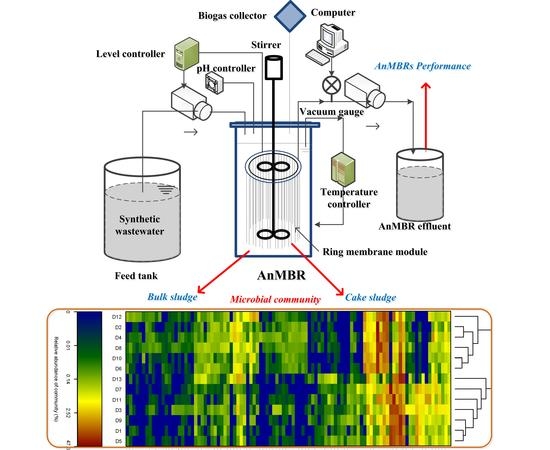
The Performance and Microbial Community Identification in Mesophilic and Atmospheric Anaerobic Membrane Bioreactor for Municipal Wastewater Treatment Associated with Different Hydraulic Retention Times
Abstract
1. Introduction
2. Materials and Methods
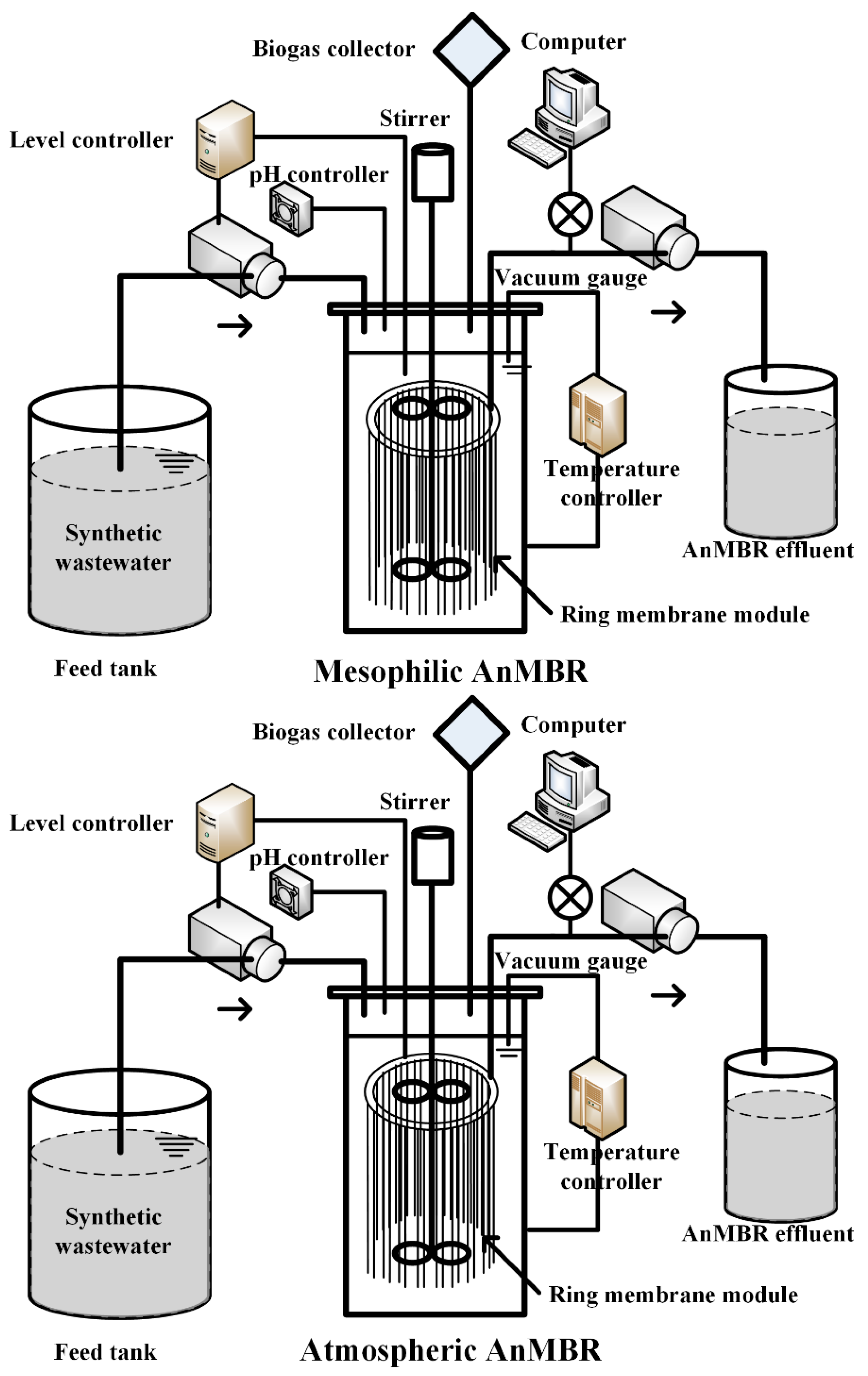
2.1. AnMBRs Operation with Ring Membrane Modules
2.2. Sludge Sample Collection
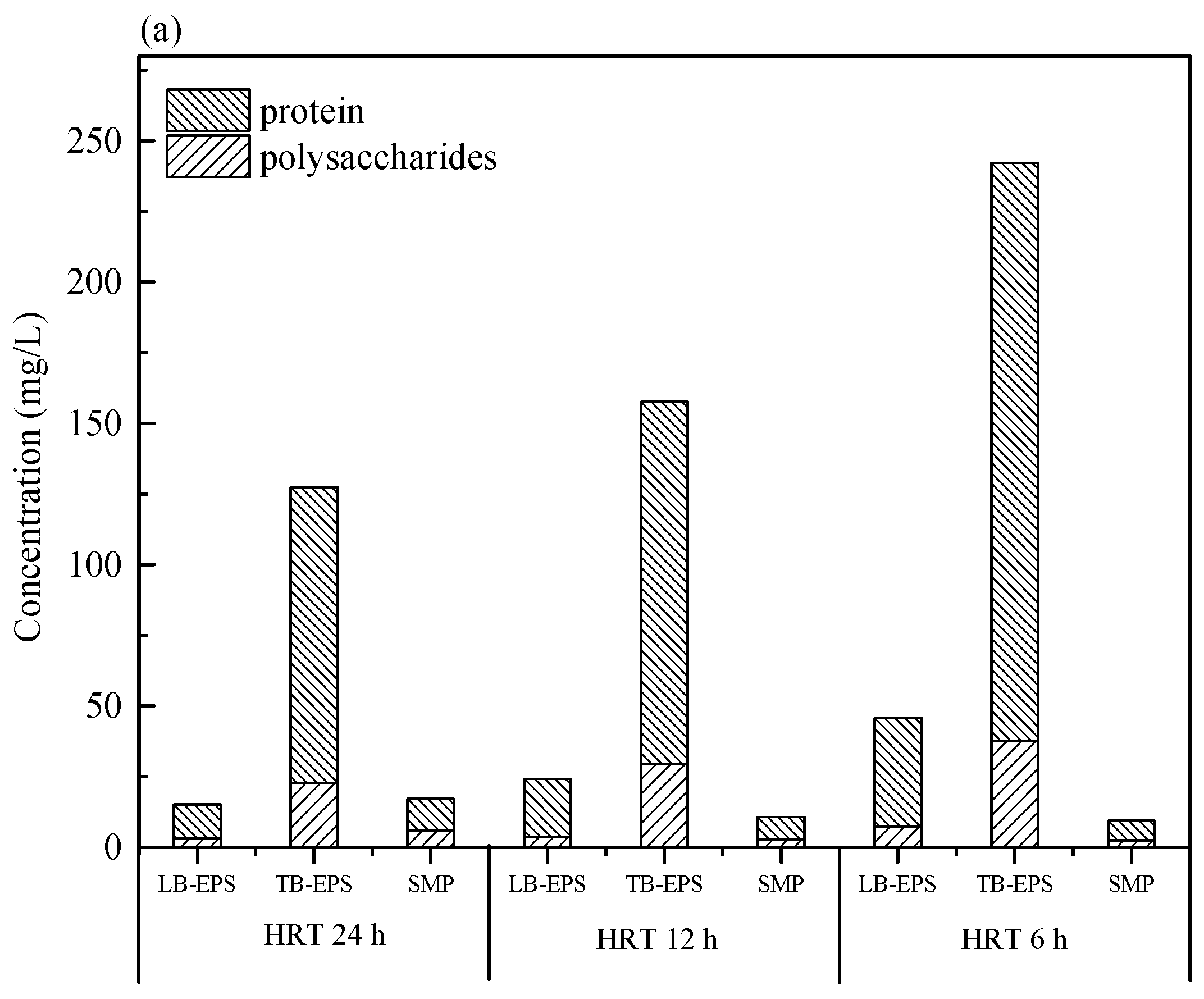
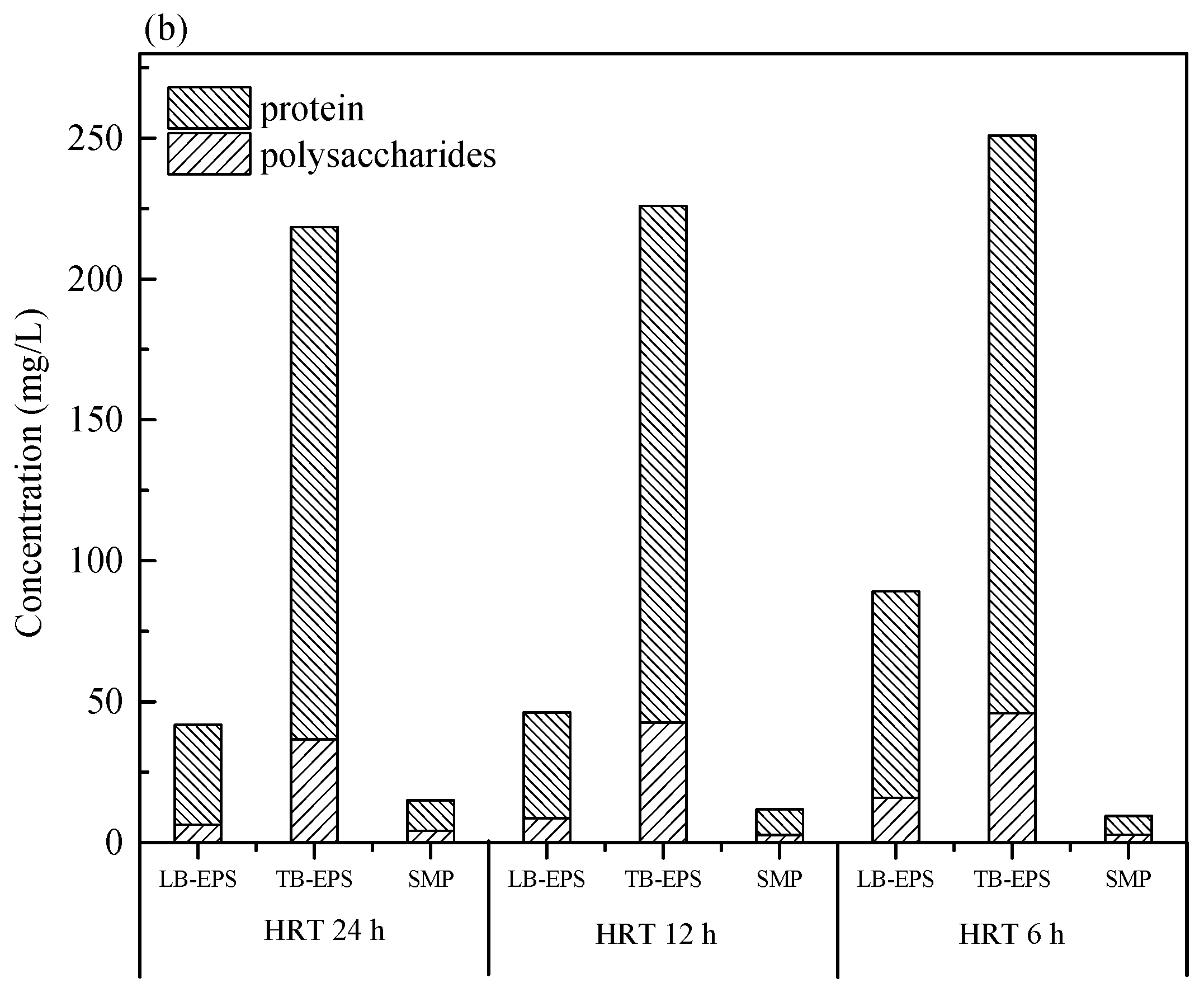
2.3. Extraction of SMP and EPS
2.4. Microbial Community Analysis
2.5. Other Item Analysis
3. Results and Discussion
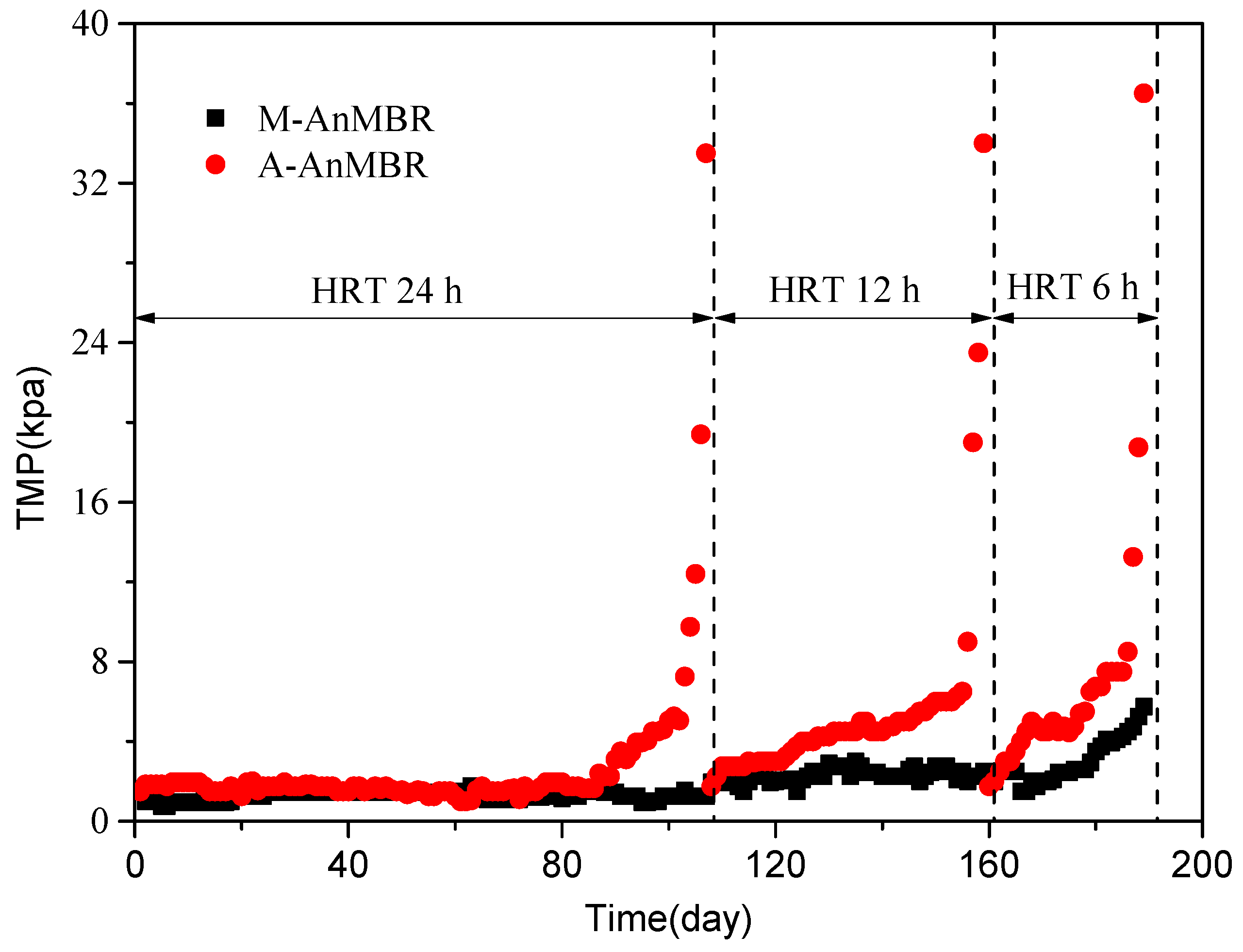
3.1. Performance of the AnMBRs
3.2. Microbial Richness and Diversity Evolution with the Variation of HRT in M-AnMBR and A-AnMBR
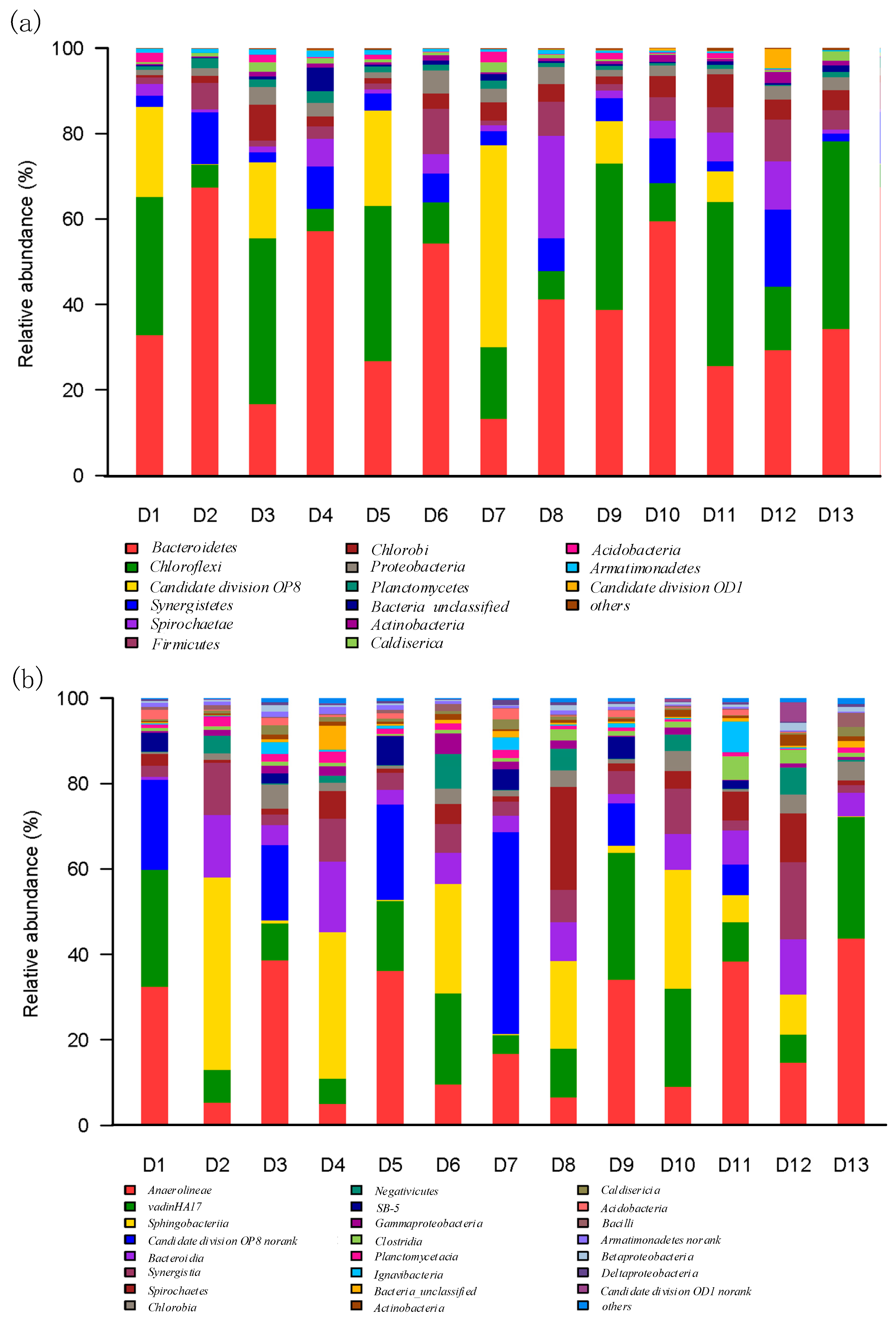
3.3. Microbial Phylum and Class Identification with Different HRTs in M-AnMBR and A-AnMBR
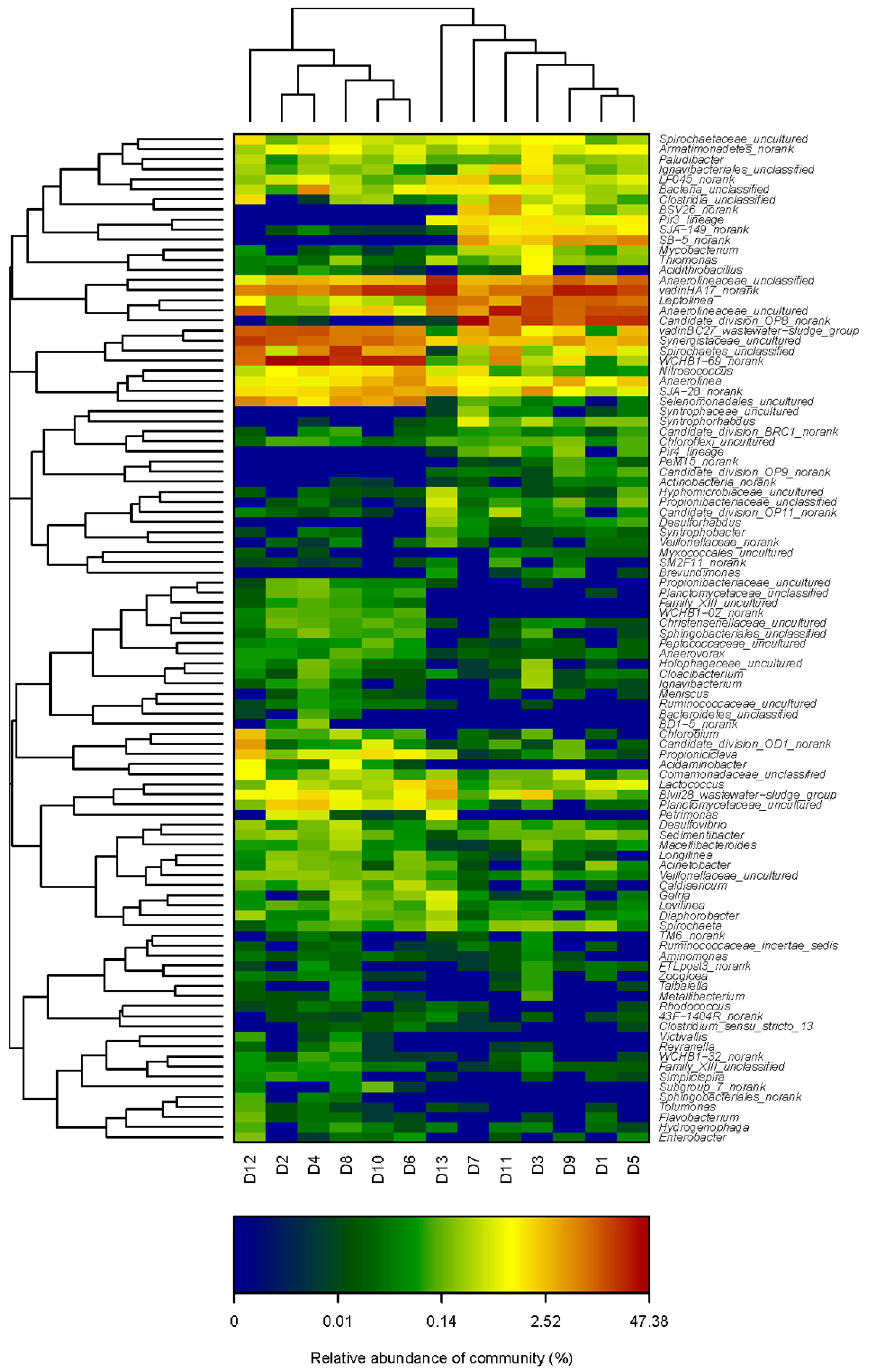
3.4. Microbial Genera Evolution with the Variation of HRT in M-AnMBR and A-AnMBR
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Luo, G.; Wang, W.; Angelidaki, I. Anaerobic Digestion for Simultaneous Sewage Sludge Treatment and CO Biomethanation: Process Performance and Microbial Ecology. Environ. Sci. Technol. 2013, 47, 10685–10693. [Google Scholar] [CrossRef] [PubMed]
- Hwang, B.-K.; Kim, J.-H.; Ahn, C.H.; Lee, C.-H.; Song, J.-Y.; Ra, Y.-H. Effect of disintegrated sludge recycling on membrane permeability in a membrane bioreactor combined with a turbulent jet flow ozone contactor. Water Res. 2010, 44, 1833–1840. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; Chae, S.-R.; Drews, A.; Kraume, M.; Shin, H.-S.; Yang, F. Recent advances in membrane bioreactors (MBRs): Membrane fouling and membrane material. Water Res. 2009, 43, 1489–1512. [Google Scholar] [CrossRef] [PubMed]
