H1.0 Linker Histone as an Epigenetic Regulator of Cell Proliferation and Differentiation
Abstract
1. Introduction
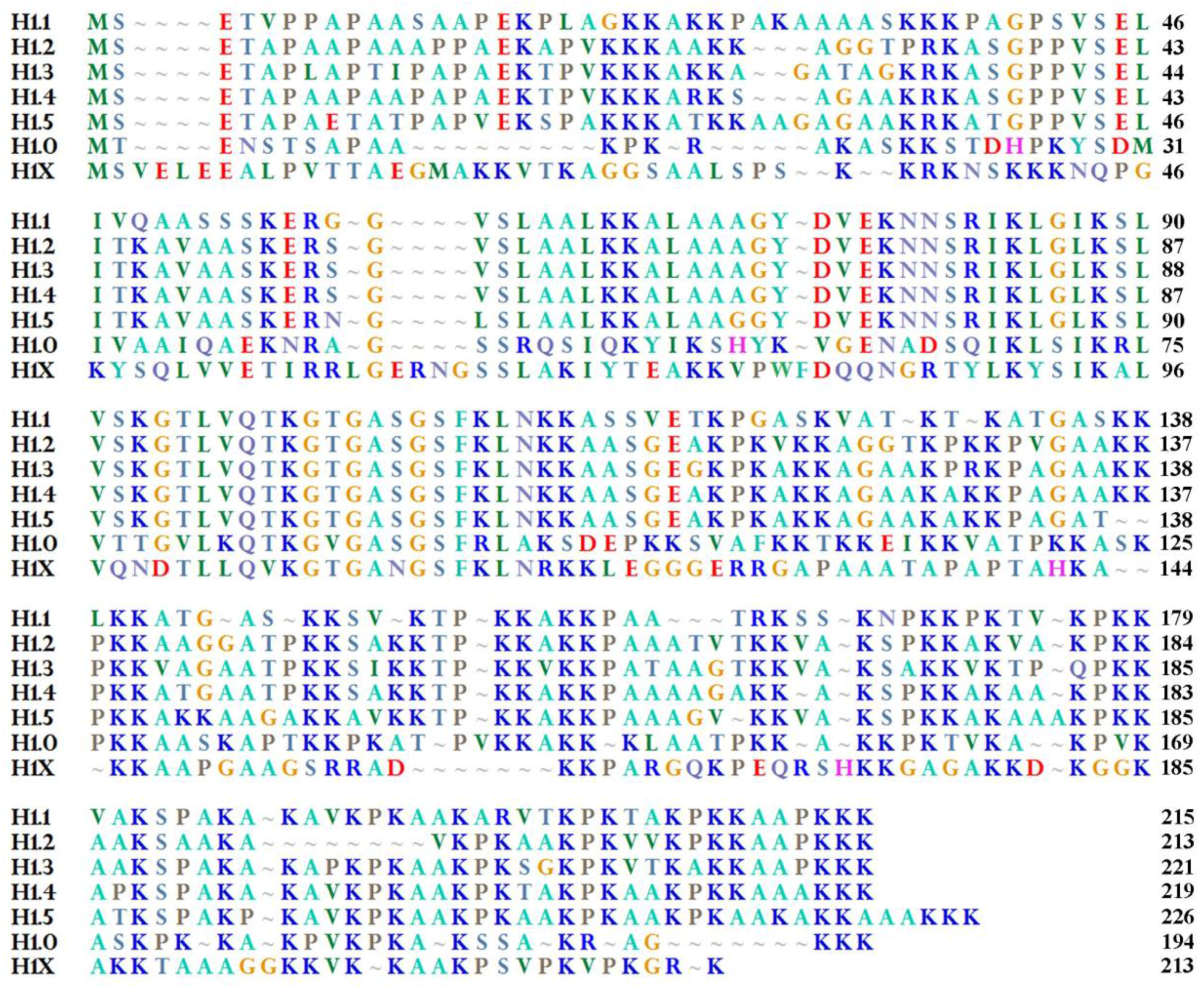
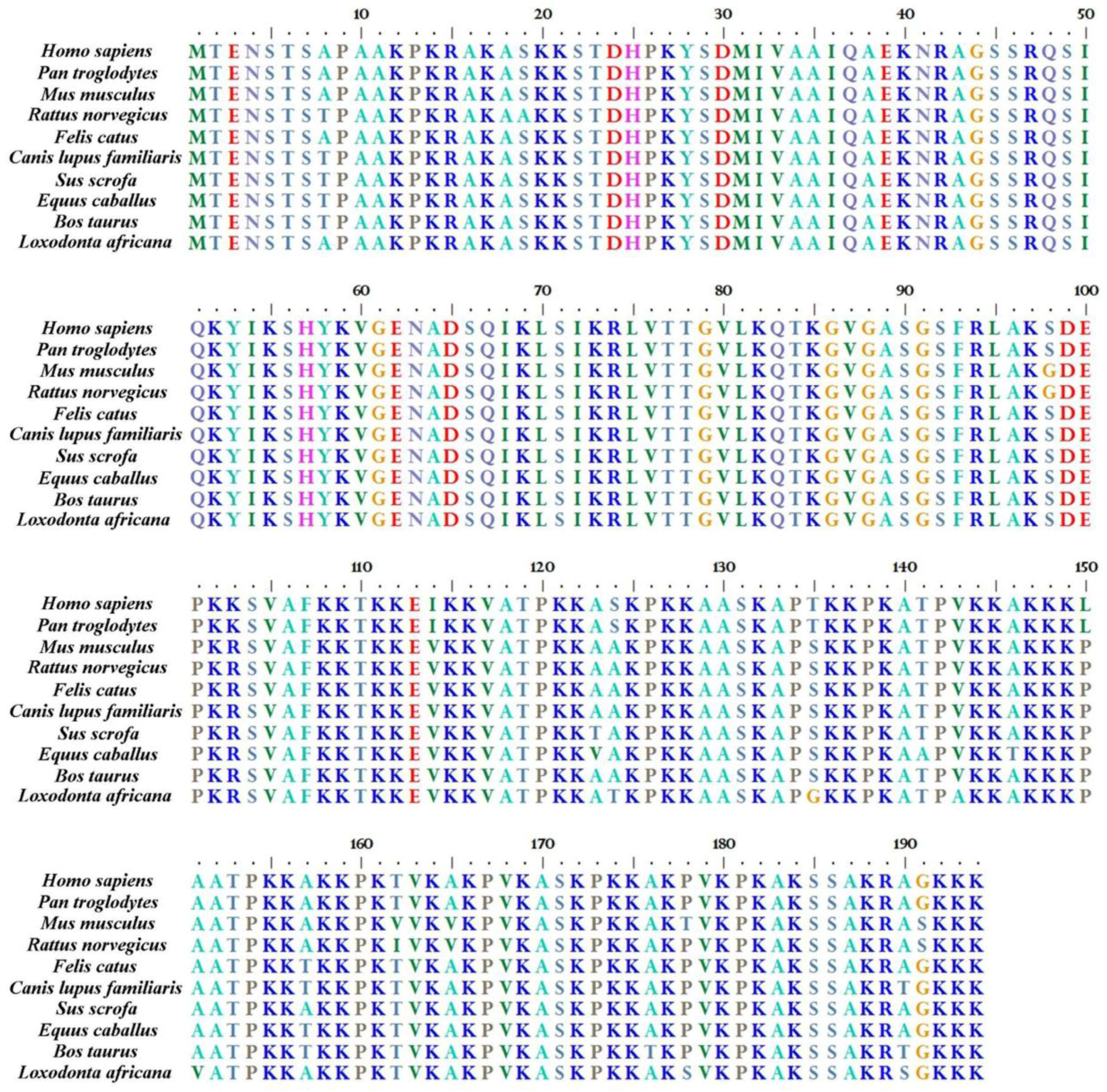
2. H1.0 Linker Histone in Mammals: Structural Peculiarities in Comparison with the Other Linker Histones and its Localization in Chromatin
3. H1.0 Expression in Development and Differentiation
4. H1.0 in Stem Cell Pluripotency Regulation and in Cancer
H1.0 and Pluripotency
5. H1.0 Protein and mRNA as Cargoes of Extracellular Vesicles
5.1 Sorting of H1.0 Protein and mRNA to EVs
5.2. H1.0 RNA as Carrier of Proteins
6. Conclusions and Perspectives
Acknowledgments
Conflicts of Interest
References
- Kornberg, R.D. Chromatin structure: a repeating unit of histones and DNA. Science 1974, 184, 868–871. [Google Scholar] [CrossRef] [PubMed]
- Kornberg, R.D.; Thomas, J.O. Chromatin structure; oligomers of the histones. Science 1974, 184, 865–868. [Google Scholar] [CrossRef] [PubMed]
- Olins, A.L.; Olins, D.E. Spheroid chromatin units (v Bodies). Science 1974, 183, 330–332. [Google Scholar] [CrossRef] [PubMed]
- McGhee, J.D.; Felsenfeld, G. Nucleosome structure. Annu. Rev. Biochem. 1980, 49, 1115–1156, Review. [Google Scholar] [CrossRef] [PubMed]
- Luger, K.; Mäder, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 1997, 389, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Noll, M.; Kornberg, R.D. Action of micrococcal nuclease on chromatin and the location of histone H1. J. Mol. Biol. 1977, 109, 393–404. [Google Scholar] [CrossRef]
- Simpson, R.T. Structure of the chromatosome, a chromatin particle containing 160 base pairs of DNA and all the histones. Biochemistry 1978, 17, 5524–5531. [Google Scholar] [CrossRef] [PubMed]
- Thoma, F.; Koller, T.; Klug, A. Involvement of histone H1 in the organization of the nucleosome and of the salt-dependent superstructures of chromatin. J. Cell. Biol. 1979, 83, 403–427. [Google Scholar] [CrossRef] [PubMed]
- Allan, J.; Hartman, P.G.; Crane-Robinson, C.; Aviles, F.X. The structure of histone H1 and its location in chromatin. Nature 1980, 288, 675–679. [Google Scholar] [CrossRef] [PubMed]
- McGhee, J.D.; Rau, D.C.; Charney, E.; Felsenfeld, G. Orientation of the nucleosome within the higher order structure of chromatin. Cell 1980, 22, 87–96. [Google Scholar]
- Izzo, A.; Schneider, R. H1 gets the genome in shape. Genome Biol. 2016, 17, 8. [Google Scholar] [CrossRef] [PubMed]
- Fyodorov, D.V.; Zhou, B.R.; Skoultchi, A.I.; Bai, Y. Emerging roles of linker histones in regulating chromatin structure and function. Nat. Rev. Mol. Cell. Biol. 2018, 19, 192–206. [Google Scholar] [CrossRef] [PubMed]
- Visser, A.E.; Jaunin, F.; Fakan, S.; Aten, J.A. High resolution analysis of interphase. chromosome domains. J. Cell. Sci. 2000, 113, 2585–2593. [Google Scholar] [PubMed]
- Lieberman-Aiden, E.; van Berkum, N.L.; Williams, L.; Imakaev, M.; Ragoczy, T.; Telling, A.; Amit, I.; Lajoie, B.R.; Sabo, P.J.; Dorschner, M.O.; et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 2009, 326, 289–293. [Google Scholar] [CrossRef] [PubMed]
- Cremer, T.; Cremer, M. Chromosome territories. Cold Spring Harb. Perspect. Biol. 2010, 2, a003889. [Google Scholar] [CrossRef] [PubMed]
- Sexton, T.; Cavalli, G. The role of chromosome domains in shaping the functional genome. Cell. 2015, 160, 1049–1059. [Google Scholar] [CrossRef] [PubMed]
- Ciabrelli, F.; Cavalli, G. Chromatin-driven behavior of topologically associating domains. J. Mol. Biol. 2015, 427, 608–625. [Google Scholar] [CrossRef] [PubMed]
- Bannister, A.J.; Kouzarides, T. Regulation of chromatin by histone modifications. Cell. Res. 2011, 21, 381–395. [Google Scholar] [CrossRef] [PubMed]
- Bowman, G.D.; Poirier, M.G. Post-translational modifications of histones that influence nucleosome dynamics. Chem. Rev. 2015, 115, 2274–2295. [Google Scholar] [CrossRef] [PubMed]
- Piatti, P.; Zeilner, A.; Lusser, A. ATP-dependent chromatin remodeling factors and their roles in affecting nucleosome fiber composition. Int. J. Mol. Sci. 2011, 12, 6544–6565. [Google Scholar] [CrossRef] [PubMed]
- Becker, P.B.; Workman, J.L. Nucleosome remodeling and epigenetics. Cold Spring Harb. Perspect. Biol. 2013, 5, pii: a017905. [Google Scholar] [CrossRef]
- Volle, C.; Dalal, Y. Histone variants: the tricksters of the chromatin world. Curr. Opin. Genet. Dev. 2014, 25, 8–14. [Google Scholar] [CrossRef] [PubMed]
- Cheema, M.S.; Ausió, J. The Structural Determinants behind the epigenetic role of histone variants. Genes 2015, 6, 685–713. [Google Scholar] [CrossRef] [PubMed]
- Henikoff, S.; Smith, M.M. Histone variants and epigenetics. Cold Spring Harb. Perspect. Biol. 2015, 7, a019364. [Google Scholar] [CrossRef] [PubMed]
- Strahl, B.D.; Allis, C.D. The language of covalent histone modifications. Nature 2000, 403, 41–45. [Google Scholar] [CrossRef] [PubMed]
