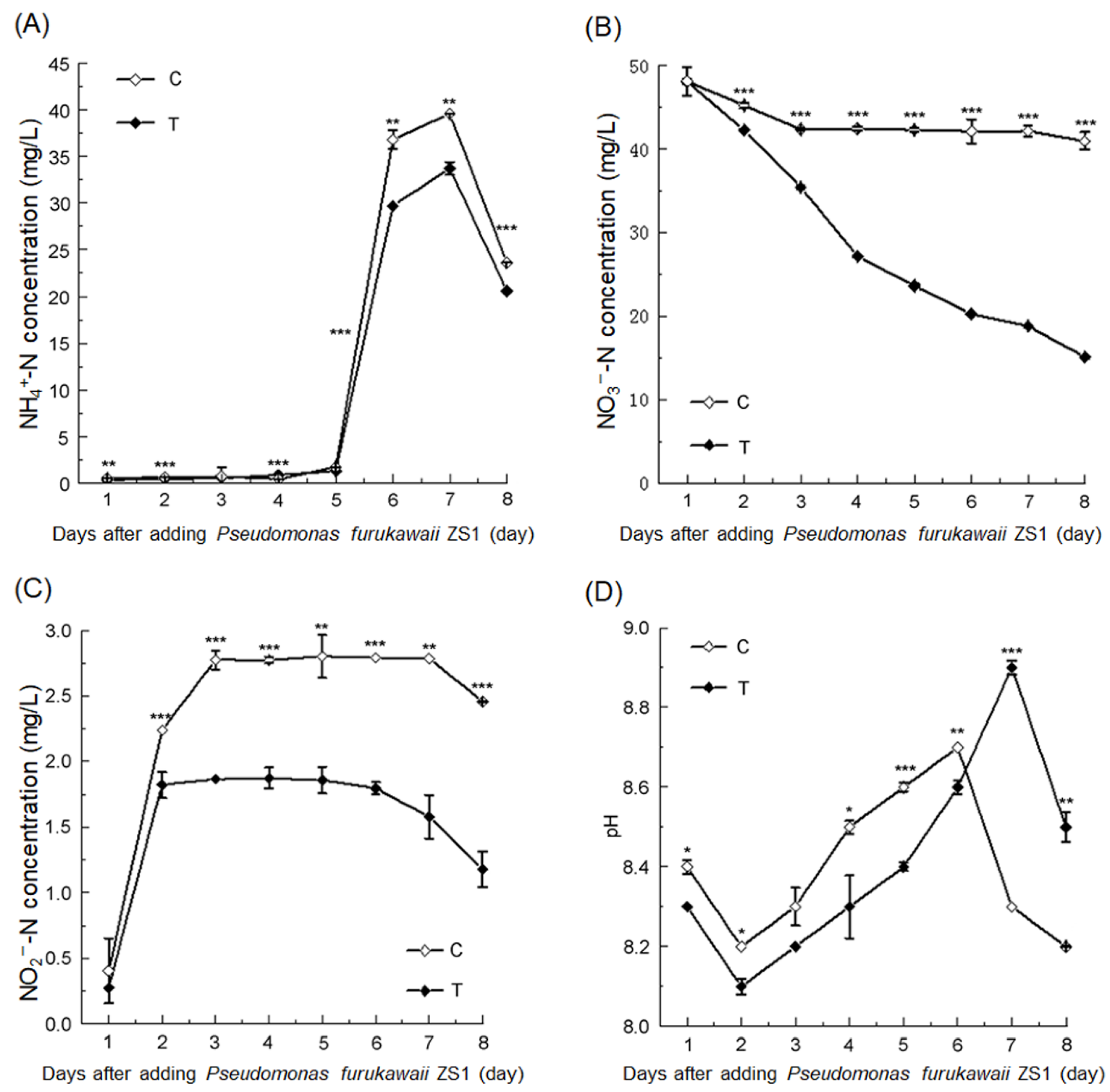
3.1. P. furukawaii ZS1 Promoted Nitrogen Removal from Aquaculture Water
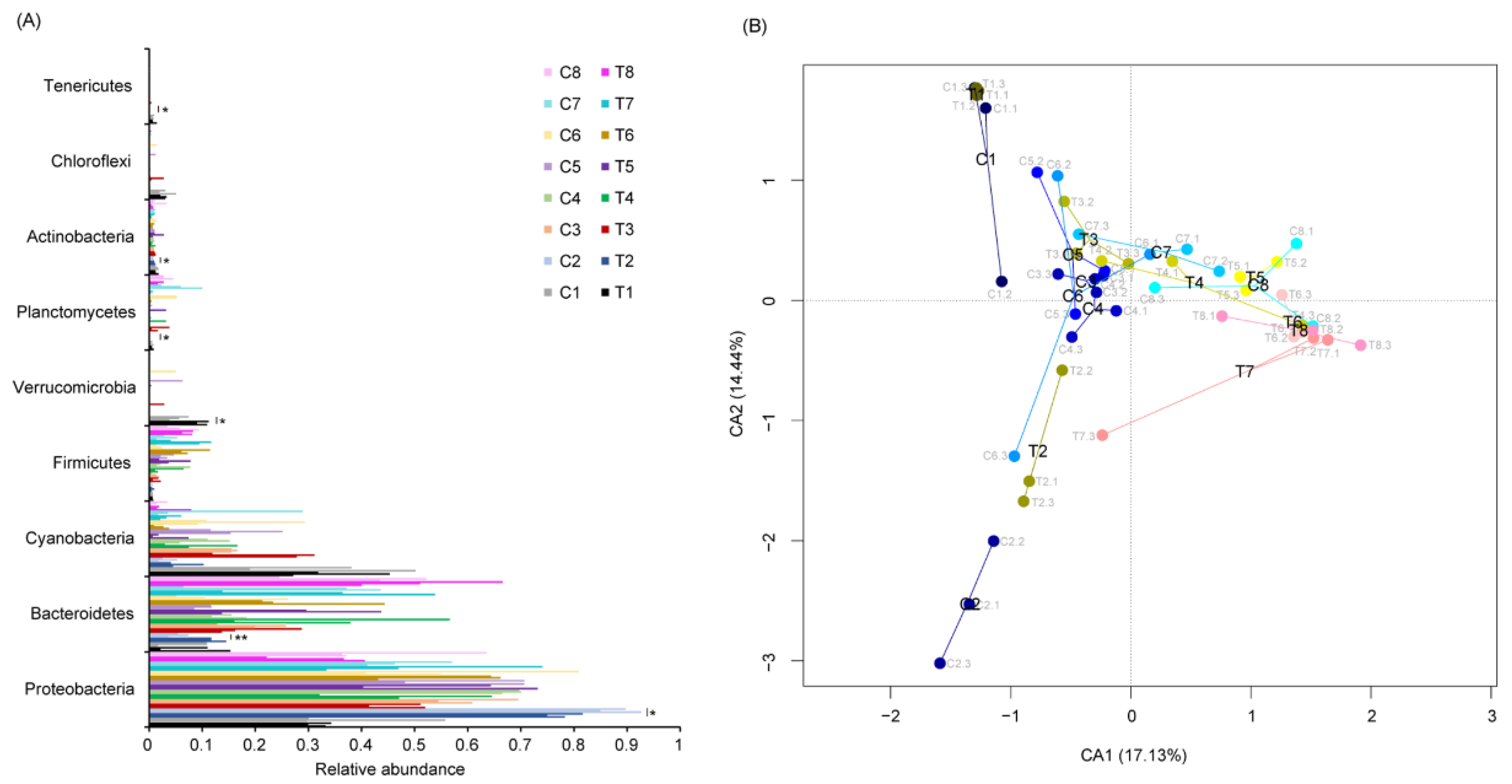
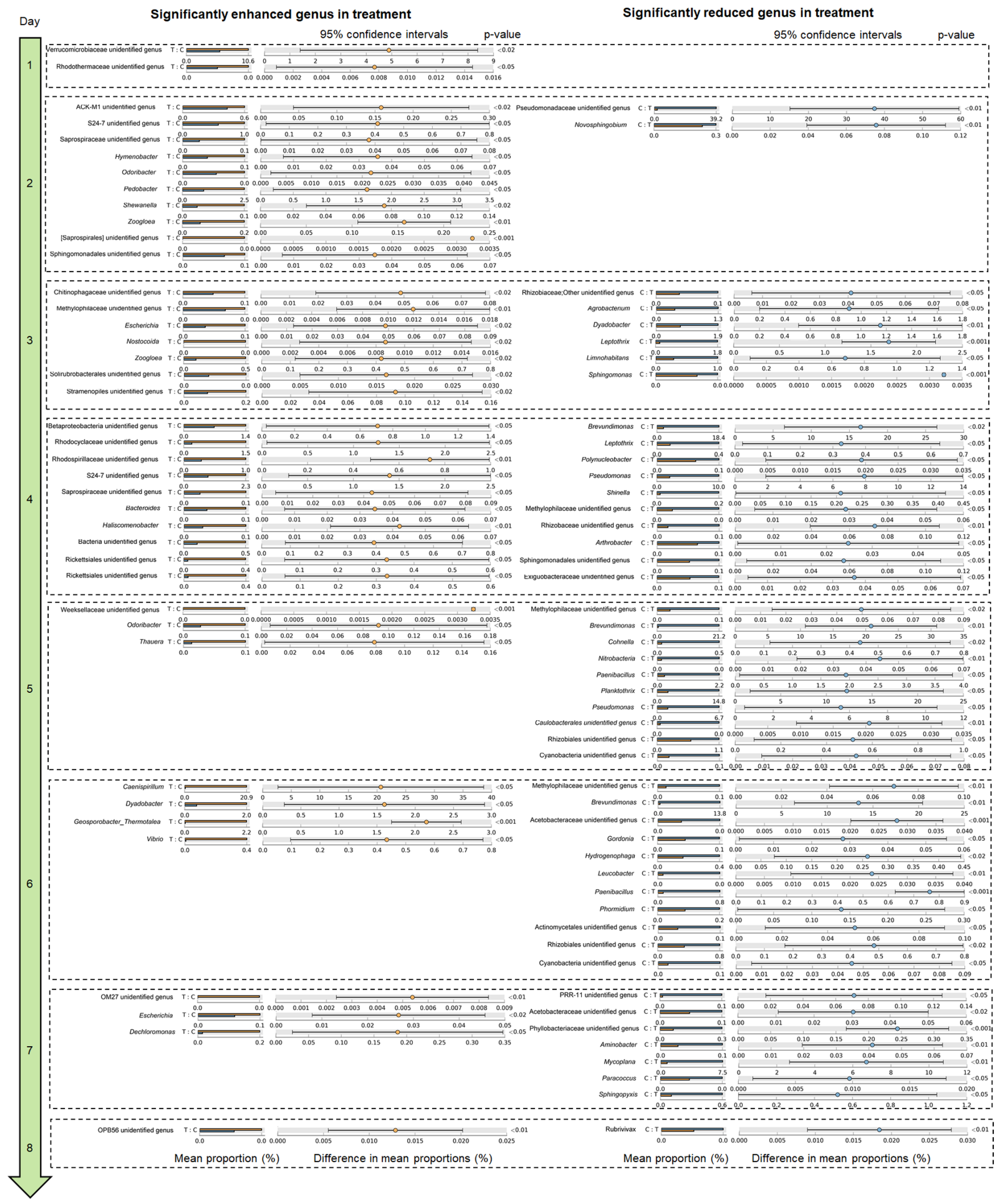
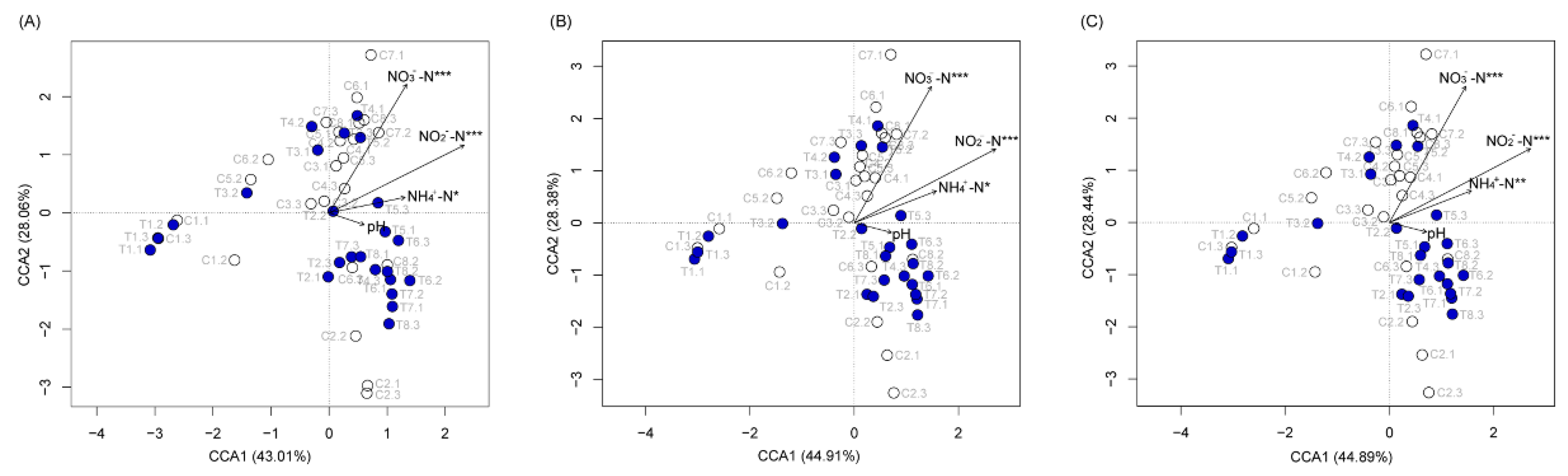
3.2. P. furukawaii ZS1 Temporarily Altered the Microbiota Composition in Aquaculture Water
4. Discussion
Practical Implications of this Study
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- China National Bureau of Statistics. China Fishery Statistical Yearbook 2019; China Agricultural Press: Beijing, China, 2019; pp. 1–138. [Google Scholar]
- Li, Z.; Che, J.; Xie, J.; Wang, G.; Yu, E.; Xia, Y.; Yu, D.; Zhang, K. Microbial succession in biofilms growing on artificial substratum in subtropical freshwater aquaculture ponds. Fems Microbiol. Lett. 2017, 364, fnx017. [Google Scholar] [CrossRef]
- Ni, J.J.; Li, X.J.; Chen, F.; Wu, H.H.; Xu, M.Y. Community structure and potential nitrogen metabolisms of subtropical aquaculture pond microbiota. Appl. Ecol. Environ. Res. 2018, 16, 7687–7697. [Google Scholar] [CrossRef]
- Li, Z.; Wang, G.; Yu, E.; Zhang, K.; Yu, D.; Gong, W.; Xie, J. Artificial substrata increase pond farming density of grass carp (Ctenopharyngodon idella) by increasing the bacteria that participate in nitrogen and phosphorus cycles in pond water. Peerj 2019, 7, e7906. [Google Scholar] [CrossRef]
- Piedrahita, R.H. Reducing the potential environmental impact of tank aquaculture effluents through intensification and recirculation. Aquaculture 2003, 226, 35–44. [Google Scholar] [CrossRef]