- Gao, W.J.; Qu, X.; Leung, K.T.; Liao, B.Q. Influence of temperature and temperature shock on sludge properties, cake layer structure, and membrane fouling in a submerged anaerobic membrane bioreactor. J. Membr. Sci. 2012, 421–422, 131–144. [Google Scholar] [CrossRef]
- Smith, A.L.; Stadler, L.B.; Love, N.G.; Skerlos, S.J.; Raskin, L. Perspectives on anaerobic membrane bioreactor treatment of domestic wastewater: A critical review. Bioresour. Technol. 2012, 122, 149–159. [Google Scholar] [CrossRef]
- Wang, W.; Yang, Q.; Zheng, S.S.; Wu, D.L. Anaerobic membrane bioreactor (AnMBR) for bamboo industry wastewater treatment. Bioresour. Technol. 2013, 149, 292–300. [Google Scholar] [CrossRef]
- Lin, H.; Chen, J.; Wang, F.; Ding, L.; Hong, H. Feasibility evaluation of submerged anaerobic membrane bioreactor for municipal secondary wastewater treatment. Desalination 2011, 280, 120–126. [Google Scholar] [CrossRef]
- Mcmahon, K.D.; Stroot, P.G.; Mackie, R.I.; Raskin, L. Anaerobic codigestion of municipal solid waste and biosolids under various mixing conditions—II: Microbial population dynamics. Water Res. 2001, 35, 1804–1816. [Google Scholar] [CrossRef]
- Gao, D.W.; Tao, Y.; An, R. Digested sewage treatment using membrane-based process at different hydraulic retention times. Desalination 2012, 286, 187–192. [Google Scholar] [CrossRef]
- Martinez-Sosa, D.; Helmreich, B.; Netter, T.; Paris, S.; Bischof, F.; Horn, H. Anaerobic submerged membrane bioreactor (AnSMBR) for municipal wastewater treatment under mesophilic and psychrophilic temperature conditions. Bioresour. Technol. 2011, 102, 10377–10385. [Google Scholar] [CrossRef]
- Gao, W.J.J.; Lin, H.J.; Leung, K.T.; Liao, B.Q. Influence of elevated pH shocks on the performance of a submerged anaerobic membrane bioreactor. Process Biochem. 2010, 45, 1279–1287. [Google Scholar] [CrossRef]
- Huang, X.; Liu, R.; Qian, Y. Behaviour of soluble microbial products in a membrane bioreactor. Process Biochem. 2000, 36, 401–406. [Google Scholar] [CrossRef]