- Torres, I.O.; Fujimori, D.G. Functional coupling between writers, erasers and readers of histone and DNA methylation. Curr. Opin. Struct. Biol. 2015, 35, 68–75. [Google Scholar] [CrossRef] [PubMed]
- Koopmans, W.J.A.; Buning, R.; Schmidt, T.; van Noort, J. spFRET using alternating excitation and FCS reveals progressive DNA unwrapping in nucleosomes. Biophys. J. 2009, 97, 195–204. [Google Scholar] [CrossRef] [PubMed]
- Bernier, M.; Luo, Y.; Nwokelo, K.C.; Goodwin, M.; Dreher, S.J.; Zhang, P.; Parthun, M.R.; Fondufe-Mittendorf, Y.; Ottesen, J.J.; Poirier, M.G. Linker histone H1 and H3K56 acetylation are antagonistic regulators of nucleosome dynamics. Nat. Commun. 2015, 6, 10152. [Google Scholar] [CrossRef] [PubMed]
- Marzluff, W.F.; Gongidi, P.; Woods, K.R.; Jin, J.; Maltais, L.J. The human and mouse replication-dependent histone genes. Genomics 2002, 80, 487–498. [Google Scholar] [CrossRef] [PubMed]
- Pehrson, J.R.; Cole, R.D. Histone H1 subfractions and H10 turnover at different rates in nondividing cells. Biochemistry 1982, 21, 456–460. [Google Scholar] [CrossRef] [PubMed]
- Thiriet, C.; Hayes, J.J. Replication-independent core histone dynamics at transcriptionally active loci in vivo. Genes Dev. 2005, 19, 677–682. [Google Scholar] [CrossRef] [PubMed]
- Ponte, I.; Romero, D.; Yero, D.; Suau, P.; Roque, A. Complex evolutionary history of the mammalian histone H1.1-H1.5 Gene Family. Mol. Biol. Evol. 2017, 34, 545–558. [Google Scholar] [CrossRef] [PubMed]
- Pan, C.; Fan, Y. Role of H1 linker histones in mammalian development and stem cell differentiation. Biochim. Biophys. Acta 2016, 1859, 496–509. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.F.; Sirotkin, A.M.; Buchold, G.M.; Skoultchi, A.I.; Marzluff, W.F. The mouse histone H1 genes: gene organization and differential regulation. J. Mol. Biol. 1997, 271, 124–138. [Google Scholar] [CrossRef] [PubMed]
- Khochbin, S. Histone H1 diversity: bridging regulatory signals to linker histone function. Gene 2001, 271, 1–12. [Google Scholar] [CrossRef]
- Godde, J.S.; Ura, K. Dynamic alterations of linker histone variants during development. Int. J. Dev. Biol. 2009, 53, 215–224. [Google Scholar] [CrossRef] [PubMed]
- George, E.M.; Izard, T.; Anderson, S.D.; Brown, D.T. Nucleosome interaction surface of linker histone H1c is distinct from that of H1(0). J. Biol. Chem. 2010, 285, 20891–20896. [Google Scholar] [CrossRef] [PubMed]
- Orrego, M.; Ponte, I.; Roque, A.; Buschati, N.; Mora, X.; Suau, P. Differential affinity of mammalian histone H1 somatic subtypes for DNA and chromatin. BMC Biol. 2007, 5, 22. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.R.; Feng, H.; Ghirlando, R.; Li, S.; Schwieters, C.D.; Bai, Y. A small number of residues can determine if linker histones are bound on or off dyad in the chromatosome. J. Mol. Biol. 2016, 428, 3948–3959. [Google Scholar] [CrossRef] [PubMed]
- Clausell, J.; Happel, N.; Hale, T.K.; Doenecke, D.; Beato, M. Histone H1 subtypes differentially modulate chromatin condensation without preventing ATP-dependent remodeling by SWI/SNF or NURF. PLoS ONE 2009, 4, e0007243. [Google Scholar] [CrossRef] [PubMed]
- Mayor, R.; Izquierdo-Bouldstridge, A.; Millán-Ariño, L.; Bustillos, A.; Sampaio, C.; Luque, N.; Jordan, A. Genome distribution of replication-independent histone H1 variants shows H1.0 associated with nucleolar domains and H1X associated with RNA polymerase II-enriched regions. J. Biol. Chem. 2015, 290, 7474–7491. [Google Scholar] [CrossRef] [PubMed]
- Happel, N.; Doenecke, D. Histone H1 and its isoforms: contribution to chromatin structure and function. Gene 2009, 431, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Vicent, G.P.; Wright, R.H.; Beato, M. Linker histones in hormonal gene regulation. Biochim. Biophys. Acta 2016, 1859, 520–525. [Google Scholar] [CrossRef] [PubMed]
- Bhan, S.; May, W.; Warren, S.L.; Sittman, D.B. Global gene expression analysis reveals specific and redundant roles for H1 variants, H1c and H1(0), in gene expression regulation. Gene 2008, 414, 10–18. [Google Scholar] [CrossRef] [PubMed]
- Allan, J.; Mitchell, T.; Harborne, N.; Bohm, L.; Crane-Robinson, C. Roles of H1 domains in determining higher order chromatin structure and H1 location. J. Mol. Biol. 1986, 187, 591–601. [Google Scholar] [CrossRef]
- Vyas, P.; Brown, D.T. N- and C-terminal domains determine differential nucleosomal binding geometry and affinity of linker histone isotypes H1(0) and H1c. J. Biol. Chem. 2012, 287, 11778–11787. [Google Scholar] [CrossRef] [PubMed]
- Roque, A.; Iloro, I.; Ponte, I.; Arrondo, J.L.; Suau, P. DNA-induced secondary structure of the carboxyl-terminal domain of histone H1. J. Biol. Chem. 2005, 280, 32141–32147. [Google Scholar] [CrossRef] [PubMed]
- Caterino, T.L.; Fang, H.; Hayes, J.J. Nucleosome linker DNA contacts and induces specific folding of the intrinsically disordered H1 carboxyl-terminal domain. Mol. Cell. Biol. 2011, 11, 2341–2348. [Google Scholar] [CrossRef] [PubMed]
