Fungal Screening on Olive Oil for Extracellular Triacylglycerol Lipases: Selection of a Trichoderma harzianum Strain and Genome Wide Search for the Genes
Abstract
1. Introduction
2. Materials and Methods
2.1. Isolation of Lipolytic Microorganisms
2.2. Screening of Lipolytic Fungi
2.3. Lipase Production by Submerged Fermentation
2.4. Molecular Identification of Selected Lipolytic Fungi Isolated Here
2.5. Searching of Genes of True Lipases in Trichoderma harzianum Genome
2.6. In Silico Analysis of Trichoderma harzianum Putative True Lipases
2.7. Phylogenetic Analysis
2.8. In Silico Modeling
2.9. RT-PCR Amplification of Lipases
3. Results and Discussion
3.1. Screening of Lipolytic Fungi Using Olive Oil
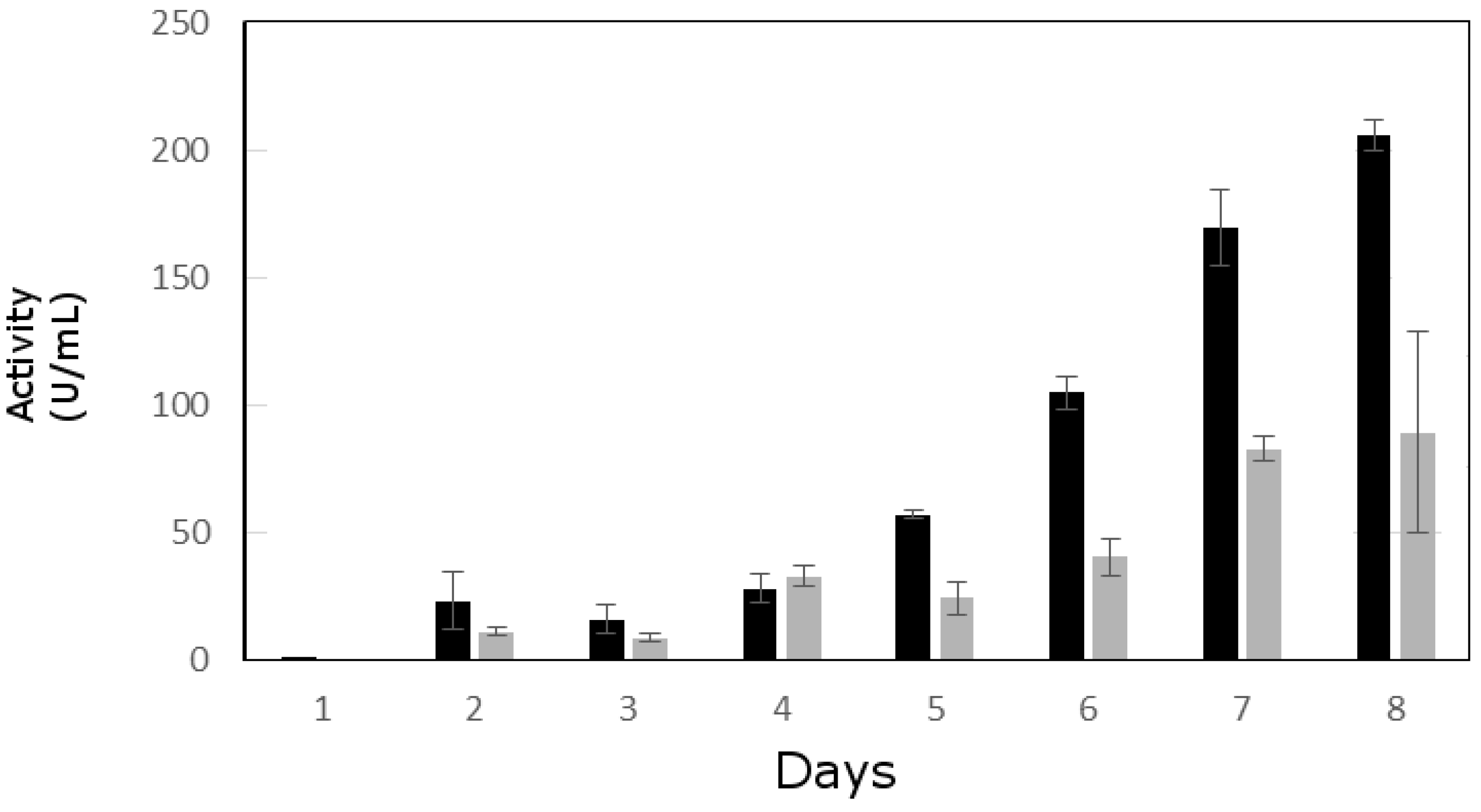
3.2. Lipolytic Activity Assay
3.3. Searching Genes of True Lipases in Trichoderma harzianum
3.4. In Silico Characterization of Trichoderma harzianum True Lipases
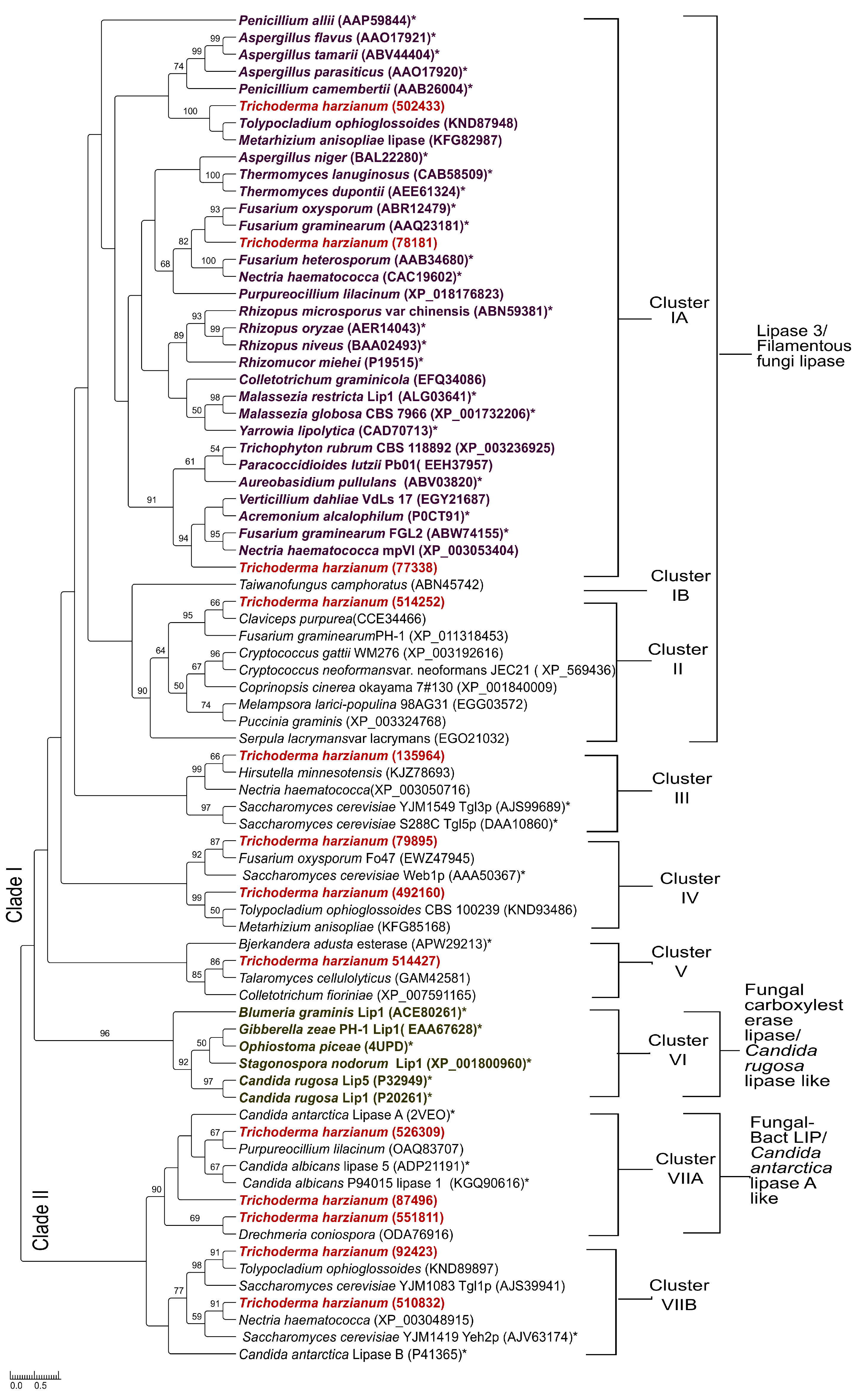
3.5. Phylogenetic Analysis
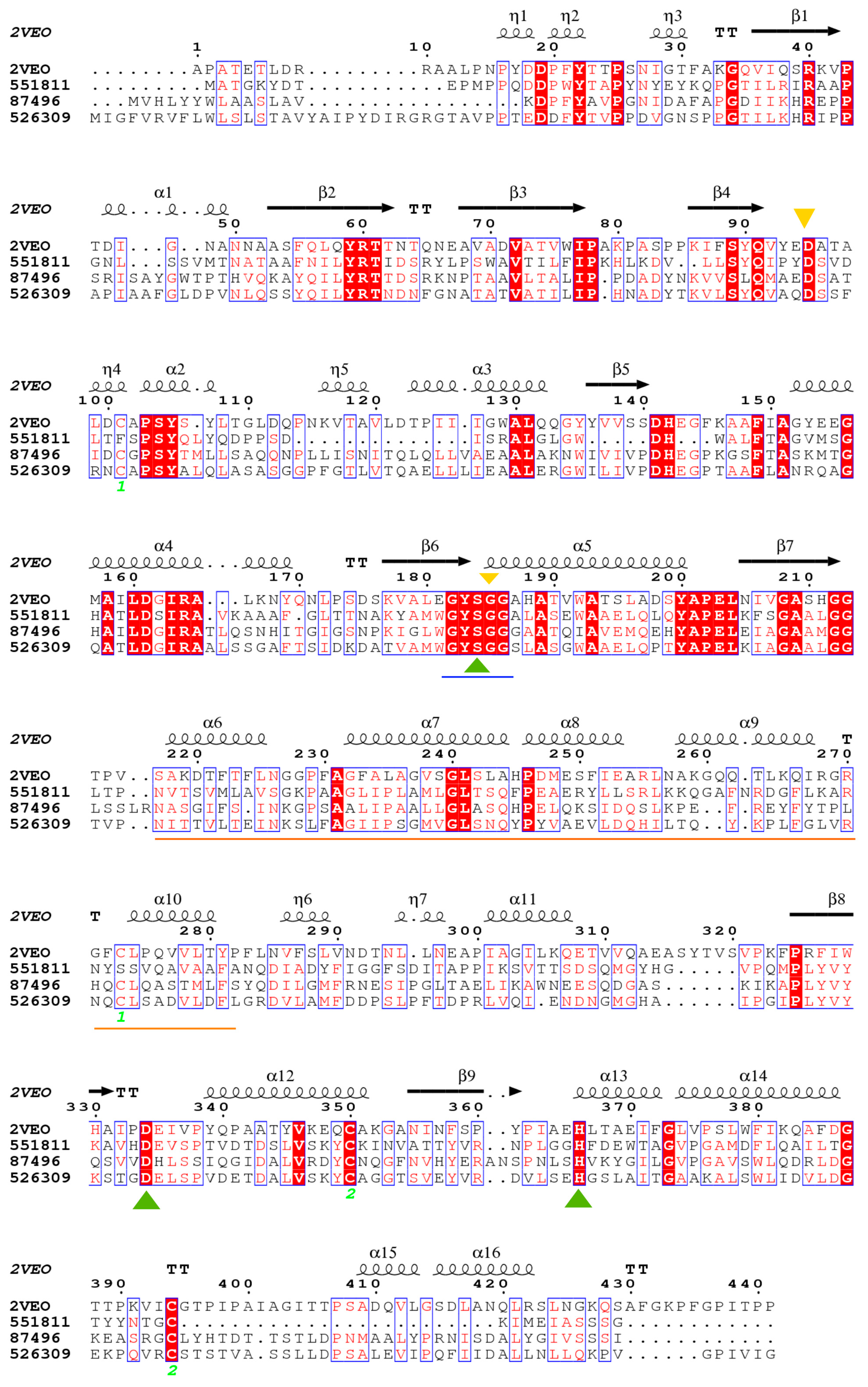
3.6. Structural Modeling
3.7. Functional Prediction of Candidates
3.8. Refining of Selection of Candidates
3.9. RT-PCR Analysis for Validation of Predicting Results
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gopinath, S.C.B.; Anbu, P.; Lakshmipriya, T.; Hilda, A. Strategies to characterize fungal lipases for applications in medicine and dairy industry. BioMed Res. Int. 2013, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Verger, R. ‘Interfacial activation’ of lipases: Facts and artifacts. Trends Biotechnol. 1997, 15, 32–38. [Google Scholar] [CrossRef]
- Nwuche, C.O.; Ogbonna, J.C. Isolation of lipase producing fungi from palm oil Mill effluent (POME) dump sites at Nsukka. Braz. Arch. Biol. Technol. 2011, 54, 113–116. [Google Scholar] [CrossRef]
