Neo-Domestication of an Interspecific Tetraploid Helianthus annuus × Helianthus tuberous Population That Segregates for Perennial Habit
Abstract
:1. Introduction
2. Materials and Methods
2.1. Populations Used and Selection within the Perennial Sunflower Breeding Program
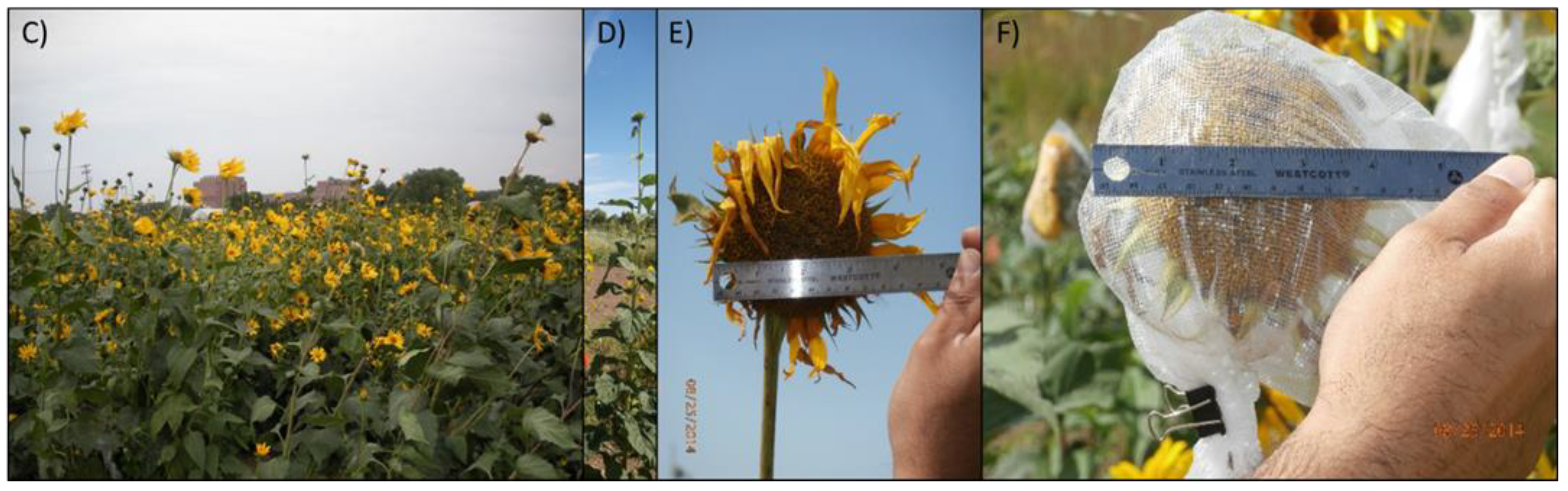
2.2. Tuber Phenotyping
2.3. DNA Extraction and Genotyping-by-Sequencing Library Preparation
2.4. Variant Identification and Marker Discovery
2.5. Quantitative Trait Loci Mapping
3. Results
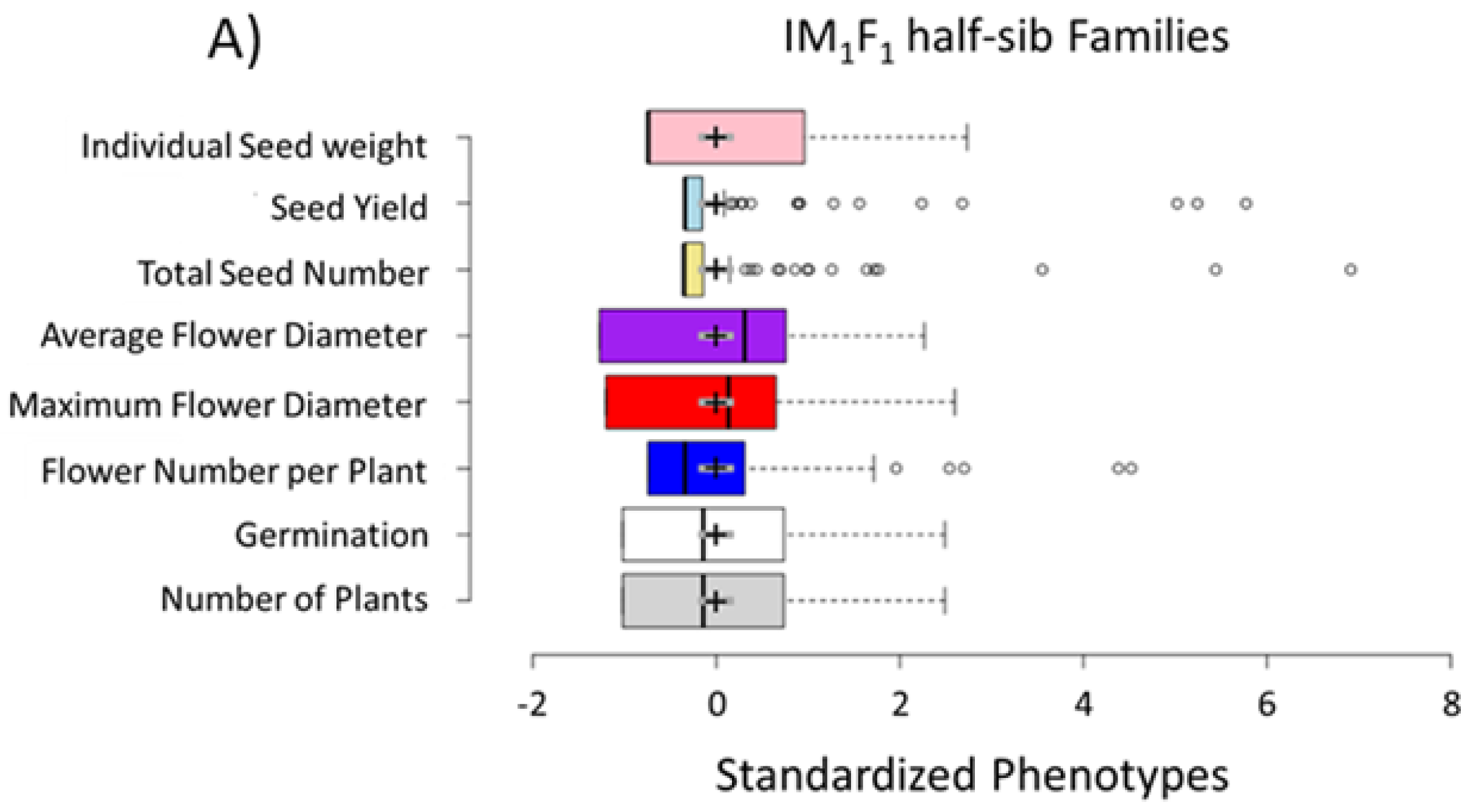
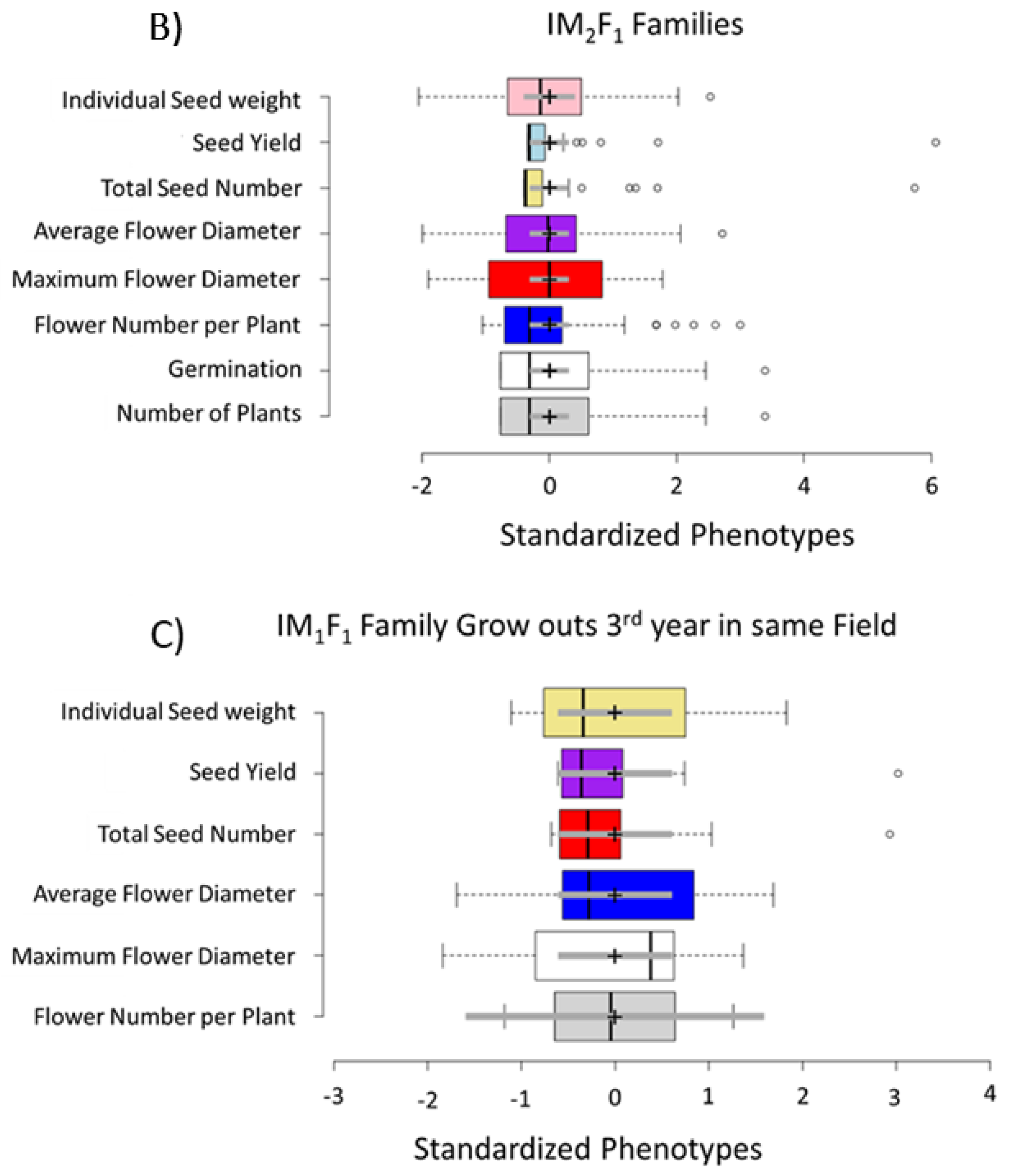
3.1. Selection for Domestication Syndrome Traits
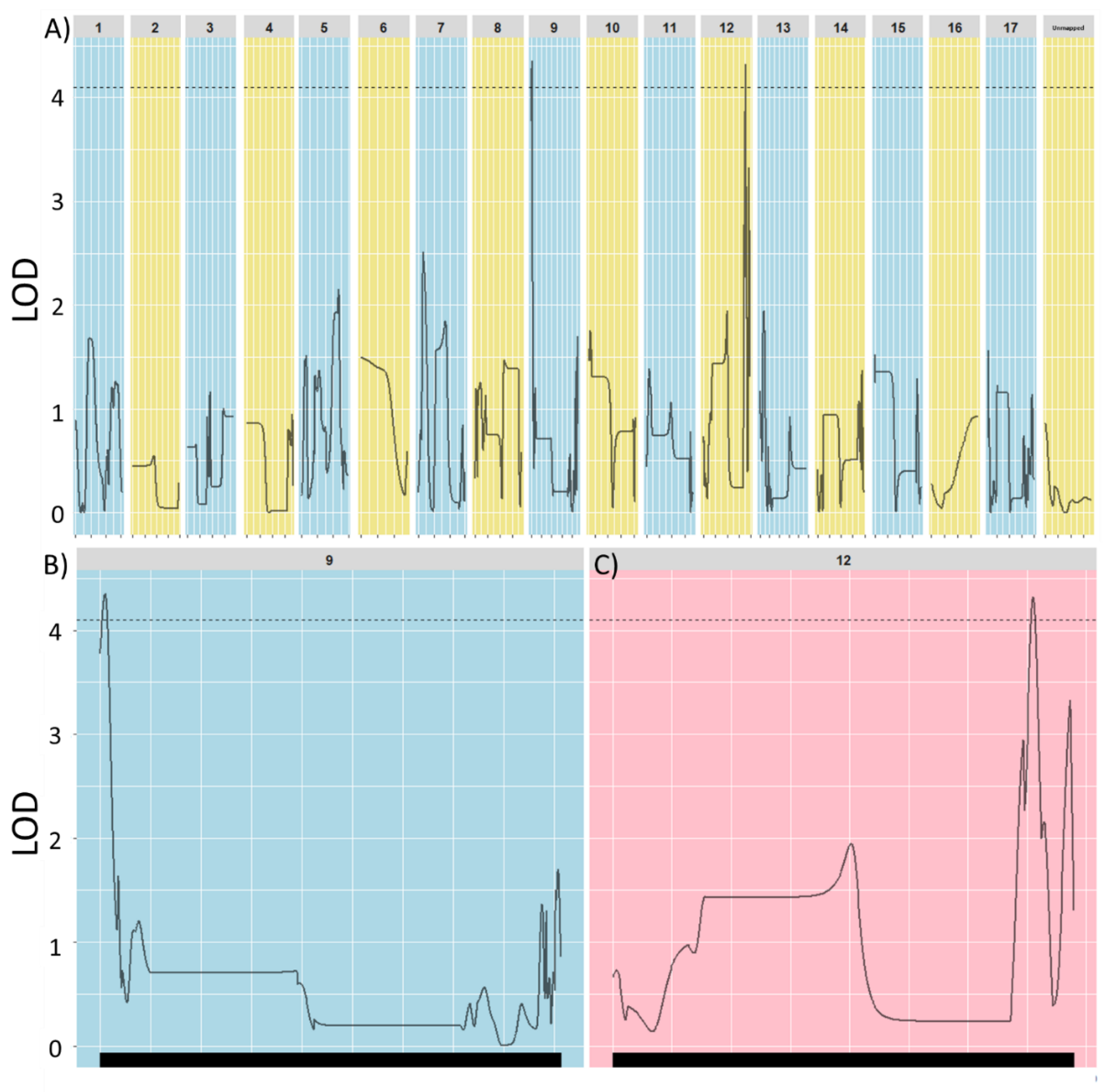
3.2. Perenniality Mapping in the Interspecific Population
4. Discussion
4.1. Modified Domestication
4.2. Genetics of Tuber Formation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Data Archiving Statement
References
- Glover, J.D.; Reganold, J.P.; Bell, L.W.; Borevitz, J.; Brummer, E.C.; Buckler, E.S.; Cox, C.M.; Cox, T.S.; Crews, T.E.; Culman, S.W.; et al. Increased food and ecosystem security via perennial grains. Science 2010, 328, 1638–1639. [Google Scholar] [CrossRef] [PubMed]
- Godfray, H.C.J.; Garnett, T. Food security and sustainable intensification. Phil. Trans. R. Soc. B Biol. Sci. 2014, 369, 20120273. [Google Scholar] [CrossRef] [PubMed]