- Roque, A.; Teruel, N.; López, R.; Ponte, I.; Suau, P. Contribution of hydrophobic interactions to the folding and fibrillation of histone H1 and its carboxy-terminal domain. J. Struct. Biol. 2012, 180, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Bolton, S.J.; Russelakis-Carneiro, M.; Betmouni, S.; Perry, V.H. Non-nuclear histone H1 is upregulated in neurones and astrocytes in prion and Alzheimer’s diseases but not in acute neurodegeneration. Neuropathol. Appl. Neurobiol. 1999, 25, 425–432. [Google Scholar] [CrossRef]
- Duce, J.A.; Smith, D.P.; Blake, R.E.; Crouch, P.J.; Li, Q.X.; Masters, C.L.; Trounce, I.A. Linker histone H1 binds to disease associated amyloid-like fibrils. J. Mol. Biol. 2006, 361, 493–505. [Google Scholar] [CrossRef] [PubMed]
- Roque, A.; Sortino, R.; Ventura, S.; Ponte, I.; Suau, P. Histone H1 favors folding and parallel fibrillar aggregation of the 1–42 Amyloid-β Peptide. Langmuir 2015, 31, 6782–6790. [Google Scholar] [CrossRef] [PubMed]
- Strom, A.; Wang, G.S.; Picketts, D.J.; Reimer, R.; Stuke, A.W.; Scott, F.W. Cellular prion protein localizes to the nucleus of endocrine and neuronal cells and interacts with structural chromatin components. Eur. J. Cell. Biol. 2011, 90, 414–419. [Google Scholar] [CrossRef] [PubMed]
- Zlatanova, J.S.; Srebreva, L.N.; Banchev, T.B.; Tasheva, B.T.; Tsanev, R.G. Cytoplasmic pool of histone H1 in mammalian cells. J. Cell. Sci. 1990, 96, 461–468. [Google Scholar] [PubMed]
- Ye, X.; Feng, C.; Gao, T.; Mu, G.; Zhu, W.; Yang, Y. Linker Histone in Diseases. Int. J. Biol. Sci. 2017, 13, 1008–1018. [Google Scholar] [CrossRef] [PubMed]
- Kowalski, A. Nuclear and nucleolar activity of linker histone variant H1.0. Cell. Mol. Biol. Lett. 2016, 21, 15. [Google Scholar] [CrossRef] [PubMed]
- Hansen, J.C.; Lu, X.; Ross, E.D.; Woody, R.W. Intrinsic protein disorder, amino acid composition, and histone terminal domains. J. Biol. Chem. 2006, 281, 1853–1856. [Google Scholar] [CrossRef] [PubMed]
- Roque, A.; Ponte, I.; Suau, P. Post-translational modifications of the intrinsically disordered terminal domains of histone H1: effects on secondary structure and chromatin dynamics. Chromosoma 2017, 126, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Sarg, B.; Helliger, W.; Talasz, H.; Forg, B.; Lindner, H.H. Histone H1 phosphorylation occurs site-specifically during interphase and mitosis: identification of a novel phosphorylation site on histone H1. J. Biol. Chem. 2006, 281, 6573–6580. [Google Scholar] [CrossRef] [PubMed]
- Miki, B.L.; Neelin, J.M. Comparison of the histones from fish erythrocytes. Can. J. Biochem. 1977, 55, 1220–1227. [Google Scholar] [CrossRef] [PubMed]
- Rutledge, R.G.; Shay, C.E.; Brown, G.L.; Neelin, J.M. The similarity of histones from turtle erythrocytes and liver. Can. J. Biochem. 1981, 59, 273–279. [Google Scholar] [CrossRef] [PubMed]
- Moorman, A.F.; de Boer, P.A.; Smit-Vis, J.H.; Lamers, W.H.; Charles, R. Immunological evidence for an H1(0) type of histone protein in chicken liver. Differentiation. 1986, 32, 44–48. [Google Scholar] [CrossRef] [PubMed]
- Srebreva, L.; Zlatanova, J. Occurrence of histone H10-related protein fraction in trout liver. Biochim. Biophys. Acta 1983, 740, 163–168. [Google Scholar] [CrossRef]
- Osley, M.A. The regulation of histone synthesis in the cell cycle. Annu. Rev. Biochem. 1991, 60, 827–861. [Google Scholar] [CrossRef] [PubMed]
- Castiglia, D.; Gristina, R.; Scaturro, M.; Di Liegro, I. Cloning and analysis of cDNA for rat histone H1(0). Nucleic Acids Res. 1993, 21, 1674. [Google Scholar] [CrossRef] [PubMed]
- Millán-Ariño, L.; Izquierdo-Bouldstridge, A.; Jordan, A. Specificities and genomic distribution of somatic mammalian histone H1 subtypes. Biochim. Biophys. Acta. 2016, 1859, 510–519. [Google Scholar] [CrossRef] [PubMed]
- Kalashnikova, A.A.; Winkler, D.D.; McBryant, S.J.; Henderson, R.K.; Herman, J.A.; DeLuca, J.G.; Luger, K.; Prenni, J.E.; Hansen, J.C. Linker histone H1.0 interacts with an extensive network of proteins found in the nucleolus. Nucleic Acids Res. 2013, 41, 4026–4035. [Google Scholar] [CrossRef] [PubMed]
- Szerlong, H.J.; Herman, J.A.; Krause, C.M.; DeLuca, J.G.; Skoultchi, A.; Winger, Q.A.; Prenni, J.E.; Hansen, J.C. Proteomic characterization of the nucleolar linker histone H1 interaction network. J. Mol. Biol. 2015, 427, 2056–2071. [Google Scholar] [CrossRef] [PubMed]
- Okuwaki, M.M.; Abe, M.; Hisaoka, M.; Nagata, K. Regulation of cellular dynamics and chromosomal binding site preference of linker histones H1.0 and H1.X. Mol. Cell. Biol. 2016, 36, 2681–2696. [Google Scholar] [CrossRef] [PubMed]
- Scaturro, M.; Nastasi, T.; Raimondi, L.; Bellafiore, M.; Cestelli, A.; Di Liegro, I. H1(0) RNA-binding proteins specifically expressed in the rat brain. J. Biol. Chem. 1998, 273, 22788–22791. [Google Scholar] [CrossRef] [PubMed]
