Seasonal Variability in the Microbial Community and Pathogens in Wastewater Final Effluents
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling Procedure
2.2. DNA Extraction and Purification
2.3. 454 Pyrosequencing
3. Results
3.1. Community Richness and Diversity of Wastewater Final Effluents in Four Seasons
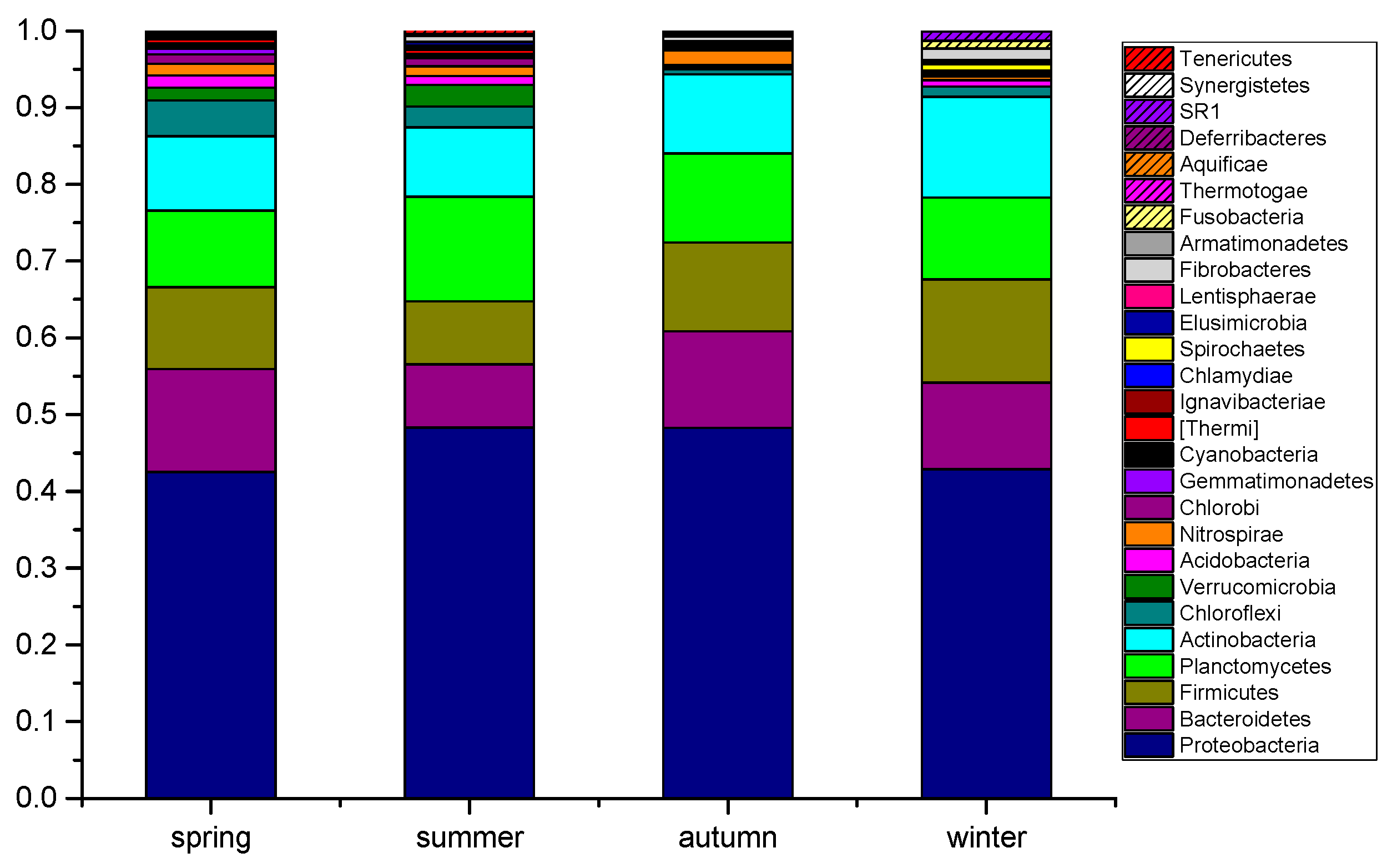
3.2. Identification of the Bacterial Community in the Wastewater Final Effluents
3.3. Seasonal Variability in the Dominant Species in Wastewater Final Effluents
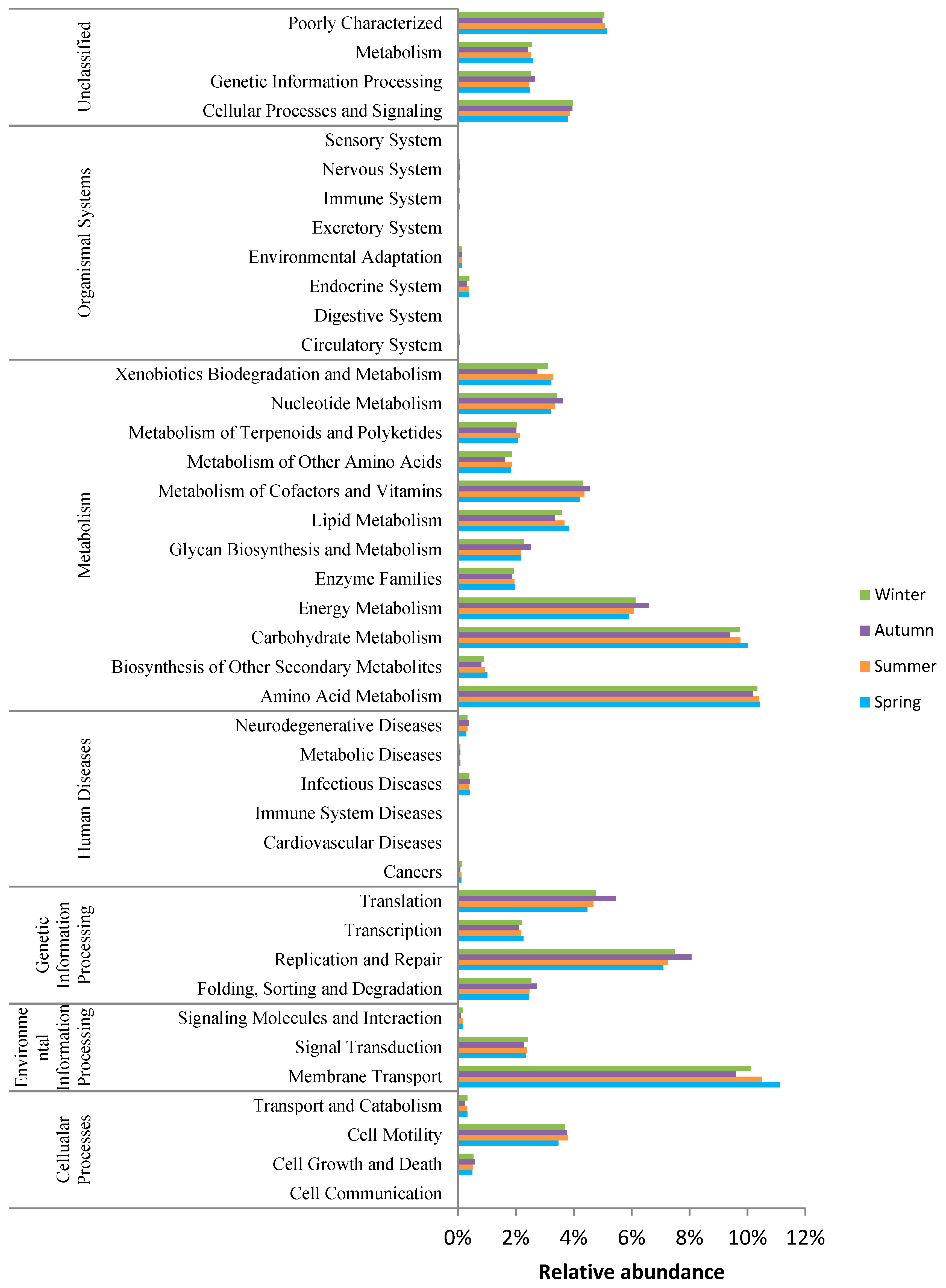
3.4. Functional Profiling of Bacteria in the Wastewater Final Effluents
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Omar, K.B.; Barnard, T.G. The occurrence of pathogenic Escherichia coli in South African wastewater treatment plants as detected by multiplex PCR. Water SA 2010, 36, 172–176. [Google Scholar] [CrossRef]
- Wani, D.; Pandit, A.K.; Kamili, A.N. Microbial assessment and effect of seasonal change on the removal efficiency of FAB based sewage treatment plant. J. Environ. Eng. Ecol. Sci. 2013, 2, 1–4. [Google Scholar] [CrossRef]
- Naidoo, S.; Olaniran, A.O. Treated Wastewater Effluent as a Source of Microbial Pollution of Surface Water Resources. Int. J. Environ. Res. Public Health 2014, 11, 249–270. [Google Scholar] [CrossRef] [PubMed]
- Osuolale, O.; Anthony, O. Assessment of the Physicochemical Qualities and Prevalence of Escherichia coli and Vibrios in the Final Effluents of Two Wastewater Treatment Plants in South Africa: Ecological and Public Health Implications. Int. J. Environ. Res. Public Health 2015, 12, 13399–13412. [Google Scholar] [CrossRef] [PubMed]
- Mema, V. Impact of poorly maintained waste water and sewage treatment plants: Lessons from South Africa. ReSource 2010, 12, 60–65. [Google Scholar]
- Muhammetoglu, A.; Yalcin, O.B.; Ozcan, T. Prediction of wastewater dilution and indicator bacteria concentrations for marine outfall systems. Mar. Environ. Res. 2012, 78, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Ashbolt, N.J. Microbial contamination of drinking water and disease outcomes in developing regions. Toxicology 2004, 198, 229–238. [Google Scholar] [CrossRef] [PubMed]
- Wakelin, S.A.; Colloff, M.J.; Kookanal, R.S. Effect of wastewater treatment plant effluent on microbial function and community structure in the sediment of a freshwater stream with variable seasonal flow. Appl. Environ. Microbiol. 2008, 74, 2659–2668. [Google Scholar] [CrossRef]
- Odjadjare, E.E.O.; Obi, L.C.; Okoh, A.I. Municipal Wastewater Effluents as a Source of Listerial Pathogens in the Aquatic Milieu of the Eastern Cape Province of South Africa: A Concern of Public Health Importance. Int. J. Environ. Res. Public Health 2010, 7, 2376–2394. [Google Scholar] [CrossRef]