- Gao, D.W.; Wen, Z.D.; Li, B.; Liang, H. Microbial community structure characteristics associated membrane fouling in A/O-MBR system. Bioresour. Technol. 2014, 154, 87–93. [Google Scholar] [CrossRef] [PubMed]
- Yeon, K.M.; Cheong, W.S.; Oh, H.S.; Lee, W.N.; Hwang, B.K.; Lee, C.H.; Beyenal, H.; Lewandowski, Z. Quorum Sensing: A New Biofouling Control Paradigm in a Membrane Bioreactor for Advanced Wastewater Treatment. Environ. Sci. Technol. 2009, 43, 380–385. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.; Kim, S.; Yeon, K.M.; Sang, B.I.; Chun, J.; Lee, C.H. Correlation between microbial community structure and biofouling in a laboratory scale membrane bioreactor with synthetic wastewater. Desalination 2012, 287, 209–215. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, Z.; Wu, Z.; Wei, T.; Lu, F.; Tong, J.; Mai, S. Membrane fouling in an anaerobic dynamic membrane bioreactor (AnDMBR) for municipal wastewater treatment: Characteristics of membrane foulants and bulk sludge. Process Biochem. 2011, 46, 1538–1544. [Google Scholar] [CrossRef]
- Miura, Y.; Watanbe, Y.; Okabe, S. Membrane biofouling in pilot-scale membrane bioreactors (MBRs) treating municipal wastewater: Impact of biofilm formation. Environ. Sci. Technol. 2007, 41, 632–638. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.N.; De Wever, H.; Diels, L. Diverse and Distinct Bacterial Communities Induced Biofilm Fouling in Membrane Bioreactors Operated under Different Conditions. Environ. Sci. Technol. 2008, 42, 8360–8366. [Google Scholar] [CrossRef]
- Ma, J.X.; Wang, Z.W.; Zou, X.X.; Feng, J.J.; Wu, Z.C. Microbial communities in an anaerobic dynamic membrane bioreactor (AnDMBR) for municipal wastewater treatment: Comparison of bulk sludge and cake layer. Process Biochem. 2013, 48, 510–516. [Google Scholar] [CrossRef]
