Mapping Grain Iron and Zinc Content Quantitative Trait Loci in an Iniadi-Derived Immortal Population of Pearl Millet
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. DNA Extraction and Marker Analysis
2.3. Construction of Linkage Map
2.4. Experimental Conditions and Phenotyping of the Population
2.5. Statistical Analysis
2.6. Quantitative Trait Loci Detection
3. Results and Discussion
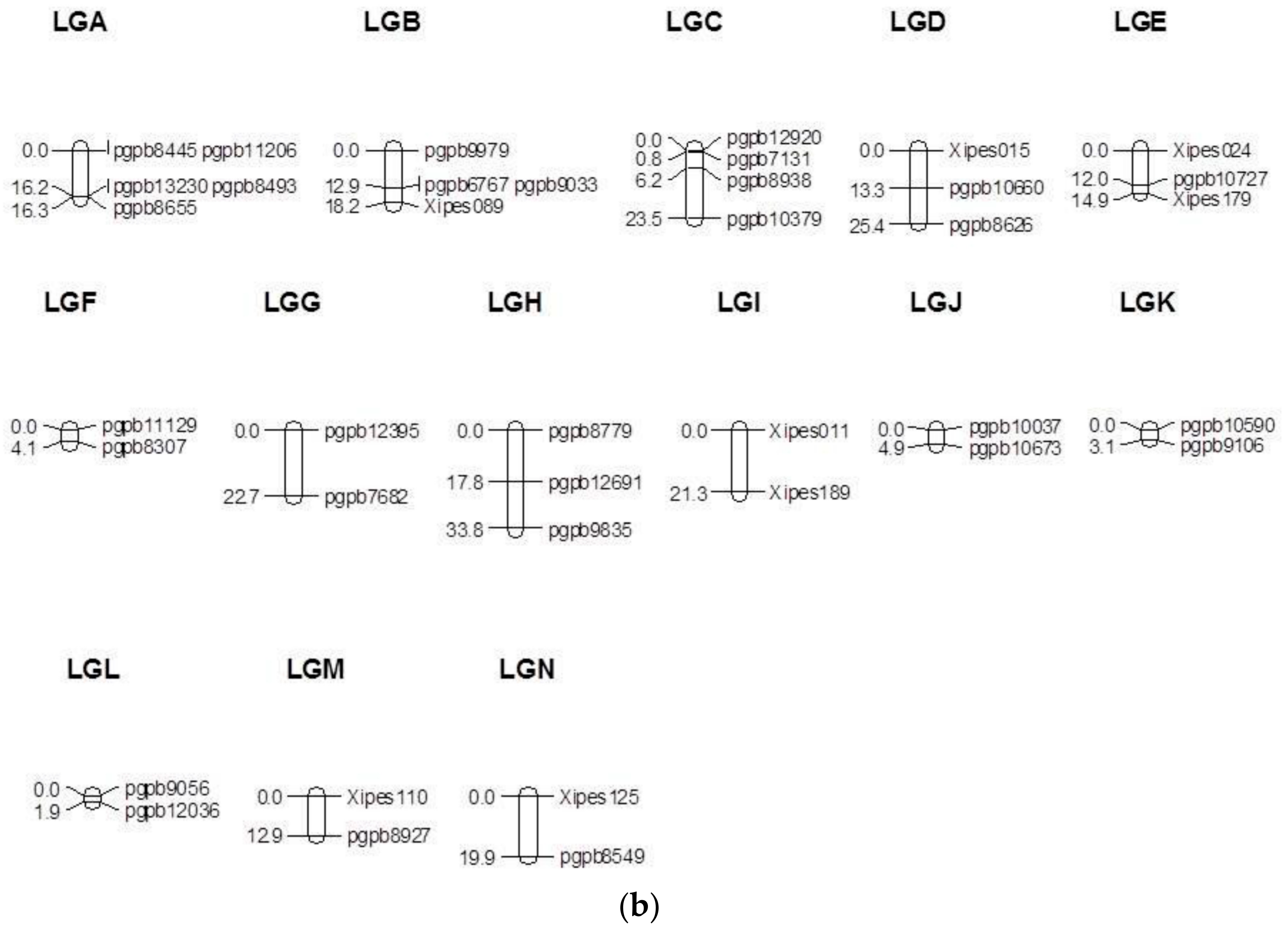
3.1. Linkage Map
3.2. Segregation Distortion and Inter-Marker Gaps
3.3. Variance Components and Heritability
3.4. Correlation Analysis
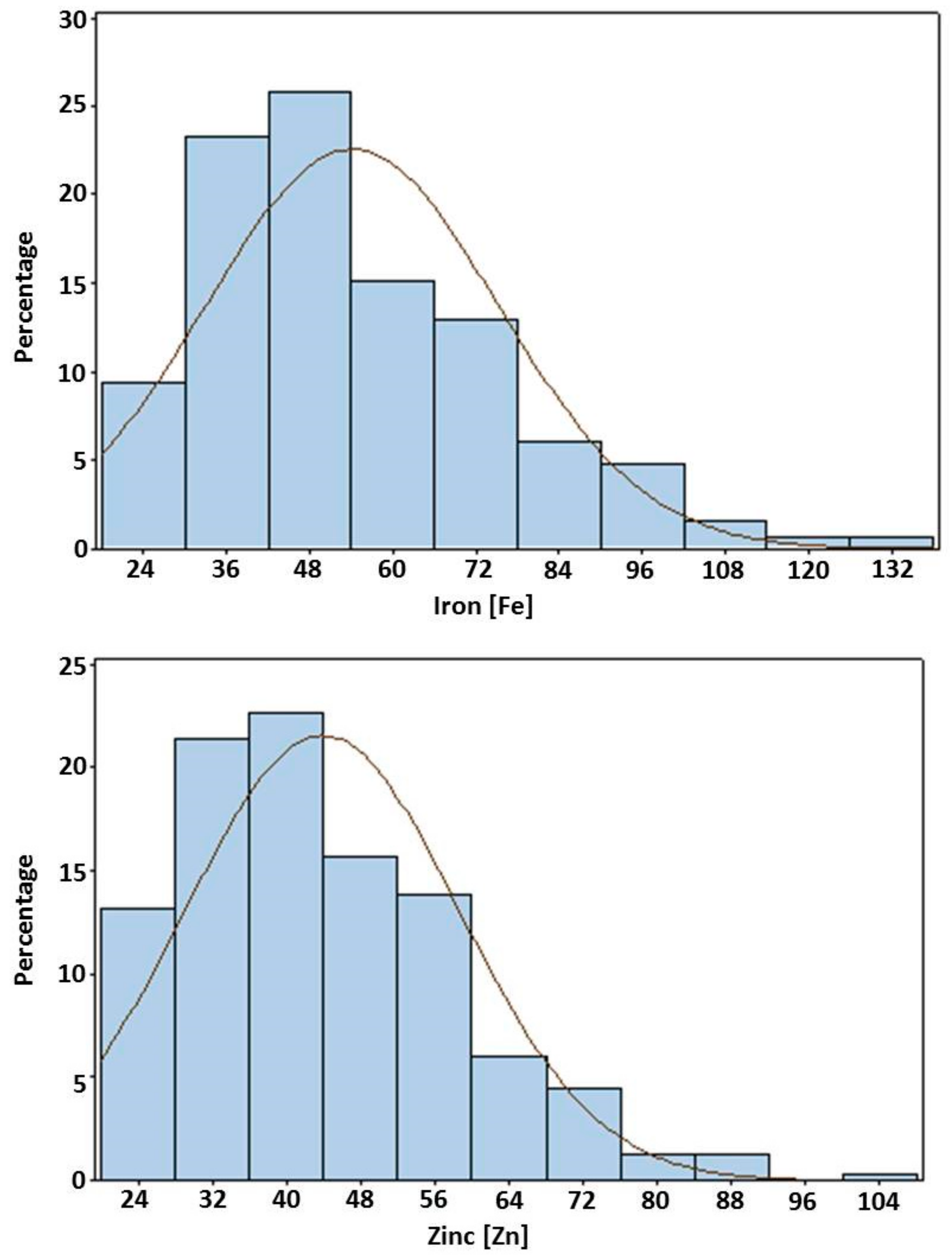
3.5. Frequency Distributions
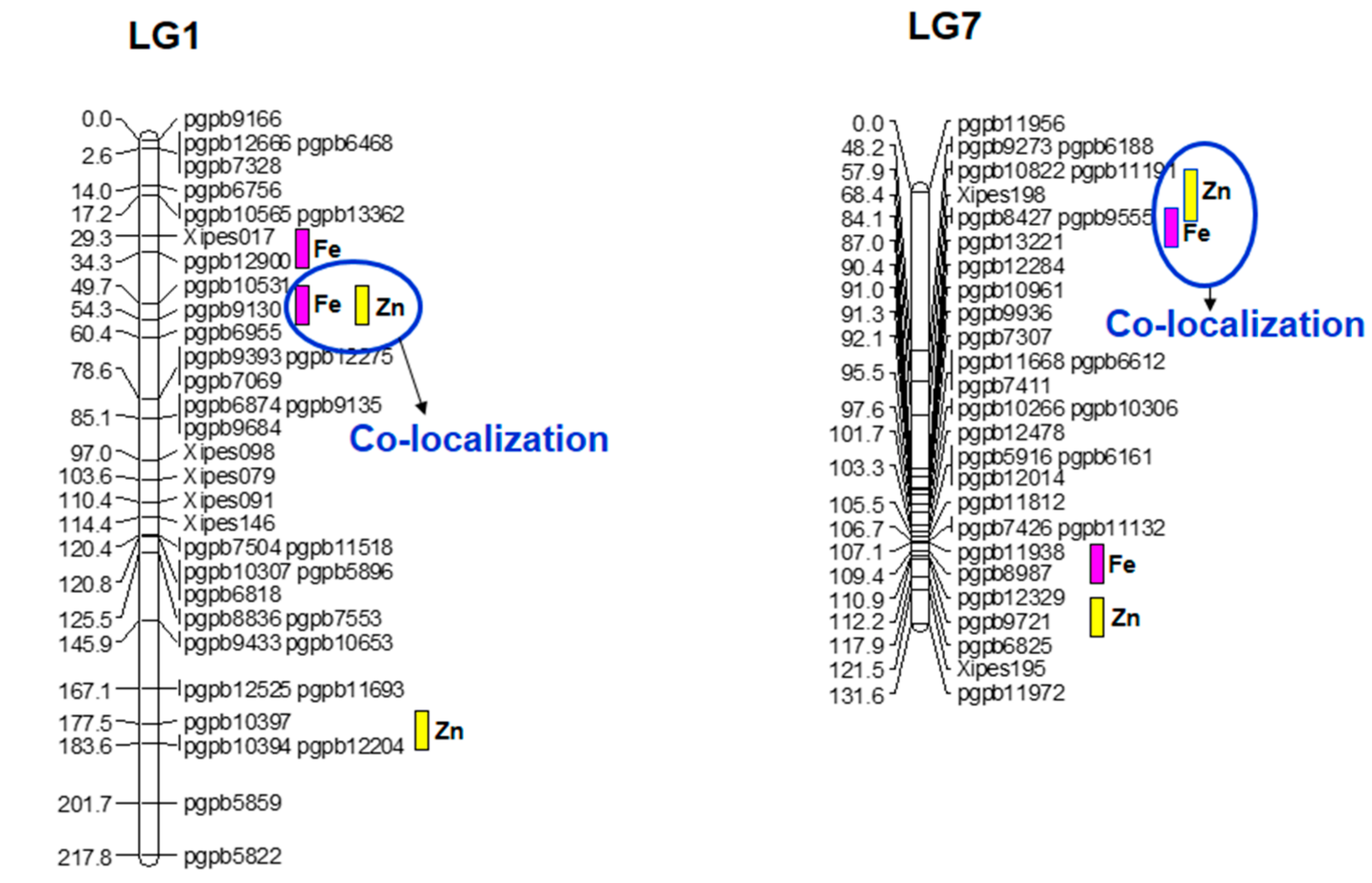
3.6. Quantitative Trait Loci for Grain Iron (Fe) and Zinc (Zn) Content and Epistasis
4. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Ramya, A.R.; Ahamed, M.L.; Satyavathi, C.T.; Rathore, A.; Katiyar, P.; Raj, A.G.B.; Kumar, S.; Gupta, R.; Mahendrakar, M.D.; Yadav, R.S.; et al. Towards defining heterotic gene pools in pearl millet [Pennisetum glaucum (L.) R. Br.]. Front. Plant Sci. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Anuradha, N.; Satyavathi, C.T.; Bharadwaj, C.; Nepolean, T.; Sankar, S.M.; Singh, S.P.; Meena, M.C.; Singhal, T.; Srivastava, R.K. Deciphering genomic regions for high grain iron and zinc content using association mapping in pearl millet. Front. Plant Sci. 2017, 8, 412. [Google Scholar] [CrossRef] [PubMed]
- Gangashetty, P.I.; Motagi, B.N.; Pavan, R.; Roodagi, M.B. Breeding Crop Plants for Improved Human Nutrition through Biofortification: Progress and Prospects. In Advances in Plant Breeding Strategies: Agronomic, Abiotic and Biotic Stress Traits, 1st ed.; Al-Khayri Jameel, M., Jain, S.M., Johnson, D., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 35–76. ISBN 978-3-319-22518-0. [Google Scholar]
- Liu, C.J.; Witcombe, J.R.; Pittaway, T.S.; Nash, M.; Hash, C.T.; Busso, C.S.; Gale, M.D. An RFLP-based genetic map of pearl millet (Pennisetum glaucum). Theor. Appl. Genet. 1994, 4, 481–487. [Google Scholar] [CrossRef] [PubMed]
- Gale, K.R.; Ma, W.; Zhang, W.; Rampling, L.; Hill, A.S.; Appels, R.; Morris, P.; Morrel, M. Simple high-throughput DNA markers for genotyping in wheat. In Proceedings 10th Australian Wheat Breeders Assembly, Mildura, Australia, 16–21 September 2001; CSIRO: Canberra, Australia, 2001; pp. 26–31. [Google Scholar]
- Vom Brocke, K.; Christinck, A.; Weltzien, E.; Presterl, T.; Geiger, H.H. Farmers’ seed systems and management practices determine pearl millet genetic diversity patterns in semiarid regions of India. Crop Sci. 2003, 5, 1680–1689. [Google Scholar] [CrossRef]