- Cai, L.; Ju, F.; Zhang, T. Tracking human sewage microbiome in a municipal wastewater treatment plant. Appl. Microbiol. Biotechnol. 2014, 98, 3317–3326. [Google Scholar] [CrossRef]
- Benedek, T.; Táncsics, A.; Szilágyi, N.; Tóth, I.; Farkas, M.; Szoboszlay, S.; Krifaton, C.; Hartman, M.; Kriszt, B. Analysis of biofilm bacterial communities responsible for carbon removal through a reactor cascade treating wastewater. World J. Microbiol. Biotechnol. 2014, 30, 977–987. [Google Scholar] [CrossRef] [PubMed]
- Benedek, T.; Táncsics, A.; Szilágyi, N.; Tóth, I.; Farkas, M.; Szoboszlay, S.; Krifaton, C.; Hartman, M.; Kriszt, B. Occurrence of Vibrio cholerae non-O1 in three wastewater treatment plants in Agadir (Morocco). World J. Microbiol. Biotechnol. 2011, 27, 1099–1108. [Google Scholar]
- Carducci, A.; Battistini, R.; Rovini, E.; Verani, M. Viral Removal by Wastewater Treatment: Monitoring of Indicators and Pathogens. Food Environ. Virol. 2009, 1, 85–91. [Google Scholar] [CrossRef]
- Asli, K.; Fatma, G. Comparison of fecal coliform bacteria before and after wastewater treatment plant in the Izmir Bay (Eastern Aegean Sea). Environ. Monit. Assess. 2010, 162, 355–363. [Google Scholar]
- Pruesse, E.; Quast, C.; Knittel, K.; Fuchs, B.M.; Ludwig, W.; Peplies, J. SILVA: A comprehensive onlineresource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 2007, 35, 7188–7196. [Google Scholar] [CrossRef]
- Ye, L.; Zhang, T. Pathogenic bacteria in sewage treatment plants as revealed by 454 pyrosequencing. Environ. Sci. Technol. 2011, 45, 7173–7179. [Google Scholar] [CrossRef]
- Cai, L.; Zhang, T. Detecting human bacterial pathogens in wastewater treatment plants by a high-throughput shotgun sequencing technique. Environ. Sci. Technol. 2013, 47, 5433–5441. [Google Scholar] [CrossRef]
- Liu, Y.; Jin, J.H.; Liu, H.C.; Liu, Z.P. Dokdonella immobilis sp. nov., isolated from a batch reactor for the treatment of triphenylmethane dye effluent. Int. J. Syst. Evol. Microbiol. 2013, 63, 1557–1561. [Google Scholar] [CrossRef]
- Amarger, N.; Macheret, V.; Laguerre, G. Rhizobium gallicum sp. nov. And Rhizobium giardinii sp. nov., from Phaseolus vulgaris Nodules. Int. J. Syst. Bacteriol. 1997, 47, 996–1006. [Google Scholar] [CrossRef]
- Carro, L.; Rivas, R.; León-Barrios, M.; González-Tirante, M.; Velázquez, E.; Valverde, A. Herbaspirillum canariense sp. nov., Herbaspirillum aurantiacum sp. nov. and Herbaspirillum soli sp. nov., isolated from volcanic mountain soil, and emended description of the genus Herbaspirillum. Int. J. Syst. Evol. Microbiol. 2012, 62, 1300–1306. [Google Scholar] [CrossRef]
