Hydrochemical Conditions for Aerobic/Anaerobic Biodegradation of Chlorinated Ethenes—A Multi-Site Assessment
Abstract
1. Introduction
2. Materials and Methods
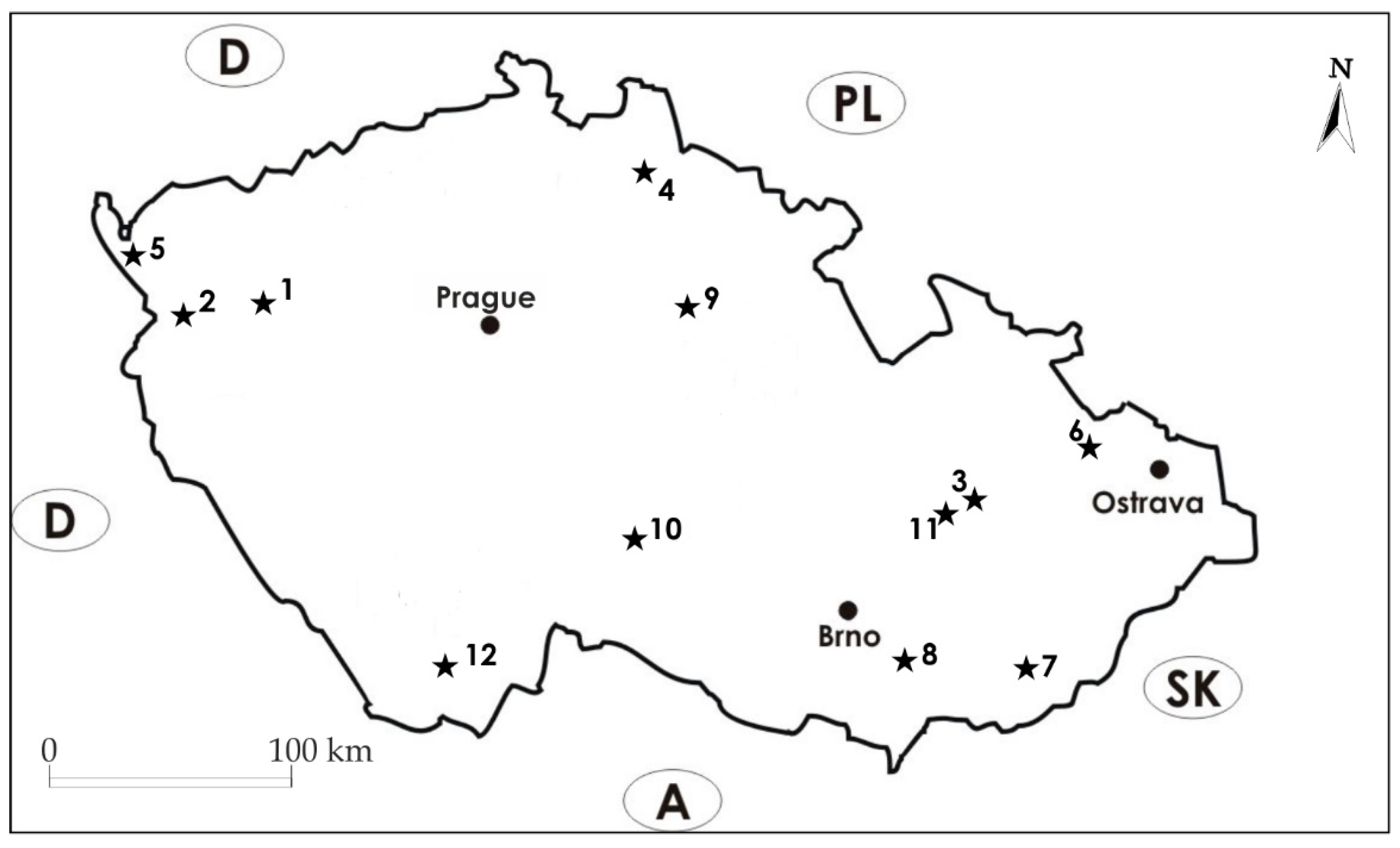
2.1. Test Sites
2.2. Groundwater Sampling and Scope of Laboratory Analysis
2.3. DNA Extraction and Real-Time Quantitative PCR
2.4. Physical and Chemical Parameters of the Groundwater
2.5. Data Analysis
3. Results and Discussion
3.1. Results of Chemical Analyses
3.2. Results of qPCR
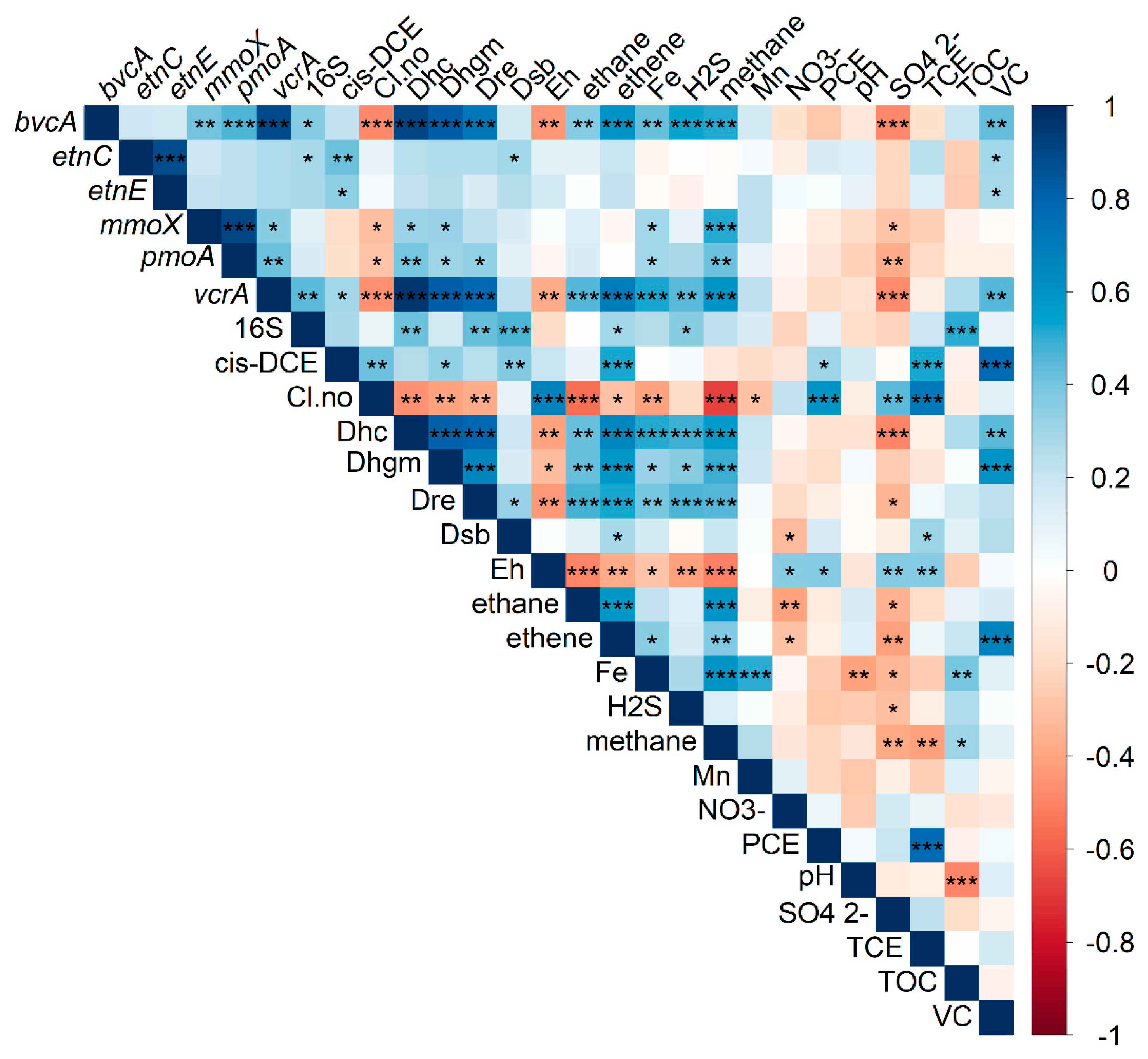
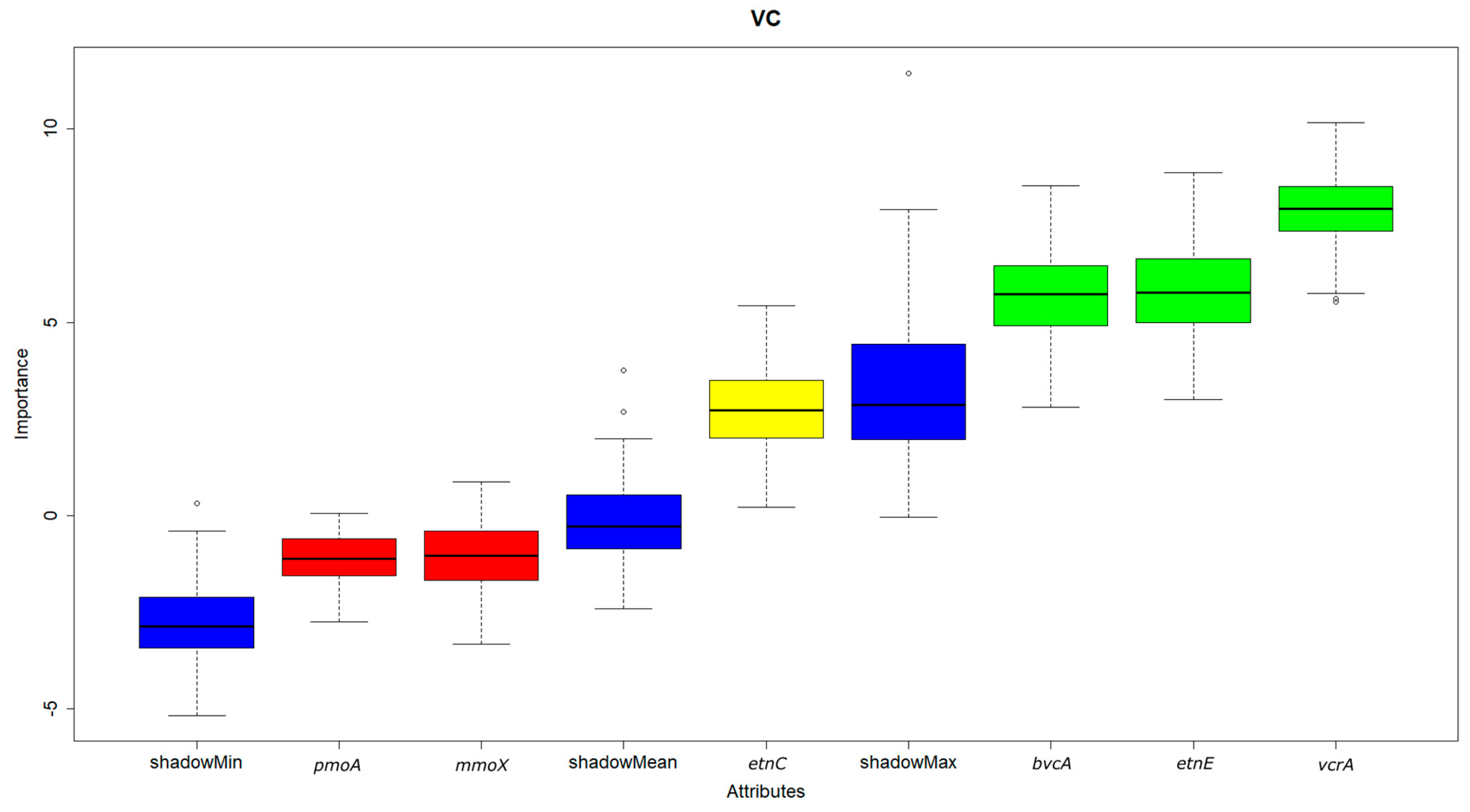
3.3. Correlation and Feature Selection Analysis
3.4. Hydrochemical Conditions for Aerobic Oxidation
4. Conclusions
- Both the ethenotroph functional genes etnC and etnE and methanotroph functional genes mmoX and pmoA were identified in 90% of groundwater samples, while all functional genes (etnC, etnE, mmoX, pmoA, vcrA and bvcA) coexisted in 78% of samples, in actively biostimulated sites in 88% of samples.