- Bertin, I.; Zhu, J.H.; Gale, M.D. SSCP-SNP in pearl millet—A new marker system for comparative genetics. Theor. Appl. Genet. 2005, 8, 1467–1472. [Google Scholar] [CrossRef] [PubMed]
- Qi, X.; Lindup, S.; Pittaway, T.S.; Allouis, S.; Gale, M.D.; Devos, K.M. Development of simple sequence repeat markers from bacterial artificial chromosomes without subcloning. Biotechnol. 2001, 2, 355–358. [Google Scholar]
- Qi, X.; Pittaway, T.S.; Lindup, S.; Liu, H.; Waterman, E.; Padi, F.K.; Hash, C.T.; Zhu, J.; Gale, M.D.; Devos, K.M. An integrated genetic map and a new set of simple sequence repeat markers for pearl millet, Pennisetum glaucum. Theor. Appl. Genet. 2004, 7, 1485–1493. [Google Scholar] [CrossRef] [PubMed]
- Budak, H.; Pedraza, F.; Cregan, P.B.; Baenziger, P.S.; Dweikat, I. Development and utilization of SSRs to estimate the degree of genetic relationships in a collection of pearl millet germplasm. Crop Sci. 2003, 6, 2284–2290. [Google Scholar] [CrossRef]
- Mariac, C.; Luong, V.; Kapran, I.; Mamadou, A.; Sagnard, F.; Deu, M.; Chantereau, J.; Gerard, B.; Ndjeunga, J.; Bezançon, G.; et al. Diversity of wild and cultivated pearl millet accessions (Pennisetum glaucum [L.] R. Br.) in Niger assessed by microsatellite markers. Theor. Appl. Genet. 2006, 1, 49–58. [Google Scholar] [CrossRef] [PubMed]
- Senthilvel, S.; Jayashree, B.; Mahalakshmi, V.; Kumar, P.S.; Nakka, S.; Nepolean, T.; Hash, C. Development and mapping of simple sequence repeat markers for pearl millet from data mining of expressed sequence tags. BMC Plant Biol. 2008, 8, 119. [Google Scholar] [CrossRef] [PubMed]
- Rajaram, V.; Nepolean, T.; Senthilvel, S.; Varshney, R.K.; Vadez, V.; Rakesh, K.; Shah, T.M.; Supriya, A.; Kumar, S.; Kumari, B.R.; et al. Pearl millet [Pennisetum glaucum (L.) R. Br.] consensus linkage map constructed using four RIL mapping populations and newly developed EST-SSRs. BMC Genomics 2013, 14, 159. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Hash, C.T.; Thirunavukkarasu, N.; Singh, G.; Rajaram, V.; Rathore, A.; Senapathy, S.; Mahendrakar, M.D.; Yadav, R.S.; Srivastava, R.K. Mapping quantitative trait loci controlling high iron and zinc content in self and open pollinated grains of pearl millet [Pennisetum glaucum (L.) R. Br.]. Front. Plant Sci. 2016, 7, 1636. [Google Scholar] [CrossRef] [PubMed]
- Supriya, A.; Senthilvel, S.; Nepolean, T.; Eshwar, K.; Rajaram, V.; Shaw, R.; Hash, C.T.; Kilian, A.; Yadav, R.C.; Narasu, M.L. Development of a molecular linkage map of pearl millet integrating DArT and SSR markers. Theor. Appl. Genet. 2011, 2, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Varshney, R.K.; Thundi, M.; May, G.D.; Jackson, S.A. Legume Genomics and Breeding. In Plant Breed Reviews; Wiley Publisher: Hoboken, NJ, USA, 2010; Volume 33, pp. 257–304. ISBN 9780470535486. [Google Scholar]