- Attramadal, K.J.K.; Salvesen, I.; Xue, R.; Øie, G.; Størseth, T.R.; Vadstein, O.; Olsen, Y. Recirculation as a possible microbial control strategy in the production of marine larvae. Aquac. Eng. 2012, 46, 27–39. [Google Scholar] [CrossRef]
- Feng, Y.; Zhang, R.; Shi, C.; Li, L.; Ma, Y. Study of microbial community participating in water cycle. J. Anhui. Agricul. Sci. 2016, 44, 132–134. [Google Scholar]
- Zheng, H.; Zhou, Q. Function of microorganisms in bioremediation of eutrophic water. Chin. J. Eco-Agricul. 2009, 17, 197–202. [Google Scholar] [CrossRef]
- Chen, J.; Zheng, J.; Mao, Y.; Su, Y.; Wang, J. Effects of probiotics and compound nutrients on water quality and community structure of bacteria in fish-shrimp farming pond. J. Trop. Biol. 2019, 10, 6–13. [Google Scholar]
- Hong, P.; Huang, Y.; Chen, M.; Xiao, B. Efficacy of inorganic nitrogen removal by a salt-tolerant aerobic denitrifying bacterium, Pseudomonas Sihuiensis LK-618. Bioprocess Biosyst. Eng. 2021. [Google Scholar] [CrossRef]
- Jaiswal, S.K.; Saxena, R.; Mittal, P.; Gupta, A.; Sharma, V.K. Draft genome sequence of Pseudomonas hussainii strain MB3, a denitrifying aerobic bacterium isolated from the rhizospheric region of mangrove trees in the Andaman Islands, India. Genome Announc. 2017, 5, e01527-16. [Google Scholar] [CrossRef]
- Cai, X.; He, T.; Ye, Q.; Li, Z. Identification and characterization of a hypothermic alkaliphilic aerobic denitrifying bacterium Pseudomonas monteilii strain H97. Environ. Sci. 2018, 39, 3314–3320. [Google Scholar]