- The reductive dechlorinator Dehalogenimonas spp., only recently identified as capable of cis-1,2-DCE and VC degradation, was detected by qPCR in 94% of samples. A positive correlation between Dehalogenimonas spp. abundance and cis-1,2-DCE and VC concentration indicates a potential contribution to reductive dechlorination at the sites tested.
- Presence of both cis-1,2-DCE and VC was positively correlated with the reductive functional gene vcrA, and the ethenotroph functional genes etnC and etnE, and VC additionally with the reductive functional gene bvcA.
- Of all the functional genes tested, only the reductive dehalogenase functional gene vcrA was found to be significant for cis-1,2-DCE degradation by feature selection using the random forest algorithm. On the hand, both the dehalogenase functional genes vcrA and bvcA and the ethenotroph functional gene etnE were indicated as significant for VC.
- No significant relationship was observed between cis-1,2-DCE and VC concentration and abundance of the methanotroph functional genes mmoX and pmoA, because methanotrophs oxidise these contaminants cometabolically only, without any energetic benefit.
- Cluster analysis revealed that aerobic oxidation of chloroethenes can take place under a broad range of apparent redox conditions, even under apparently methanogenic conditions, probably due to a high redox microheterogeneity of the aquifer environment, ability of ethenotrophs to survive in environments with very limited oxygen contents and high concentrations of ethene and VC (as electron donors in aerobic oxidation of chloroethenes), these being the most important hydrochemical parameters affecting abundance of ethenotrophs.