- Lai, C.C.; Tan, C.K.; Chou, C.H.; Hsu, H.L.; Liao, C.H.; Huang, Y.T.; Yang, P.C.; Luh, K.T.; Hsueh, P.R. Increasing incidence of nontuberculous mycobacteria, Taiwan, 2000–2008. Emerg. Infect. Dis. 2010, 16, 294–296. [Google Scholar] [CrossRef] [PubMed]
- Tomohiro, I.; Tomoyuki, H.; Hidenobu, A.; Hiroshi, H. Architecture, component, and microbiome of biofilm involved in the fouling of membrane bioreactors. Biofilms Microbiomes 2017, 3, 5. [Google Scholar] [CrossRef]
- Lloret, J.; Bolaños, L.; Lucas, M.M.; Peart, J.M.; Brewin, N.J.; Bonilla, I.; Rivilla, R. Ionic stress and osmotic pressure induce different alterations in the lipopolysaccharide of a Rhizobium meliloti strain. Appl. Environ. Microbiol. 1995, 61, 3701–3704. [Google Scholar] [PubMed]
- Ahmad, M.; Zahir, Z.A.; Nazli, F.; Akram, F.; Arshad, M.; Khalid, M. Effectiveness of halo-tolerant, auxin producing Pseudomonas and Rhizobium strains to improve osmotic stress tolerance in mung bean (Vigna radiata L.). Braz. J. Microbiol. 2013, 44, 1341–1348. [Google Scholar] [CrossRef] [PubMed]
- Mnasri, B.; Tajini, F.; Trabelsi, M.; Aouani, M.E.; Mhamdi, R. Rhizobium gallicum as an efficient symbiont for bean cultivation. Agron. Sustain. 2007, 27, 331–336. [Google Scholar] [CrossRef]
- Hahn, M.W. Description of seven candidate species affiliated with the phylum Actinobacteria, representing planktonic freshwater bacteria. Int. J. Syst. Evol. Microbiol. 2009, 59, 112–117. [Google Scholar] [CrossRef] [PubMed]
- Newton, R.J.; Jones, S.E.; Eiler, A.; McMahon, K.D.; Bertilsson, S. A Guide to the Natural History of Freshwater Lake Bacteria. Am. Soc. Microbiol. 2011, 1, 14–49. [Google Scholar] [CrossRef]
- Bajerski, F.; Ganzert, L.; Mangelsdorf, K.; Lipski, A.; Busse, H.J.; Padur, L.; Wagner, D. Herbaspirillum psychrotolerans sp. nov., a member of the family Oxalobacteraceae from a glacier forefield. Int. J. Syst. Evol. Microbiol. 2013, 63, 3197–3203. [Google Scholar] [CrossRef]
- Kampfer, P.; Glaeser, S.P.; Lodders, N.; Busse, H.J.; Falsen, E. Herminiimonas contaminans sp. nov., isolated as a contaminant of biopharmaceuticals. Int. J. Syst. Evol. Microbiol. 2012, 63, 412–420. [Google Scholar] [CrossRef]
- Lin, S.Y.; Hameed, A.; Arun, A.B.; Liu, Y.C.; Hsu, Y.H.; Lai, W.A.; Rekha, P.D.; Young, C.C. Description of Noviherbaspirillum malthae gen. nov., sp. nov., isolated from an oil-contaminated soil, and proposal to reclassify Herbaspirillum soli, Herbaspirillum aurantiacum, Herbaspirillum canariense and Herbaspirillum psychrotolerans as Noviherbaspirillum soli comb. nov., Noviherbaspirillum aurantiacum comb. nov., Noviherbaspirillum canariense comb. nov. and Noviherbaspirillum psychrotolerans comb. nov. based on polyphasic analysis. Int. J. Syst. Evol. Microbiol. 2013, 63, 4100–4107. [Google Scholar]
