Abstract
The effects of hydraulic retention time (HRT) and influent nitrate-N concentration on nitrogen removal and the microbial community composition of an aerobic denitrification reactor treating recirculating marine aquaculture system effluent were evaluated. Results showed that over 98% of nitrogen was removed and ammonia-N and nitrite-N levels were below 1 mg/L when influent nitrate-N was below 150 mg/L and HRT over 5 h. The maximum nitrogen removal efficiency and nitrogen removal rate were observed at HRT of 6 or 7 h when influent nitrate-N was 150 mg/L. High-throughput DNA sequencing analysis revealed that the microbial phyla Proteobacteria and Bacteroidetes were predominant in the reactor, with an average relative total abundance above 70%. The relative abundance of denitrifying bacteria of genera Halomonas and Denitratisoma within the reactor decreased with increasing influent nitrate-N concentrations. Our results show the presence of an aerobically denitrifying microbial consortium with both expected and unexpected members, many of them relatively new to science. Our findings provide insights into the biological workings and inform the design and operation of denitrifying reactors for marine aquaculture systems.
1. Introduction
Nitrogenous contamination of surface waters is a serious environmental problem, causing eutrophication of rivers, lakes, and near-shore marine environments [1,2] and raising the likelihood of toxic algal blooms in receiving waters [3,4]. Accumulation of nitrogenous wastes is also a problem in aquaculture systems [5], posing toxicity to cultured organisms [6,7,8,9]. Microbially mediated nitrification in biofilters is commonly used to convert ammonia-N to nitrite-N and further to nitrate-N [10]. In recirculating aquaculture systems (RASs), however, the low rate of water replacement may lead to the accumulation of nitrate-N in rearing water, to concentrations as high as 400–500 mg/L [11], bringing about the need for conversion of nitrate-N to N2 via denitrification.
The denitrification process in biological treatment of wastewater is affected by environmental conditions [12,13,14]. In particular, dissolved oxygen (DO) concentration, temperature, C/N ratio, carbon source, pH, nitrate loading rate, and hydraulic retention time (HRT) affect the performance of the aerobic denitrification process [15,16,17,18]. Although it was long thought that denitrification would not occur in the presence of oxygen, subsequent experiments have since demonstrated that aerobic denitrification is indeed mediated by a broad range of microbes, especially in environments with fluctuating oxygen concentrations and available reduced carbon [19]. These factors have also been linked to nitrite-N and ammonia-N generation and their accumulation in the denitrification reactor [20,21]. Because most denitrifying bacteria are heterotrophic, an external carbon source is often applied as a substrate and an electron donor to facilitate cell growth and nitrate-N reduction [11,22,23]. Methanol, ethanol, and sodium acetate are commonly used in laboratory- and industrial-scale biological wastewater treatment processes [24], and can be directly applied into the treatment process [25]. Despite considerable advances in promoting effective denitrification in freshwater systems [26,27,28,29], there is less information on the performance of denitrification systems in water with high salinity, including wastewater from marine aquaculture systems. Both nitrification and denitrification were achieved by the sequential use of a batch reactor treating low-salinity shrimp-production wastewater under aerobic and anaerobic conditions [30,31]. Borges et al. [32] showed both the aerobic and anaerobic denitrification ability of a Pseudomonas bacterium isolated from a marine RAS. Gutierrez-Wing et al. [33] showed effective denitrification using four polyhydroxybutyrate media at different levels of salinity, dissolved oxygen and nitrate-N. Zhu et al. [34] suggested that the presence of salinity (25% vs. 0%) could improve the nitrate-N removal efficiency and stability of a poly-butanediol succinate (PBS)-supported aquaculture denitrification system. With increasing development of the marine aquaculture sector, further investigation of the effects of environmental factors on nitrogen removal, accumulation of intermediates, and characterization of the microbial community in high-salinity recirculating aquaculture wastewater is timely.
The microbiota active within biofilms attached to the media within biofilters mediate the denitrification process. Our understanding of the functional dynamics of the microbial communities, while still incomplete, is advancing with the application of molecular microbial identification techniques [35]. Due to its presence in most microbes and its slow rate of evolution [36], the 16S rRNA gene is widely used for the classification and molecular identification of microbes. Using the approach, Michaud et al. [37] showed that the microbial community in a biofilter differed from that in the rearing water of an aquaculture system. Keuter et al. [38] showed that the nitrite-oxidizing community in a biofilter consists of at least two representatives of genus Nitrospira. Huang et al. [39] collected samples from nine biofilters at five commercial recirculating aquaculture facilities with three different biofilter types, and showed the high abundance and diversity of nitrifying bacteria across these systems. Rud et al. [40] showed that microbial communities differed between culture water and bioreactor biofilms and between salinities in salmon production systems. Brailo et al. [41] showed distinct, diverse microbial communities in a commercial inoculant and in nitrifying and denitrifying biofilters. Reviewing the literature, Ortiz-Estrada et al. [35] suggested the use of metagenomic 16s rRNA data to gain a deeper understanding of the functional profiles of microbial communities in aquaculture systems. With the ongoing development of biofiltration technology and increased knowledge of microbial diversity, there have been reports of new microbes utilizing different nitrogen removal pathways, including nitratation, nitrifier denitrification, anaerobic ammonium oxidation (anammox), and combined pathways [42], including aerobic denitrification [15,16,17,18,19,20]. Using the metagenomics of the 16S rRNA gene for microbial community analysis of autotrophic denitrification, Zhang et al. [43] showed that the abundances of genera Azoarcus, Thauera, and Aliidiomorina were positively related to the aeration rate.
Hypothesizing that biofilter loading and hydraulic retention time (HRT) would directly affect the microbial community structure and function and hence the biofilter efficacy, the objectives of the present study were: (1) to evaluate the nitrogen removal performance of an aerobic denitrification reactor treating recirculating marine aquaculture system effluent under different influent nitrate-N concentrations and HRTs, and (2) to characterize the microbial community in the aerobic denitrification reactor using 16S rDNA sequencing technology. Our findings offer useful information about microbial responses to operational factors and enhance our understanding of the denitrification processes for the treatment of marine aquaculture wastewater.
2. Materials and Methods
2.1. The Aerobic Denitrification Reactor
A schematic diagram of the experimental marine recirculating aquaculture system is shown in Figure 1. The bioreactor column was fabricated with an internal diameter of 12 cm and height of 25 cm, with a total volume of 2.8 L. The working volume in the bioreactor was 2.2 L, with a height of 19.5 cm. The Kaldness-1 packing (Dynamic Aqua Science, Inc., Laguna Beach, CA, USA) used in the bioreactors had a density of 0.96 kg/m3 and a specific surface area of 850 m2/m3, occupying 40% of the reactor volume. Wastewater was fed to the reactor from the bottom of the reactor using a peristaltic pump. The hydraulic retention time (HRT) was adjusted according to the requirements of a particular experimental phase. An air compressor supplied air through an air stone placed at the bottom of the reactor, with dissolved oxygen (DO) concentrations ranging from 5.0 to 7.0 mg/L. The temperature of the reactor was controlled at 25 ± 1 °C by a water bath. The pH value was approximately 7.8, and the salinity was about 35%.
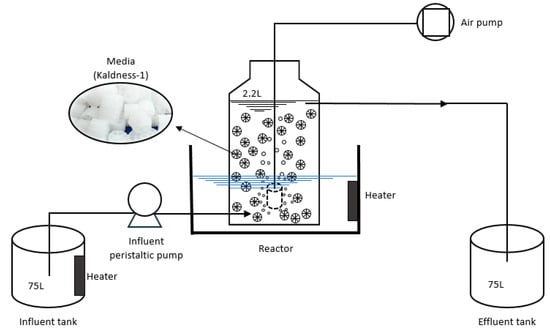
Figure 1.
Schematic diagram (not to scale) of the denitrification assembly.