- Nastasi, T.; Scaturro, M.; Bellafiore, M.; Raimondi, L.; Beccari, S.; Cestelli, A.; Di Liegro, I. PIPPin is a brain-specific protein that contains a cold-shock domain and binds specifically to H1 degrees and H3.3 mRNAs. J. Biol. Chem. 1999, 274, 24087–24093. [Google Scholar] [CrossRef] [PubMed]
- Schiera, G.; Di Liegro, C.M.; Puleo, V.; Colletta, O.; Fricano, A.; Cancemi, P.; Di Cara, G.; Di Liegro, I. Extracellular vesicles shed by melanoma cells contain a modified form of H1.0 linker histone and H1.0 mRNA-binding proteins. Int. J. Oncol. 2016, 49, 1807–1814. [Google Scholar] [CrossRef] [PubMed]
- Di Liegro, C.M.; Schiera, G.; Di Liegro, I. Extracellular vesicle-associated RNA as a carrier of epigenetic information. Genes (Basel) 2017, 8, pii: E240. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl. Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Clarke, H.J.; Bustin, M.; Oblin, C. Chromatin modifications during oogenesis in the mouse: removal of somatic subtypes of histone H1 from oocyte chromatin occurs post-natally through a post-transcriptional mechanism. J. Cell. Sci. 1997, 110, 477–487. [Google Scholar] [PubMed]
- Sirotkin, A.M.; Edelmann, W.; Cheng, G.; Klein-Szanto, A.; Kucherlapati, R.; Skoultchi, A.I. Mice develop normally without the H1(0) linker histone. Proc. Natl. Acad. Sci. USA. 1995, 92, 6434–6438. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Sirotkin, A.; Russell, R.G.; Ayala, J.; Skoultchi, A.I. Individual somatic H1 subtypes are dispensable for mouse development even in mice lacking the H1(0) replacement subtype. Mol. Cell. Biol. 2001, 21, 7933–7943. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Nikitina, T.; Morin-Kensicki, E.M.; Zhao, J.; Magnuson, T.R.; Woodcock, C.L.; Skoultchi, A.I. H1 linker histones are essential for mouse development and affect nucleosome spacing in vivo. Mol. Cell. Biol. 2003, 23, 4559–4572. [Google Scholar] [CrossRef] [PubMed]
- Alami, R.; Fan, Y.; Pack, S.; Sonbuchner, T.M.; Besse, A.; Lin, Q.; Greally, J.M.; Skoultchi, A.I.; Bouhassira, E.E. Mammalian linker-histone subtypes differentially affect gene expression in vivo. Proc. Natl. Acad. Sci. USA 2003, 100, 5920–5925. [Google Scholar] [CrossRef] [PubMed]
- Sancho, M.; Diani, E.; Beato, M.; Jordan, A. Depletion of human histone H1 variants uncovers specific roles in gene expression and cell growth. PLoS Genet. 2008, 4, e1000227. [Google Scholar] [CrossRef] [PubMed]
- Gjerset, R.; Gorka, C.; Hasthorpe, S.; Lawrence, J.J.; Eisen, H. Developmental and hormonal regulation of protein H1 degrees in rodents. Proc. Natl. Acad. Sci. U.S.A. 1982, 79, 2333–2337. [Google Scholar] [CrossRef] [PubMed]
- Lennox, R.W.; Cohen, L.H. The histone H1 complements of dividing and nondividing cells of the mouse. J. Biol. Chem. 1983, 258, 262–268. [Google Scholar] [PubMed]
- Piña, B.; Martinez, P.; Simon, L.; Suau, P. Differential kinetics of histone H1(0) accumulation in neuronal and glial cells from rat cerebral cortex during postnatal development. Biochem. Biophys. Res. Commun. 1984, 123, 697–702. [Google Scholar] [CrossRef]
- Di Liegro, I.; Cestelli, A. The relative proportion of H1(0) and A24 is reversed in oligodendrocytes during rat brain development. Cell. Mol. Neurobiol. 1990, 10, 267–274. [Google Scholar] [CrossRef] [PubMed]
- García-Segura, L.M.; Luquín, S.; Martínez, P.; Casas, M.T.; Suau, P. Differential expression and gonadal hormone regulation of histone H1(0) in the developing and adult rat brain. Brain Res. Dev. Brain Res. 1993, 73, 63–70. [Google Scholar] [CrossRef]
- Zlatanova, J.; Doenecke, D. Histone H1 zero: a major player in cell differentiation? FASEB J. 1994, 8, 1260–1268. [Google Scholar] [CrossRef] [PubMed]
- Gabrilovich, D.I.; Cheng, P.; Fan, Y.; Yu, B.; Nikitina, E.; Sirotkin, A.; Shurin, M.; Oyama, T.; Adachi, Y.; Nadaf, S.; et al. H1(0) histone and differentiation of dendritic cells. A molecular target for tumor-derived factors. J. Leukoc. Biol. 2002, 72, 285–296. [Google Scholar] [PubMed]
- Popova, E.Y.; Grigoryev, S.A.; Fan, Y.; Skoultchi, A.I.; Zhang, S.S.; Barnstable, C.J. Developmentally regulated linker histone H1c promotes heterochromatin condensation and mediates structural integrity of rod photoreceptors in mouse retina. J. Biol. Chem. 2013, 288, 17895–17907. [Google Scholar] [CrossRef] [PubMed]
- Domínguez, V.; Piña, B.; Suau, P. Histone H1 subtype synthesis in neurons and neuroblasts. Development. 1992, 115, 181–185. [Google Scholar] [PubMed]
- Castiglia, D.; Cestelli, A.; Scaturro, M.; Nastasi, T.; Di Liegro, I. H1(0) and H3.3B mRNA levels in developing rat brain. Neurochem. Res. 1994, 19, 1531–1537. [Google Scholar] [CrossRef] [PubMed]