- Bernal-Guadarrama, M.J.; Fernández-Gallardo, N.; Zamora-Padrón, R.; Pacheco, V.; Reyes-Batlle, M.; Valladares, B.; Lorenzo-Morales, J.; Martínez-Carretero, E. Evaluation of Two Commercially Available Immunological Kits for the Diagnosis of Helicobacter spp. in Bottlenose dolphins (Tursiops truncatus). Curr. Microbiol. 2015, 70, 685–689. [Google Scholar] [CrossRef] [PubMed]
- Jogler, C.; Glöckner, F.O.; Roberto, K. Characterization of Planctomyces limnophilus and Development of Genetic Tools for Its Manipulation Establish It as a Model Species for the Phylum Planctomycetes. Appl. Environ. Microbiol. 2011, 77, 5826–5829. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, S.; Silva, F.; Queiroz, J.A.; Oleastro, M.; Domingues, F.C. Resveratrol against Arcobacter butzleri and Arcobacter cryaerophilus: Activity and effect on cellular functions. Int. J. Food Microbiol. 2014, 180, 62–68. [Google Scholar] [CrossRef] [PubMed]
- Giacometti, F.; Lucchi, A.; Di Francesco, A.; Delogu, M.; Grilli, E.; Guarniero, I.; Stancampiano, L.; Manfreda, G.; Merialdi, G.; Serraino, A. Arcobacter butzleri, Arcobacter cryaerophilus, and Arcobacter skirrowii Circulation in a Dairy Farm and Sources of Milk Contamination. Appl. Environ. Microbiol. 2015, 15, 5055–5063. [Google Scholar] [CrossRef]
- Jiang, Z.D.; Dupont, H.L.; Brown, E.L.; Nandy, R.K.; Ramamurthy, T.; Sinha, A.; Ghosh, S.; Guin, S.; Gurleen, K.; Rodrigues, S.; et al. Microbial etiology of travelers’ diarrhea in Mexico, Guatemala and India importance of enterotoxigenic Bacteroides fragilis and Arcobacter species. J. Clin. Microbiol. 2010, 48, 1417–1419. [Google Scholar] [CrossRef]
- Collado, L.; Figueras, M.J. Taxonomy, Epidemiology, and Clinical Relevance of the Genus Arcobacter. Clin. Microbiol. Rev. 2011, 24, 174–192. [Google Scholar] [CrossRef]
- Mercante, J.W.; Winchel, J.M. Current and Emerging Legionella Diagnostics for Laboratory and Outbreak Investigations. Clin. Microbiol. Rev. 2015, 1, 95–133. [Google Scholar] [CrossRef]
- McDade, J.E.; Shepard, C.C.; Fraser, D.W.; Tsai, T.R.; Redus, M.A.; Dowdle, W.R. Legionnaires’ disease: Isolation of abacterium and demonstration of its role in other respiratorydisease. N. Engl. J. Med. 1977, 297, 1197–1203. [Google Scholar] [CrossRef]
- Newton, H.J.; Ang, D.K.; van Driel, I.R.; Hartland, E.L. Molecular pathogenesis of infections caused by Legionella pneumophila. Clin. Microbiol. Rev. 2010, 23, 274–298. [Google Scholar] [CrossRef]
- Radomski, N.; Roguet, A.; Lucas, F.S.; Veyrier, F.J.; Cambau, E.; Accrombessi, H.; Moilleron, R.; Behr, M.A.; Moulin, L. atpE gene as a new useful specific molecular target to quantify Mycobacterium in environmental samples. BMC Microbiol. 2013, 13, 277. [Google Scholar] [CrossRef]