2.2. Experimental Design
The reactor was inoculated with aerobic denitrifying bacteria, Halomonas venusta, isolated by Jiang et al. [44]. Prior to the trial, the reactor, with a theoretical hydraulic retention time (HRT) of 4 h, was operated for two weeks to achieve stable performance. Generally, after the effluent nitrate-N and total nitrogen concentration were stable for at least for three days, the operational mode was changed to meet the requirements of the following experimental phase (Table 1). Recirculating marine aquaculture wastewater was from Laizhou Mingbo Aquatic Co., Ltd. (Laizhou, China). The desired nitrate-N input concentration was produced by adding analytical grade NaNO3 (Sinopharm Chemical Reagent Co., Ltd., Shanghai, China). Preliminary tests of alternative carbon sources (glucose, sodium succinate, and trisodium citrate; unpublished data) upon the performance of Halomonas venusta [45] showed that the highest denitrification rate was obtained using acetate. Hence, approximately 0.5, 1.0, or 1.5 g/L analytical grade sodium acetate (Sinopharm Chemical Reagent Co., Ltd., Shanghai, China) was added as the main source of organic carbon, to make sure the C/N ratio was around 6/1.

Table 1.
Operational parameters for the respective experimental phases.
2.3. Water Samples and Quality Analysis
Ammonia-N, nitrite-N, nitrate-N, and total nitrogen concentrations in the influent and effluent were measured every other day according to Standard Methods [46]. A YSI 85 probe (DO200, YSI, Inc., Yellow Springs, OH, USA) was used to determine temperature, and dissolved oxygen (DO). The pH was measured using a pH 100 m (YSI, Inc.).
2.4. DNA Sample and DNA Extraction
For DNA extraction and microbial community analysis, 50 g of Kaldness K1 polyethylene media with biofilm was collected from the aerobic denitrification reactor at the end of each experimental phase. Huang et al. [39] evaluated several methods—such as vortex, shaker, and sonication—to collect the biofilm. Following those findings, the Kaldness K1 polyethylene media was sliced into small pieces, shaken in a shaker, and centrifuged [39]. Deoxyribonucleic acid was extracted using the Fast DNA Spin Kit for Soil (MP Biomedicals, Santa Ana, CA, USA) according to the manufacturer’s instructions. The V3–V4 hypervariable region of the 16S rRNA gene was amplified from genomic DNA using the primers 338F (5′-ACTCCTACGGGAGGCAGCAG-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′) for bacteria and archaea [47,48]. The PCR amplification conditions for bacterial 16S rRNA genes were as follow: initial denaturation at 95 °C for 3 min; 35 cycles of denaturation (95 °C; 30 s), annealing (55 °C; 30 s), and extension (72 °C; 1 min); and a final elongation (72 °C, 10 min). The PCR products (3 mL) were purified by 2% agarose gel electrophoresis. After purification, the products were quantified, and a barcoded library comprised of the products of the respective treatments was prepared. Amplicons from the samples were sequenced using the Illumina-MiSeq platform at Shanghai Majorbio Bio-Pharm Technology Co., Ltd. (Shanghai, China). Raw sequencing reads were deposited in the NCBI Sequence Read Archive database with accession no. SUB5923142.
2.5. Processing of Sequencing Data
Raw FASTQ files were quality-filtered using Trimmomatic [49] and merged using the Fast Length Adjustment of Short Reads [50] computational tool. The reads were first truncated at sites receiving an average quality score of <20 over a 50-bp sliding window. Sequences whose overlap was longer than 10 base-pairs (bp) were merged according to their overlap with no more than a 2-bp mismatch. Sequences of each sample were separated according to barcodes (exactly matching) and primers (allowing 2 nucleotide mismatches), and reads containing ambiguous bases were removed.
Operational taxonomic units (OTUs) were first clustered with a 97% similarity cutoff using UPARSE version 7.1 [51] with a novel ‘greedy’ algorithm that performs chimera filtering and OTU clustering simultaneously, and then the chimeric sequences were identified and removed using UCHIME [52]. Trimmed reads were subsampled according to the minimum reads number of the 10 data groups for bacteria or archaea, and the OTU numbers were recalculated. The phylogenetic affiliation of each 16S rRNA gene sequence was analyzed using the RDP Classifier (version 2.2) algorithm [53] against the Silva (Release 132, http://www.arb-silva.de) 16S rRNA database using a confidence threshold of 70%. In order to compare the variability of each sample fairly at the same sequence depth, reads were normalized from each high-throughput dataset for further analysis [54]; read numbers were normalized to the 21,333 reads obtained from sample H4_N50 for further analysis of data from each high-throughput data run.
2.6. Statistical Analysis
Alpha diversity indices—including the observed number of Operational Taxonomic Units (OTUs), community richness indices (observed richness-Sobs, abundance-based coverage estimator—ACE, and Chao1), community diversity indices (Shannon and Simpson), community evenness indices (Shannoneven and Simpsoneven) and Good’s community coverage indices (coverage)—were calculated using MOTHUR project software [55]. Heatmap analysis was conducted using the R package “heatmap”. A weighted UniFrac distance metric [56] was calculated for β-diversity comparison across microbial communities; principal components analysis (PCoA) was used based on the Bray‒Curtis dissimilarity using the vegan package in R (v. 3.5.1).
Detrended correspondence analysis (DCA) [57] was applied to indicate the best methodology for finding the main factors that influence community composition. Redundancy analysis (RDA) [58] was carried out using the vegan package in R (v. 3.5.1). The environmental variables with variance inflation factors (VIF) < 10 were selected. Both DCA and RDA were performed using CANOCO 4.5 for Windows software (Microcomputer Power Co., Ithaca, NY, USA) to reveal the correlations between the microbial community and operational parameters. Gephi0.9.1 software (https://gephi.org/) and the Fruchterman and Reingold [59] algorithm were used to create a graphical network showing microbial interactions and correlations of the abundances of microorganisms with process parameters [47,48].
2.7. Statistical Analyses
The volumetric nitrate-N conversion rate (NRR; mg·N·m−3·d−1), and nitrate-N removal efficiency (NRE; %) were calculated [25,60] as follows:
where nitrate-Ninf (mg N/L) is the nitrate-N concentration in the influent, nitrate-Neff (mg N/L) is the nitrate-N concentration in the effluent, Q is total flow passing through the biofilter (L·min-1), and Vmedia is the media volume (m3). We determined the nitrate removal rate only after a new stable condition was established for a particular set of operating conditions. Repeat measurements were performed when we calculated NNR and NRE. While a nitrogen balance approach is often used for the characterization of the denitrification performance of a bacterium, it is not suitable for characterizing the performance of an aerobic bioreactor, as it is problematic to sample N2 when an air diffuser is used in the reactor.
All data were analyzed statistically via one-way ANOVA using SPSS 22.0 (IBM Corporation, Armonk, NY, USA). A probability level of 0.05 was used to assess the statistical significance of differences among means.
3. Results
3.1. Performance of Nitrogen Removal
The effects of four HRTs and three nitrate-N loadings on performance of the denitrification reactor are shown in Table 2 and Figure 2. After 40 days of biofilter acclimation, the effluent nitrate-N concentrations decreased from 4.04 ± 0.49 to 0.31 ± 0.24 mg/L in the denitrification reactor (Table 2). When influent nitrate-N concentrations were approximately 50 mg/L (as in phases H4-7_N50), over 92% of nitrate-N was removed. Increasing influent nitrate-N to 100 mg/L (phases H5-7_N100) resulted in the effluent nitrate-N sharply increasing to 10.63 ± 0.91 mg/L, but nitrate-N removal efficiency changed indistinctively. However, when influent nitrate-N was above 150 mg/L (phases H5-7_150), the effluent NO2−N concentration increased substantially, and the total nitrogen and nitrate-N removal efficiencies were only 81.3% and 93.5%, respectively.

Table 2.
Long-term performance of the denitrification process under different experimental conditions.1 TN = total nitrogen, NRE = nitrate-N removal efficiency, and NRR = volumetric nitrate-N conversion rate.
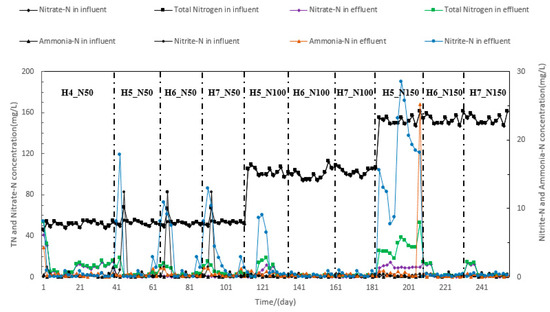
Figure 2.