- Liu, L.; Li, T.; Wei, X.; Jiang, B.; Fang, P. Effects of a nutrient additive on the density of functional bacteria and the microbial community structure of bioorganic fertilizer. Bioresour. Technol. 2014, 172, 328–334. [Google Scholar] [CrossRef]
- Liang, S.; Gliniewicz, K.; Mendes-Soares, H.; Settles, M.L.; Forney, L.J.; Coats, E.R.; McDonald, A.G. Comparative analysis of microbial community of novel lactic acid fermentation inoculated with different undefined mixed cultures. Bioresour. Technol. 2015, 179, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Tian, Y.; Li, Z.P.; Liu, F.; You, H. Characterization of organic membrane foulants in a forward osmosis membrane bioreactor treating anaerobic membrane bioreactor effluent. Bioresour. Technol. 2014, 167, 137–143. [Google Scholar] [CrossRef] [PubMed]
- Li, X.Y.; Yang, S.F. Influence of loosely bound extracellular polymeric substances (EPS) on the flocculation, sedimentation and dewaterability of activated sludge. Water Res. 2007, 41, 1022–1030. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Tian, Y.; Liu, J.; Li, N.; Zhang, J.; Zuo, W.; Li, Z. Investigation of microbial structure and composition involved in membrane fouling in the forward osmosis membrane bioreactor treating anaerobic bioreactor effluent. Chem. Eng. J. 2016, 286, 198–207. [Google Scholar] [CrossRef]
- CEPB (Chinese Environmental Science Press). Standard Methods for Examination of Water and Wastewater, 4th ed.; Chinese Environmental Science Press: Beijing, China, 2002. [Google Scholar]
- Wang, Z.W.; Wu, Z.C. A Review of Membrane Fouling in MBRs: Characteristics and Role of Sludge Cake Formed on Membrane Surfaces. Sep. Sci. Technol. 2009, 44, 3571–3596. [Google Scholar] [CrossRef]
- Flemming, H.C.; Wingender, J. Relevance of microbial extracellular polymeric substances (EPSs)—Part I: Structural and ecological aspects. Water Sci. Technol. 2001, 43, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.M.; Sheng, G.P.; Luo, H.W.; Zhang, F.; Yuan, S.J.; Xu, J.; Zeng, R.J.; Wu, J.G.; Yu, H.Q. Contribution of Extracellular Polymeric Substances (EPS) to the Sludge Aggregation. Environ. Sci. Technol. 2010, 44, 4355–4360. [Google Scholar] [CrossRef]