- Kantar, M.B.; Tyl, C.E.; Dorn, K.M.; Zhang, X.; Jungers, J.M.; Kaser, J.M.; Schendel, R.R.; Eckberg, J.O.; Runck, B.C.; Bunzel, M.; et al. Perennial grain and oilseed crops. Annu. Rev. Plant Biol. 2016, 67, 703–729. [Google Scholar] [CrossRef] [PubMed]
- Runck, B.C.; Kantar, M.B.; Jordan, N.R.; Anderson, J.A.; Wyse, D.L.; Eckberg, J.O.; Barnes, R.J.; Lehman, C.L.; DeHaan, L.R.; Stupar, R.M.; et al. The Reflective Plant Breeding Paradigm: A Robust System of Germplasm Development to Support Strategic Diversification of Agroecosystems. Crop Sci. 2014, 54, 1939–1948. [Google Scholar] [CrossRef] [Green Version]
- DeHaan, L.R.; Van Tassel, D.L. Useful insights from evolutionary biology for developing perennial grain crops. Am. J. Bot. 2014, 101, 1801–1819. [Google Scholar] [CrossRef] [PubMed]
- Henry, R.J. Genomics strategies for germplasm characterization and the development of climate resilient crops. Front. Plant Sci. 2014, 5, 68. [Google Scholar] [CrossRef] [PubMed]
- Henry, R.J. Next-generation sequencing for understanding and accelerating crop domestication. Brief. Funct. Genom. 2012, 11, 51–56. [Google Scholar] [CrossRef] [PubMed]
- Allaby, R.G.; Kitchen, J.; Fuller, D.Q. Surprisingly low limits of selection in plant domestication. Evol. Bioinform. 2016, 2015, 41–51. [Google Scholar] [CrossRef] [PubMed]
- Allaby, R.G.; Brown, T.A.; Fuller, D.Q. A simulation of the effect of inbreeding on crop domestication genetics with comments on the integration of archaeobotany and genetics: A reply to Honne and Heun. Veg. Hist. Archaeobot. 2010, 19, 151–158. [Google Scholar] [CrossRef]