- Daiha, K.; Angeli, R.; de Oliveira, S.D.; Almeida, R.V. Are Lipases Still Important Biocatalysts? A Study of Scientific Publications and Patents for Technological Forecasting. PLoS ONE 2015, 10, e0131624. [Google Scholar] [CrossRef] [PubMed]
- Andualema, B.; Gessesse, A. Microbial lipases and their industrial applications. Biotechnology 2012, 11, 100–118. [Google Scholar] [CrossRef]
- Yadav, S.; Dubey, A.K.; Yadav, S.; Bisht, D.; Singh-Darmwal, N.; Yadav, D. Amino acid sequences based phylogenetic and motif assessment of lipases from different organisms. Online J. Bioinform. 2012, 13, 400–417. [Google Scholar]
- Lanka, S.; Latha, J.N.L. Purification and characterization of a new cold active lipase, EnL A from Emericella nidulans NFCCI 3643. Afr. J. Biotechnol. 2015, 14, 1897–1909. [Google Scholar] [CrossRef]
- Ortiz-Lechuga, E.G.; Quintero-Zapata, I.; Niño, K.A. Detection of extracellular enzymatic activity in microorganisms isolated from waste vegetable oil contaminated soil using plate methodologies. Afr. J. Biotechnol. 2016, 15, 408–416. [Google Scholar] [CrossRef]
- Coradi, G.V.; da Visitação, V.L.; de Lima, E.A.; Saito, L.Y.T.; Palmieri, D.A.; Takita, M.A.; Neto, P.O.; de Lima, V.M.G. Comparing submerged and solid-state fermentation of agro-industrial residues for the production and characterization of lipase by Trichoderma harzianum. Ann. Microbiol. 2013, 63, 533–540. [Google Scholar] [CrossRef]
- Falony, G.; Armas, J.C.; Mendoza, J.C.D.; Hernández, J.L.M. Production of extracellular lipase from Aspergillus niger by solid-state fermentation. Food Technol. Biotechnol. 2006, 44, 235–240. [Google Scholar]