- Thomson, R.M. NTM working group at Queensland TB Control Centre and Queensland Mycobacterial Reference Laboratory. Changing epidemiology of pulmonary nontuberculous mycobacteria infections. Emerg. Infect. Dis. 2010, 16, 1576–1583. [Google Scholar] [CrossRef] [PubMed]
- Dai, J.L.; Chen, Y.S.; Lauzardo, M. Web-accessible database of hsp65 sequences from mycobacterium reference strains. J. Clin. Microbiol. 2011, 6, 2296–2303. [Google Scholar] [CrossRef] [PubMed]
- Higgins, J.; Camp, P.; Farrell, D.; Bravo, D.; Pate, M.; Robbe-Austerman, S. Identification of Mycobacterium spp. of veterinary importance using rpoB gene sequencing. BMC Vet. Res. 2011, 7, 77–81. [Google Scholar] [CrossRef] [PubMed]
- Hamid, R.; Ahmad, A.; Usup, G. Pathogenicity of Aeromonas hydrophila isolated from the Malaysian Sea against coral (Turbinaria sp.) and sea bass (Lates calcarifer). Environ. Sci. Pollut. Res. 2016, 23, 1614–7499. [Google Scholar] [CrossRef] [PubMed]
- Van den Abeele, A.M.; Vogelaers, D.; Vanlaere, E.; Houf, K. Antimicrobial susceptibility testing of Arcobacter butzleri and Arcobacter cryaerophilus strains isolated from Belgian patients. J. Antimicrob. Chemotherpy 2016, 71, 1241–1244. [Google Scholar] [CrossRef] [PubMed]
- Fisher, J.C.; Levican, A.; Figueras, M.J.; Mclellan, S.L. Population dynamics and ecology of Arcobacter in sewage. Front. Microbiol. 2014, 5, 525–530. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, S.; Queiroz, J.A.; Oleastro, M.; Domingues, F.C. Insights in the pathogenesis and resistance of Arcobacter. Crit. Rev. Microbiol. 2016, 42, 364–383. [Google Scholar]
- Dumler, J.S.; Barbet, A.F.; Bekker, C.P.; Dasch, G.A.; Palmer, G.H. Arcobacter trophiarum sp. nov., isolated from fattening pigs. Int. J. Syst. Evol. Microbiol. 2011, 61, 356–361. [Google Scholar]
- Levican, A.; Collado, L.; Aguilar, C.; Yustes, C.; Diéguez, A.L. Arcobacter bivalviorum sp. nov. and Arcobacter venerupis sp. nov., new species isolated from shellfish. Syst. Appl. Microbiol. 2012, 35, 133–138. [Google Scholar] [CrossRef]
- Wetsch, N.M.; Somani, K.; Tyrrell, G.J.; Gebhart, C.; Bailey, R.J. Campylobacter Curvus-Associated Hepatic Abscesses: A Case Report. J. Clin. Microbol. 2006, 5, 1909–1911. [Google Scholar] [CrossRef]
- Almuzara, M.N.; Cittadini, R.; Vera, O.C.; Bakai, R.; Traglia, G. Intra-Abdominal Infections Due to Comamonas kerstersii. J. Clin. Microbiol. 2013, 6, 1998–2000. [Google Scholar] [CrossRef] [PubMed]
- Vasoo, S.; Mason, E.L.; Gustafson, D.R.; Cunningham, S.A.; Cole, N.C. Desulfovibrio legallii Prosthetic Shoulder Joint Infection and Review of Antimicrobial Susceptibility and Clinical Characteristics of Desulfovibrio Infections. J. Clin. Microbiol. 2014, 52, 3105–3110. [Google Scholar] [CrossRef] [PubMed]