Long-term nitrogen removal performance of the denitrification reactor at different hydraulic retention times (H, in hours) and influent nitrate-N concentrations (N, in mg/L).
The nitrate-N removal performance of the CH3COONa-supported denitrification reactor under hydraulic retention times of 4, 5, 6, or 7 h with different influent nitrate-N concentrations is shown in Figure 2. As expected, increasing HRT led to decreasing effluent nitrate-N concentrations and increasing nitrate-N removal efficiencies. Through the whole operating period, the majority of effluent NH4−N concentrations were under 1 mg/L and large nitrite-N accumulations were observed in H4_N50, H5_N100 and H5_N150. To maximize nitrogen removals in the aerobic denitrification process, the results showed an optimum HRT of 5 h at 50 mg/L and 6 h at 100 and 150 mg/L influent nitrate-N concentrations. After increases in nitrate-N loading, the biofilters generally took several days to bring effluent nitrogen species to equilibrium conditions; the one exception was treatment H5_N150.
3.2. Microbial Community Analysis
3.2.1. Richness and Alpha Diversity Analyses of Bacterial Communities
To assess the biological response of the biofilters to different HRTs and nitrate-N loadings, we characterized the microbial communities of each experimental phase after the effluent of the reactor at that phase had been stable for at least three days. The average read length of high-quality sequence tags was 395 bp. The DNA sequences obtained (Table 3) captured most of the bacteria present, based on high values of Good’s coverage index (>99%), indicating that almost all the OTUs of the reactor were observed in the sequencing results. Rarefaction curves of all samples arrived at the plateau phase, supporting this interpretation. We conclude that the results of the diversity analysis had high reliability.

Table 3.
Analysis of adequacy of sampling and richness and diversity of the microbial community.
We evaluated three metrics of species richness. The mean numbers of species observed, Sobs, was 215 for treatments with nitrate-N of 50 mg/L, 213 for treatments of 100 mg/L, and 179 for treatments of 150 mg/L. Sobs was 174 for the one treatment with an HRT of four hours; mean Sobs values were 213 for treatments of 5 h and 204 for those of 6 or 7 h. The mean abundance-based coverage estimator, ACE, was 247.5 for treatments with nitrate-N of 50 mg/L and 245.6 for those of 100 mg/L, and 211.6 for those of 150 mg/L. ACE was 198 for the one treatment of HRT = 4, with means of 248.9, 244.5, and 234.3 for those of HRT of 5, 6, or 7 h, respectively. The mean Chao1 index was 251.0 for treatments of 150 mg/L nitrate-N, 242.2 for those of 100 mg/L, and 210.0 for those of 150 mg/L. Chao1 was 214.6 for the one HRT = 4 treatment, a mean of 246.1 for HRT = 5, 238.9 for HRT = 6, and 230.3 for HRT = 7 h. The total numbers of species and bacterial diversity differed among the experimental phases. The Chao 1 estimator indicated that the biofilm in phase H6_N50 had the greatest species richness, especially in comparison to that in phase H6_N150, which showed the smallest.
Among two metrics of species diversity, i.e., accounting for differences in relative abundance of the respective species, the Shannon index was higher (about 3.63) under 50 or 100 mg/L influent nitrate-N than that (about 3.35) when influent nitrate-N was 150 mg/L, suggesting that the microbial community diversity decreased with increasing influent nitrate-N concentration. Simpson diversity indices (about 0.05) were somewhat lower for phases with 50 or 100 mg/L than for those with 150 mg/L (about 0.069). Shannon diversity indices were lower at lower HRTs (about 3.6) than at HRT = 7 h (about 3.4). Similarly, Simpson indices were lower at lower HRTs (about 0.05) than at HRT = 7 (0.067). Both the Shannoneven and Simpsoneven indices were higher for H4_N50 than for other treatments. Clearly, changes in the mode of operation of the biofilter affected the microbial diversity.
3.2.2. Bacterial Community Composition at the Phylum and Genus Levels
The microbial communities present in each experimental phase—characterized at the phylum, class, family, and genus levels—are shown in Figure 3. Fourteen main phyla were detected in the DNA sequences recovered from biofilms from the respective experimental phases. Phyla detected (Figure 3a) included Proteobacteria (53–69%), Bacteroidetes (5.6–28%), Epsilonbacteraeota (0.6–18%), Cloacimonetes (0.2–16.4%), Spirochaetes (1.7–5.4%) and Firmicutes (1.3–10.7%). The phylum Proteobacteria dominated each sample, while the relative abundances of the other respective phyla differed. The relative abundance of Firmicutes was high in phase H4_N50, and decreased with longer HRT. Meanwhile, other phyla with members that participate in the nitrogen cycle, such as Planctomycetes [54], increased in relative abundance in the reactor.
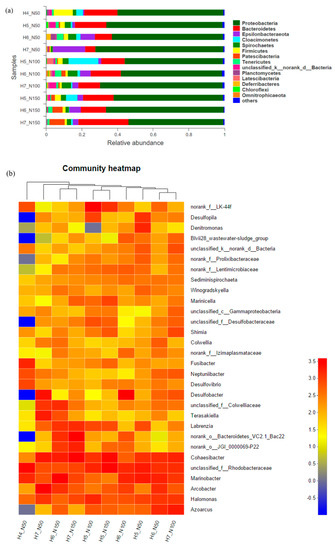
Figure 3.
(a) Taxonomic classification of microbes at the phylum level within the biofilter in the respective experimental phases. (b) Heat map of the top 30 genera in each phase, representing the relative abundance and distribution of the representative 16S rRNA sequences at the genus level. Heat map colors correspond with the relative abundance of each phylogenetic group, ranging from blue (low abundance), to yellow, to red (high abundance).
To gain a deeper understanding of the effects of changing bioreactor operating conditions, the microbial community was characterized at the genus level. As illustrated in Figure 3b, heatmap analysis showed distinctions in the community structure among the respective experimental phases. In the first phase (H4_N50), the bacterial community was rather distinctive, as indicated by its location in the most basal branch of the heatmap cluster analysis. This community was distinct in terms of low relative abundance of genera Desulfopila, Desulfobacter, and Azoarctus, families Prolixibacteraceae, Lentimicrobiaceae, and Desulfobacteriaceae, and phylum Bacteriodetes; and higher relative abundance of genera Fusibacter and Desulfovibrio. That is, its sulfur-reducing and nitrogen-fixing communities in H4_N50 were distinct from those in other experimental phases. The overall community then changed, but remained comparable with subsequent operating conditions. Focusing on bacteria affecting nitrogen processing, Halomonas and Azoarcus were abundant and clustered in the community heatmap analysis across all phases. The genera most frequently identified among the 10 experimental phases were Cohaesibacter, Marinobacter, and Halomonas, which have been found in marine environments [61,62]. The most dominant genera in phase H4_N50 were Rhodobacteraceae (13.7%), Fusibacter (8.0%), and Halomonas (6.6%), while the dominant ones changed to Cohaesibacter (17.7%) and Labrenzia (7.1%) in H7_N150. Marine Rhodobacteraceae are key players in biogeochemical cycling and show a versatile physiology to survive in a highly variable marine habitat [63] and account for a large percentage of bacterioplankton communities [64,65,66]. Fusibacter plays an important role in a process that reduces elemental sulfur, but not sulfate, thiosulfate, or sulfite into sulfide [67]. Sulfate-reducing bacteria, such as Desulfovibrio, Desulfobacter, Desulfobacteraceae, and Desulfotignum, were present in all experimental phases, presumably because of the high sulfate concentration in seawater. The proportion of Labrenzia increased with HRT and nitrate-N loading, which suggested the stability of the biofilm. Priyanka et al. [68] proposed the production of a sulfated exopolysaccharide (EPS) by Labrenzia sp. PRIM-30; many marine bacteria produce EPS as a strategy for adhering to solid surfaces, survival, and growth under adverse conditions. Halomonas, which we inoculated into the reactor, accounted for only 2% of observations at the end of the first phase, H4_N50, and its relative abundance increased with nitrate-N removal. In addition, some Halomonas species have been shown to be able to perform denitrification to gain energy through the processing of nitrate-N to nitrogen [45,69,70].