- García-Iglesias, M.J.; Ramirez, A.; Monzo, M.; Steuer, B.; Martínez, J.M.; Jorcano, J.L.; Alonso, A. Specific expression in adult mice and post-implantation embryos of a transgene carrying the histone H1(0) regulatory region. Differentiation 1993, 55, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Miguel-Hidalgo, J.J.; Puckett Robinson, C. Histone H1(o) expression in the developing cat retina. Brain Res. Dev. Brain Res. 1999, 117, 39–45. [Google Scholar] [CrossRef]
- Khochbin, S.; Wolffe, A.P. Developmental regulation and butyrate-inducible transcription of the Xenopus histone H1(0) promoter. Gene 1993, 128, 173–180. [Google Scholar] [CrossRef]
- Bouterfa, H.L.; Piedrafita, F.J.; Doenecke, D.; Pfahl, M. Regulation of H1(0) gene expression by nuclear receptors through an unusual response element: implications for regulation of cell proliferation. DNA Cell. Biol. 1995, 14, 909–919. [Google Scholar] [CrossRef] [PubMed]
- Lemercier, C.; Duncliffe, K.; Boibessot, I.; Zhang, H.; Verdel, A.; Angelov, D.; Khochbin, S. Involvement of retinoblastoma protein and HBP1 in histone H1(0) gene expression. Mol. Cell. Biol. 2000, 20, 6627–6637. [Google Scholar] [CrossRef] [PubMed]
- Duncliffe, K.N.; Rondahl, M.E.; Wells, J.R. A H1 histone gene-specific AC-box-related element influences transcription from a major chicken H1 promoter. Gene 1995, 163, 227–232. [Google Scholar] [CrossRef]
- Ramsey-Ewing, A.; Van Wijnen, A.J.; Stein, G.S.; Stein, J.L. Delineation of a human histone H4 cell cycle element in vivo: the master switch for H4 gene transcription. Proc. Natl. Acad. Sci. U.S.A. 1994, 91, 4475–4479. [Google Scholar] [CrossRef] [PubMed]
- Peretti, M.; Khochbin, S. The evolution of the differentiation-specific histone H1 gene basal promoter. J. Mol. Evol. 1997, 44, 128–134. [Google Scholar] [CrossRef] [PubMed]
- Bauer-Hofmann, R.; Alonso, A. Thyroid hormone receptors bind to the promoter of the mouse histone H10 gene and modulate its transcription. Nucleic Acids Res. 1995, 23, 5034–5040. [Google Scholar] [CrossRef] [PubMed]
- Azuara, V.; Perry, P.; Sauer, S.; Spivakov, M.; Jørgensen, H.F.; John, R.M.; Gouti, M.; Casanova, M.; Warnes, G.; Merkenschlager, M.; et al. Chromatin signatures of pluripotent cell lines. Nat. Cell. Biol. 2006, 8, 532–538. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, B.E.; Mikkelsen, T.S.; Xie, X.; Kamal, M.; Huebert, D.J.; Cuff, J.; Fry, B.; Meissner, A.; Wernig, M.; Plath, K.; et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell. 2006, 125, 315–326. [Google Scholar] [CrossRef] [PubMed]
- Terme, J.M.; Sese, B.; Millan-Arino, L.; Mayor, R.; Izpisua Belmonte, J.C.; Barrero, M.J.; Jordan, A. Histone H1 variants are differentially expressed and incorporated into chromatin during differentiation and reprogramming to pluripotency. J. Biol. Chem. 2011, 286, 35347–35357. [Google Scholar] [CrossRef] [PubMed]
- Panyim, S.; Chalkley, R. A new histone found only in mammalian tissues with little cell division. Biochem. Biophys. Res. Commun. 1969, 37, 1042–1049. [Google Scholar] [CrossRef]
- Seyedin, S.M.; Kistler, W.S. Levels of chromosomal protein high mobility group 2 parallel the proliferative activity of testis, skeletal muscle, and other organs. J. Biol. Chem. 1979, 254, 11264–112671. [Google Scholar] [PubMed]
- Benjamin, W.B. Selective in vitro methylation of rat chromatin associated histone after partial hepatectomy. Nat. New Biol. 1971, 234, 18–20. [Google Scholar] [CrossRef] [PubMed]
- Marsh, W.H.; Fitzgerald, P.J. Pancreas acinar cell regeneration. 13. Histone synthesis and modification. Fed. Proc. 1973, 32, 2119–2125. [Google Scholar] [PubMed]
- Gorka, C.; Lawrence, J.J.; Khochbin, S. Variation of H1(0) content throughout the cell cycle in regenerating rat liver. Exp. Cell. Res. 1995, 217, 528–533. [Google Scholar] [CrossRef] [PubMed]
- Happel, N.; Warneboldt, J.; Hänecke, K.; Haller, F.; Doenecke, D. H1 subtype expression during cell proliferation and growth arrest. Cell. Cycle 2009, 8, 2226–2232. [Google Scholar] [CrossRef] [PubMed]
- Scaturro, M.; Cestelli, A.; Castiglia, D.; Nastasi, T.; Di Liegro, I. Posttranscriptional regulation of H1 zero and H3.3B histone genes in differentiating rat cortical neurons. Neurochem. Res. 1995, 20, 969–976. [Google Scholar] [CrossRef] [PubMed]
- Cuisset, L.; Tichonicky, L.; Delpech, M. Quantitative analysis of histone H1 degrees protein synthesis in HTC cells. Eur. J. Biochem. 1999, 261, 593–599. [Google Scholar] [CrossRef] [PubMed]
- Cao, K.; Lailler, N.; Zhang, Y.; Kumar, A.; Uppal, K.; Liu, Z.; Lee, E.K.; Wu, H.; Medrzycki, M.; Pan, C.; et al. High-resolution mapping of H1 linker histone variants in embryonic stem cells. PLoS Genet. 2013, 9, e1003417. [Google Scholar] [CrossRef] [PubMed]
- Medrzycki, M.; Zhang, Y.; Cao, K.; Fan, Y. Expression analysis of mammalian linker-histone subtypes. J. Vis. Exp. 2012, pii: 3577. [Google Scholar] [CrossRef]
- Izzo, A.; Ziegler-Birling, C.; Hill, P.W.S.; Brondani, L.; Hajkova, P.; Torres-Padilla, M.E.; Schneider, R. Dynamic changes in H1 subtype composition during epigenetic reprogramming. J. Cell. Biol. 2017, pii: jcb.201611012. [Google Scholar] [CrossRef]