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bradley, P.M.; Chapelle, F.H. In Situ Remediation of Chlorinated Solvent Plumes; Stroo, H., Ward, C., Eds.; Springer: New York, NY, USA, 2010; Chapter 3; pp. 39–67. ISBN 978-1-4419-1400-2. [Google Scholar]
- Stroo, H.F.; West, M.R.; Kueper, B.H.; Borden, R.C.; Major, D.W.; Ward, C.H. Chlorinated Solvent Source Zone Remediation; Kueper, B.H., Stroo, H.F., Vogel, C.M., Ward, C.H., Eds.; Springer: New York, NY, USA, 2014; Chapter 12; pp. 395–457. ISBN 978-1-4614-6921-6. [Google Scholar]
- Holliger, C.; Hahn, D.; Harmsen, H.; Ludwig, W.; Schumacher, W.; Tindall, B.; Vazquez, F.; Weiss, N.; Zehnder, A.J. Dehalobacter restrictus gen. nov. and sp. nov., a strictly anaerobic bacterium that reductively dechlorinates tetra-and trichloroethene in an anaerobic respiration. Arch. Microbiol. 1998, 169, 313–321. [Google Scholar] [CrossRef] [PubMed]
- Holliger, C.; Schraa, G.; Stams, A.; Zehnder, A. A highly purified enrichment culture couples the reductive dechlorination of tetrachloroethene to growth. Appl. Environ. Microbiol. 1993, 59, 2991–2997. [Google Scholar] [CrossRef]
- Neumann, A.; Scholz-Muramatsu, H.; Diekert, G. Tetrachloroethene metabolism of Dehalospirillum multivorans. Arch. Microbiol. 1994, 162, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Krumholz, L.R.; Sharp, R.; Fishbain, S.S. A freshwater anaerobe coupling acetate oxidation to tetrachloroethylene dehalogenation. Appl. Environ. Microbiol. 1996, 62, 4108–4113. [Google Scholar] [CrossRef] [PubMed]
- Sung, Y.; Ritalahti, K.M.; Sanford, R.A.; Urbance, J.W.; Flynn, S.J.; Tiedje, J.M.; Löffler, F.E. Characterization of two tetrachloroethene-reducing, acetate-oxidizing anaerobic bacteria and their description as Desulfuromonas michiganensis sp. nov. Appl. Environ. Microbiol. 2003, 69, 2964–2974. [Google Scholar] [CrossRef] [PubMed]
- Sung, Y.; Fletcher, K.E.; Ritalahti, K.M.; Apkarian, R.P.; Ramos-Hernández, N.; Sanford, R.A.; Mesbah, N.M.; Löffler, F.E. Geobacter lovleyi sp. nov. strain SZ, a novel metal-reducing and tetrachloroethene-dechlorinating bacterium. Appl. Environ. Microbiol. 2006, 72, 2775–2782. [Google Scholar] [CrossRef] [PubMed]
- Luijten, M.L.; de Weert, J.; Smidt, H.; Boschker, H.T.; de Vos, W.M.; Schraa, G.; Stams, A.J. Description of Sulfurospirillum halorespirans sp. nov., an anaerobic, tetrachloroethene-respiring bacterium, and transfer of Dehalospirillum multivorans to the genus Sulfurospirillum as Sulfurospirillum multivorans comb. nov. Int. J. Syst. Evol. Microbiol. 2003, 53, 787–793. [Google Scholar] [CrossRef]
- Maillard, J.; Regeard, C.; Holliger, C. Isolation and characterization of Tn-Dha1, a transposon containing the tetrachloroethene reductive dehalogenase of Desulfitobacterium hafniense strain TCE1. Environ. Microbiol. 2005, 7, 107–117. [Google Scholar] [CrossRef]
- Löffler, F.E.; Ritalahti, K.M.; Zinder, S.H. Bioaugmentation for Groundwater Remediation; Stroo, H., Leeson, A., Ward, C., Eds.; Springer: New York, NY, USA, 2013; Chapter 2; pp. 39–88. ISBN 978-1-4614-4114-4. [Google Scholar]
- Maymó-Gatell, X.; Nijenhuis, I.; Zinder, S.H. Reductive dechlorination of cis-1, 2-dichloroethene and vinyl chloride by “Dehalococcoides ethenogenes”. Environ. Sci. Technol. 2001, 35, 516–521. [Google Scholar] [CrossRef]
- Yang, Y.; Higgins, S.A.; Yan, J.; Şimşir, B.; Chourney, K.; Iyer, R.; Hettlich, R.L.; Baldwin, B.; Ogles, D.M.; Löffler, F.E. Grape pomace compost harbors organohalide-respiring Dehalogenimonas species with novel reductive dehalogenase genes. ISME J. 2017, 11, 2767–2780. [Google Scholar] [CrossRef]
- Judger, B.-E.; Ertan, H.; Lee, M.; Manefield, M.; Marquis, C.P. Reductive dehalogenases come of age in biological destruction of organohalides. Trends Biotechnol. 2015, 33, 596–608. [Google Scholar] [CrossRef]
- Tratnyek, P.G.; Johnson, R.L.; Lowry, G.V.; Brown, R.A. Chlorinated Solvent Source Zone Remediation; Kueper, B.H., Stroo, H.F., Vogel, C.M., Ward, C.H., Eds.; Springer: New York, NY, USA, 2014; Chapter 10; pp. 307–351. ISBN 978-1-4614-6921-6. [Google Scholar]
- Thiem, A.; Schmidt, K.R. Sequential anaerobic biodegradation of chloroethenes—Aspects of field application. Curr. Opin. Biotechnol. 2011, 22, 415–421. [Google Scholar] [CrossRef] [PubMed]
- Coleman, N.V.; Mattes, T.E.; Gossett, J.M.; Spain, J.C. Phylogenetic and kinetic diversity of aerobic vinyl chloride-assimilating bacteria from contaminated sites. Appl. Environ. Microbiol. 2002, 68, 6162–6171. [Google Scholar] [CrossRef] [PubMed]
- Mattes, T.E.; Alexander, A.K.; Coleman, N.V. Aerobic biodegradation of the chloroethenes: Pathways, enzymes, ecology, and evolution. FEMS Microbiol. Rev. 2010, 34, 445–475. [Google Scholar] [CrossRef]