3.2.3. Community Composition and Response to Environmental Variables
Principal coordinates analysis (PCoA) of the relative abundance of OTUs at the genus level showed how the composition of bacterial communities changed among experimental phases. Proximity of points representing the respective treatments in principal coordinates space (Figure 4a) indicates that their phylogenetic compositions were more similar to one another than to those of treatments shown at more distant locations. The first (PC1) and the second (PC2) components accounted for 29.94% and 18.94% of the variability in community composition, respectively, explaining just less than half of the variability observed. These results showed four distinct microbial community assemblages during the succession through experimental phases. As shown in Figure 4a, these communities were more similar if they shared the same HRT as opposed to the same N-loading, especially in experimental phases that showed stable nitrate-N removal. These observations imply that microbial community composition was related not only to HRT and nitrate-N loading, but also to having attained equilibrium in nitrogen removal performance.
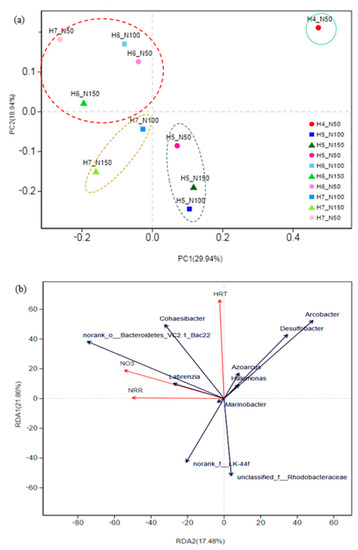
Figure 4.
Correlation analysis for microbial communities. (a) Principal coordinates analysis (PCoA) of microbial communities of 10 samples. The operational taxonomic units (OTU) were identified by 16S rRNA gene sequencing. Phylogenetic distances between samples were determined using the Bray‒Curtis algorithm. (b) Redundancy analysis (RDA) illustrating relationships between operational variables (HRT, nitrate concentration, and NRR), and dominant genera.
RDA analysis was further used to analyze the relationship between the relative abundance of predominant bacteria (at the genus level) and the operational variables (HRT, nitrate concentration and NRR) (Figure 4b). The pattern of relative abundances of genera within the temporal succession of experimental phases in the RDA results were similar to those shown by PCoA. The lengths of the arrows show the extent to which the relative abundance of dominant genera is explained by operational variables. Variation in the operational variables drove the presence of ten most dominant genera in the bacterial communities of the respective samples. The angles of the vectors indicated that the genera were closely related with HRT, nitrate concentration and NRR. HRTs were associated with high relative abundances of Halomonas, Cohaesibacter, Marinobacter, Arcobacter, and Desulfobacter, and with relatively good bioreactor performance. Figure 4b again shows that nitrogen removal was most affected by HRT, and that process efficiency was negatively correlated with influent nitrate-N concentration.
4. Discussion
4.1. Process Performance
We tested the effects of four hydraulic loading times and three nitrate-N loadings on the effectiveness and microbial community composition of an aerobic denitrification reactor. We noted that, with a prolonged operation time, nitrogen removal became more complete and relatively stable, a result similar to those of earlier studies. Using a moving bed biofilm reactor, Zinatizadeh and Ghaytooli [71] showed that the reactor could achieve higher TN removal efficiency when HRT was increased from 4 to 8 h. Wang and Wang [72] and Ovez et al. [73] showed that lower HRT caused effluent nitrate-N and nitrite-N to rise, results that are similar to those of our study. Zhu et al. [74] showed that increasing HRT led to increased organic loading, hydraulic loading, and liquid shear, which benefit biofilm formation and the growth of aerobic denitrifying bacteria. Although an increase in HRT might allow the bacterial population to degrade organic substrates and lead to high nitrate-N removal efficiency, it is also responsible for the high release of dissolved organic carbon and ammonium [26]. Hence, it is crucial to choose an appropriate hydraulic retention time to complete the aerobic denitrification process.
Meanwhile, Qiu et al. [75] found that when the influent nitrate-N concentration increased with limited HRT, the result was insufficient nitrate-N removal and the accumulation of nitrite-N. In our study, massive accumulation of nitrite-N was found with 150 mg/L influent nitrate-N concentration with a limited HRT of 4 h. Körner et al. [76] believed that the presence of high nitrate-N concentration inhibited the activity of nitrite-N reductase (Nirs) in Pseudomonas stutzeri, resulting in accumulation of nitrite-N in a chemostat. Higher effluent nitrate-N and nitrite-N might result from shorter HRT and higher influent nitrate-N concentration due to the dissimilatory nitrate-to-ammonia process (DNRA). Effluent ammonia-N generated by DNRA was observed in the reduction of nitrate-N during the denitrification process [77,78]. Denitrovibrio sp. was isolated as a representative of a microbial population reducing nitrate-N to nitrite-N and then to ammonia-N [79]. In our experiments, however, the frequency of occurrence of Denitrovibrio was <0.001% in low-HRT treatments H4_N50, H5_N100 and H5_N150, but more than 1% in high-HRT treatments H6_N50 and H6_N100, where it presumably helped reduce nitrite-N accumulation and promote nitrate-N removal. At a practical level, it would be preferable to operate a biofilter with a short HRT, as it could save space and time. Alternatively, HRT could be lengthened by increasing the size of the biofilter, and some previous studies [80,81] have used that approach for RAS systems operated in areas where regulatory standards for discharge of nitrogen is regulated.
The characteristics of carbon sources have major effects upon important aspects of the denitrification process, including denitrification rate and accumulations of intermediate products [82]. Among possible substrates, CH3COONa can be used directly and without modification by most denitrifying bacteria [83,84]. Our results revealed maximum nitrogen removal efficiency in treatment H7_N150 and nitrogen removal rate in H6_N150 (Table 2). Moreover, we found that the proportion of the bacterium Labrenzia sp., which can produce exopolysaccharides to improve stability of biofilm and increase its growth [68], was significantly higher in treatments H6_N150 and H7_N150. Carbon and nitrogen sources have to cross the biofilm–liquid interface and be transported through the biofilm to reach the microbial cells and to be consumed [85]. Other research has reported decreased permeability with increasing biofilm thickness [77,86]; although the rise of Labrenzia may increase biofilm thickness, we did not find correspondingly decreased biofilm activity.
4.2. Microbial Community Structure
To understand the biological factors underlying the relative effectiveness observed in our denitrifier, we characterized the microbial communities within the respective experimental treatments. At the phylum level, our DNA sequencing results revealed that different microbial community compositions are related to changes in system operating conditions, including relative abundances of key denitrifying bacteria and other functionally important microorganisms. More than 70% of the sequences that we identified were members of the phyla Proteobacteria and Bacteroidetes, which predominated in the reactor with average relative abundances of 61.9% and 15.3%, respectively. Both of these are widely reported as including denitrifying bacteria [87,88], predominant phyla commonly found in activated sludge systems [89], as well as microbes with many other functions. Our findings parallel those of Chu and Wang [77], who found that Proteobacteria contributed over 80% of the total bacterial sequences in a fixed bed bioreactor filled with the biopolymer polycaprolactone (PCL). Proteobacteria and Bacteroidetes accounted for 62.7% and 20% in an integrated solid-phase denitrification biofilter (SDNF) using KNO3 and PCL as the main sources of nitrate-N and organic carbon, respectively [27]. The major functional role of Bacteroidetes is to break down macromolecules, which is important for hydrolyzation and the utilization of solid carbon sources [90].
