Evolutionary Mechanisms of Varying Chromosome Numbers in the Radiation of Erebia Butterflies
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection
2.2. Phylogenetic Analyses
3. Results
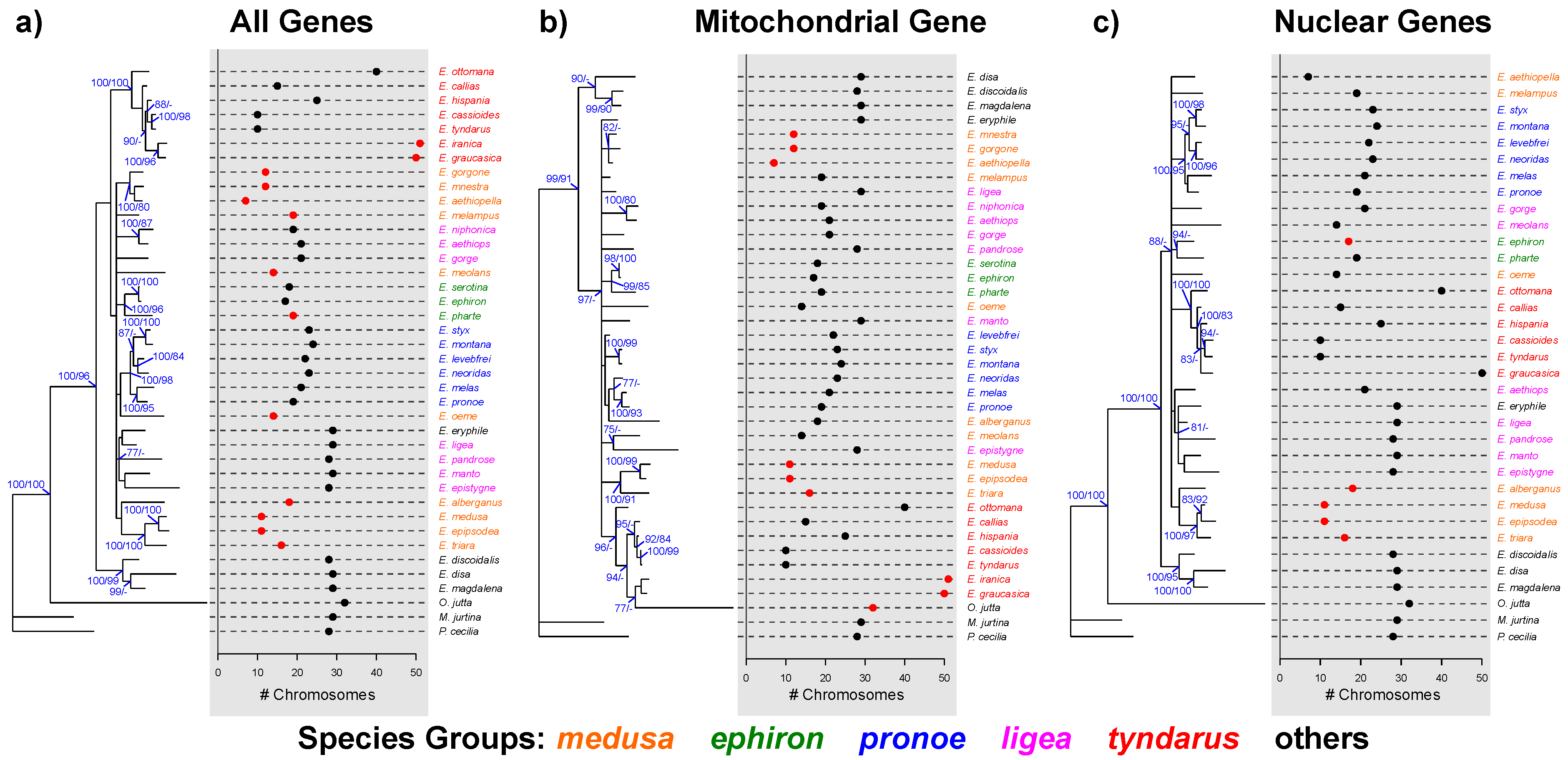
3.1. Phylogenetic Reconstruction
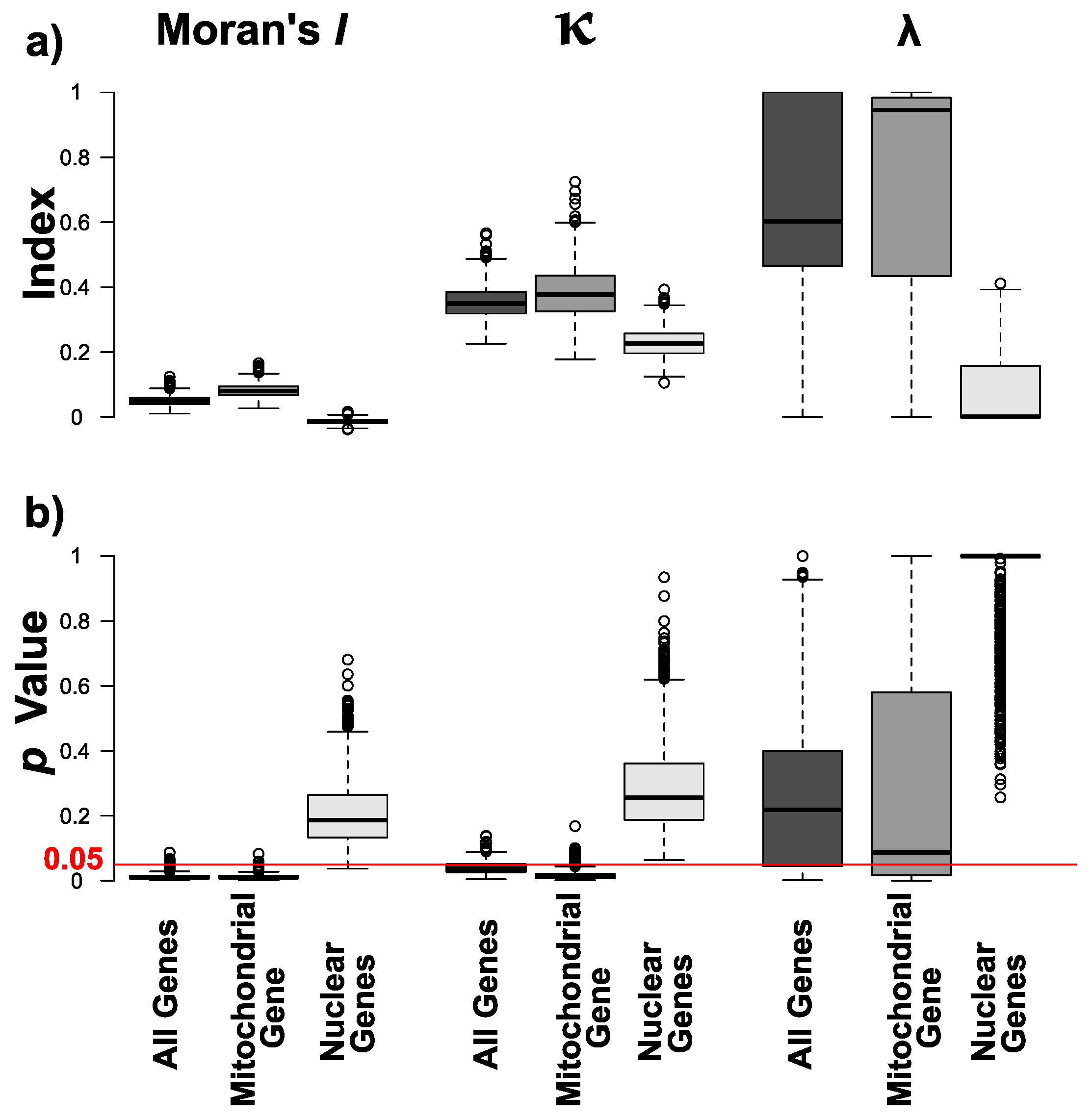
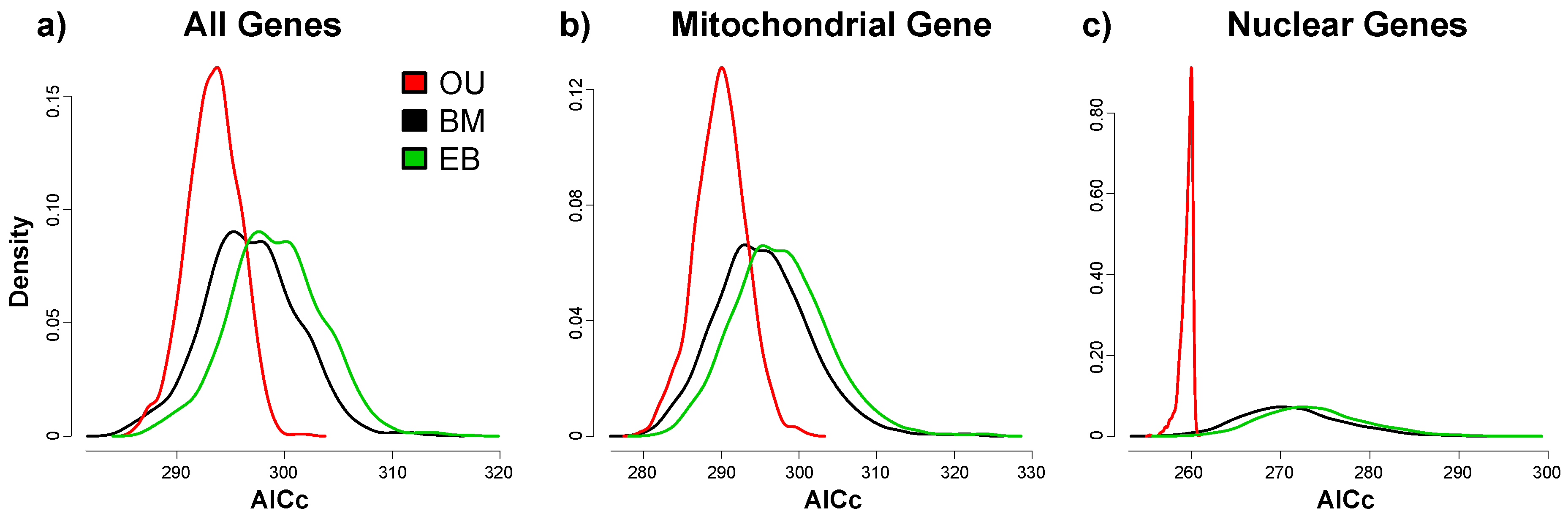
3.2. Phylogenetic Signals
4. Discussion
Supplementary Materials
Acknowledgments
Conflict of interest
References
- Hendry, A.P. Ecological speciation! Or the lack thereof? Can. J. Fish. Aquat. Sci. 2009, 66, 1383–1398. [Google Scholar] [CrossRef]
- Nosil, P. Ecological Speciation; Oxford University Press: Oxford, UK, 2012. [Google Scholar]
- Seehausen, O.; Butlin, R.K.; Keller, I.; Wagner, C.E.; Boughman, J.W.; Hohenlohe, P.A.; Peichel, C.L.; Saetre, G.-P.; Bank, C.; Brännström, Å.; et al. Genomics and the origin of species. Nat. Rev. Genet. 2014, 15, 176–192. [Google Scholar] [CrossRef] [PubMed]
- Nosil, P.; Harmon, L.J.; Seehausen, O. Ecological explanations for (incomplete) speciation. Trends Ecol. Evol. 2009, 24, 145–156. [Google Scholar] [CrossRef] [PubMed]
- Butlin, R.K. Recombination and speciation. Mol. Ecol. 2005, 14, 2621–2635. [Google Scholar] [CrossRef] [PubMed]
- Kirkpatrick, M.; Barton, N.J. Chromosome inversions, local adaptation and speciation. Genetics 2006, 173, 419–434. [Google Scholar] [CrossRef] [PubMed]
- Via, S. Natural selection in action during speciation. Proc. Natl. Acad. Sci. USA 2009, 106 (Suppl. 1), 9939–9946. [Google Scholar] [CrossRef] [PubMed]
- Kulmuni, J.; Westram, A.M. Intrinsic incompatibilities evolving as a by-product of divergent ecological selection: Considering them in empirical studies on divergence with gene flow. Mol. Ecol. 2017, 26, 3093–3103. [Google Scholar] [CrossRef] [PubMed]