- Xie, Z.F.; Wang, Z.W.; Wang, Q.Y.; Zhu, C.W.; Wu, Z.C. An anaerobic dynamic membrane bioreactor (AnDMBR) for landfill leachate treatment: Performance and microbial community identification. Bioresour. Technol. 2014, 161, 29–39. [Google Scholar] [CrossRef]
- Schmidt, I.; Sliekers, O.; Schmid, M.; Bock, E.; Fuerst, J.; Kuenen, J.G.; Jetten, M.S.M.; Strous, M. New concepts of microbial treatment processes for the nitrogen removal in wastewater. FEMS Microbiol. Rev. 2003, 27, 481–492. [Google Scholar] [CrossRef]
- Nogueira, R.; Melo, L.F.; Purkhold, U.; Wuertz, S.; Wagner, M. Nitrifying and heterotrophic population dynamics in biofilm reactors: Effects of hydraulic retention time and the presence of organic carbon. Water Res. 2002, 36, 469–481. [Google Scholar] [CrossRef]
- Gao, D.W.; Zhang, T.; Tang, C.Y.Y.; Wu, W.M.; Wong, C.Y.; Lee, Y.H.; Yeh, D.H.; Criddle, C.S. Membrane fouling in an anaerobic membrane bioreactor: Differences in relative abundance of bacterial species in the membrane foulant layer and in suspension. J. Membr. Sci. 2010, 364, 331–338. [Google Scholar] [CrossRef]
- Yu, Z.Y.; Wen, X.H.; Xu, M.L.; Huang, X. Characteristics of extracellular polymeric substances and bacterial communities in an anaerobic membrane bioreactor coupled with online ultrasound equipment. Bioresour. Technol. 2012, 117, 333–340. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.D.; Li, J.; Guo, P.; Li, P.P.; Suo, Y.L.; Wang, X.F.; Cui, Z.J. Dynamic transition of microbial communities in response to acidification in fixed-bed anaerobic baffled reactors (FABR) of two different flow directions. Bioresour. Technol. 2011, 102, 4703–4711. [Google Scholar] [CrossRef] [PubMed]