- Zhang, W.; Yan, C.; Shen, J.; Wei, R.; Gao, Y.; Miao, A.; Xiao, L.; Yang, L. Characterization of aerobic denitrifying bacterium Pseudomonas mendocina strain GL6 and its potential application in wastewater treatment plant effluent. Int. J. Environ. Res. Public Health 2019, 16, 364. [Google Scholar] [CrossRef]
- Zhu, Y.; Gong, W.; Xie, J.; Wang, G.; Yu, D.; Li, Z.; Zhang, K.; Tian, J. Identification and denitrification characteristics of the aerobic denitrifying bacteria Pseudomonas furukawaii ZS1. Acta Hydrobiol. Sin. 2020, 44, 895–903. [Google Scholar]
- Huang, X.F. Survey, Observation and Analysis of Lake Ecology; China Standard Press: Beijing, China, 2000. [Google Scholar]
- Ni, J.; Yu, Y.; Feng, W.; Yang, Q.; Pan, G.; Yang, B.; Zhang, X.; Li, X. Impacts of algal blooms removal by chitosan-modified soils on zooplankton community in Taihu Lake. China J. Environ. Sci. 2010, 22, 1500–1507. [Google Scholar] [CrossRef]
- Ni, J.; Li, X.; He, Z.; Xu, M. A novel method to determine the minimum number of sequences required for reliable microbial community analysis. J. Microbiol. Methods 2017, 139, 196–201. [Google Scholar] [CrossRef]
- Ni, J.; Huang, R.; Zhou, H.; Xu, X.; Li, Y.; Cao, P.; Zhong, K.; Ge, M.; Chen, X.; Hou, B.; et al. Analysis of the relationship between the degree of dysbiosis in gut microbiota and prognosis at different stages of primary hepatocellular carcinoma. Front. Microbiol. 2019, 10, 1458. [Google Scholar] [CrossRef] [PubMed]