- Fox, G.J.; Taylor, N.S.; Howe, S. Helicobacter anseris sp. nov. and Helicobacter brantae sp. nov., Isolated from Feces of Resident Canada Geese in the Greater Boston Area. Appl. Environ. Microbiol. 2006, 72, 4633–4637. [Google Scholar] [CrossRef] [PubMed]
- Adeleke, A.A.; Fields, B.S.; Benson, R.F.; Daneshvar, M.I.; Pruckler, J.M. Legionella drozanskii sp. nov., Legionella rowbothamii sp. nov. and Legionella fallonii sp.nov.: Three unusual new Legionella species. Int. J. Syst. Evol. Microbiol. 2001, 51, 1151–1160. [Google Scholar] [CrossRef] [PubMed]
- Daniel, G.; David, T.; Kathryn, H.; Deborah, T. Cutaneous Legionella longbeachae Infection in Immunosuppressed Woman, United Kingdom. Emerg. Infect. Dis. 2015, 21, 1426–1428. [Google Scholar]
- Lo, P.F.; Riffard, S.; Jarraud, S.; Le, G.F.; Richet, H. Isolation of Legionella oakridgensis from Two Patients with Pleural Effusion Living in the Same Geographical Area. J. Clin. Microbiol. 2000, 38, 3128–3130. [Google Scholar]
- Huth, R.G.; Brown-Elliott, B.A.; Wallace, R.J., Jr. Mycobacterium mageritense Pulmonary Disease in Patient with Compromised Immune System. Emerg. Infect. Dis. 2011, 17, 556–558. [Google Scholar] [CrossRef]
- Pinkos, A.C.; Friedrichs, K.R.; Monaghan, K.; Sample, N.S.H.; Trepanier, L.A. Transient cold agglutinins associated with Mycoplasma cynos pneumonia in a dog. Vet. Clin. Pathol. 2015, 44, 498–502. [Google Scholar] [CrossRef]
- Ilha, M.R.S.; Rajeev, S.; Watson, C.; Woldemeskel, M. Meningoencephalitis caused by Mycoplasma edwardii in a dog. J. Vet. Diagn. Investig. 2010, 22, 805–808. [Google Scholar] [CrossRef]
- Solary, J.; Socolovschi, C.; Aubry, C.; Brouqui, P.; Raoult, D.; Parola, P. Detection of Rickettsia sibirica mongolitimonae by Using Cutaneous Swab Samples and Quantitative PCR. Emerg. Infect. Dis. 2014, 20, 716–718. [Google Scholar] [CrossRef]
- Best, M.G.; Stout, J.E.; Yu, V.L.; Muder, R.R. Tatlockia micdadei (Pittsburgh Pneumonia Agent) Growth Kinetics May Explain Its Infrequent Isolation from Water and the Low Prevalence of Pittsburgh Pneumonia. Appl. Environ. Microbiol. 1985, 49, 1521–1522. [Google Scholar] [PubMed]
| Season | Date | T (°C) | BOD5 (mg/L) | CODcr (mg/L) | SS (mg/L) | TN (mg/L) | TP (mg/L) | pH Values |
|---|---|---|---|---|---|---|---|---|
| Spring | 2017.04.23 | 15.3 | 9 | 47 | 8 | 15 | 0.6 | 6.8 |
| Summer | 2017.08.17 | 25.1 | 8 | 42 | 6 | 12 | 0.5 | 6.8 |
| Autumn | 2016.10.24 | 20.0 | 11 | 56 | 10 | 17 | 0.7 | 6.9 |
| Winter | 2017.01.25 | 1.0 | 10 | 52 | 7 | 16 | 0.6 | 6.9 |
| Sample | Chao | Ace | Chao/ACE | Shannon |
|---|---|---|---|---|
| Sample in spring | 1732 | 1940 | 0.89 | 8.56 |
| Sample in summer | 5381 | 5397 | 0.99 | 8.54 |