- Jaccoud, D.; Peng, K.; Feinstein, D.; Kilian, A. Diversity Arrays: A solid state technology for sequence information independent genotyping. Nucleic Acids Res. 2001, 29, E25. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.Y.; Saxena, R.K.; Kulwal, P.L.; Ash, G.J.; Dubey, A.; Harper, J.D.; Upadhyaya, H.D.; Gothalwal, R.; Kilian, A.; Varshney, R.K. The first genetic map of pigeon pea based on diversity arrays technology (DArT) markers. J. Genet. 2011, 1, 103–109. [Google Scholar] [CrossRef]
- Mace, E.S.; Xia, L.; Jordan, D.R.; Halloran, K.; Parh, D.K.; Huttner, E.; Wenzl, P.; Kilian, A. DArT markers: Diversity analyses and mapping in Sorghum bicolor. BMC Genom. 2008, 9, 26. [Google Scholar] [CrossRef] [PubMed]
- Jing, H.C.; Bayon, C.; Kanyuka, K.; Berry, S.; Wenzl, P.; Huttner, E.; Kilian, A.; Hammond-Kosack, K.E. DArT markers: Diversity analyses, genomes comparison, mapping and integration with SSR markers in Triticum monococcum. BMC Genom. 2009, 10, 458. [Google Scholar] [CrossRef] [PubMed]
- Soller, M.; Brody, T.; Genizi, A. On the power of experimental designs for the detection of linkage between marker loci and quantitative loci in crosses between inbred lines. Theor. Appl. Genet. 1976, 1, 35–39. [Google Scholar]
- Lander, E.S.; Botstein, D. Mapping mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 1989, 121, 185–199. [Google Scholar] [PubMed]
- Lynch, M.; Walsh, B. Genetics Analysis of Quantitative Traits, 1st ed.; Sinauer Associates: Sunderland, MA, USA, 1998; pp. 469–476. ISBN 0878934812. [Google Scholar]
- Darvasi, A.; Weinreb, A.; Minke, V.; Weller, J.I.; Soller, M. Detecting marker-QTL linkage and estimating QTL gene effect and map location using a saturated genetic map. Genetics 1993, 3, 943–951. [Google Scholar]
- Bouis, H.E. Micronutrient fortification of plants through plant breeding: Can it improve nutrition in man at low cost? Proc. Nutr. Soc. 2003, 2, 403–411. [Google Scholar] [CrossRef]
- Welch, R.M.; Graham, R.D. Breeding for micronutrients in staple food crops from a human nutrition perspective. J. Exp. Bot. 2004, 396, 353–364. [Google Scholar] [CrossRef] [PubMed]
- White, P.J.; Broadley, M.R. Biofortification of crops with seven mineral elements often lacking in human diets–iron, zinc, copper, calcium, magnesium, selenium and iodine. New Phytol. 2009, 1, 49–84. [Google Scholar] [CrossRef] [PubMed]
- Graham, R.D.; Welch, R.M.; Bouis, H.E. Addressing micronutrient malnutrition through enhancing the nutritional quality of staple foods: Principles, perspectives and knowledge gaps. Adv. Agron. 2001, 70, 77–142. [Google Scholar]