- Martinez, P.A.; Zurano, J.P.; Amado, T.F.; Penone, C.; Betancur-R, R.; Bidau, C.J.; Jacobina, U.P. Chromosomal diversity in tropical reef fishes is related to body size and depth range. Mol. Phylogenet. Evol. 2015, 93, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Hooper, D.M.; Price, T.D. Chromosomal inversion differences correlate with range overlap in passerine birds. Nat. Ecol. Evol. 2017, 1, 1526–1534. [Google Scholar] [CrossRef] [PubMed]
- White, M.J.D. Modes of Speciation; W.H. Freeman: San Francisco, CA, USA, 1978. [Google Scholar]
- King, M. Species Evolution; Cambridge University Press: Cambridge, UK, 1995. [Google Scholar]
- Faria, R.; Navarro, A. Chromosomal speciation revisited: Rearranging theory with pieces of evidence. Trends Ecol. Evol. 2010, 25, 660–669. [Google Scholar] [CrossRef] [PubMed]
- Dion-Côté, A.-M.; Barbash, D.A. Beyond speciation genes: An overview of genome stability in evolution and speciation. Curr. Opin. Genet. Dev. 2017, 47, 17–23. [Google Scholar] [CrossRef] [PubMed]
- Cohuet, A.; Dia, I.; Simard, F.; Raymond, M.; Rousset, F.; Antonio-Nkondjio, C.; Awono-Ambene, P.H.; Wondji, C.S.; Fontenille, D. Gene flow between chromosomal forms of the malaria vector Anopheles funestus in Cameroon, Central Africa, and its relevance in malaria fighting. Genetics 2005, 169, 301–311. [Google Scholar] [CrossRef] [PubMed]
- Polyakov, A.V.; White, T.A.; Jones, R.M.; Borodin, P.M.; Searle, J.B. Natural hybridization between extremely divergent chromosomal races of the common shrew (Sorex araneus, Soricidae, Soricomorpha): Hybrid zone in European Russia. J. Evol. Biol. 2011, 24, 573–583. [Google Scholar] [CrossRef] [PubMed]
- Descimon, H.; Mallet, J. Bad Species. In Ecology of Butterflies in Europe; Settle, R., Shreeve, T., Konvicka, M., van Dyck, H., Eds.; Cambridge University Press: Cambridge, UK, 2009. [Google Scholar]