- Narihiro, T.; Terada, T.; Ohashi, A.; Kamagata, Y.; Nakamura, K.; Sekiguchi, Y. Quantitative detection of previously characterized syntrophic bacteria in anaerobic wastewater treatment systems by sequence-specific rRNA cleavage method. Water Res. 2012, 46, 2167–2175. [Google Scholar] [CrossRef]
- Chen, C.L.; Macarie, H.; Ramirez, I.; Olmos, A.; Ong, S.L.; Monroy, O.; Liu, W.T. Microbial community structure in a thermophilic anaerobic hybrid reactor degrading terephthalate. Microbiology 2004, 150, 3429–3440. [Google Scholar] [CrossRef]
- Fang, H.H.P.; Liang, D.W.; Zhang, T.; Liu, Y. Anaerobic treatment of phenol in wastewater under thermophilic condition. Water Res. 2006, 40, 427–434. [Google Scholar] [CrossRef] [PubMed]
- Mbadinga, S.M.; Li, K.P.; Zhou, L.; Wang, L.Y.; Yang, S.Z.; Liu, J.F.; Gu, J.D.; Mu, B.Z. Analysis of alkane-dependent methanogenic community derived from production water of a high-temperature petroleum reservoir. Appl. Microbiol. Biotechnol. 2012, 96, 531–542. [Google Scholar] [CrossRef]
- Chen, S.Y.; Dong, X.Z. Proteiniphilum acetatigenes gen. nov., sp nov., from a UASB reactor treating brewery wastewater. Int. J. Syst. Evol. Microbiol. 2005, 55, 2257–2261. [Google Scholar] [CrossRef]
- Riviere, D.; Desvignes, V.; Pelletier, E.; Chaussonnerie, S.; Guermazi, S.; Weissenbach, J.; Li, T.; Camacho, P.; Sghir, A. Towards the definition of a core of microorganisms involved in anaerobic digestion of sludge. ISME J. 2009, 3, 700–714. [Google Scholar] [CrossRef]
- Wirth, R.; Kovács, E.; Maróti, G.; Bagi, Z.; Rákhely, G.; Kovács, K.L. Characterization of a biogas-producing microbial community by short-read next generation DNA sequencing. Biotechnol. Biofuels 2012, 5, 1–16. [Google Scholar] [CrossRef]