Within the microbial communities that we characterized, the presence of some taxa would be expected (e.g., Halomonas, Denitromonas, Denitrovibrio, Azoarcus) due to their utilization of nitrogen species for metabolism. Yet, many taxa metabolized substrates other than nitrogen species or with unknown function, reflecting the workings of a microbial consortium. Observation of some taxa within the community was surprising, e.g., Colwellia (an extremophile for cold temperatures), Arcobacter (a common sulfur-oxidizing and dissimilatory nitrate reduction to ammonia bacterium [91,92]; most members of the genus are not halophiles), and Fusibacter and Acholeplasma (some members of the genera are pathogens). Both aerobic (e.g., Halomonas, Arcobacter, and Flavobacteriaceae) and anaerobic taxa (e.g., Neptunomonas, Desulfotignum, and Clostridiales) were present within the same biofilter, the anaerobes presumably within anaerobic microhabitats. Our ability to interpret interactions among members of the microbial community was limited by the developing state of knowledge of marine microbiology. Many species (e.g., Celeribacter marinus [93]), genera (e.g., Neptuniibacter, gen. nov. [94], Xanthomarina, gen. nov. [95]), and even families (Prolixibacteraceae, fam. nov. [96], Salinivirgaceae, fam. nov. [97]) are recently described, and some sequences in our dataset cannot be linked to formally described species (e.g., HTCC5015 and several “unclassified” microbes). For many taxa, substrates and metabolic pathways are poorly known or unknown (e.g., Marincella, Shimia, and Izimiaplasmataceae). Applied research needs to clarify the correlations between microbial interactions and process stability, and reveal the mechanisms underlying stability of biofilter function. We observed that species richness and diversity declined at higher N loadings and increased with HRT (Table 3). We presume that more diverse, species-rich microbial communities with greater functional redundancy might prove more robust to changes in operating conditions, and hence would more reliably process nitrogenous and other aquaculture waste products. This view would suggest design of denitrification biofilters to operate at nitrate-N loadings of ≤100 mg/L and HRTs of as much as 5 h. That time is too long for many practical applications, and shortening of HRT has been achieved by use of granules [98], a technical possibility that we did not evaluate.
At the genus level, the relative abundances of key microorganisms clearly differed among reactors in our and previous studies. Many Desulfovibrio, Tolumonas, Brevinema, and Ideonella species are efficient degraders of high molecular-weight organic substrates into short-chain fatty acids and other low molecular-weight soluble compounds [99]. Marinobacter are hydrolyzing and fermenting bacteria, which degrade aliphatic and polycyclic aromatic hydrocarbons as well as acyclic isoprenoid compounds [100]. We found 18% and 9% frequencies of occurrence of Marinobacter and Desulfovibrio species, respectively, which can incompletely oxidize compounds such as volatile fatty acids and lactate to acetate [100,101]. Genera Desulfobacter and Desulfopila are sulfate-reducing bacteria, some species of which can reduce nitrate-N or nitrite-N to ammonia-N [102], driving the dissimilatory nitrate-to-ammonia process in the reactor. Desulfobacter and Desulfopila were detected in a constructed wetland to which starch/polylactic acid blends were added to support denitrification [103]. Denitratisoma was found capable of complete denitrification of wastewater [104,105]. We observed both Halomonas and Denitratisoma in our reactor, whose frequencies decreased with increasing influent nitrate-N concentration. Further characterization of the metabolic functions of particular microbes would advance insights into the significance of the presence and relative abundances of particular microbes in biofiltration systems.
5. Conclusions
We quantified the effects of hydraulic retention time (HRT) and influent nitrate-N concentration on nitrogen removal and microbial community composition of an aerobic denitrification reactor at a bench scale. Our results showed that increased HRT and the reduction of influent nitrate-N concentration promoted the denitrification process, with HRT having the stronger effect on aerobic denitrification process and microbial community composition. Our results show that it is critical to choose a suitably long hydraulic retention time in order to complete the aerobic denitrification process. To assess the functional biological underpinnings of biofilter function, we employed DNA sequencing analyses, which revealed the complexity of community structure and the presence of key denitrifying bacteria, explaining the high nitrate-removal performance and the low nitrite-N accumulation displayed by the bioreactor. These results suggest that greater community species diversity and richness promote more complete degradation of nitrogenous wastes. Our findings highlight the role of environmental variables upon denitrification, operator control of which is critical for managing bacterial community structure and fostering process efficiency. Assessment of a commercial-scale aerobic denitrification reactor operated under practical conditions is needed in future study. Our results will inform the design, evaluation, and operation of practical aquaculture biofiltration systems, especially those for saltwater production.
Author Contributions
Conceptualization, X.S. and Z.H.; Data curation, X.Y. and Y.J.; Formal analysis, X.Y. and Z.H.; Funding acquisition and Project administration, Z.H.; Writing—original draft, X.S., and X.Y.; Writing—review and editing, E.H., and Z.H. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Science Foundation of China (31502212) and the National Key Research and Development Program of China (2017YFD0701701, 2019YFD0900503).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Vollenweider, R.A. Scientific Fundamentals of The Eutrophication of Lakes and Flowing Waters, with Particular Reference to Nitrogen and Phosphorus as Factors in Eutrophication; OECD: Paris, France, 1970. [Google Scholar]
- Ryther, J.H.; Dunstan, W.M. Nitrogen, phosphorus, and eutrophication in the coastal marine environment. Science 1971, 171, 1008–1013. [Google Scholar] [CrossRef] [PubMed]
- Heisler, J.; Gilbert, P.M.; Burkholder, J.M.; Anderson, D.M.; Cochlan, W.; Dennison, W.C.; Dortch, Q.; Gobler, C.J.; Heil, C.A.; Humphries, E.; et al. Eutrophication and harmful algal blooms: A scientific consensus. Harmful Algae 2008, 8, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Gobler, C.J.; Burkholder, J.M.; Davis, T.W.; Harke, M.J.; Johengen, T.; Stow, C.A.; Van de Waal, D.B. The dual role of nitrogen supply in controlling the growth and toxicity of cyanobacterial blooms. Harmful Algae 2016, 54, 87–97. [Google Scholar] [CrossRef] [PubMed]
- Boyd, C.E. Water Quality Management for Pond Fish Culture; Elsevier Scientific Publishing Co.: Amsterdam, The Netherlands, 1982. [Google Scholar]
- Camargo, J.A.; Alonso, A.; Salamanca, A. Nitrate toxicity to aquatic animals: A review with new data for freshwater invertebrates. Chemosphere 2005, 58, 1255–1267. [Google Scholar] [CrossRef]
- Davidson, J.; Good, C.; Welsh, C.; Summerfelt, S.T. Comparing the effects of high vs. low nitrate on the health, performance, and welfare of juvenile rainbow trout Oncorhynchus mykiss within water recirculating aquaculture systems. Aquacult. Eng. 2014, 59, 30–40. [Google Scholar] [CrossRef]