- Welch, R.M. Importance of seed mineral nutrient reserves in crop growth and development. In Mineral Nutrition of Crops: Fundamental Mechanisms and Implications, 1st ed.; Rengel, Z., Ed.; CRC Press: New York, NY, USA, 1999; pp. 205–226. ISBN 978-1560228806. [Google Scholar]
- Velu, G.; Kulkarni, V.N.; Muralidharan, V.; Rai, K.N.; Longvah, T.; Sahrawat, K.L.; Raveendran, T.S. A rapid screening method for grain iron content in pearl millet. International Sorghum and Millets Newsletter. 2006. Available online: http://agropedia.iitk.ac.in/sites/default/files/ISMN-47%20158-161.pdf (accessed on 15 December 2017).
- Allouis, S.; Qi, X.; Lindup, S.; Gale, M.D.; Devos, K.M. Construction of a BAC library of pearl millet, Pennisetum glaucum. Theor. Appl. Genet. 2001, 102, 1200–1205. [Google Scholar] [CrossRef]
- Lander, E.S.; Green, P.; Abrahamson, J.; Barlow, A.; Daly, M.J.; Lincoln, S.E.; Newberg, L.A. MAPMAKER: An interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1987, 2, 174–181. [Google Scholar] [CrossRef]
- Voorrips, R.E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Heredity 2002, 93, 77–78. [Google Scholar] [CrossRef]
- Falconer, D.S. Introduction to Quantitative Genetics, 3rd ed.; John Wiley & Sons: New York, NY, USA, 1989; pp. 261–266. ISBN 0321042472. [Google Scholar]
- Zeng, Z.B. Precision mapping of quantitative trait loci. Genetics 1994, 4, 1457–1468. [Google Scholar]
- Utz, H.F.; Melchinger, A.E. PLABQTL: A program for composite interval mapping of QTL. J. Quant. Trait Loci 1996, 2, 1–5. [Google Scholar]
- Ferreira, A.; da Silva, M.F.; Silva, L.C.; Cruz, C.D. Estimating the effects of population size and type on the accuracy of genetic maps. Genet. Mol. Biol. 2006, 29, 187–192. [Google Scholar] [CrossRef]
- Yadav, R.S.; Hash, C.T.; Bidinger, F.R.; Devos, K.M.; Howarth, C.J. Genomic regions associated with grain yield and aspects of post-flowering drought tolerance in pearl millet across stress environments and tester background. Euphytica 2004, 136, 265–277. [Google Scholar] [CrossRef]
- Gulia, S.K. QTL Mapping for improvement of downy mildew [Sclerospora Graminicola (Sacc.) J. Schroet.] resistance (DMR) in pearl millet [Pennisetum glaucum (L.) R. Br.] hybrid parental line ICMB 89111. Ph.D. Thesis, Chaudhary Charan Singh Haryana Agricultural University, Hisar, India, 2004. [Google Scholar]
- Sehgal, D.; Rajaram, V.; Armstead, I.P.; Vadez, V.; Yadav, Y.P.; Hash, C.T.; Yadav, R.S. Integration of gene-based markers in a pearl millet genetic map for identification of candidate genes underlying drought tolerance quantitative trait loci. BMC Plant Bio. 2012, 12, 9. [Google Scholar] [CrossRef] [PubMed]