- Lima, V.M.G.; Krieger, N.; Sarquis, M.I.M.; Mitchell, D.A.; Ramos, L.P.; Fontana, J.D. Effect of Nitrogen and Carbon Sources on Lipase Production by Penicillium aurantiogriseum. FTB J. 2003, 41, 105–110. [Google Scholar]
- Ülker, S.; Özel, A.; Colak, A.; Karaoğlu, Ş.A. Isolation, production, and characterization of an extracellular lipase from Trichoderma harzianum isolated from soil. Turk. J. Biol. 2011, 35, 543–550. [Google Scholar] [CrossRef]
- Mier, T.; Toriello, C.; Ulloa, M. Hongos Microscópicos Saprobios y Parásitos: Métodos de Laboratorio; Universidad Autónoma Metropolitana, Unidad Xochimilco: México D.F., Mexico, 2002; pp. 1–90. ISBN 978-970-654-609-8. [Google Scholar]
- Panuthai, T.; Sihanonth, P.; Piapukiew, J.; Sooksai, S.; Sangvanich, P.; Karnchanatat, A. An extracellular lipase from the endophytic fungi Fusarium oxysporum isolated from the Thai medicinal plant, Croton oblongifolius Roxb. Afr. J. Microbiol. Res. 2012, 6, 2622–2638. [Google Scholar] [CrossRef]
- Carissimi, M.; de Souza, T.F.; Corbellini, V.A.; Scroferneker, M.L. Comparison of lipolytic activity of Sporothrix schenckii strains utilizing olive oil-rhodamine B and Tween 80. Tecno-Lógica 2007, 11, 33–36. [Google Scholar]
- Maia, M.d.M.D.; Morais, M.M.C.d.; Morais, M.A.d., Jr.; Melo, E.H.M.; de Lima Filho, J.L. Production of extracellular lipase by the phytopathogenic fungus Fusarium solani FS1. Rev. Microbiol. 1999, 30, 304–309. [Google Scholar] [CrossRef]
- Winkler, U.K.; Stuckmann, M. Glycogen, hyaluronate, and some other polysaccharides greatly enhance the formation of exolipase by Serratia marcescens. J. Bacteriol. 1979, 138, 663–670. [Google Scholar] [PubMed]