- Richards, P.M.; Liang, Y.; Johnson, R.; Mattes, T.E. Cryogenic soil coring reveals coexistence of aerobic and anaerobic vinyl chloride degrading bacteria in a chlorinated ethene contaminated aquifer. Water Res. 2019, 157, 281–291. [Google Scholar] [CrossRef]
- Liang, Y.; Liu, X.; Singletary, M.A.; Wang, K.; Mattes, T.E. Relationships between the abundance and expression of functional genes from vinyl chloride (VC)-degrading bacteria and geochemical parameters at VC-contaminated sites. Environ. Sci. Technol. 2017, 51, 12164–12174. [Google Scholar] [CrossRef]
- Wymore, R.A.; Lee, M.H.; Keener, W.K.; Miller, A.R.; Colwell, F.S.; Watwood, M.E.; Sorenson, K.S. Field evidence for intrinsic aerobic chlorinated cometabolism by methanotrophs expressing soluble methane monooxygenase. Biorem. J. 2007, 11, 125–139. [Google Scholar] [CrossRef][Green Version]
- Paszynski, A.J.; Paidisetti, R.; Johnson, A.K.; Crawford, R.L.; Colwell, F.S.; Green, T.; Delwiche, M.; Lee, H.; Newby, D.; Brodie, E.L.; et al. Proteomic and targeted qPCR analyses of ubsurface microbial communities for presence of methane monooxygenase. Biodegradation 2011, 22, 1045–1059. [Google Scholar] [CrossRef]
- Coleman, N.V.; Spain, J.C. Epoxyalkane: Coenzyme M transferase in the ethene and vinyl chloride biodegradation pathways of mycobacterium strain JS60. J. Bacteriol. 2003, 185, 5536–5545. [Google Scholar] [CrossRef]
- Jin, Y.O.; Mattes, T.E. A quantitative PCR assay for aerobic, vinyl chloride- and ethene-assimilating microorganisms in groundwater. Environ. Sci. Technol. 2010, 44, 9036–9041. [Google Scholar] [CrossRef]
- Abe, Y.; Aravena, R.; Zopfi, J.; Shouakar-Stash, O.; Cox, E.; Roberts, J.D.; Hunkeler, D. Carbon and chlorine isotope fractionation during aerobic oxidation and reductive dechlorination of vinyl chloride and cis-1, 2-dichloroethene. Environ. Sci. Technol. 2009, 43, 101–107. [Google Scholar] [CrossRef] [PubMed]
- Jennings, L.K.; Chartrand, M.M.; Lacrampe-Couloume, G.; Lollar, B.S.; Spain, J.C.; Gossett, J.M. Proteomic and transcriptomic analyses reveal genes upregulated by cis-dichloroethene in Polaromonas sp. strain JS666. Appl. Environ. Microbiol. 2009, 75, 3733–3744. [Google Scholar] [CrossRef] [PubMed]
- Dolinová, I.; Štrojsová, M.; Černík, M.; Němeček, J.; Macháčková, M.; Ševců, A. Microbial degradation of chloroethenes: A review. Environ. Sci. Pollut. Res. 2017, 24, 13262–13283. [Google Scholar] [CrossRef] [PubMed]
- ČSN ISO 5667-11. Water Quality—Sampling—Part 11: Guidance on Sampling of Groundwaters; International Organization for Standardization: Geneva, Switzerland, 1993. [Google Scholar]
- Němeček, J.; Dolinová, I.; Macháčková, J.; Špánek, R.; Ševců, A.; Lederer, T.; Černík, M. Stratification of Chlorinated Ethenes Natural Attenuation in an Alluvial Aquifer Assessed by Hydrochemical and Biomolecular Tools. Chemosphere 2017, 184, 1157–1167. [Google Scholar] [CrossRef]
- ČSN EN ISO 11885. Water Quality—Determination of Selected Elements by Inductively Coupled Plasma Optical Emission Spectrometry (ICP-OES); International Organization for Standardization: Geneva, Switzerland, 2007. [Google Scholar]
- ČSN 83 0530-31. Chemical and Physical Analysis of Surface Water. Determination of Sulfide and Hydrogen Sulfide; International Organization for Standardization: Geneva, Switzerland, 1980. [Google Scholar]
- ČSN EN 1484. Water AnalysisGuidelines for the Determination of Total Organic Carbon (TOC) and Dissolved Organic Carbon (DOC); International Organization for Standardization: Geneva, Switzerland, 1998. [Google Scholar]
- ČSN EN ISO 10304-1. Water Quality—Determination of Dissolved Anions by Liquid Chromatography of Ions—Part 1: Determination of Bromide, Chloride, Fluoride, Nitrate, Nitrite, Phosphate and Sulfate; International Organization for Standardization: Geneva, Switzerland, 2007. [Google Scholar]
- Bewley, R.; Hick, P.; Rawcliffe, A. Meeting the challenges for bioremediation of chlorinated solvents at operational sites: A comparison of case studies. In Proceeding of the 13th International UFZ-Deltarez Conference on Sustainable Use and Management of Soil, Sediment and Water Resources, Copenhagen, Denmark, 9–12 June 2015. [Google Scholar]
- Chapelle, F.H.; Bradley, P.M.; Thomas, M.A.; McMahon, P.B. Distinguishing iron-reducing from sulfate-reducing conditions. Ground Water 2009, 47, 300–3058. [Google Scholar] [CrossRef]
- RStudio Team. RStudio: Integrated Development for R; RStudio, Inc.: Boston, MA, USA, 2015; Available online: http://www.rstudio.com/ (accessed on 28 November 2019).