- Devos, K.M.; Pittaway, T.S.; Reynolds, A.; Gale, M.D. Comparative mapping reveals a complex relationship between the pearl millet genome and those of foxtail millet and rice. Theor. Appl. Genet. 2000, 100, 190–198. [Google Scholar] [CrossRef]
- Singh, K.; Ghai, M.; Garg, M.; Chhuneja, P.; Kaur, P.; Schnurbusch, T.; Keller, B.; Dhaliwal, H.S. An integrated molecular linkage map of diploid wheat based on a Triticum boeoticum × T. monococcum RIL population. Theor. Appl. Genet. 2007, 115, 301–312. [Google Scholar] [CrossRef] [PubMed]
- Graner, A.; Jahoor, A.; Schondelmaier, J.; Siedler, H.; Pillen, K.; Fischbeck, G.; Wenzel, G.; Herrmann, R.G. Construction of a RFLP map of barley. Theor. Appl. Genet. 1991, 83, 250–256. [Google Scholar] [CrossRef] [PubMed]
- Devaux, P.; Kilian, A.; Kleinhofs, A. Comparative mapping of the barley genome with male and female recombination-derived, doubled haploid populations. Mol. Gen. Genet. 1995, 6, 600–608. [Google Scholar] [CrossRef]
- Causse, M.A.; Fulton, T.M.; Cho, Y.G.; Ahn, S.N.; Chunwongse, J.; Wu, K.S.; Xiao, J.H.; Yu, Z.H.; Ronald, P.C.; Harrington, S.E.; et al. Saturated molecular map of the rice genome based on an interspecific backcross population. Genetics 1994, 138, 1251–1274. [Google Scholar] [PubMed]
- Xu, Y.; Zhu, L.; Xiao, J.; Huang, N.; McCouch, S.R. Chromosomal regions associated with segregation distortion of molecular markers in F2, backcross, doubled haploid, and recombinant inbred populations in rice (Oryza sativa L.). Mol. Gen. Genet. 1997, 5, 535–545. [Google Scholar] [CrossRef]
- Wendel, J.F.; Edwards, M.D.; Stuber, C.W. Evidence for multilocus genetic control of preferential fertilization in maize. Heredity 1987, 2, 297–301. [Google Scholar] [CrossRef]
- Lu, H.; Romero-Severson, J.; Bernardo, R. Chromosomal regions associated with segregation distortion in maize. Theor. Appl. Genet. 2002, 105, 622–628. [Google Scholar] [PubMed]
- Blanco, A.; Simeone, R.; Cenci, A.; Gadaleta, A.; Tanzarella, O.A.; Porceddu, E.; Salvi, S.; Tuberosa, R.; Figliuolo, G.; Spagnoletti, P.; et al. Extension of the Messapia × dicoccoides linkage map of Triticum turgidum (L.) Thell. Cell Mol. Biol. Lett. 2004, 9, 529–541. [Google Scholar] [PubMed]
- Peng, J.; Korol, A.B.; Fahima, T.; Röder, M.S.; Ronin, Y.I.; Li, Y.C.; Nevo, E. Molecular genetic maps in wild emmer wheat, Triticum dicoccoides: Genome-wide coverage, massive negative interference, and putative quasi-linkage. Genome Res. 2000, 10, 1509–1531. [Google Scholar] [CrossRef] [PubMed]
- Quarrie, S.A.; Steed, A.; Calestani, C.; Semikhodskii, A.; Lebreton, C.; Chinoy, C.; Steele, N.; Pljevljakusić, D.; Waterman, E.; Weyen, J.; et al. A high-density genetic map of hexaploid wheat (Triticum aestivum L.) from the cross Chinese Spring × SQ1 and its use to compare QTLs for grain yield across a range of environments. Theor. Appl. Genet. 2005, 5, 865–880. [Google Scholar] [CrossRef] [PubMed]