- Pacheco, S.; Júnior, A.; Morgado, A.; Júnior, A.; Amadi, O.; Guisán, J.; Pessela, B. Isolation and Screening of Filamentous Fungi Producing Extracellular Lipase with Potential in Biodiesel Production. Adv. Enzyme Res. 2015, 03, 101–114. [Google Scholar] [CrossRef]
- Johanson, A.; Jeger, M.J. Use of PCR for detection of Mycosphaerella fijiensis and M. musicola, the causal agents of Sigatoka leaf spots in banana and plantain. Mycol. Res. 1993, 97, 670–674. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J.L. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protoc. Guide Methods Appl. 1990, 18, 315–322. [Google Scholar]
- Hall, T. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Okonechnikov, K.; Golosova, O.; Fursov, M. Unipro UGENE: A unified bioinformatics toolkit. Bioinformatics 2012, 28, 1166–1167. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- NCBI Resource Coordinators. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2016, 44, D7–D19. [Google Scholar] [CrossRef]
- Druzhinina, I.S.; Kopchinskiy, A.G.; Komoń, M.; Bissett, J.; Szakacs, G.; Kubicek, C.P. An oligonucleotide barcode for species identification in Trichoderma and Hypocrea. Fungal Genet. Biol. 2005, 42, 813–828. [Google Scholar] [CrossRef] [PubMed]
- Nordberg, H.; Cantor, M.; Dusheyko, S.; Hua, S.; Poliakov, A.; Shabalov, I.; Smirnova, T.; Grigoriev, I.V.; Dubchak, I. The genome portal of the Department of Energy Joint Genome Institute: 2014 updates. Nucleic Acids Res. 2014, 42, D26–D31. [Google Scholar] [CrossRef] [PubMed]
- Voigt, C.A.; Schäfer, W.; Salomon, S. A secreted lipase of Fusarium graminearum is a virulence factor required for infection of cereals: Lipase as a virulence factor. Plant J. 2005, 42, 364–375. [Google Scholar] [CrossRef] [PubMed]