- Hajkova, P.; Ancelin, K.; Waldmann, T.; Lacoste, N.; Lange, U.C.; Cesari, F.; Lee, C.; Almouzni, G.; Schneider, R.; Surani, M.A. Chromatin dynamics during epigenetic reprogramming in the mouse germ line. Nature 2008, 452, 877–881. [Google Scholar] [CrossRef] [PubMed]
- Hajkova, P. Epigenetic reprogramming--taking a lesson from the embryo. Curr. Opin. Cell. Biol. 2010, 22, 342–350. [Google Scholar] [CrossRef] [PubMed]
- Keppel, F.; Allet, B.; Eisen, H. Appearance of a chromatin protein during the erythroid differentiation of Friend virus-transformed cells. Proc. Natl. Acad. Sci. U.S.A. 1977, 74, 653–656. [Google Scholar] [CrossRef] [PubMed]
- Rousseau, D.; Khochbin, S.; Gorka, C.; Lawrence, J.J. Induction of H1(0)-gene expression in B16 murine melanoma cells. Eur. J. Biochem. 1992, 208, 775–779. [Google Scholar] [CrossRef] [PubMed]
- Helliger, W.; Lindner, H.; Grübl-Knosp, O.; Puschendorf, B. Alteration in proportions of histone H1 variants during the differentiation of murine erythroleukaemic cells. Biochem. J. 1992, 288, 747–751. [Google Scholar] [CrossRef] [PubMed]
- Boix, J.; Ruiz-Castillo, A. Increased Histone H1° expression in differentiating mouse erythroleukemia cells is related to decreased cell proliferation. Exp. Cell. Res. 1992, 201, 531–534. [Google Scholar] [CrossRef]
- Yellajoshyula, D.; Brown, D.T. Global modulation of chromatin dynamics mediated by dephosphorylation of linker histone H1 is necessary for erythroid differentiation. Proc. Natl. Acad. Sci. U.S.A. 2006, 103, 18568–18573. [Google Scholar] [CrossRef] [PubMed]
- Laitinen, J.; Sistonen, L.; Alitalo, K.; Hölttä, E. Cell transformation by c-Ha-rasVal12 oncogene is accompanied by a decrease in histone H1 zero and an increase in nucleosomal repeat length. J. Cell. Biochem. 1995, 57, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.M.; Kim, B.J.; Norwood Toro, L.; Skoultchi, A.I. H1 linker histone promotes epigenetic silencing by regulating both DNA methylation and histone H3 methylation. Proc. Natl. Acad. Sci. U.S.A. 2013, 110, 1708–1713. [Google Scholar] [CrossRef] [PubMed]
- Scaffidi, P. Histone H1 alterations in cancer. Biochim. Biophys. Acta. 2016, 1859, 533–539. [Google Scholar] [CrossRef] [PubMed]
- Kostova, N.N.; Srebreva, L.N.; Milev, A.D.; Bogdanova, O.G.; Rundquist, I.; Lindner, H.H.; Markov, D.V. Immunohistochemical demonstration of histone H1(0) in human breast carcinoma. Histochem. Cell. Biol. 2005, 124, 435–443. [Google Scholar] [CrossRef] [PubMed]
- Medrzycki, M.; Zhang, Y.; McDonald, J.F.; Fan, Y. Profiling of linker histone variants in ovarian cancer. Front. Biosci. (Landmark Ed.) 2012, 17, 396–406. [Google Scholar] [CrossRef] [PubMed]
- Gabrovsky, N.; Georgieva, M.; Laleva, M.; Uzunov, K.; Miloshev, G. Histone H1.0--a potential molecular marker with prognostic value for patients with malignant gliomas. Acta Neurochir. (Wien.) 2013, 155, 1437–1442. [Google Scholar] [CrossRef] [PubMed]
- Orsi, G.A.; Naughtin, M.; Almouzni, G. The epigenome and cancer stem cell fate: Connected by a linker histone variant. Cell. Stem Cell. 2016, 19, 567–568. [Google Scholar] [CrossRef] [PubMed]
- Gambara, G.; Gaebler, M.; Keilholz, U.; Regenbrecht, C.R.A.; Silvestri, A. From chemotherapy to combined targeted therapeutics: in vitro and in vivo models to decipher intra-tumor heterogeneity. Front. Pharmacol. 2018, 9, 77. [Google Scholar] [CrossRef] [PubMed]
- Torres, C.M.; Biran, A.; Burney, M.J.; Patel, H.; Henser-Brownhill, T.; Cohen, A.S.; Li, Y.; Ben-Hamo, R.; Nye, E.; Spencer-Dene, B.; et al. The linker histone H1.0 generates epigenetic and functional intratumor heterogeneity. Science 2016, 353, pii: aaf1644. [Google Scholar] [CrossRef]
- Scaffidi, P.; Misteli, T. In vitro generation of human cells with cancer stem cell properties. Nat. Cell Biol. 2011, 13, 1051–1061. [Google Scholar] [CrossRef] [PubMed]
- Mashburn-Warren, L.M.; Whiteley, M. Special delivery: Vesicle trafficking in prokaryotes. Mol. Microbiol. 2006, 61, 839–846. [Google Scholar] [CrossRef] [PubMed]
- Tsatsaronis, J.A.; Franch-Arroyo, S.; Resch, U.; Charpentier, E. Extracellular vesicle RNA: A universal mediator of microbial communication? Trends Microbiol. 2018, 26, 401–410. [Google Scholar] [CrossRef] [PubMed]
- Schiera, G.; Di Liegro, C.M.; Di Liegro, I. Extracellular membrane vesicles as vehicles for brain cell-to-cell interactions in physiological as well as pathological conditions. Biomed. Res. Int. 2015, 152926. [Google Scholar] [CrossRef] [PubMed]
- Yáñez-Mó, M.; Siljander, P.R.; Andreu, Z.; Zavec, A.B.; Borràs, F.E.; Buzas, E.I.; Buzas, K.; Casal, E.; Cappello, F.; Carvalho, J.; et al. Biological properties of extracellular vesicles and their physiological functions. J. Extracell. Vesicles 2015, 4, 27066. [Google Scholar] [CrossRef] [PubMed]