- Mallet, J.; Besansky, N.; Hahn, M.W. How reticulated are species? Bioessays 2016, 38, 140–149. [Google Scholar] [CrossRef] [PubMed]
- Charron, G.; Leducq, J.-B.; Landry, C.R. Chromosomal variation segregates within incipient species and correlates with reproductive isolation. Mol. Ecol. 2014, 23, 4362–4372. [Google Scholar] [CrossRef] [PubMed]
- Potter, S.; Bragg, J.G.; Blom, M.P.K.; Deakin, J.E.; Kirkpatrick, M.; Eldridge, M.D.B.; Moritz, C. Chromosomal speciation in the genomics era: Disentangling phylogenetic evolution of rock-wallabies. Front. Genet. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Vershinina, A.O.; Lukhtanov, V.A. Evolutionary mechanisms of runaway chromosome number change in Agrodiaetus butterflies. Sci. Rep. 2017, 7, 8199. [Google Scholar] [CrossRef] [PubMed]
- Robinson, R. Lepidoptera Genetics; Pergamon Press: Oxford, UK, 1971. [Google Scholar]
- Lukhtanov, V.A.; Kandul, N.P.; Plotkin, J.B.; Dantchenko, A.V.; Haig, D.; Pierce, N.E. Reinforcement of pre-zygotic isolation and karyotype evolution in Agrodiaetus butterflies. Nature 2005, 436, 385–389. [Google Scholar] [CrossRef] [PubMed]
- Talavera, G.; Lukhtanov, V.A.; Rieppel, L.; Pierce, N.E.; Vila, R. In the shadow of phylogenetic uncertainty: The recent diversification of Lysandra butterflies through chromosomal change. Mol. Phylogenet. Evol. 2013, 69, 469–478. [Google Scholar] [CrossRef] [PubMed]
- Lorkovic, Z. Some peculiarities of spatially and sexually restricted gene exchange in the Erebia tyndarus group. Cold Spring Harb. Symp. Quant. Biol. 1958, 23, 319–325. [Google Scholar] [CrossRef] [PubMed]