- Nagao, T.; Shimada, Y.; Sugihara, A.; Tominaga, Y. Cloning and Nucleotide Sequence of cDNA Encoding a Lipase from Fusarium heterosporum. J. Biochem. 1994, 116, 536–540. [Google Scholar] [CrossRef] [PubMed]
- Haraldsson, G.G.; Kristinsson, B. Process for Separating Polyunsaturated Fatty Acids from Long Chain Unsaturated or Less Saturated Fatty Acids. U.S. Patent 9476008, 25 October 2016. [Google Scholar]
- Marchler-Bauer, A.; Bo, Y.; Han, L.; He, J.; Lanczycki, C.J.; Lu, S.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.-R.; et al. CDD/SPARCLE: Functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2017, 45, D200–D203. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef] [PubMed]
- Alva, V.; Nam, S.Z.; Söding, J.; Lupas, A.N. The MPI bioinformatics Toolkit as an integrative platform for advanced protein sequence and structure analysis. Nucleic Acids Res. 2016, 44, W410–W415. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Attwood, T.K.; Babbitt, P.C.; Bateman, A.; Bork, P.; Bridge, A.J.; Chang, H.-Y.; Dosztányi, Z.; El-Gebali, S.; Fraser, M.; et al. InterPro in 2017—beyond protein family and domain annotations. Nucleic Acids Res. 2017, 45, D190–D199. [Google Scholar] [CrossRef] [PubMed]
- Petersen, T.N.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785. [Google Scholar] [CrossRef] [PubMed]
- Krogh, A.; Larsson, B.; von Heijne, G.; Sonnhammer, E.L. Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. J. Mol. Biol. 2001, 305, 567–580. [Google Scholar] [CrossRef] [PubMed]
- Horton, P.; Park, K.-J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef] [PubMed]
- Protein Molecular Weight Calculator. Available online: http://www.sciencegateway.org/tools/proteinmw.htm (accessed on 27 December 2017).