- Cloutier, S.; Cappadocia, M.; Landry, B.S. Analysis of RFLP mapping inaccuracy in Brassica napus L. Theor. Appl. Genet. 1997, 95, 83–91. [Google Scholar] [CrossRef]
- Livingstone, K.D.; Lackney, V.K.; Blauth, J.R.; Van Wijk, R.I.K.; Jahn, M.K. Genome mapping in capsicum and the evolution of genome structure in the Solanaceae. Genetics 1999, 152, 1183–1202. [Google Scholar] [PubMed]
- Yadav, R.S.; Sehgal, D.; Vadez, V. Using genetic mapping and genomics approaches in understanding and improving drought tolerance in pearl millet. J. Exp. Bot. 2011, 2, 397–408. [Google Scholar] [CrossRef] [PubMed]
- Varshney, R.K.; Hoisington, D.A.; Tyagi, A.K. Advances in cereal genomics and applications in crop breeding. Trends Biotechnol. 2006, 11, 490–499. [Google Scholar] [CrossRef] [PubMed]
- Mace, E.S.; Rami, J.F.; Bouchet, S.; Klein, P.E.; Klein, R.R.; Kilian, A.; Wenzl, P.; Xia, L.; Halloran, K.; Jordan, D.R. A consensus genetic map of sorghum that integrates multiple component maps and high-throughput Diversity Array Technology (DArT) markers. BMC Plant Biol. 2009, 9, 13. [Google Scholar] [CrossRef] [PubMed]
- Van Os, H.; Andrzejewski, S.; Bakker, E.; Barrena, I.; Bryan, G.J.; Caromel, B.; Ghareeb, B.; Isidore, E.; de Jong, W.; Van Koert, P.; et al. Construction of a 10,000-marker ultra-dense genetic recombination map of potato: Providing a framework for accelerated gene isolation and a genome wide physical map. Genetics 2006, 2, 1075–1087. [Google Scholar] [CrossRef] [PubMed]
- Vreugdenhil, D.; Aarts, M.G.M.; Koornneef, M.; Nelissen, H.; Ernst, W.H.O. Natural variation and QTL analysis for cationic mineral content in seeds of Arabidopsis thaliana. Plant Cell Environ. 2004, 27, 828–839. [Google Scholar] [CrossRef]
- Ghandilyan, A.; Vreugdenhil, D.; Mark, G.M.; Aart, M. Progress in the genetic understanding of plant iron and zinc nutrition. Physiol. Plant. 2006, 126, 407–417. [Google Scholar] [CrossRef]
- Khairwal, I.S.; Rai, K.N.; Andrew, D.J.; Harinarayana, G. Pearl Millet Breeding; Oxford and IBH Publishing Co.: New Delhi, India, 1999; p. 511. ISBN 81-204-1305-9. [Google Scholar]
- Ezeaku, I.E.; Mohammed, S.G. Character association and path analysis in grain sorghum. Afr. J. Biotechnol. 2006, 14, 1337–340. [Google Scholar]
- Jinks, J.L.; Pooni, H.S. Predicting the properties of pure breeding lines extractable from a cross in the presence of linkage. Heredity 1982, 49, 265–270. [Google Scholar] [CrossRef]
- Rieseberg, L.H.; Widmer, A.; Arntz, A.M.; Burke, J.M. The genetic architecture necessary for transgressive segregation is common in both natural and domesticated populations. Philos. Trans. R. Soc. B 2003, 358, 1141–1147. [Google Scholar] [CrossRef] [PubMed]