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018; Available online: http://www.R-project.org/ (accessed on 28 November 2019).
- Myers, J.L.; Well, A.D. Research Design and Statistical Analysis, 2nd ed.; Lawrence Erlbaum Associates: Mahwah, NJ, USA, 2003; p. 760. ISBN 0805840370. [Google Scholar]
- Kursa, M.B.; Rudnicki, W.R. Feature Selection with the Boruta Package. J. Stat. Soft. 2010, 36, 1–13. [Google Scholar] [CrossRef]
- Brock, E.; Pihur, V.; Datta, S.; Datta, S. clValid: An R Package for Cluster Validation. J. Stat. Soft. 2008, 25, 1–22. [Google Scholar] [CrossRef]
- Roberts, A.L.; Totten, L.A.; Arnold, W.A.; Burris, D.R.; Campbell, T.J. Reductive elimination of chlorinated ethylenes by zero valent metals. Environ. Sci. Technol. 1996, 30, 2654–2659. [Google Scholar] [CrossRef]
- Wild, A.; Hermann, R.; Leisinger, T. Isolation of an anaerobic bacterium which reductively dechlorinates tetrachloroethene and trichloroethene. Biodegradation 1996, 7, 507–511. [Google Scholar] [CrossRef]
- Bouwer, E.J. Bioremediation of chlorinated solvents using alternate electron acceptors. In Handbook of Bioremediation; Norris, R., Hinchee, R., Brown, R., McCarty, P., Semprini, L., Wilson, J., Kampbell, D., Reinhard, M., Borden, R., Eds.; Lewis Publishers: Boca Raton, FL, USA, 1994; pp. 149–175. [Google Scholar]
- Ehrlich, H.L.; Newman, D.K. Geomicrobiology, 5th ed.; CRC Press, Taylor & Francis Group: Boca Raton, FL, USA, 2009; p. 628. ISBN 978-0-8493-7906-2. [Google Scholar]
- Schmidt, K.R.; Tiehm, A. Natural attenuation of chloroethenes: Identification of sequential reductive/oxidative biodegradation by microcosm studies. Water Sci. Technol. 2008, 58, 1137–1145. [Google Scholar] [CrossRef]
- Gossett, J.M. Sustained aerobic oxidation of vinyl chloride at low oxygen concentrations. Environ. Sci. Technol. 2010, 44, 1405–1411. [Google Scholar] [CrossRef] [PubMed]
- Atashgahi, S.; Maphosa, F.; Doğan, E.; Smidt, H.; Springael, D.; Dejonghe, W. Small-scale oxygen distribution determines the vinyl chloride biodegradation pathway in surficial sediments of riverbed hyporheic zones. FEMS Microbiol. Ecol. 2013, 84, 133–142. [Google Scholar] [CrossRef] [PubMed]
- Atashgahi, S.; Lu, Y.; Ramiro-Garcia, J.; Peng, P.; Maphosa, F.; Sipkema, D.; Dejonghe, W.; Smidt, H.; Springael, D. Geochemical parameters and reductive dechlorination determine aerobic cometabolic vs. aerobic metabolic vinyl chloride biodegradation at oxic/anoxic interface of hyporheic zones. Environ. Sci. Technol. 2017, 51, 1626–1634. [Google Scholar] [CrossRef] [PubMed]


| Predominant Redox Process | NO3− (mg/L) | Mn2+ (mg/L) | Fe2+ (mg/L) | SO42− (mg/L) | Fe/H2S | Methane (mg/L) | |
|---|---|---|---|---|---|---|---|
| Oxic | O2 reduction | - | <0.05 | <0.1 | - | - | - |
| Anoxic | NO3− reduction | ≥1.0 | <0.05 | <0.1 | - | - | - |
| Mn(IV) reduction | <1.0 | ≥0.05 | <0.1 | - | - | - | |
| Fe(III) reduction | <1.0 | - | ≥0.1 | ≥0.5 | >10 | - | |
| Mix Fe(III)/SO42− reduction | <1.0 | - | ≥0.1 | ≥0.5 | 3–10 | - | |
| SO42− reduction | <1.0 | - | ≥0.1 | ≥0.5 | <3 | - | |
| Methanogenesis | <1.0 | - | ≥0.1 | <0.5 | - | ≥0.5 |