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server. In The Proteomics Protocols Handbook; Walker, J.M., Ed.; Humana Press: Totowa, NJ, USA, 2005; pp. 571–607. ISBN 978-1-59259-890-8. [Google Scholar]
- Marchler-Bauer, A.; Derbyshire, M.K.; Gonzales, N.R.; Lu, S.; Chitsaz, F.; Geer, L.Y.; Geer, R.C.; He, J.; Gwadz, M.; Hurwitz, D.I.; et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 2015, 43, D222–D226. [Google Scholar] [CrossRef] [PubMed]
- Fischer, M.; Pleiss, J. The Lipase Engineering Database: A navigation and analysis tool for protein families. Nucleic Acids Res. 2003, 31, 319–321. [Google Scholar] [CrossRef] [PubMed]
- Hotelier, T.; Renault, L.; Cousin, X.; Negre, V.; Marchot, P.; Chatonnet, A. ESTHER, the database of the α/β-hydrolase fold superfamily of proteins. Nucleic Acids Res. 2004, 32, D145–D147. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Schmidt-Dannert, C. Recombinant microbial lipases for biotechnological applications. Bioorg. Med. Chem. 1999, 7, 2123–2130. [Google Scholar] [CrossRef]
- Feng, J.; Guosheng, L.; Gopalan, S.; Geoffrey, R.; Wei, Y. A secreted lipase encoded by LIP1 is necessary for efficient use of saturated triglyceride lipids in Fusarium graminearum. Microbiology 2005, 151, 3911–3921. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Wang, F.; Liu, G.; Greenshields, D.; Shen, W.; Kaminskyj, S.; Hughes, G.R.; Peng, Y.; Selvaraj, G.; Zou, J.; et al. Analysis of a Blumeria graminis-secreted lipase reveals the importance of host epicuticular wax components for fungal adhesion and development. Mol. Plant-Microbe Interact. MPMI 2009, 22, 1601–1610. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Hwang, R.; Hwang, S.-F.; Gaudet, D.; Strelkov, S.E. Molecular characterization of a Stagonospora nodorum lipase gene LIP1: A triglyceride lipase from Stagonospora nodorum. Plant Pathol. 2011, 60, 698–708. [Google Scholar] [CrossRef]
- Park, M.; Jung, W.H.; Han, S.H.; Lee, Y.H.; Lee, Y.W. Characterisation and Expression Analysis of MrLip1, a Class 3 Family Lipase of Malassezia restricta. Mycoses 2015, 58, 671–678. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Saunders, C.W.; Hu, P.; Grant, R.A.; Boekhout, T.; Kuramae, E.E.; Kronstad, J.W.; DeAngelis, Y.M.; Reeder, N.L.; Johnstone, K.R.; et al. Dandruff-associated Malassezia genomes reveal convergent and divergent virulence traits shared with plant and human fungal pathogens. Proc. Natl. Acad. Sci. USA 2007, 104, 18730–18735. [Google Scholar] [CrossRef] [PubMed]
- Pereira, E.; Tsang, A.; McAllister, T.A.; Menassa, R. The production and characterization of a new active lipase from Acremonium alcalophilum using a plant bioreactor. Biotechnol. Biofuels 2013, 6, 111. [Google Scholar] [CrossRef] [PubMed]
- Klein, I.; Klug, L.; Schmidt, C.; Zandl, M.; Korber, M.; Daum, G.; Athenstaedt, K. Regulation of the yeast triacylglycerol lipases Tgl4p and Tgl5p by the presence/absence of nonpolar lipids. Mol. Biol. Cell 2016, 27, 2014–2024. [Google Scholar] [CrossRef] [PubMed]
- Cedillo, V.; Plou, F.J.; Martínez, M. Recombinant sterol esterase from Ophiostoma piceae: An improved biocatalyst expressed in Pichia pastoris. Microb. Cell Factories 2012, 11, 73. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Carbente, M.R.; Batista-García, R.A.; Sánchez-Reyes, A.; Escudero-Garcia, A.; Morales-Herrera, C.; Cuervo-Soto, L.I.; French-Pacheco, L.; Fernández-Silva, A.; Amero, C.; Castillo, E.; et al. The first description of a hormone-sensitive lipase from a basidiomycete: Structural insights and biochemical characterization revealed Bjerkandera adusta Ba EstB as a novel esterase. MicrobiologyOpen 2017, 6, e00463. [Google Scholar] [CrossRef]
- Katoh, K.; Toh, H. Recent developments in the MAFFT multiple sequence alignment program. Brief. Bioinform. 2008, 9, 286–298. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef] [PubMed]
- Biasini, M.; Bienert, S.; Waterhouse, A.; Arnold, K.; Studer, G.; Schmidt, T.; Kiefer, F.; Cassarino, T.G.; Bertoni, M.; Bordoli, L.; et al. SWISS-MODEL: Modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 2014, 42, W252–W258. [Google Scholar] [CrossRef] [PubMed]
- Söding, J.; Biegert, A.; Lupas, A.N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 2005, 33, W244–W248. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER Suite: Protein structure and function prediction. Nat. Methods 2015, 12, 7–8. [Google Scholar] [CrossRef] [PubMed]
- Gelly, J.-C.; Joseph, A.P.; Srinivasan, N.; de Brevern, A.G. iPBA: A tool for protein structure comparison using sequence alignment strategies. Nucleic Acids Res. 2011, 39, W18–W23. [Google Scholar] [CrossRef] [PubMed]
- DeLano, W.L. The PyMol Molecular Graphics System; DeLano Scientific LLC: Palo Alto, CA, USA, 2002. [Google Scholar]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Robert, X.; Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 2014, 42, W320–W324. [Google Scholar] [CrossRef] [PubMed]
- PrimerQuest Tool. Available online: https://www.idtdna.com/Primerquest/Home/Index (accessed on 27 December 2017).