- Lu, K.; Li, L.; Zheng, X.; Zhang, Z.; Mou, T.; Hu, Z. Quantitative trait loci controlling Cu, Ca, Zn, Mn and Fe content in rice grains. J. Genet. 2008, 87, 305–310. [Google Scholar] [CrossRef] [PubMed]
- Sankaran, R.P.; Huguet, T.; Grusak, M.A. Identification of QTL affecting seed mineral concentrations and content in the model legume Medicago truncatula. Theor. Appl. Genet. 2009, 119, 241–253. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Hash, C.T.; Nepolean, T.; Satyavathi, C.T.; Singh, G.; Mahendrakar, M.D.; Yadav, R.S.; Srivastava, R.K. Mapping QTLs controlling flowering time and important agronomic traits in pearl millet. Front. Plant Sci. 2017, 8, 1731. [Google Scholar] [CrossRef] [PubMed]

| Linkage Group | SSR Marker Loci | Skewed SSR Loci | DArT Marker Loci | Skewed DArT Loci | Total Marker Loci | Total Skewed Marker Loci | Skewed Loci (%) | Total Length (cM) | Average Inter-Marker Distance (cM) |
|---|---|---|---|---|---|---|---|---|---|
| 1 | 5 | 4 | 33 | 23 | 38 | 27 | 71 | 218 | 5.9 |
| 2 | 2 | 2 | 28 | 23 | 30 | 25 | 83 | 114 | 3.9 |
| 3 | 3 | 3 | 19 | 14 | 22 | 17 | 77 | 84 | 4.0 |
| 4A | 3 | 3 | 35 | 23 | 38 | 26 | 68 | 178 | 4.8 |
| 4B | 0 | 0 | 5 | 2 | 5 | 2 | 40 | 26 | 6.6 |
| 5 | 3 | 0 | 15 | 6 | 18 | 6 | 33 | 166 | 9.7 |
| 6 | 1 | 1 | 12 | 3 | 13 | 4 | 31 | 47 | 3.9 |
| 7 | 2 | 1 | 30 | 24 | 32 | 25 | 78 | 132 | 4.3 |
| A | 0 | 0 | 5 | 4 | 5 | 4 | 80 | 16.3 | 4.1 |
| B | 1 | 1 | 3 | 3 | 4 | 4 | 100 | 18.2 | 6.1 |
| C | 0 | 0 | 4 | 2 | 4 | 2 | 50 | 23.5 | 7.8 |
| D | 1 | 0 | 1 | 1 | 2 | 1 | 50 | 25.4 | 25.4 |
| E | 2 | 2 | 1 | 1 | 3 | 3 | 100 | 14.9 | 7.5 |
| F | 0 | 0 | 2 | 2 | 2 | 2 | 100 | 4.1 | 4.1 |
| G | 0 | 0 | 2 | 0 | 2 | 0 | 0 | 22.7 | 22.7 |
| H | 0 | 0 | 4 | 3 | 4 | 3 | 75 | 33.8 | 11.3 |
| I | 2 | 2 | 0 | 0 | 2 | 2 | 100 | 21.3 | 21.3 |
| J | 0 | 0 | 2 | 2 | 2 | 2 | 100 | 4.9 | 4.9 |
| K | 0 | 0 | 2 | 2 | 2 | 2 | 100 | 3.1 | 3.1 |
| L | 0 | 0 | 2 | 2 | 2 | 2 | 100 | 1.9 | 1.9 |
| M | 1 | 0 | 1 | 1 | 2 | 1 | 50 | 12.9 | 12.9 |
| N | 1 | 0 | 1 | 1 | 2 | 1 | 50 | 19.9 | 19.9 |
| Total | 27 (19) # | 19 (14) | 207 (177) | 142 (118) | 234 (196) | 161 (132) | 69 (68) | 1187 (964) | 5.0 (4.9) |
| 8511 (P1) | AIMP (P2) | RILs | P1 vs. P2 | P1 vs. RILs | P2 vs. RILs | ||
|---|---|---|---|---|---|---|---|
| Trait | Mean | Mean | Mean | Range | Pr > F | Pr > F | Pr > F |
| Fe | 29.8 ± 3.3 | 124.1 ± 2.9 | 54.0 ± 0.3 | 20.0–131.0 | ** | ** | ** |
| Zn | 27.7 ± 2.7 | 86.3 ± 2.4 | 43.9 ± 0.3 | 18.2–109.8 | ** | ** | ** |
| Trait | σ2g | SE | H2 |
|---|---|---|---|
| Fe | 417.98 | 35.90 | 0.88 |