- Huang, R.; Li, T.; Ni, J.; Bai, X.; Gao, Y.; Li, Y.; Zhang, P.; Gong, Y. Different sex-based responses of gut microbiota during the development of hepatocellular carcinoma in liver-specific Tsc1-knockout mice. Front. Microbiol. 2018, 9, 1008. [Google Scholar] [CrossRef]
- Magoč, T.; Salzberg, S.L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011, 27, 2957–2963. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Xiang, J.; He, T.; Wang, P.; Xie, M.; Xiang, J.; Ni, J. Opportunistic pathogens are abundant in the gut of cultured giant spiny frog (Paa spinosa). Aquacul. Res. 2018, 49, 2033–2041. [Google Scholar] [CrossRef]
- Edgar, R.C. UPASE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996–998. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef] [PubMed]
- Anderson, M.J. A new method for non-parametric multivariate analysis of variance. Austral. Ecol. 2001, 26, 32–46. [Google Scholar]
- Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 2003, 14, 927–930. [Google Scholar] [CrossRef]
- Parks, D.H.; Beiko, R.G. Identifying biologically relevant differences between metagenomic communities. Bioinformatics 2010, 26, 715–721. [Google Scholar] [CrossRef] [PubMed]
- Hargreaves, J.A. Nitrogen biogeochemistry of aquaculture ponds. Aquaculture 1998, 166, 181–212. [Google Scholar] [CrossRef]
- Hu, Z.; Lee, J.W.; Chandran, K.; Kim, S.; Sharma, K.; Brotto, A.C.; Khanal, S.K. Nitrogen transformations in intensive aquaculture system and its implication to climate change through nitrous oxide emission. Bioresour. Technol. 2013, 130, 314–320. [Google Scholar] [CrossRef]