- Rifaat, H.M.; El-Mahalawy, A.A.; El-Menofy, H.A.; Donia, S.A. Production, optimization and partial purification of lipase from Fusarium oxysporum. J. Appl. Sci. Environ. Sanit. 2010, 5, 39–53. [Google Scholar]
- Savitha, J.; Srividya, S.; Jagat, R.; Payal, P.; Priyanki, S.; Rashmi, G.W.; Roshini, K.T.; Shantala, Y.M. Identification of potential fungal strain (s) for the production of inducible, extracellular and alkalophilic lipase. Afr. J. Biotechnol. 2007, 6, 564. [Google Scholar]
- Winayanuwattikun, P.; Kaewpiboon, C.; Piriyakananon, K.; Chulalaksananukul, W.; Yongvanich, T.; Svasti, J. Immobilized lipase from potential lipolytic microbes for catalyzing biodiesel production using palm oil as feedstock. Afr. J. Biotechnol. 2011, 10, 1666–1673. [Google Scholar]
- Jayaprakash, A.; Ebenezer, P. Investigation on Extracellular Lipase Production by Aspergillus japonicus Isolated from the Paper Nest of Ropalidia marginata. Indian J. Sci. Technol. 2010, 3, 113–117. [Google Scholar] [CrossRef]
- Peil, G.H.S.; Kuss, A.V.; Rave, A.F.G.; Villarreal, J.P.V.; Hernandes, Y.M.L.; Nascente, P.S. Bioprospecting of lipolytic microorganisms obtained from industrial effluents. An. Acad. Bras. Ciênc. 2016, 88, 1769–1779. [Google Scholar] [CrossRef] [PubMed]
- Rabbani, M.; Shafiee, F.; Shayegh, Z.; Sadeghi, H.M.M.; Shariat, Z.S.; Etemadifar, Z.; Moazen, F. Isolation and Characterization of a New Thermoalkalophilic Lipase from Soil Bacteria. Iran. J. Pharm. Res. IJPR 2015, 14, 901–906. [Google Scholar] [PubMed]
- Shafei, M.S.; Allam, R.F. Production and immobilization of partially purified lipase from Penicillium chrysogenum. Malays. J. Microbiol. 2010, 6, 196–202. [Google Scholar]
- Hiol, A.; Jonzo, M.D.; Rugani, N.; Druet, D.; Sarda, L.; Comeau, L.C. Purification and characterization of an extracellular lipase from a thermophilic Rhizopus oryzae strain isolated from palm fruit. Enzyme Microb. Technol. 2000, 26, 421–430. [Google Scholar] [CrossRef]
- Kantak, J.B.; Bagade, A.V.; Mahajan, S.A.; Pawar, S.P.; Shouche, Y.S.; Prabhune, A.A. Isolation, Identification and Optimization of a New Extracellular Lipase Producing Strain of Rhizopus sp. Appl. Biochem. Biotechnol. 2011, 164, 969–978. [Google Scholar] [CrossRef] [PubMed]
- Mehta, A.; Kumar-Saun, N.; Gupta, R. Purification and Characterization of Lipase from thermophilic Geobacillus sp. Curr. Biotechnol. 2016, 5, 81–89. [Google Scholar] [CrossRef]
- Sharma, D.; Sharma, B.; Shukla, A.K. Biotechnological approach of microbial lipase: A review. Biotechnology 2011, 10, 23–40. [Google Scholar] [CrossRef]
- Barriuso, J.; Martínez, M.J. Evolutionary history of versatile-lipases from Agaricales through reconstruction of ancestral structures. BMC Genom. 2017, 18. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.; Kumari, A.; Syal, P.; Singh, Y. Molecular and functional diversity of yeast and fungal lipases: Their role in biotechnology and cellular physiology. Prog. Lipid Res. 2015, 57, 40–54. [Google Scholar] [CrossRef] [PubMed]
- Zan, X.; Tang, X.; Chu, L.; Zhao, L.; Chen, H.; Chen, Y.Q.; Chen, W.; Song, Y. Lipase genes in Mucor circinelloides: Identification, sub-cellular location, phylogenetic analysis and expression profiling during growth and lipid accumulation. J. Ind. Microbiol. Biotechnol. 2016, 43, 1467–1480. [Google Scholar] [CrossRef] [PubMed]