- Mateescu, B.; Kowal, E.J.; van Balkom, B.W.; Bartel, S.; Bhattacharyya, S.N.; Buzás, E.I.; Buck, A.H.; de Candia, P.; Chow, F.W.; Das, S.; et al. Obstacles and opportunities in the functional analysis of extracellular vesicle RNA - an ISEV position paper. J. Extracell. Vesicles. 2017, 6, 1286095. [Google Scholar] [CrossRef] [PubMed]
- Cocucci, E.; Meldolesi, J. Ectosomes and exosomes: shedding the confusion between extracellular vesicles. Trends Cell. Biol. 2015, 25, 364–372. [Google Scholar] [CrossRef] [PubMed]
- Mineo, M.; Garfield, S.H.; Taverna, S.; Flugy, A.; De Leo, G.; Alessandro, R.; Kohn, E.C. Exosomes released by K562 chronic myeloid leukemia cells promote angiogenesis in a Src-dependent fashion. Angiogenesis 2012, 15, 33–45. [Google Scholar] [CrossRef] [PubMed]
- Azmi, A.S.; Bao, B.; Sarkar, F.H. Exosomes in cancer development, metastasis, and drug resistance: a comprehensive review. Cancer Metastasis Rev. 2013, 32, 623–642. [Google Scholar] [CrossRef] [PubMed]
- Raimondo, S.; Saieva, L.; Corrado, C.; Fontana, S.; Flugy, A.; Rizzo, A.; De Leo, G.; Alessandro, R. Chronic myeloid leukemia-derived exosomes promote tumor growth through an autocrine mechanism. Cell. Commun. Signal. 2015, 13, 8. [Google Scholar] [CrossRef] [PubMed]
- Schiera, G.; Di Liegro, C.M.; Saladino, P.; Pitti, R.; Savettieri, G.; Proia, P.; Di Liegro, I. Oligodendroglioma cells synthesize the differentiation-specific linker histone H1° and release it into the extracellular environment through shed vesicles. Int. J. Oncol. 2013, 43, 1771–1776. [Google Scholar] [CrossRef] [PubMed]
- Lo Cicero, A.; Schiera, G.; Proia, P.; Saladino, P.; Savettieri, G.; Di Liegro, C.M.; Di Liegro, I. Oligodendroglioma cells shed microvesicles which contain TRAIL as well as molecular chaperones and induce cell death in astrocytes. Int. J. Oncol. 2011, 39, 1353–1357. [Google Scholar] [CrossRef] [PubMed]
- Ostrowski, M.; Carmo, N.B.; Krumeich, S.; Fanget, I.; Raposo, G.; Savina, A.; Moita, C.F.; Schauer, K.; Hume, A.N.; Freitas, R.P.; et al. Rab27a and Rab27b control different steps of the exosome secretion pathway. Nat. Cell. Biol. 2010, 12, 19–30; sup pp 1–13. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.; Morohashi, Y.; Yoshimura, S.; Manrique-Hoyos, N.; Jung, S.; Lauterbach, M.A.; Bakhti, M.; Grønborg, M.; Möbius, W.; Rhee, J.; et al. Regulation of exosome secretion by Rab35 and its GTPase-activating proteins TBC1D10A-C. J. Cell. Biol. 2010, 189, 223–232. [Google Scholar] [CrossRef] [PubMed]
- Théry, C.; Ostrowski, M.; Segura, E. Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 2009, 9, 581–593. [Google Scholar] [CrossRef] [PubMed]
- Subra, C.; Grand, D.; Laulagnier, K.; Stella, A.; Lambeau, G.; Paillasse, M.; De Medina, P.; Monsarrat, B.; Perret, B.; Silvente-Poirot, S.; et al. Exosomes account for vesicle-mediated transcellular transport of activatable phospholipases and prostaglandins. J. Lipid Res. 2010, 51, 2105–2120. [Google Scholar] [CrossRef] [PubMed]
- Rashed, M.H.; Bayraktar, E.; Helal, G.K.; Abd-Ellah, M.F.; Amero, P.; Chavez-Reyes, A.; Rodriguez-Aguayo, C. Exosomes: From Garbage Bins to Promising Therapeutic Targets. Int. J. Mol. Sci. 2017, 18, pii: E538. [Google Scholar] [CrossRef] [PubMed]
- Vyas, N.; Dhawan, J. Exosomes: mobile platforms for targeted and synergistic signaling across cell boundaries. Cell. Mol. Life Sci. 2017, 74, 1567–1576. [Google Scholar] [CrossRef] [PubMed]
- Rajendran, L.; Honsho, M.; Zahn, T.R.; Keller, P.; Geiger, K.D.; Verkade, P.; Simons, K. Alzheimer’s disease β-amyloid peptides are released in association with exosomes. Proc. Natl. Acad. Sci. U.S.A. 2006, 103, 11172–11177. [Google Scholar] [CrossRef] [PubMed]
- Vella, L.J.; Sharples, R.A.; Nisbet, R.M.; Cappai, R.; Hill, A.F. The role of exosomes in the processing of proteins associated with neurodegenerative diseases. Eur. Biophys. J. 2008, 37, 323–332. [Google Scholar] [CrossRef] [PubMed]
- Emmanouilidou, E.; Melachroinou, K.; Roumeliotis, T.; Garbis, S.D.; Ntzouni, M.; Margaritis, L.H.; Stefanis, L.; Vekrellis, K. Cell-produced α-synuclein is secreted in a calcium-dependent manner by exosomes and impacts neuronal survival. J. Neurosci. 2010, 30, 6838–6851. [Google Scholar] [CrossRef] [PubMed]
- Saman, S.; Kim, W.; Raya, M.; Visnick, Y.; Miro, S.; Saman, S.; Jackson, B.; McKee, A.C.; Alvarez, V.E.; Lee, N.C.; et al. Exosome-associated tau is secreted in tauopathy models and is selectively phosphorylated in cerebrospinal fluid in early Alzheimer disease. J. Biol. Chem. 2012, 287, 3842–3849. [Google Scholar] [CrossRef] [PubMed]
- Record, M.; Carayon, K.; Poirot, M.; Silvente-Poirot, S. Exosomes as new vesicular lipid transporters involved in cell-cell communication and various pathophysiologies. Biochim. Biophys. Acta, 2014, 1841, 108–120. [Google Scholar] [CrossRef] [PubMed]
- Bieri, G.; Gitler, A.D.; Brahic, M. Internalization, axonal transport and release of fibrillar forms of α-synuclein. Neurobiol. Dis. 2017, pii: S0969–9961(17)30055–4. [Google Scholar] [CrossRef]