- Peña, C.; Witthauer, H.; Kleckova, I.; Fric, Z. Adaptive radiations in butterflies: Evolutionary history of the genus Erebia (Nymphalidae: Satyrinae). Biol. J. Linn. Soc. 2015, 116, 449–467. [Google Scholar] [CrossRef]
- Martin, J.F.; Gilles, A.; Lortscher, M.; Descimon, H. Phylogenetics and differentiation among the western taxa of the Erebia tyndarus group (Lepidoptera: Nymphalidae). Biol. J. Linn. Soc. 2002, 75, 319–332. [Google Scholar] [CrossRef]
- Albre, J.; Gers, C.; Legal, L. Molecular phylogeny of the Erebia tyndarus (Lepidoptera, Rhopalocera, Nymphalidae, Satyrinae) species group combining CoxII and ND5 mitochondrial genes: A case study of a recent radiation. Mol. Phylogenet. Evol. 2008, 47, 196–210. [Google Scholar] [CrossRef] [PubMed]
- Gratton, P.; Trucchi, E.; Trasatti, A.; Riccarducci, G.; Marta, S.; Allegrucci, G.; Cesaroni, D.; Sbordoni, V. Testing classical species properties with contemporary data: How “Bad Species” in the brassy ringlets (Erebia tyndarus complex, Lepidoptera) turned good. Syst. Biol. 2016, 65, 292–303. [Google Scholar] [CrossRef] [PubMed]
- Lorkovic, Z. Die Speziationsstufen in der Erebia tyndarus Gruppe. Biol. Glas. 1957, 10, 61–110. [Google Scholar]
- Kleckova, I.; Konvicka, M.; Klecka, J. Thermoregulation and microhabitat use in mountain butterflies of the genus Erebia: Importance of fine-scale habitat heterogeneity. J. Therm. Biol. 2014, 41, 50–58. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, J. Maximum-likelihood estimation of evolutionary trees from continuous characters. Am. J. Hum. Genet. 1973, 25, 471–492. [Google Scholar] [PubMed]
- Harmon, L.J.; Losos, J.B.; Jonathan Davies, T.; Gillespie, R.G.; Gittleman, J.L.; Bryan Jennings, W.; Kozak, K.H.; McPeek, M.A.; Moreno-Roark, F.; Near, T.J.; et al. Early bursts of body size and shape evolution are rare in comparative data. Evolution 2010, 64, 2385–2396. [Google Scholar] [CrossRef] [PubMed]