- Allaby, R.G.; Fuller, D.Q.; Brown, T.A. The genetic expectations of a protracted model for the origins of domesticated crops. Proc. Natl. Acad. Sci. USA 2008, 105, 13982–13986. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cox, T.S.; Bender, M.; Picone, C.; Van Tassel, D.L.; Holland, J.B.; Brummer, E.C.; Zoeller, B.E.; Paterson, A.H.; Jackson, W. Breeding perennial grain crops. Crit. Rev. Plant Sci. 2002, 21, 59–91. [Google Scholar] [CrossRef]
- Cox, T.S.; Van Tassel, D.L.; Cox, C.M.; DeHaan, L.R. Progress in breeding perennial grains. Crop Pasture Sci. 2010, 61, 513–521. [Google Scholar] [CrossRef]
- Fuller, D.Q. Contrasting patterns in crop domestication and domestication rates: Recent archaeobotanical insights from the Old World. Ann. Bot. 2007, 100, 903–924. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.Z.; Innan, H. Nonindependent domestication of the two rice subspecies, Oryza sativa ssp. indica and ssp. japonica, demonstrated by multilocus microsatellites. Genetics 2008, 179, 965–976. [Google Scholar] [CrossRef] [PubMed]
- Hillman, G.C.; Davies, M.S. Domestication rates in wild-type wheats and barley under primitive cultivation. Biol. J. Linn. Soc. 1990, 39, 39–78. [Google Scholar] [CrossRef]
- Gepts, P. Crop domestication as a long-term selection experiment. Plant Breed. Rev. 2004, 24, 1–44. [Google Scholar]