- Ito, T.; Yoshiguchi, K.; Ariesyady, H.D.; Okabe, S. Identification of a novel acetate-utilizing bacterium belonging to Synergistes group 4 in anaerobic digester sludge. ISME J. 2011, 5, 1844–1856. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.M.; Kim, J.H.; Ha, J.H.; Park, J.M. Bacterial and methanogenic archaeal communities during the single-stage anaerobic digestion of high-strength food wastewater. Bioresour. Technol. 2014, 165, 174–182. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.M.; Chon, D.H.; Kim, H.S.; Park, C. Investigation of bacterial community in activated sludge with an anaerobic side-stream reactor (ASSR) to decrease the generation of excess sludge. Water Res. 2012, 46, 4292–4300. [Google Scholar] [CrossRef] [PubMed]
- Shin, S.G.; Han, G.; Lim, J.; Lee, C.; Hwang, S. A comprehensive microbial insight into two-stage anaerobic digestion of food waste-recycling wastewater. Water Res. 2010, 44, 4838–4849. [Google Scholar] [CrossRef] [PubMed]
- Delbes, C.; Moletta, R.; Godon, J.J. Monitoring of activity dynamics of an anaerobic digester bacterial community using 16S rRNA polymerase chain reaction—Single-strand conformation polymorphism analysis. Environ. Microbiol. 2000, 2, 506–515. [Google Scholar] [CrossRef] [PubMed]
| Items | HRT | Influent COD (mg/L) | Supernate COD (mg/L) | Microbial Removal Rate (%) | Permeate COD (mg/L) | Total Treatment Efficiency (%) |
|---|---|---|---|---|---|---|
| M-AnMBR | 24 h | 570.41 | 152.53 | 73.26 | 50.81 | 91.09 |
| A-AnMBR | 24 h | 570.41 | 259.97 | 54.42 | 137.06 | 75.91 |
| M-AnMBR | 12 h | 630.49 | 87.44 | 86.13 | 40.94 | 93.51 |
| A-AnMBR | 12 h | 630.49 | 171.86 | 72.74 | 109.84 | 82.58 |
| M-AnMBR | 6 h | 578.15 | 63.00 | 89.11 | 44.00 | 92.39 |
| A-AnMBR | 6 h | 578.15 | 209.50 | 63.79 | 170.00 | 71.63 |
| Sample | Reads | 0.97 | |||||
|---|---|---|---|---|---|---|---|
| OTU | ACE a | Chao1 b | Shannon c | Simpson d | Coverage e | ||
| D1 | 11549 | 161 | 219 | 205 | 2.56 | 0.14 | 0.995584 |
| D2 | 8399 | 152 | 221 | 236 | 2.54 | 0.2129 | 0.994166 |
| D3 | 10960 | 240 | 286 | 290 | 3.26 | 0.0883 | 0.994708 |
| D4 | 20442 | 253 | 286 | 288 | 3.22 | 0.1161 | 0.997750 |