| Sample in autumn | 6075 | 6121 | 0.99 | 9.38 |
| Sample in winter | 3588 | 5079 | 0.71 | 6.37 |
| Serial Number | Similar Species of Geneticrelationship | Accession Number in NCBI | Spring (%) | Summer (%) | Autumn (%) | Winter (%) |
|---|---|---|---|---|---|---|
| 1 | Acidisoma tundrae | NR_042705 | — | — | 2.27 | — |
| 2 | Anaerolinea thermophila | NR_074383 | 2.40 | — | — | — |
| 3 | Arcobacter cryaerophilus | NR_025905 | — | 1.76 | 0.67 | 3.59 |
| 4 | Blastopirellula cremea | NR_118153 | — | 0.99 | 1.19 | 1.89 |
| 5 | Candidatus Flaviluna lacus | NR_125496 | — | 2.86 | 5.50 | 0.54 |
| 6 | Candidatus Nitrospira defluvii | NR_074700 | 1.16 | — | 0.69 | 0.23 |
| 7 | Comamonas terrigena | NR_114856 | — | — | — | 2.20 |
| 8 | Corynebacterium deserti | NR_118005 | — | — | 1.98 | 0.62 |
| 9 | Demequina flava | NR_113566 | — | — | 1.85 | 0.26 |
| 10 | Dokdonella immobilis | NR_108377 | 4.97 | — | — | — |
| 11 | Ferruginibacter alkalilentus | NR_044588 | 0.56 | — | — | 2.08 |
| 12 | Flavobacter iumanseonense | NR_132690 | 2.51 | — | — | — |
| 13 | Geobacter anodireducens | NR_126282 | — | 2.50 | — | — |
| 14 | Gracilimonas rosea | NR_109751 | — | 1.31 | — | — |
| 15 | Helicobacter cetorum | NR_074476 | — | 1.01 | 5.15 | 1.59 |
| 16 | Lactococcus piscium | NR_043739 | 1.65 | — | — | — |
| 17 | Levilinea saccharolytica | NR_040972 | 2.40 | 0.37 | — | — |
| 18 | Limisphaera ngatamarikiensis | NR_134756 | — | — | — | 1.17 |
| 19 | Limnobacter thiooxidans | NR_025421 | — | 1.59 | 0.51 | — |
| 20 | Massili aflava | NR_117901 | 0.35 | 2.03 | — | 1.15 |
| 21 | Mycoplasma cynos | NR_102477 | — | — | — | 1.15 |
| 22 | Nautilia abyssi | NR_042690 | — | — | 2.15 | 0.60 |
| 23 | Nevskia ramosa | NR_025269 | — | — | 0.59 | 1.47 |
| 24 | Noviherbaspirillum aurantiacum | NR_118040 | — | — | 6.06 | — |
| 25 | Noviherbaspirillum soli | NR_118041 | 1.22 | — | — | — |
| 26 | Pedobacter oryzae | NR_116174 | 1.45 | 2.00 | — | — |
| 27 | Pirellula staleyi | NR_074521 | — | 1.76 | — | — |
| 28 | Planctomyces limnophilus | NR_074670 | 0.27 | 0.22 | 1.88 | 3.99 |
| 29 | Rhizobium gallicum | NR_036785 | — | 4.67 | — | — |
| 30 | Salimicrobium flavidum | NR_104548 | — | 1.86 | — | 1.40 |
| 31 | Schlesneria paludicola | NR_042466 | — | 1.65 | 1.03 | 1.76 |
| 32 | Sphingomonas starnbergensis | NR_109485 | — | 1.02 | — | — |
| 33 | Sulfuricurvum kujiense | NR_074398 | — | — | 0.27 | 1.11 |
| 34 | Thermogutta hypogea | NR_134825 | — | — | — | 1.34 |
| 35 | Thermogutta terrifontis | NR_134826 | — | — | 2.22 | 1.94 |
| Serial Number | Similar Species of Geneticrelationship | Spring (%) | Summer (%) | Autumn (%) | Winter (%) | Total | Disease Category |
|---|---|---|---|---|---|---|---|
| 1 | Aeromonas hydrophila | — | 0.61 | — | — | 0.61 | dermatosis/diarrhea [44] |