| Zn | 197.29 | 17.46 | 0.83 |
| Trait | QTL 1 | Marker Interval | Support Interval | LOD 2 | R2 (%) | Additive Effects 3 |
|---|---|---|---|---|---|---|
| Fe | 1/30 | Xipes017-pgpb12900 | 26–36 | 6.22 | 9.0 | 4.0 |
| 1/54 | pgpb10531-pgpb9130 | 52–56 | 25.36 | 31.9 | 9.7 | |
| 3/20 | Xipes188-pgpb6069 | 8–26 | 6.59 | 9.5 | 0.4 | |
| 4B/8 | pgpb9502-pgpb6039 | 4–12 | 6.87 | 10.4 | −0.6 | |
| 7/16 | pgpb11956-pgpb9273 | 0–30 | 7.25 | 12.5 | −1.9 | |
| 7/86 | pgpb8427-pgpb13221 | 84–90 | 8.58 | 12.2 | 5.3 | |
| 7/108 | pgpb11938-pgpb8987 | 106–110 | 8.83 | 12.5 | 4.9 | |
| 7/120 | pgpb6825-Xipes195 | 118–122 | 9.70 | 14.0 | 0.1 | |
| A/0 | pgpb8445-pgpb11206 | 0–2 | 7.67 | 12.4 | 4.0 | |
| D/20 | pgpb10660-pgpb8626 | 14–24 | 7.00 | 11.6 | 1.2 | |
| E/14 | pgpb10727-Xipes179 | 12–14 | 9.36 | 14.3 | 3.1 | |
| Zn | 1/54 | pgpb10531-pgpb9130 | 52–56 | 23.93 | 30.4 | 6.7 |
| 1/182 | pgpb10397-pgpb10394 | 178–186 | 6.50 | 9.4 | 1.7 | |
| 4B/8 | pgpb9502-pgpb6039 | 4–10 | 7.33 | 11.1 | −0.6 | |
| 4B/22 | pgpb10483-pgpb11463 | 14–26 | 6.68 | 11.6 | −2.2 | |
| 5/112 | pgpb13229-pgpb12681 | 108–118 | 8.17 | 11.6 | 2.7 | |
| 7/82 | Xipes198-pgpb8427 | 74–88 | 7.16 | 10.2 | 2.7 | |
| 7/112 | pgpb12329-pgpb9721 | 110–116 | 7.58 | 10.9 | 2.8 | |
| H/16 | pgpb8779-pgpb12691 | 10–22 | 6.68 | 11.6 | 2.1 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kumar, S.; Hash, C.T.; Nepolean, T.; Mahendrakar, M.D.; Satyavathi, C.T.; Singh, G.; Rathore, A.; Yadav, R.S.; Gupta, R.; Srivastava, R.K. Mapping Grain Iron and Zinc Content Quantitative Trait Loci in an Iniadi-Derived Immortal Population of Pearl Millet. Genes 2018, 9, 248. https://doi.org/10.3390/genes9050248
Kumar S, Hash CT, Nepolean T, Mahendrakar MD, Satyavathi CT, Singh G, Rathore A, Yadav RS, Gupta R, Srivastava RK. Mapping Grain Iron and Zinc Content Quantitative Trait Loci in an Iniadi-Derived Immortal Population of Pearl Millet. Genes. 2018; 9(5):248. https://doi.org/10.3390/genes9050248
Chicago/Turabian StyleKumar, Sushil, Charles Tom Hash, Thirunavukkarasu Nepolean, Mahesh D. Mahendrakar, Chellapilla Tara Satyavathi, Govind Singh, Abhishek Rathore, Rattan S. Yadav, Rajeev Gupta, and Rakesh K. Srivastava. 2018. "Mapping Grain Iron and Zinc Content Quantitative Trait Loci in an Iniadi-Derived Immortal Population of Pearl Millet" Genes 9, no. 5: 248. https://doi.org/10.3390/genes9050248
APA StyleKumar, S., Hash, C. T., Nepolean, T., Mahendrakar, M. D., Satyavathi, C. T., Singh, G., Rathore, A., Yadav, R. S., Gupta, R., & Srivastava, R. K. (2018). Mapping Grain Iron and Zinc Content Quantitative Trait Loci in an Iniadi-Derived Immortal Population of Pearl Millet. Genes, 9(5), 248. https://doi.org/10.3390/genes9050248