| D5 | 11562 | 160 | 202 | 196 | 2.91 | 0.0987 | 0.996281 |
| D6 | 20943 | 216 | 264 | 252 | 3.02 | 0.0977 | 0.997565 |
| D7 | 19287 | 196 | 233 | 245 | 2.66 | 0.2062 | 0.997615 |
| D8 | 13563 | 248 | 284 | 287 | 3.23 | 0.1008 | 0.996387 |
| D9 | 11678 | 166 | 213 | 224 | 3.05 | 0.0818 | 0.996061 |
| D10 | 19954 | 189 | 226 | 232 | 2.89 | 0.1183 | 0.997845 |
| D11 | 16000 | 195 | 249 | 239 | 2.97 | 0.117 | 0.996687 |
| D12 | 12428 | 217 | 336 | 289 | 3.37 | 0.0607 | 0.994689 |
| D13 | 16176 | 184 | 239 | 248 | 2.81 | 0.1278 | 0.996847 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ding, Y.; Liang, Z.; Guo, Z.; Li, Z.; Hou, X.; Jin, C. The Performance and Microbial Community Identification in Mesophilic and Atmospheric Anaerobic Membrane Bioreactor for Municipal Wastewater Treatment Associated with Different Hydraulic Retention Times. Water 2019, 11, 160. https://doi.org/10.3390/w11010160
Ding Y, Liang Z, Guo Z, Li Z, Hou X, Jin C. The Performance and Microbial Community Identification in Mesophilic and Atmospheric Anaerobic Membrane Bioreactor for Municipal Wastewater Treatment Associated with Different Hydraulic Retention Times. Water. 2019; 11(1):160. https://doi.org/10.3390/w11010160
Chicago/Turabian StyleDing, Yi, Zhenlin Liang, Zhansheng Guo, Zhipeng Li, Xuguang Hou, and Chao Jin. 2019. "The Performance and Microbial Community Identification in Mesophilic and Atmospheric Anaerobic Membrane Bioreactor for Municipal Wastewater Treatment Associated with Different Hydraulic Retention Times" Water 11, no. 1: 160. https://doi.org/10.3390/w11010160
APA StyleDing, Y., Liang, Z., Guo, Z., Li, Z., Hou, X., & Jin, C. (2019). The Performance and Microbial Community Identification in Mesophilic and Atmospheric Anaerobic Membrane Bioreactor for Municipal Wastewater Treatment Associated with Different Hydraulic Retention Times. Water, 11(1), 160. https://doi.org/10.3390/w11010160