- Yang, X.; Hu, F.; Guo, W.; Hallerman, E.; Song, X.; Huang, Z. Acute and chronic toxicity of nitrate to greenlings (Hexagrammos otakii). J. World Aquac. Soc. in press. [CrossRef]
- Yang, X.; Song, X.; Peng, L.; Hallerman, E.; Huang, Z. Effects of nitrate on aquaculture production, blood and histological markers and liver transcriptome of Oplegnathus punctatus (Temminck et Schlegel, 1846). Aquaculture 2019, 501, 387–396. [Google Scholar] [CrossRef]
- Lekang, O.-I. Ammonia removal. In Aquaculture Engineering; Blackwell Publishing: Oxford, UK, 2007; pp. 121–132. [Google Scholar]
- Van Rijn, J.; Tal, Y.; Schreier, H.J. Denitrification in recirculating systems: Theory and applications. Aquac. Eng. 2006, 34, 364–376. [Google Scholar] [CrossRef]
- Luo, G.; Xu, G.; Gao, J.; Tan, H. Effect of dissolved oxygen on nitrate removal using polycaprolactone as an organic carbon source and biofilm carrier in fixed-film denitrifying reactors. J. Environ. Sci. 2016, 43, 147–152. [Google Scholar] [CrossRef]
- Ding, A.; Zhao, D.; Ding, F.; Du, S.; Lu, H.; Zhang, M.; Zheng, P. Effect of inocula on performance of bio-cathode denitrification and its microbial mechanism. Chem. Eng. J. 2018, 343, 399–407. [Google Scholar] [CrossRef]
- Xu, Z.; Dai, X.; Chai, X. Effect of influent pH on biological denitrification using biodegradable PHBV/PLA blends as electron donor. Biochem. Eng. J. 2018, 131, 24–30. [Google Scholar] [CrossRef]
- Liu, Y.; Li, G.; Chen, Z.; Megharaj, M.; Naidu, R. Removal of nitrate using Paracoccus sp. Yf1 immobilized on bamboo carbon. J. Hazard. Mater. 2012, 229, 419–425. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Wang, X.; Zang, M.; Zhou, J.; Wang, J.; Guo, W. Degradation pathways and kinetics of anthraquinone compounds along with nitrate removal by a newly isolated Rhodococcus pyridinivorans Gf3 under aerobic conditions. Bioresour. Technol. 2019, 285, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Su, J.; Shi, J.; Ma, F. Aerobic denitrification and biomineralization by a novel heterotrophic bacterium, Acinetobacter sp. H36. Mar. Pollut. Bull. 2017, 116, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Wang, T.; Ye, J.; Zhao, J.; Yang, L.; Wu, P.; Duan, J.L.; Ye, G. Effects of carbon sources and operation modes on the performances of aerobic denitrification process and its microbial community shifts. J. Environ. Manag. 2019, 239, 299–305. [Google Scholar] [CrossRef]
- Ji, B.; Yang, K.; Zhu, L.; Jiang, Y.; Wang, H.; Zhou, J.; Zhang, H. Aerobic denitrification: A review of important advances of the last 30 years. Biotechnol. Bioprocess Eng. 2015, 20, 643–651. [Google Scholar] [CrossRef]
- Hu, Z.; Zhang, J.; Xie, H.; Li, S.; Wang, J.; Zhang, T. Effect of anoxic/aerobic phase fraction on N2O emission in a sequencing batch reactor under low temperature. Bioresour. Technol. 2011, 102, 5486–5491. [Google Scholar] [CrossRef]
- Vacková, L.; Srb, M.; Stloukal, R.; Wanner, J. Comparison of denitrification at low temperature using encapsulated Paracoccus denitrificans, Pseudomonas fluorescens and mixed culture. Bioresour. Technol. 2011, 102, 4661–4666. [Google Scholar] [CrossRef]
- Lee, K.C.; Rittmann, B.E. Effects of pH and precipitation on autohydrogenotrophic denitrification using the hollow-fiber membrane-biofilm reactor. Water Res. 2003, 37, 1551–1556. [Google Scholar] [CrossRef]
- Grommen, R.; Verhaege, M.; Verstraete, W. Removal of nitrate in aquaria by means of electrochemically generated hydrogen gas as electron donor for biological denitrification. Aquac. Eng. 2006, 34, 33–39. [Google Scholar] [CrossRef]
- Lee, N.M.; Welander, T. The effect of different carbon sources on respiratory denitrification in biological wastewater treatment. J. Ferment. Bioeng. 1996, 82, 277–285. [Google Scholar] [CrossRef]
- Yang, W.; He, S.; Han, M.; Wang, B.; Niu, Q.; Xu, Y.; Chen, Y.; Wang, H. Nitrogen removal performance and microbial community structure in the start-up and substrate inhibition stages of an anammox reactor. J. Biosci. Bioeng. 2018, 126, 88–95. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chu, L. Biological nitrate removal from water and wastewater by solid-phase denitrification process. Biotechnol. Adv. 2016, 34, 1103–1112. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Zuo, J.; Wang, Y.; Zhao, J.; Tang, L.; Li, Z. Tertiary nitrogen removal for municipal wastewater using a solid-phase denitrifying biofilter with polycaprolactone as the carbon source and filtration medium. Water Res. 2016, 93, 74–83. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Song, L.; Dai, X.; Chai, X. PHBV polymer supported denitrification system efficiently treated high nitrate concentration wastewater: Denitrification performance, microbial community structure evolution and key denitrifying bacteria. Chemosphere 2018, 197, 96–104. [Google Scholar] [CrossRef] [PubMed]
- Qiu, T.; Xu, Y.; Gao, M.; Han, M.; Wang, X. Bacterial community dynamics in a biodenitrification reactor packed with polylactic acid/poly(3-hydroxybutyrate-co-3-hydroxyvalerate) blend as the carbon source and biofilm carrier. J. Biosci. Bioeng. 2017, 123, 606–612. [Google Scholar] [CrossRef]
- Boopathy, R.; Bonvillain, C.; Fontenot, Q.; Kilgen, M. Biological treatment of low-salinity shrimp aquaculture wastewater using sequencing batch reactor. Int. Biodeter. Biodegrad. 2007, 59, 16–19. [Google Scholar] [CrossRef]
- Fontenot, Q.; Bonvillain, C.; Kilgen, M.; Boopathy, R. Effects of temperature, salinity, and carbon: Nitrogen ratio on sequencing batch reactor treating shrimp aquaculture wastewater. Bioresour. Technol. 2007, 98, 1700–1703. [Google Scholar] [CrossRef]
- Borges, M.T.; Sousa, A.; De Marco, P.; Matos, A.; Hönigová, P.; Castro, P.M. Aerobic and anoxic growth and nitrate removal capacity of a marine denitrifying bacterium isolated from a recirculation aquaculture system. Microb. Ecol. 2008, 55, 107–118. [Google Scholar] [CrossRef]
- Gutierrez-Wing, M.T.; Malone, R.F.; Rusch, K.A. Evaluation of polyhydroxybutyrate as a carbon source for recirculating aquaculture water denitrification. Aquac. Eng. 2012, 51, 36–43. [Google Scholar] [CrossRef]
- Zhu, S.M.; Deng, Y.L.; Ruan, Y.J.; Guo, X.S.; Shi, M.M.; Shen, J.Z. Biological denitrification using poly (butylene succinate) as carbon source and biofilm carrier for recirculating aquaculture system effluent treatment. Bioresour. Technol. 2015, 192, 603–610. [Google Scholar] [CrossRef] [PubMed]
- Ortiz-Estrada, A.M.; Gollas-Galvan, T.; Martinez-Cordova, L.R.; Martinez-Porchas, M. Predictive functional profiles using metagenomics 16S rRNA data: A novel functional approach to understanding the microbial ecology or aquaculture systems. Rev. Aquacult. 2019, 11, 234–245. [Google Scholar] [CrossRef]
- Woese, C.R.; Fox, G.E. Phylogenetic structure of the prokaryotic domain: The primary kingdoms. Proc. Nat. Acad. Sci. USA 1977, 74, 5088–5090. [Google Scholar] [CrossRef] [PubMed]
- Michaud, L.; Lo Giudice, A.; Troussellier, M.; Smedile, F.; Bruni, V.; Blancheton, J.P. Phylogenetic characterization of the heterotrophic bacterial communities inhabiting a marine recirculating aquaculture system. J. Appl. Microbiol. 2009, 107, 1935–1946. [Google Scholar] [CrossRef] [PubMed]
- Keuter, S.; Kruse, M.; Lipski, A.; Spieck, E. Relevance of Nitrospira for nitrite oxidation in a marine recirculation aquaculture system and physiological features of a Nitrospira marina-like isolate. Environ. Microbiol. 2011, 13, 2536–2547. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Wan, R.; Song, X.; Liu, Y.; Hallerman, E.; Dong, D.; Zhai, J.M.; Zhang, H.S.; Sun, L.Y. Metagenomic analysis shows diverse, distinct bacterial communities in biofilters among different marine recirculating aquaculture systems. Aquac. Int. 2016, 24, 1393–1408. [Google Scholar] [CrossRef]