- Wang, X.; Wu, Y.; Yao, X.; Liu, Y.; Sun, H.; Tang, J. Effect of probiotics on water quality and bacterial community in shrimp pools at later stage of cultivation. Acta. Agricul. Zhejiangensis 2014, 26, 1–47. [Google Scholar]
- Li, Z.; Wang, G.; Yu, D.; Lu, B.; Xie, J.; Yu, E.; Xia, Y.; Chong, X. Effect of bioflocs on water quality and metabolic functions of microbial community in zero-water exchange tanks. J. Shanghai Fish. Univ. 2015, 24, 503–512. [Google Scholar]
- Li, Z.; Lin, L.; Yang, Y.; Lin, X. Influence of Bacillus on the microbial communities in shrimp ponds. J. Agro-Environ. Sci. 2007, 26, 1183–1189. [Google Scholar]
- Xiong, H. Preliminary Study on the Effect of External Bacteria on Water Quality and Microbial Community Structure of Aquaculture Water; Huazhong Agricultural University: Wuhan, China, 2011. [Google Scholar]
- Wu, L.; Shu, F.; Ou, Z.; Chen, Q.; Wang, L.; Wang, H.; Xu, Z. Compositions of prokaryote communities and their relationship to physiochemical factors in December in Chaohu Lake and three urban rivers in China. Appl. Ecol. Environ. Res. 2019, 17, 7265–7281. [Google Scholar] [CrossRef]
- Betiku, O.C.; Sarjeant, K.C.; Ngatia, L.W.; Aghimien, M.O.; Odewumi, C.O.; Latinwo, L.M. Evaluation of microbial diversity of three recreational water bodies using 16S rRNA metagenomic approach. Sci. Total Environ. 2021, 771, 144773. [Google Scholar] [CrossRef]