- Donald, C.M.T. The breeding of crop ideotypes. Euphytica 1968, 17, 385–403. [Google Scholar] [CrossRef]
- Shapter, F.M.; Cross, M.; Ablett, G.; Malory, S.; Chivers, I.H.; Graham, J.K.; Robert, J.H. High throughput sequencing and mutagenesis to accelerate the domestication of Microlaena stipoides as a new food crop. PLoS ONE 2013, 8, e82641. [Google Scholar] [CrossRef] [PubMed]
- Harlan, J.R.; De Wet, J.M.J.; Price, E.G. Comparative evolution of cereals. Evolution 1973, 27, 311–325. [Google Scholar] [CrossRef] [PubMed]
- Doebley, J.; Stec, A. Genetic analysis of the morphological differences between maize and teosinte. Genetics 1991, 129, 285–295. [Google Scholar] [PubMed]
- Sang, T. Genes and mutations underlying domestication transitions in grasses. Plant Physiol. 2009, 149, 63–70. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Zhou, A.; Sang, T. Genetic analysis of rice domestication syndrome with the wild annual species, Oryza nivara. New Phytol. 2006, 170, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Cuevas, H.E.; Das, S.; Sezen, U.U.; Zhou, C.; Guo, H.; Valorie, H.G.; Ge, Z.; Clemente, T.E.; Paterson, A.H. Seed shattering in a wild sorghum is conferred by a locus unrelated to domestication. Proc. Natl. Acad. Sci. USA 2013, 110, 15824–15829. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ray, D.K.; Mueller, N.D.; West, P.C.; Foley, J.A. Yield trends are insufficient to double global crop production by 2050. PLoS ONE 2013, 8, e66428. [Google Scholar] [CrossRef] [PubMed]