- Kawamura, Y.; Yamamoto, Y.; Sato, T.A.; Ochiya, T. Extracellular vesicles as trans-genomic agents: Emerging roles in disease and evolution. Cancer Sci. 2017, 108, 824–830. [Google Scholar] [CrossRef] [PubMed]
- Roque, A.; Ponte, I.; Suau, P. The subtype-specific role of histone H1.0 in cancer cell differentiation and intratumor heterogeneity. Transl. Cancer Res. 2017, 6 (Suppl. 2), S414–S417. [Google Scholar] [CrossRef]
- Castello, A.; Fischer, B.; Frese, C.K.; Horos, R.; Alleaume, A.M.; Foehr, S.; Curk, T.; Krijgsveld, J.; Hentze, M.W. Comprehensive identification of RNA-binding domains in human cells. Mol. Cell. 2016, 63, 696–710. [Google Scholar] [CrossRef] [PubMed]
- Varela-Eirin, M.; Varela-Vazquez, A.; Rodríguez-Candela Mateos, M.; Vila-Sanjurjo, A.; Fonseca, E.; Mascareñas, J.L.; Eugenio Vázquez, M.; Mayan, M.D. Recruitment of RNA molecules by connexin RNA-binding motifs: Implication in RNA and DNA transport through microvesicles and exosomes. Biochim. Biophys. Acta. 2017, 1864, 728–736. [Google Scholar] [CrossRef] [PubMed]
- Genz, C.; Fundakowski, J.; Hermesh, O.; Schmid, M.; Jansen, R.P. Association of the yeast RNA-binding protein She2p with the tubular endoplasmic reticulum depends on membrane curvature. J. Biol. Chem. 2013, 288, 32384–32393. [Google Scholar] [CrossRef] [PubMed]
- Holers, V.M.; Kotzin, B.L. Human peripheral blood monocytes display surface antigens recognized by monoclonal antinuclear antibodies. J. Clin. Invest. 1985, 76, 991–998. [Google Scholar] [CrossRef] [PubMed]
- Bolton, S.J.; Perry, V.H. Histone H1; a neuronal protein that binds bacterial lipopolysaccharide. J. Neurocytol. 1997, 26, 823–831. [Google Scholar] [CrossRef] [PubMed]
- Brix, K.; Summa, W.; Lottspeich, F.; Herzog, V. Extracellularly occurring histone H1 mediates the binding of thyroglobulin to the cell surface of mouse macrophages. J. Clin. Invest. 1998, 102, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Kunadt, M.; Eckermann, K.; Stuendl, A.; Gong, J.; Russo, B.; Strauss, K.; Rai, S.; Kügler, S.; Falomir Lockhart, L.; Schwalbe, M.; et al. Extracellular vesicle sorting of α-Synuclein is regulated by sumoylation. Acta Neuropathol. 2015, 129, 695–713. [Google Scholar] [CrossRef] [PubMed]
- Villarroya-Beltri, C.; Gutiérrez-Vázquez, C.; Sánchez-Cabo, F.; Pérez-Hernández, D.; Vázquez, J.; Martin-Cofreces, N.; Martinez-Herrera, D.J.; Pascual-Montano, A.; Mittelbrunn, M.; Sánchez-Madrid, F. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat. Commun. 2013, 4, 2980. [Google Scholar] [CrossRef] [PubMed]
- Khvorova, A.; Kwak, Y.G.; Tamkun, M.; Majerfeld, I.; Yarus, M. RNAs that bind and change the permeability of phospholipid membranes. Proc. Natl. Acad. Sci. U.S.A. 1999, 96, 10649–10654. [Google Scholar] [CrossRef] [PubMed]
- Vlassov, A.; Khvorova, A.; Yarus, M. Binding and disruption of phospholipid bilayers by supramolecular RNA complexes. Proc. Natl. Acad. Sci. U.S.A. 2001, 98, 7706–7711. [Google Scholar] [CrossRef] [PubMed]
- Janas, T.; Janas, T.; Yarus, M. Specific RNA binding to ordered phospholipid bilayers. Nucleic Acids Res. 2006, 34, 2128–21236. [Google Scholar] [CrossRef] [PubMed]
- Haas, S.; Steplewski, A.; Siracusa, L.D.; Amini, S.; Khalili, K. Identification of a sequence-specific single-stranded DNA binding protein that suppresses transcription of the mouse myelin basic protein gene. J. Biol. Chem. 1995, 270, 12503–12510. [Google Scholar] [CrossRef] [PubMed]
- Muralidharan, V.; Tretiakova, A.; Steplewski, A.; Haas, S.; Amini, S.; Johnson, E.; Khalili, K. Evidence for inhibition of MyEF-2 binding to MBP promoter by MEF-1/Pur α. J. Cell. Biochem. 1997, 66, 524–531. [Google Scholar] [CrossRef]
- van Riel, B.; Pakozdi, T.; Brouwer, R.; Monteiro, R.; Tuladhar, K.; Franke, V.; Bryne, J.C.; Jorna, R.; Rijkers, E.J.; van Ijcken, W.; et al. A novel complex, RUNX1-MYEF2, represses hematopoietic genes in erythroid cells. Mol. Cell. Biol. 2012, 32, 3814–3822. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Di Liegro, C.M.; Schiera, G.; Di Liegro, I. H1.0 Linker Histone as an Epigenetic Regulator of Cell Proliferation and Differentiation. Genes 2018, 9, 310. https://doi.org/10.3390/genes9060310
Di Liegro CM, Schiera G, Di Liegro I. H1.0 Linker Histone as an Epigenetic Regulator of Cell Proliferation and Differentiation. Genes. 2018; 9(6):310. https://doi.org/10.3390/genes9060310
Chicago/Turabian StyleDi Liegro, Carlo Maria, Gabriella Schiera, and Italia Di Liegro. 2018. "H1.0 Linker Histone as an Epigenetic Regulator of Cell Proliferation and Differentiation" Genes 9, no. 6: 310. https://doi.org/10.3390/genes9060310
APA StyleDi Liegro, C. M., Schiera, G., & Di Liegro, I. (2018). H1.0 Linker Histone as an Epigenetic Regulator of Cell Proliferation and Differentiation. Genes, 9(6), 310. https://doi.org/10.3390/genes9060310