- Huang, H.K.; Tseng, S.K. Nitrate reduction by Citrobacter diversus under aerobic environment. Appl. Microbiol. Biotechnol. 2001, 55, 90–94. [Google Scholar] [CrossRef] [PubMed]
- Bernat, K.; Wojnowska-Baryła, I. Carbon source in aerobic denitrification. Biochem. Eng. J. 2007, 36, 116–122. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gong, W.; Gao, S.; Zhu, Y.; Wang, G.; Zhang, K.; Li, Z.; Yu, E.; Tian, J.; Xia, Y.; Xie, J.; et al. Effect of the Aerobic Denitrifying Bacterium Pseudomonas furukawaii ZS1 on Microbiota Compositions in Grass Carp Culture Water. Water 2021, 13, 1329. https://doi.org/10.3390/w13101329
Gong W, Gao S, Zhu Y, Wang G, Zhang K, Li Z, Yu E, Tian J, Xia Y, Xie J, et al. Effect of the Aerobic Denitrifying Bacterium Pseudomonas furukawaii ZS1 on Microbiota Compositions in Grass Carp Culture Water. Water. 2021; 13(10):1329. https://doi.org/10.3390/w13101329
Chicago/Turabian StyleGong, Wangbao, Shuwei Gao, Yun Zhu, Guangjun Wang, Kai Zhang, Zhifei Li, Ermeng Yu, Jingjing Tian, Yun Xia, Jun Xie, and et al. 2021. "Effect of the Aerobic Denitrifying Bacterium Pseudomonas furukawaii ZS1 on Microbiota Compositions in Grass Carp Culture Water" Water 13, no. 10: 1329. https://doi.org/10.3390/w13101329
APA StyleGong, W., Gao, S., Zhu, Y., Wang, G., Zhang, K., Li, Z., Yu, E., Tian, J., Xia, Y., Xie, J., & Ni, J. (2021). Effect of the Aerobic Denitrifying Bacterium Pseudomonas furukawaii ZS1 on Microbiota Compositions in Grass Carp Culture Water. Water, 13(10), 1329. https://doi.org/10.3390/w13101329