- Khoury, C.K.; Bjorkman, A.D.; Dempewolf, H.; Ramirez-Villegas, J.; Guarino, L.; Jarvis, A.; Rieseberg, L.H.; Struik, P.C. Increasing homogeneity in global food supplies and the implications for food security. Proc. Natl. Acad. Sci. USA 2014, 111, 4001–4006. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baulcombe, D.; Crute, I.; Davies, B.; Dunwell, J.; Gale, M.; Jones, J.; Pretty, J.; Sutherland, W.; Toulmin, C. Reaping the Benefits: Science and the Sustainable Intensification of Global Agriculture; The Royal Society: London, UK, 2009. [Google Scholar]
- Laurance, W.F.; Sayer, J.; Cassman, K.G. Agricultural expansion and its impacts on tropical nature. Trends Ecol. Evol. 2014, 29, 107–116. [Google Scholar] [CrossRef] [PubMed]
- Tilman, D.; Balzer, C.; Hill, J.; Befort, B.L. Global food demand and the sustainable intensification of agriculture. Proc. Natl. Acad. Sci. USA 2011, 108, 20260–20264. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Foley, J.A.; Ramankutty, N.; Brauman, K.A.; Cassidy, E.S.; Gerber, J.S.; Johnston, M.; Mueller, N.D.; O’Connell, C.; Ray, D.K.; West, P.C.; et al. Solutions for a cultivated planet. Nature 2011, 478, 337–342. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miller, A.J.; Gross, B.L. From Forest to Field: Perennial Fruit Crop Domestication. Am. J. Bot. 2011, 98, 1389–1414. [Google Scholar] [CrossRef] [PubMed]