| 2 | Arcobacter cryaerophilus | — | 1.76 | 0.67 | 3.59 | 6.02 | diarrhea [45] |
| 3 | Arcobacter defluvii | — | 0.26 | — | — | 0.26 | enteropatia [46] |
| 4 | Arcobacter skirrowii | 0.15 | 0.24 | — | — | 0.39 | enterogastritis [47] |
| 5 | Arcobacter trophiarum | — | — | — | 0.35 | 0.35 | septicemia [48] |
| 6 | Arcobacter venerupis | 0.32 | — | 0.87 | — | 1.19 | enteropatia [49] |
| 7 | Campylobacter curvus | — | — | — | 0.53 | 0.53 | enterogastritis [50] |
| 8 | Chlamydophila pneumoniae | 0.21 | — | — | — | 0.21 | pneumonia |
| 9 | Comamonas kerstersii | — | 0.73 | — | — | 0.73 | diarrhea [51] |
| 10 | Desulfovibriolegallii | — | 0.41 | — | — | 0.41 | Dermatosis [52] |
| 11 | Helicobacter brantae | — | — | 0.39 | — | 0.39 | Gastrohelcoma [53] |
| 12 | Helicobacter mastomyrinus | 0.24 | — | — | — | 0.24 | Gastrohelcoma [53] |
| 13 | Legionella fallonii | 0.98 | 0.55 | — | — | 1.53 | Legionellosis [54] |
| 14 | Legionella longbeachae | 0.63 | 0.44 | — | — | 1.07 | Dermatosis [55] |
| 15 | Legionella oakridgensis | — | 0.7 | — | — | 0.7 | fever and dyspnea [56] |
| 16 | Mycobacterium mageritense | 0.29 | 0.2 | 0.89 | — | 1.38 | Dermatosis [57] |
| 17 | Mycoplasma cynos | — | — | — | 1.15 | 1.15 | pneumonia [58] |
| 18 | Mycoplasma edwardii | — | 0.22 | — | — | 0.22 | pneumonia/septicemia [59] |
| 19 | Pseudomonas putida | 0.42 | — | — | — | 0.42 | urinary infections/dermatosis |
| 20 | Prevotella copri | 0.25 | — | — | — | 0.25 | pneumonia |
| 21 | Rickettsia sibirica | — | 0.24 | — | — | 0.24 | Dermatosis [60] |
| 22 | Staphylococcus Equorum | 0.55 | — | — | — | 0.55 | Dermatosis/septicemia |
| 23 | Tatlockiamic dadei | — | 0.22 | — | — | 0.22 | pittsburgh pneumonia [61] |
| total | 4.04 | 6.58 | 2.82 | 5.62 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, X.; Lang, X.L.; Xu, A.-L.; Song, Z.-W.; Yang, J.; Guo, M.-Y. Seasonal Variability in the Microbial Community and Pathogens in Wastewater Final Effluents. Water 2019, 11, 2586. https://doi.org/10.3390/w11122586
Chen X, Lang XL, Xu A-L, Song Z-W, Yang J, Guo M-Y. Seasonal Variability in the Microbial Community and Pathogens in Wastewater Final Effluents. Water. 2019; 11(12):2586. https://doi.org/10.3390/w11122586
Chicago/Turabian StyleChen, Xiang, Xiu Lu Lang, Ai-Ling Xu, Zhi-Wen Song, Juan Yang, and Ming-Yue Guo. 2019. "Seasonal Variability in the Microbial Community and Pathogens in Wastewater Final Effluents" Water 11, no. 12: 2586. https://doi.org/10.3390/w11122586
APA StyleChen, X., Lang, X. L., Xu, A.-L., Song, Z.-W., Yang, J., & Guo, M.-Y. (2019). Seasonal Variability in the Microbial Community and Pathogens in Wastewater Final Effluents. Water, 11(12), 2586. https://doi.org/10.3390/w11122586