- Rud, I.; Kolarevic, J.; Holan, A.B.; Berget, I.; Calabrese, S.; Terjesen, B.F. Deep-sequencing of the bacterial microbiota in commercial-scale recirculating and semi-closed aquaculture systems for Atlantic salmon post-smolt production. Aquac. Eng. 2017, 78, 50–62. [Google Scholar] [CrossRef]
- Brailo, M.; Schreier, H.J.; McDonald, R.; Maršić-Lučić, J.; Gavrilović, A.; Pećarević, M.; Jug-Dujaković, J. Bacterial community analysis of marine recirculating aquaculture system bioreactors for complete nitrogen removal established from a commercial inoculum. Aquaculture 2019, 503, 198–206. [Google Scholar] [CrossRef]
- Ahn, Y.H. Sustainable nitrogen elimination biotechnologies: A review. Process Biochem. 2006, 41, 1709–1721. [Google Scholar] [CrossRef]
- Zhang, R.C.; Xu, X.J.; Chen, C.; Shao, B.; Zhou, X.; Yuan, Y.; Lee, D.J.; Ren, N.Q. Bioreactor performance and microbial community analysis of autotrophic denitrification under micro-aerobic condition. Sci. Total Environ. 2019, 647, 914–922. [Google Scholar] [CrossRef]
- Jiang, Y.L.; Huang, Z.T.; Song, X.F. Carbon metabolism characteristics of microbial community in aerobic denitrification reactor. Ocean. Limnol. 2018, 49, 331–339. (In Chinese) [Google Scholar]
- Huang, Z.T.; Song, X.; Jiang, Y.; Hallerman, E. Characterization of a novel haloduric heterotrophic nitrifying and aerobically denitrifying bacterium, Halomonas sp. Z8 as a potential bioagent for wastewater treatment. J. Environ. Biol. 2020, 41, 43–52. [Google Scholar] [CrossRef]
- APHA (American Public Health Association), American Water Works Association, Water Environment Federation. Standard Methods for the Examination of Water and Wastewater; American Public Health Association: Washington, DC, USA, 2005; Volume 21, pp. 258–259. [Google Scholar]
- Peng, X.; Zhang, S.; Li, L.; Zhao, X.; Ma, Y.; Shi, D. Long-term high-solids anaerobic digestion of food waste: Effects of ammonia on process performance and microbial community. Bioresour. Technol. 2018, 262, 148–158. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Ji, X.; Wang, J.; Yang, D.; Shen, Y.; Chen, C.; Qian, F.Y.; Wu, P. Microbial community response to influent shift and lowering temperature in a two-stage mainstream deammonification process. Bioresour. Technol. 2018, 262, 132–140. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Magoč, T.; Salzberg, S.L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011, 27, 2957–2963. [Google Scholar] [CrossRef]
- Edgar, R.C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Meth. 2013, 10, 996. [Google Scholar] [CrossRef]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2014, 27, 2194–2200. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- Cheng, C.; Zhou, Z.; Pang, H.; Zheng, Y.; Chen, L.; Jiang, L.M.; Zhao, X. Correlation of microbial community structure with pollutants removal, sludge reduction and sludge characteristics in micro-aerobic side-stream reactor coupled membrane bioreactors under different hydraulic retention times. Bioresour. Technol. 2018, 260, 177–185. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Lozupone, C.A.; Hamady, M.; Kelley, S.T.; Knight, R. Quantitative and qualitative β diversity measures lead to different insights into factors that structure microbial communities. Appl. Environ. Microbiol. 2007, 73, 1576–1585. [Google Scholar] [CrossRef] [PubMed]
- Hill, M.O.; Gauch, H.G. Detrended correspondence analysis: An improved ordination technique. In Classification and Ordination; Springer: Dordrecht, The Netherlands, 1980; pp. 47–58. [Google Scholar]
- Van Den Wollenberg, A.L. Redundancy analysis an alternative for canonical correlation analysis. Psychometrika 1977, 42, 207–219. [Google Scholar] [CrossRef]
- Fruchterman, T.M.J.; Reingold, E.M. Graph drawing by force-directed placement. Softw. Pract. Exper. 1991, 21, 1129–1164. [Google Scholar] [CrossRef]
- Colt, J.; Lamoureux, J.; Patterson, R.; Rogers, G. Reporting standards for biofilter performance studies. Aquac. Eng. 2006, 34, 377–388. [Google Scholar] [CrossRef]
- Kim, B.-S.; Lim, Y.W.; Chun, J. Sphingopyxis marina sp. nov. and Sphingopyxis litoris sp. nov., isolated from seawater. Int. J. Syst. Evol. Microbiol. 2008, 58, 2415–2419. [Google Scholar] [CrossRef]
- Luján-Facundo, M.J.; Fernández-Navarro, J.; Alonso-Molina, J.L.; Amorós- Muñoz, I.; Moreno, Y.; Mendoza-Roca, J.A.; Pastor-Alcañiz, L. The role of salinity on the changes of the biomass characteristics and on the performance of an OMBR treating tannery wastewater. Water Res. 2018, 142, 129–137. [Google Scholar] [CrossRef]
- Simon, M.; Scheuner, C.; Meier-Kolthoff, J.P.; Brinkhoff, T.; Wagner-Döbler, I.; Ulbrich, M.; Klenk, H.P.; Schomburg, D.; Petersen, J.; Göker, M. Phylogenomics of Rhodobacteraceae reveals evolutionary adaptation to marine and non-marine habitats. ISME J. 2017, 11, 1483–1499. [Google Scholar] [CrossRef]
- Selje, N.; Simon, M.; Brinkhoff, T. A newly discovered Roseobacter cluster in temperate and polar oceans. Nature 2004, 427, 445–448. [Google Scholar] [CrossRef]
- Giebel, H.A.; Kalhoefer, D.; Lemke, A.; Thole, S.; Gahl-Janssen, R.; Simon, M.; Brinkhoff, T. Distribution of Roseobacter RCA and SAR11 lineages in the North Sea and characteristics of an abundant RCA isolate. ISME J. 2011, 5, 8. [Google Scholar] [CrossRef]
- Wemheuer, B.; Wemheuer, F.; Hollensteiner, J.; Meyer, F.; Voget, S.; Daniel, R. The green impact: Bacterioplankton response towards a phytoplankton spring bloom in the southern North Sea assessed by comparative metagenomic and metatranscriptomic approaches. Front. Microbiol. 2015, 6, 805. [Google Scholar] [CrossRef] [PubMed]
- Fadhlaoui, K.; Ben, H.W.; Postec, A.; Fauque, G.; Hamdi, M.; Ollivier, B.; Fardeau, M.L. Fusibacter fontis sp. nov., a sulfur-reducing, anaerobic bacterium isolated from a mesothermic Tunisian spring. Int. J. Syst. Evol. Microbiol. 2015, 65, 3501. [Google Scholar] [CrossRef] [PubMed]
- Priyanka, P.; Arun, A.B.; Rekha, P.D. Sulfated exopolysaccharide produced by Labrenzia sp. PRIM-30, characterization and prospective applications. Int. J. Biol. Macromol. 2014, 69, 290–295. [Google Scholar]
- Okamoto, T.; Maruyama, A.; Imura, S.; Takeyama, H.; Naganuma, T. Comparative phylogenetic analyses of Halomonas variabilis and related organisms based on 16S rRNA, gyrB and ectBC gene sequences. Syst. Appl. Microbiol 2004, 27, 323–333. [Google Scholar] [CrossRef] [PubMed]
- Quillaguamán, J.; Hashim, S.; Bento, F.; Mattiasson, B.; Hatti-Kaul, R. Poly(ß-hydroxybutyrate) production by a moderate halophile, Halomonas boliviensis LC1 using starch hydrolysate as substrate. J. Appl. Microbiol. 2005, 99, 151. [Google Scholar] [CrossRef]
- Zinatizadeh, A.A.L.; Ghaytooli, E. Simultaneous nitrogen and carbon removal from wastewater at different operating conditions in a moving bed biofilm reactor (MBBR): Process modeling and optimization. J. Taiwan Inst. Chem. Eng. 2015, 53, 98–111. [Google Scholar] [CrossRef]
- Wang, X.M.; Wang, J.L. Nitrate removal from groundwater using solid-phase denitrification process without inoculating with external microorganisms. Int. J. Environ. Sci. Technol. 2013, 10, 955–960. [Google Scholar] [CrossRef]
- Ovez, B.; Ozgen, S.; Yuksel, M. Biological denitrification in drinking water using Glycyrrhiza glabra and Arunda donax as the carbon source. Process Biochem. 2006, 41, 1539–1544. [Google Scholar] [CrossRef]