- Kantar, M.B.; Betts, K.; Michno, J.M.; Luby, J.J.; Morrell, P.L.; Hulke, B.S.; Stupar, R.M.; Wyse, D.L. Evaluating an interspecific Helianthus annuus × Helianthus tuberosus population for use in a perennial sunflower breeding program. Field Crops Res. 2014, 155, 254–264. [Google Scholar] [CrossRef]
- Hulke, B.S.; Wyse, D.L. Using interspecific hybrids with H. annuus L. In Proceedings of the 17th International Sunflower Conference, Cordoba, Spain, 8–12 June 2008; pp. 729–734. [Google Scholar]
- Elshire, R.J.; Glaubitz, J.C.; Sun, Q.; Poland, J.A.; Kawamoto, K.; Buckler, E.S.; Mitchell, S.E. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 2011, 6, e19379. [Google Scholar] [CrossRef] [PubMed]
- Poland, J.A.; Brown, P.J.; Sorrells, M.E.; Jannink, J.L. Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequencing approach. PLoS ONE 2012, 7, e32253. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.F.; Gulya, T.J.; Vick, B.A. Registration of two maintainer (HA 434 and HA 435) and three restorer (RHA 436–RHA 438) high oleic oilseed sunflower germplasms. Crop Sci. 2004, 44, 1034–1035. [Google Scholar] [CrossRef]
- Hockett, E.A.; Knowles, P.F. Inheritance of branching in sunflowers, Helianthus annuus L. Crop Sci. 1970, 10, 432–436. [Google Scholar] [CrossRef]
- R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2016; Available online: http://www.Rproject.org/ (accessed on 31 July 2018).