| Sample ID | Site ID | Plume ID | Date of Sampling DD.MM.YY | Affected by Whey Application Y/N | Time Elapsed after the Last Whey Application (Months) | Total Bacteria | Aerobic Biomarkers | Reductive Biomarkers | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| VC Reductive Dehalogenase Genes | Reductive Dechlorinators | |||||||||||||||
| 16S | etnC | etnE | mmoX | pmoA | bvcA | vcrA | Dhc | Dhgm | Dsb | Dre | ||||||
| AT-15 | 1 | 1_1 | 24.01.19 | Y | 1 | +++ | + | ++ | + | + | +++ | +++ | +++ | +++ | + | + |
| AT-19 | 1 | 1_1 | 01.05.19 | N | NA | +++ | ++ | +++ | + | ++ | ++ | ++ | ++ | ++ | ++ | + |
| AT-20 | 1 | 1_1 | 01.05.19 | N | NA | ++ | ++ | +++ | + | + | ND | + | + | ND | ++ | ND |
| AT-21 | 1 | 1_1 | 01.05.19 | N | NA | ++ | ++ | ++ | ++ | ++ | +- | + | + | ++ | + | +- |
| SV-10 | 2 | 2_1 | 24.01.19 | Y* | 8 | +++ | ++ | +++ | + | + | + | ++ | + | +- | +++ | ND |
| VS-5 | 2 | 2_1 | 24.01.19 | Y* | 8 | +++ | +++ | +++ | + | + | + | + | + | + | ++ | +- |
| A_I. | 3 | 3_1 | 04.02.19 | N | NA | + | ND | ND | ND | ND | ND | + | + | + | + | ND |
| B_IV. | 3 | 3_1 | 04.02.19 | N* | NA | + | ND | ND | ND | ND | ND | + | ND | ND | ++ | ND |
| SM-7D | 3 | 3_2 | 04.02.19 | Y | 16 | +++ | ++ | ++ | + | + | +++ | +++ | +++ | +++ | +++ | +++ |
| VS-7S_1 | 3 | 3_3 | 18.07.17 | N* | NA | +++ | + | + | ++ | ++ | + | ++ | + | ++ | ++ | + |
| VS-7S_2 | 3 | 3_3 | 13.10.17 | Y | 1 | +++ | + | + | ++ | +++ | +++ | +++ | +++ | +++ | ++ | ++ |
| VS-7S_3 | 3 | 3_3 | 12.02.18 | Y | 4 | ++ | ++ | ++ | + | ++ | +++ | +++ | +++ | +++ | ++ | + |
| VS-7S_4 | 3 | 3_3 | 26.03.18 | Y | 5 | ++ | ++ | ++ | ++ | ++ | +++ | +++ | +++ | +++ | ++ | + |
| VS-7S_5 | 3 | 3_3 | 04.02.19 | Y | 16 | +++ | + | + | + | ND | ++ | +++ | +++ | +++ | +++ | + |
| Studna_1 | 3 | 3_3 | 18.07.17 | N | NA | ++ | ++ | ++ | + | + | ++ | ++ | ++ | ++ | + | + |
| Studna_2 | 3 | 3_3 | 13.10.17 | Y | 1 | ++ | + | + | + | + | +++ | +++ | +++ | +++ | ++ | ++ |
| Studna_3 | 3 | 3_3 | 12.02.18 | Y | 4 | +++ | + | + | + | + | ++ | +++ | +++ | ++ | + | ++ |
| Studna_4 | 3 | 3_3 | 26.03.18 | Y | 5 | +++ | + | + | + | + | ++ | +++ | +++ | ++ | + | + |
| SM-8_1 | 3 | 3_3 | 18.07.17 | N | NA | ++ | ++ | ++ | ++ | ++ | +++ | +++ | +++ | +++ | ++ | ++ |
| SM-8_2 | 3 | 3_3 | 13.10.17 | Y | 1 | +++ | ++ | ++ | + | + | +++ | +++ | +++ | +++ | ++ | +++ |
| SM-8_3 | 3 | 3_3 | 12.02.18 | Y | 4 | +++ | ++ | ++ | ++ | +++ | ++ | +++ | +++ | +++ | ++ | ++ |
| SM-8_4 | 3 | 3_3 | 26.03.18 | Y | 5 | ++ | ++ | ++ | +++ | +++ | ++ | +++ | +++ | ++ | ++ | ++ |
| AP-2 | 4 | 4_1 | 05.02.19 | Y | 28 | ++ | ++ | ++ | ++ | + | ++ | ++ | ++ | +++ | ++ | ND |
| HV-16 | 4 | 4_1 | 09.04.19 | N | NA | ++ | +- | + | ++ | +++ | ++ | ++ | ++ | + | + | +- |
| HV-25 | 4 | 4_1 | 09.04.19 | Y | 50 | ++ | + | + | ++ | ++ | ++ | ++ | ++ | ++ | + | + |
| HV-8_1 | 4 | 4_1 | 03.08.16 | Y | 50 | ++ | + | ++ | +++ | +++ | +++ | ++ | ++ | ++ | + | + |
| HV-8_2 | 4 | 4_1 | 10.10.16 | Y | 1 | +++ | + | ++ | +++ | +++ | +++ | +++ | +++ | ++ | ++ | + |
| HV-8_3 | 4 | 4_1 | 16.01.17 | Y | 3 | ++ | + | + | +++ | +++ | ++ | ++ | ++ | + | ++ | +- |
| HML-4S_1 | 4 | 4_1 | 03.08.16 | Y | 50 | ++ | + | + | + | + | ++ | ++ | ++ | +++ | ++ | + |
| HML-4S_2 | 4 | 4_1 | 10.10.16 | Y | 1 | ++ | + | + | + | ++ | ++ | +++ | +++ | ++ | ++ | + |
| HML-4S_3 | 4 | 4_1 | 16.01.17 | Y | 3 | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ | + |
| HV-53D | 4 | 4_2 | 09.04.19 | Y | 30 | ++ | + | ++ | + | ++ | ++ | ++ | ++ | ++ | ++ | + |
| V-5 | 5 | 5_1 | 22.01.19 | Y | 3 | +++ | + | ++ | + | + | ++ | +++ | +++ | ++ | ND | + |
| V-11 | 5 | 5_1 | 22.01.19 | Y | 3 | +++ | + | + | + | + | + | ND | + | ++ | +++ | + |
| V-13 | 5 | 5_2 | 22.01.19 | Y* | 3 | +++ | + | ND | + | + | ND | ++ | + | + | +++ | + |
| SV-1 | 5 | 5_2 | 22.01.19 | N | NA | ++ | ++ | + | + | ++ | ++ | ++ | ++ | ++ | ++ | ND |
| HJ-4 | 6 | 6_1 | 29.01.19 | N | NA | ++ | + | ++ | + | + | ND | ND | ND | +- | + | ND |
| V-32 | 6 | 6_1 | 29.01.19 | N | NA | +++ | ++ | ++ | + | + | ND | + | +- | + | ++ | ND |
| MV - 6A | 7 | 7_1 | 20.02.19 | Y | 34 | + | +- | +- | + | + | + | + | + | ++ | + | ND |
| Z - 4 | 7 | 7_1 | 20.02.19 | Y | 34 | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ | + | ND |
| Žd - 2 | 8 | 8_1 | 20.02.19 | Y | 26 | +++ | +++ | +++ | +++ | +++ | ++ | +++ | +++ | +++ | ++ | ++ |
| Žd - 4 | 8 | 8_1 | 20.02.19 | Y | 26 | +++ | +++ | +++ | ++ | +++ | +++ | +++ | +++ | ++ | +++ | +++ |
| MR 4 | 9 | 9_1 | 21.02.19 | Y | 50 | ++ | + | + | ++ | + | ++ | ++ | ++ | ++ | ++ | + |
| ZMS 4 | 9 | 9_1 | 21.02.19 | Y | 9 | ++ | + | ++ | +++ | +++ | ++ | ++ | ++ | ++ | ++ | ND |
| HLV-5 | 10 | 10_1 | 04.03.19 | N | NA | +++ | + | ++ | +++ | ++ | ND | ++ | ++ | ++ | ++ | + |
| HV-26 | 10 | 10_1 | 04.03.19 | N | NA | + | ND | ND | ND | ND | ND | ND | ND | ND | + | ND |
| ID-2 | 11 | 11_1 | 05.03.19 | N | NA | + | ND | ND | ND | ND | + | + | + | + | ND | ND |
| HV-223 | 12 | 12_1 | 09.05.19 | Y | 2 | +++ | + | ++ | +++ | +++ | +++ | +++ | +++ | ++ | ++ | ++ |
| HV-112 | 12 | 12_1 | 09.05.19 | Y | 2 | +++ | + | +- | +++ | +++ | +++ | ++ | ++ | ++ | ++ | + |
| Legend: | * affected by application of ZVI | |||||||||||||||
| +++ | high quantity | |||||||||||||||
| ++ | medium quantity | |||||||||||||||
| + | low quantity | |||||||||||||||
| +- | close to the detection limit | |||||||||||||||
| ND | not detected | |||||||||||||||
| Categories of “high”, “medium”, and “low” quantitites represent the respective thirds of the determined range of Cq values for each biomarker | ||||||||||||||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Němeček, J.; Marková, K.; Špánek, R.; Antoš, V.; Kozubek, P.; Lhotský, O.; Černík, M. Hydrochemical Conditions for Aerobic/Anaerobic Biodegradation of Chlorinated Ethenes—A Multi-Site Assessment. Water 2020, 12, 322. https://doi.org/10.3390/w12020322
Němeček J, Marková K, Špánek R, Antoš V, Kozubek P, Lhotský O, Černík M. Hydrochemical Conditions for Aerobic/Anaerobic Biodegradation of Chlorinated Ethenes—A Multi-Site Assessment. Water. 2020; 12(2):322. https://doi.org/10.3390/w12020322
Chicago/Turabian StyleNěmeček, Jan, Kristýna Marková, Roman Špánek, Vojtěch Antoš, Petr Kozubek, Ondřej Lhotský, and Miroslav Černík. 2020. "Hydrochemical Conditions for Aerobic/Anaerobic Biodegradation of Chlorinated Ethenes—A Multi-Site Assessment" Water 12, no. 2: 322. https://doi.org/10.3390/w12020322
APA StyleNěmeček, J., Marková, K., Špánek, R., Antoš, V., Kozubek, P., Lhotský, O., & Černík, M. (2020). Hydrochemical Conditions for Aerobic/Anaerobic Biodegradation of Chlorinated Ethenes—A Multi-Site Assessment. Water, 12(2), 322. https://doi.org/10.3390/w12020322