- Cressler, C.E.; Butler, M.A.; King, A.A. Detecting adaptive evolution in phylogenetic comparative analysis using the Ornstein-Uhlenbeck model. Syst. Biol. 2015, 64, 953–968. [Google Scholar] [CrossRef] [PubMed]
- Albre, J.; Gers, C.; Legal, L. Taxonomic notes on the species of the Erebia tyndarus group (Lepidoptera, Nymphalidae, Satyrinae). Lépidoptères 2008, 17, 12–28. [Google Scholar]
- Lanfear, R.; Frandsen, P.B.; Wright, A.M.; Senfeld, T.; Calcott, B. PartitionFinder 2: New methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol. Biol. Evol. 2017, 34, 772–773. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [PubMed]
- Rambaut, A.; Suchard, M.A.; Xie, D.; Drummond, A.J. Tracer v1.6. Available online: http://tree.bio.ed.ac.uk/software/tracer (accessed on 26 October 2017).
- Keck, F.; Rimet, F.; Bouchez, A.; Franc, A. Phylosignal: An R package to measure, test, and explore the phylogenetic signal. Ecol. Evol. 2016, 6, 2774–2780. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R Foundation for Statistical Computing; R 3.3.1; R Foundation for Statistical Computing: Vienna, Austria, 2016. [Google Scholar]
- Münkemüller, T.; Lavergne, S.; Bzeznik, B.; Dray, S.; Jombart, T.; Schiffers, K.; Thuiller, W. How to measure and test phylogenetic signal. Methods Ecol. Evol. 2012, 3, 743–756. [Google Scholar] [CrossRef]
- Diniz-Filho, J.A.F.; Santos, T.; Rangel, T.F.; Bini, L.M. A comparison of metrics for estimating phylogenetic signal under alternative evolutionary models. Genet. Mol. Biol. 2012, 35, 673–679. [Google Scholar] [CrossRef] [PubMed]
- Gustavo, P.; Werner, G.; Caterina, P. SensiPhy: Sensitivity Analysis for Comparative Methods. Available online: https://github.com/paternogbc/sensiPhy (accessed on 6 December 2017).