- Schneiter, A.A.; Miller, J.F. Description of sunflower growth stages. Crop Sci. 1981, 21, 901–903. [Google Scholar] [CrossRef]
- Webb, D.M.; Knapp, S.J. DNA extraction from a previously recalcitrant plant genus. Mol. Biol. Rep. 1990, 8, 180–185. [Google Scholar] [CrossRef]
- Badouin, H.; Gouzy, J.; Grassa, C.J.; Murat, F.; Staton, S.E.; Cottret, L.; Lelandais-Brière, C.; Owens, G.L.; Carrère, S.; Mayjonade, B.; et al. The sunflower genome provides insights into oil metabolism, flowering and Asterid evolution. Nature 2017, 546, 148–152. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, H. Toward better understanding of artifacts in variant calling from high-coverage samples. Bioinformatics 2014, 30, 2843–2851. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lam, H.Y.; Pan, C.; Clark, M.J.; Lacroute, P.; Chen, R.; Haraksingh, R.; O'Huallachain, M.; Gerstein, M.B.; Kidd, J.M.; Bustamante, C.D.; et al. Detecting and annotating genetic variations using the HugeSeq pipeline. Nat. Biotechnol. 2012, 30, 226–229. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef] [PubMed]
- Warnes, G.; Gorjanc, G.; Friedrich Leisch, F.; Man, M. Genetics: Population Genetics. R Package Version 1.3.8.1. 2013. Available online: https://CRAN.R-project.org/package=genetics (accessed on 31 July 2018).
- Arends, D.; Prins, P.; Jansen, R.C.; Broman, K.W. R/qtl: High-throughput multiple QTL mapping. Bioinformatics 2010, 26, 2990–2992. [Google Scholar] [CrossRef] [PubMed]
- Broman, K.W. A Brief Tour of R/qtl. Available online: http://www.rqtl.org/tutorials/rqtltour.pdf (accessed on 31 July 2018).
- Churchill, G.A.; Doerge, R.W. Empirical threshold values for quantitative trait mapping. Genetics 1994, 138, 963–971. [Google Scholar] [PubMed]
- Falconer, D.S.; Mackay, T.F.C. Introduction to Quantitative Genetics; Longman: London, UK, 1995; Volume 19. [Google Scholar]
- Bock, D.G.; Kane, N.C.; Ebert, D.P.; Rieseberg, L.H. Genome skimming reveals the origin of the Jerusalem artichoke tuber crop species: Neither from Jerusalem nor an Artichoke. New Phytol. 2014, 201, 1021–1030. [Google Scholar] [CrossRef] [PubMed]
- Kostoff, D. A contribution to the meiosis of Helianthus tuberosus L. Z Pflanzenzuchtg 1934, 19, 423–438. [Google Scholar]
- Kostoff, D. Autosyndesis and structural hybridity in F1-hybrid Helianthus tuberosus L. × Helianthus annuus L. and their sequences. Genetica 1939, 21, 285–299. [Google Scholar] [CrossRef]
- Scibria, N. Hybrids between the Jerusalem Artichoke (Helianthus tuberosus L.) and the Sunflower (Helianthus annuus L.). C. R. Acad. Sci. URSS 1938, 2, 193–196. [Google Scholar]
- Long, R.W., Jr. Hybridization in perennial sunflowers. Am. J. Bot. 1955, 42, 769–777. [Google Scholar] [CrossRef]
- Long, R.W. Biosystematics of two perennial species of Helianthus (Compositae). I. Crossing relationships and transplant studies. Am. J. Bot. 1960, 47, 729–735. [Google Scholar] [CrossRef]
- Beavis, W.D. The power and deceit of QTL experiments: Lessons from comparative QTL studies. In Proceedings of the 49th Annual Corn and Sorghum Research Conference, Chicago, IL, USA, 9–10 December 1994; American Seed Trade Association: Washington, DC, USA, 1994; pp. 250–266. [Google Scholar]
- Linz, G.; Hulke, B.; Kantar, M.; Homan, J.; Stupar, R.; Wyse, D. Potential Use of Perennial Sunflower to Reduce Blackbird Damage to Sunflower; University of California: Davis, CA, USA, 2014. [Google Scholar]
- Klug, P.E.; Linz, G.M. Strategies for Evading Blackbird Damage. In Ecology and Management of Blackbirds (Icteridae) in North America; CRC Press: Boca Raton, FL, USA, 2017; pp. 175–190. [Google Scholar]