- Zhu, W.J.; Yang, Y.; Qiao, Y.M.; Ran, T. Effect of hydraulic retention time on purification efficiency of natural aerated biological filter bed. Ecol. Sci. 2013, 32, 224–229. [Google Scholar]
- Qiu, L.P.; Ma, J.; Zhang, L.X. Effects of hydraulic residence time on treatment efficiency and operation characteristics of biological aerated filter. Environ. Pollut. Prevent. Cont. 2004, 26, 433–436. [Google Scholar]
- Körner, H.; Zumft, W.G. Expression of denitrification enzymes in response to the dissolved oxygen level and respiratory substrate in continuous culture of Pseudomonas stutzeri. Appl. Environ. Microbiol. 1989, 55, 1670–1676. [Google Scholar] [CrossRef] [PubMed]
- Chu, L.; Wang, J. Denitrification performance and biofilm characteristics using biodegradable polymers PCL as carriers and carbon source. Chemosphere 2013, 91, 1310–1316. [Google Scholar] [CrossRef] [PubMed]
- Hamlin, H.J.; Michaels, J.T.; Beaulaton, C.M.; Graham, W.F.; Dutt, W.; Steinbach, P.; Losordo, T.M.; Schrader, K.K.; Main, K.L. Comparing denitrification rates and carbon sources in commercial scale upflow denitrification biological filters in aquaculture. Aquac. Eng. 2008, 38, 79–92. [Google Scholar] [CrossRef]
- Kiss, H.; Lang, E.; Lapidus, A.; Kiss, H.; Lang, E.; Lapidus, A.; Copeland, A.; Nolan, M.; Del Rio, T.G.; Han, C. Complete genome sequence of Denitrovibrio acetiphilus type strain (N2460 T). Stand. Genom. Sci. 2010, 2, 270. [Google Scholar] [CrossRef] [PubMed]
- Torno, J.; Naas, C.; Schroeder, J.P.; Schulz, C. Impact of hydraulic retention time, back flushing intervals, and C/N ratio on the SID-reactor denitrification performance in marine RAS. Aquaculture 2018, 496, 112–122. [Google Scholar] [CrossRef]
- Yin, H.; Yang, C.; Jia, Y.; Chen, H.; Gu, X. Dual removal of phosphate and ammonium from high concentrations of aquaculture wastewaters using an efficient two-stage infiltration system. Sci. Total Environ. 2018, 635, 936–946. [Google Scholar] [CrossRef]
- Obaja, D.; Mace, S.; Mata-Alvarez, J. Biological nutrient removal by a sequencing batch reactor (SBR) using an internal organic carbon source in digested piggery wastewater. Bioresour. Technol. 2005, 96, 7–14. [Google Scholar] [CrossRef]
- Elefsiniotis, P.; Wareham, D.G.; Smith, M.O. Use of volatile fatty acids from an acid phase digester for denitrification. J. Biotechnol. 2004, 114, 289–297. [Google Scholar] [CrossRef]
- Xu, Z.; Dai, X.; Chai, X. Effect of different carbon sources on denitrification performance, microbial community structure and denitrification genes. Sci. Total Environ. 2018, 634, 195–204. [Google Scholar] [CrossRef]
- Nicolella, C.; van Loosdrecht, M.C.M.; Heijnen, J.J. Wastewater treatment with particulate biofilm reactors. J. Biotechnol. 2000, 80, 1–33. [Google Scholar] [CrossRef]
- Cunningham, A.B.; Characklis, W.G.; Aberdeen, F.; Crawford, D. Influence of biofilm accumulation on porous media hydrodynamics. Environ. Sci. Technol. 1991, 25, 1305–1311. [Google Scholar] [CrossRef]
- Kondaveeti, S.; Lee, S.H.; Park, H.D.; Min, B. Bacterial communities in a bioelectrochemical denitrification system: The effects of supplemental electron acceptors. Water Res. 2014, 51, 25–36. [Google Scholar] [CrossRef]
- Lu, H.J.; Chandran, K.; Stensel, D. Microbial ecology of denitrification in biological wastewater treatment. Water Res. 2014, 64, 237–254. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Wang, Z.; Yang, Y.; Mei, X.; Wu, Z. Correlating microbial community structure and composition with aeration intensity in submerged membrane bioreactors by high-throughput pyrosequencing. Water Res. 2013, 47, 859–869. [Google Scholar] [CrossRef]
- Nakasaki, K.; Tran, L.T.H.; Idemoto, Y.; Abe, M.; Rollon, A.P. Comparison of organic matter degradation and microbial community during thermophilic composting of two different types of anaerobic sludge. Bioresour. Technol. 2009, 100, 676–682. [Google Scholar] [CrossRef]
- Lam, P.; Kuypers, M.M. Microbial nitrogen cycling processes in oxygen minimum zones. Annu. Rev. Mar. Sci. 2011, 15, 317–345. [Google Scholar] [CrossRef]
- Li, Y.; Tang, K.; Zhang, L.; Zhao, Z.; Xie, X.; Chen, C.T.; Wang, D.; Jiao, N.; Zhang, Y. Coupled carbon, sulfur, and nitrogen cycles mediated by microorganisms in the water column of a shallow-water hydrothermal ecosystem. Front. Microbiol. 2018, 13, 2718. [Google Scholar] [CrossRef]
- Baek, K.; Choi, A.; Kang, I.; Cho, J.C. Celeribacter marinus sp. nov., isolated from coastal seawater. Int. J. Syst. Evol. Microbiol. 2014, 64, 1323–1327. [Google Scholar] [CrossRef]
- Arahal, D.R.; Lenkunberru, I.; Gonzalez, J.M.; Pascual, J.; Pujalte, M.J.; Pedros-Alio, C.; Pinhassi, J. Neptuniibacter caesariensis gen. nov., sp. nov., a novel marine genome-sequenced gammaproteobacterium. Int. J. Syst. Evol. Microbiol. 2007, 57, 1000–1006. [Google Scholar] [CrossRef]
- Vaidya, B.; Kumar, R.; Sharma, G.; Srinivas, T.N.; Kumar, P.A. Xanthomarina gelatinilytica gen. nov., sp. nov., isolated from seawater. Int. J. Syst. Evol. Microbiol. 2015, 65, 3926–3932. [Google Scholar] [CrossRef]
- Huang, X.-F.; Liu, Y.J.; Dong, J.-D.; Qu, L.-Y.; Zhang, Y.-Y.; Wang, F.-Z.; Tian, X.-P.; Zhang, S. Mangrovibacterium diazotrophicum gen. nov., sp. nov., a nitrogen-fixing bacterium isolated from a mangrove sediment, and proposal of Prolixibacteraceae fam. nov. Int. J. Syst. Evol. Microbiol. 2014, 64, 875–881. [Google Scholar] [CrossRef] [PubMed]
- Ben Hania, W.; Joseph, M.; Bunk, B.; Sproer, C.; Klenk, H.P.; Fardeau, M.L.; Spring, S. Characterization of the first cultured representative of a Bacteroidetes clade specialized on the scavenging of cyanobacteria. Environ. Microbiol. 2017, 19, 1134–1148. [Google Scholar] [CrossRef] [PubMed]
- Letelier-Gordo, C.O.; Herreros, M.M. Denitrifying granules in a marine upflow anoxic sludge bed (UASB) reactor. Aquac. Eng. 2019, 84, 42–49. [Google Scholar] [CrossRef]
- Nielsen, J.L.; Nguyen, H.; Meyer, R.L.; Nielsen, P.H. Identification of glucose fermenting bacteria in a full-scale enhanced biological phosphorus removal plant by stable isotope probing. Microbiology 2012, 158, 1818–1825. [Google Scholar] [CrossRef]
- McGenity, T.; Van Der Meer, J.R.; de Lorenzo, V. Handbook of Hydrocarbon and Lipid Microbiology; Springer: Berlin, Germany, 2010. [Google Scholar]
- Fernandez, N.; Sierra-Alvarez, R.; Field, J.A.; Amils, R.; Sanz, J.L. Microbial community dynamics in a chemolithotrophic denitrification reactor inoculated with methanogenic granular sludge. Chemosphere 2008, 70, 462–474. [Google Scholar] [CrossRef]
- Dannenberg, S.; Kroder, M.; Dilling, W.; Cypionka, H. Oxidation of H2, organic compounds and inorganic sulfur compounds coupled to reduction of O2 or nitrate by sulfate-reducing bacteria. Arch. Microbiol. 1992, 158, 93–99. [Google Scholar] [CrossRef]
- Shen, Z.; Zhou, Y.; Liu, J.; Xiao, Y.; Cao, R.; Wu, F. Enhanced removal of nitrate using starch/PCL blends as solid carbon source in a constructed wetland. Bioresour. Technol. 2015, 175, 239–244. [Google Scholar] [CrossRef]
- Du, R.; Cao, S.B.; Li, B.K.; Niu, M.; Wang, S.Y.; Peng, Y.Z. Performance and microbial community analysis of a novel DEAMOX based on partial-denitrification and anammox treating ammonia-N and nitrate wastewaters. Water Res. 2017, 108, 46–56. [Google Scholar] [CrossRef]
- Wu, W.; Yang, L.; Wang, J. Denitrification performance and microbial diversity in a packed-bed bioreactor using PCL as carbon source and biofilm carrier. Appl. Microbiol. Biotechnol. 2013, 97, 2725–2733. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).