- Clavel, J.; Escarguel, G.; Merceron, G. mv morph: An R package for fitting multivariate evolutionary models to morphometric data. Methods Ecol. Evol. 2015, 6, 1311–1319. [Google Scholar] [CrossRef]
- Sites, J.W., Jr.; Moritz, C. Chromosomal evolution and speciation revisited. Syst. Biol. 1987, 36, 153–174. [Google Scholar] [CrossRef]
- Kandul, N.P.; Lukhtanov, V.A.; Pierce, N.E. Karyotypic diversity and speciation in Agrodiaetus butterflies. Evolution 2007, 61, 546–559. [Google Scholar] [CrossRef] [PubMed]
- Toews, D.P.L.; Brelsford, A. The biogeography of mitochondrial and nuclear discordance in animals. Mol. Ecol. 2012, 21, 3907–3930. [Google Scholar] [CrossRef] [PubMed]
- Pardo-Diaz, C.; Salazar, C.; Baxter, S.W.; Merot, C.; Figueiredo-Ready, W.; Joron, M.; Mcmillan, W.O.; Jiggins, C.D. Adaptive introgression across species boundaries in Heliconius butterflies. PLoS Genet. 2012, 8, e1002752. [Google Scholar] [CrossRef] [PubMed]
- Narita, S.; Nomura, M.; Kato, Y.; Fukatsu, T. Genetic structure of sibling butterfly species affected by Wolbachia infection sweep: Evolutionary and biogeographical implications. Mol. Ecol. 2006, 15, 1095–1108. [Google Scholar] [CrossRef] [PubMed]
- Gompert, Z.; Forister, M.L.; Fordyce, J.A.; Nice, C.C. Widespread mito-nuclear discordance with evidence for introgressive hybridization and selective sweeps in Lycaeides. Mol. Ecol. 2008, 17, 5231–5244. [Google Scholar] [CrossRef] [PubMed]
- Davies, T.J.; Kraft, N.J.B.; Salamin, N.; Wolkovich, E.M. Incompletely resolved phylogenetic trees inflate estimates of phylogenetic conservatism. Ecology 2012, 93, 242–247. [Google Scholar] [CrossRef] [PubMed]
- Cooper, N.; Thomas, G.H.; Venditti, C.; Meade, A.; Freckleton, R.P. A cautionary note on the use of Ornstein Uhlenbeck models in macroevolutionary studies. Biol. J. Linn. Soc. 2016, 118, 64–77. [Google Scholar] [CrossRef] [PubMed]
- Kandul, N.P.; Lukhtanov, V.A.; Dantchenko, A.V.; Coleman, J.W.S.; Sekercioglu, C.H.; Haig, D.; Pierce, N.E. Phylogeny of Agrodiaetus Hübner 1822 (Lepidoptera: Lycaenidae) inferred from mtDNA sequences of COI and COII and nuclear sequences of EF1α: Karyotype diversification and species radiation. Syst. Biol. 2004, 53, 278–298. [Google Scholar] [CrossRef] [PubMed]
- Butler, M.A.; King, A.A. Phylogenetic comparative analysis: A modeling approach for adaptive evolution. Am. Nat. 2004, 164, 683–695. [Google Scholar] [CrossRef]
- Molina-Venegas, R.; Rodríguez, M.Á. Revisiting phylogenetic signal; strong or negligible impacts of polytomies and branch length information? BMC Evol. Biol. 2017, 17, 53. [Google Scholar] [CrossRef] [PubMed]
- Saura, A.; Von Schoultz, B.; Saura, A.O.; Brown, K.S.J. Chromosome evolution in neotropical butterflies. Hereditas 2013, 150, 26–37. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lucek, K. Evolutionary Mechanisms of Varying Chromosome Numbers in the Radiation of Erebia Butterflies. Genes 2018, 9, 166. https://doi.org/10.3390/genes9030166
Lucek K. Evolutionary Mechanisms of Varying Chromosome Numbers in the Radiation of Erebia Butterflies. Genes. 2018; 9(3):166. https://doi.org/10.3390/genes9030166
Chicago/Turabian StyleLucek, Kay. 2018. "Evolutionary Mechanisms of Varying Chromosome Numbers in the Radiation of Erebia Butterflies" Genes 9, no. 3: 166. https://doi.org/10.3390/genes9030166
APA StyleLucek, K. (2018). Evolutionary Mechanisms of Varying Chromosome Numbers in the Radiation of Erebia Butterflies. Genes, 9(3), 166. https://doi.org/10.3390/genes9030166