- Todesco, M.; Pascual, M.A.; Owens, G.L.; Ostevik, K.L.; Moyers, B.T.; Hübner, S.; Heredia, S.M.; Hahn, M.A.; Caseys, C.; Bock, D.G.; et al. Hybridization and extinction. Evol. Appl. 2016, 9, 892–908. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hufford, K.M.; Mazer, S.J. Plant ecotypes: Genetic differentiation in the age of ecological restoration. Trends Ecol. Evol. 2003, 18, 147–155. [Google Scholar] [CrossRef]
- Rieseberg, L.H.; Archer, M.A.; Wayne, R.K. Transgressive segregation, adaptation, and speciation. Heredity 1999, 83, 363–372. [Google Scholar] [CrossRef] [PubMed]
- Reimann-Philipp, R. Breeding perennial rye. Plant Breed. Rev. 1995, 13, 265–292. [Google Scholar]
- Van Tassel, D.L.; Albrecht, K.A.; Bever, J.D.; Boe, A.A.; Brandvain, Y.; Timothy, E.C.; Markus, G.; Pedro, G.; Luciana, G.-P.; Brent, S.H.; et al. Accelerating Silphium Domestication: An Opportunity to Develop New Crop Ideotypes and Breeding Strategies Informed by Multiple Disciplines. Crop Sci. 2017, 57, 1274–1284. [Google Scholar] [CrossRef]
- Hu, F.Y.; Tao, D.Y.; Sacks, E.; Fu, B.Y.; Xu, P.; Li, J.; Yang, Y.; McNally, K.; Khush, G.S.; Paterson, A.H.; et al. Convergent evolution of perenniality in rice and sorghum. Proc. Natl. Acad. Sci. USA 2003, 100, 4050–4054. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Westerbergh, A.; Doebley, J. Quantitative trait loci controlling phenotypes related to the perennial versus annual habit in wild relatives of maize. Theor. Appl. Genet. 2004, 109, 1544–1553. [Google Scholar] [CrossRef] [PubMed]
- Moyers, B.T.; Owens, G.L.; Baute, G.J.; Rieseberg, L.H. The genetic architecture of UV floral patterning in sunflower. Ann. Bot. 2017, 120, 39–50. [Google Scholar] [CrossRef] [PubMed]
- Fuller, D.Q.; Denham, T.; Arroyo-Kalin, M.; Lucas, L.; Stevens, C.J.; Qin, L.; Allaby, R.G.; Purugganan, M.D. Convergent evolution and parallelism in plant domestication revealed by an expanding archaeological record. Proc. Natl. Acad. Sci. USA 2014, 111, 6147–6152. [Google Scholar] [CrossRef] [PubMed] [Green Version]


© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kantar, M.B.; Hüber, S.; Herman, A.; Bock, D.G.; Baute, G.; Betts, K.; Ott, M.; Brandvain, Y.; Wyse, D.; Stupar, R.M.; et al. Neo-Domestication of an Interspecific Tetraploid Helianthus annuus × Helianthus tuberous Population That Segregates for Perennial Habit. Genes 2018, 9, 422. https://doi.org/10.3390/genes9090422
Kantar MB, Hüber S, Herman A, Bock DG, Baute G, Betts K, Ott M, Brandvain Y, Wyse D, Stupar RM, et al. Neo-Domestication of an Interspecific Tetraploid Helianthus annuus × Helianthus tuberous Population That Segregates for Perennial Habit. Genes. 2018; 9(9):422. https://doi.org/10.3390/genes9090422
Chicago/Turabian StyleKantar, Michael B., Sariel Hüber, Adam Herman, Dan G. Bock, Greg Baute, Kevin Betts, Matthew Ott, Yaniv Brandvain, Donald Wyse, Robert M. Stupar, and et al. 2018. "Neo-Domestication of an Interspecific Tetraploid Helianthus annuus × Helianthus tuberous Population That Segregates for Perennial Habit" Genes 9, no. 9: 422. https://doi.org/10.3390/genes9090422
APA StyleKantar, M. B., Hüber, S., Herman, A., Bock, D. G., Baute, G., Betts, K., Ott, M., Brandvain, Y., Wyse, D., Stupar, R. M., & Rieseberg, L. H. (2018). Neo-Domestication of an Interspecific Tetraploid Helianthus annuus × Helianthus tuberous Population That Segregates for Perennial Habit. Genes, 9(9), 422. https://doi.org/10.3390/genes9090422