- Ericsson, D.J.; Kasrayan, A.; Johansson, P.; Bergfors, T.; Sandström, A.G.; Bäckvall, J.-E.; Mowbray, S.L. X-ray Structure of Candida antarctica Lipase A Shows a Novel Lid Structure and a Likely Mode of Interfacial Activation. J. Mol. Biol. 2008, 376, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Roussel, A.; Miled, N.; Berti-Dupuis, L.; Rivière, M.; Spinelli, S.; Berna, P.; Gruber, V.; Verger, R.; Cambillau, C. Crystal Structure of the Open Form of Dog Gastric Lipase in Complex with a Phosphonate Inhibitor. J. Biol. Chem. 2002, 277, 2266–2274. [Google Scholar] [CrossRef] [PubMed]
- Selvan, A.; Seniya, C.; Chandrasekaran, S.N.; Siddharth, N.; Anishetty, S.; Pennathur, G. Molecular dynamics simulations of human and dog gastric lipases: Insights into domain movements. FEBS Lett. 2010, 584, 4599–4605. [Google Scholar] [CrossRef] [PubMed]
- Dror, A.; Kanteev, M.; Kagan, I.; Gihaz, S.; Shahar, A.; Fishman, A. Structural insights into methanol-stable variants of lipase T6 from Geobacillus stearothermophilus. Appl. Microbiol. Biotechnol. 2015, 99, 9449–9461. [Google Scholar] [CrossRef] [PubMed]
- Bordes, F.; Barbe, S.; Escalier, P.; Mourey, L.; André, I.; Marty, A.; Tranier, S. Exploring the Conformational States and Rearrangements of Yarrowia lipolytica Lipase. Biophys. J. 2010, 99, 2225–2234. [Google Scholar] [CrossRef] [PubMed]
- Lou, Z.; Li, M.; Sun, Y.; Liu, Y.; Liu, Z.; Wu, W.; Rao, Z. Crystal structure of a secreted lipase from Gibberella zeae reveals a novel “double-lock” mechanism. Protein Cell 2010, 1, 760–770. [Google Scholar] [CrossRef] [PubMed]
- Casas-Godoy, L.; Duquesne, S.; Bordes, F.; Sandoval, G.; Marty, A. Lipases: An Overview. In Lipases and Phospholipases; Methods in Molecular Biology; Humana Press: New York, NY, USA, 2012; pp. 3–30. ISBN 978-1-61779-599-2. [Google Scholar]
- Kumari, A.; Gupta, R. Functional Characterisation of Novel Enantioselective Lipase TALipA from Trichosporon asahii MSR54: Sequence Comparison Revealed New Signature Sequence AXSXG Among Yeast Lipases. Appl. Biochem. Biotechnol. 2015, 175, 360–371. [Google Scholar] [CrossRef] [PubMed]
- Ham, H.J.; Rho, H.J.; Shin, S.K.; Yoon, H.-J. The TGL2 gene of Saccharomyces cerevisiae encodes an active acylglycerol lipase located in the mitochondria. J. Biol. Chem. 2010, 285, 3005–3013. [Google Scholar] [CrossRef] [PubMed]
- Najjar, A.; Robert, S.; Guérin, C.; Violet-Asther, M.; Carrière, F. Quantitative study of lipase secretion, extracellular lipolysis, and lipid storage in the yeast Yarrowia lipolytica grown in the presence of olive oil: Analogies with lipolysis in humans. Appl. Microbiol. Biotechnol. 2011, 89, 1947–1962. [Google Scholar] [CrossRef] [PubMed]
- Eddine, A.N.; Hannemann, F.; Schafer, W. Cloning and expression analysis of NhL1, a gene encoding an extracellular lipase from the fungal pea pathogen Nectria haematococca MP VI (Fusarium solani f. sp. pisi) that is expressed in planta. Mol. Genet. Genom. 2001, 265, 215–224. [Google Scholar] [CrossRef]
- Nguyen, L.N.; Bormann, J.; Le, G.T.T.; Stärkel, C.; Olsson, S.; Nosanchuk, J.D.; Giese, H.; Schäfer, W. Autophagy-related lipase FgATG15 of Fusarium graminearum is important for lipid turnover and plant infection. Fungal Genet. Biol. 2011, 48, 217–224. [Google Scholar] [CrossRef] [PubMed]
- Widmann, M.; Juhl, P.B.; Pleiss, J. Structural classification by the Lipase Engineering Database: A case study of Candida antarctica lipase A. BMC Genom. 2010, 11, 123. [Google Scholar] [CrossRef] [PubMed]
- Jaeger, K.E.; Ransac, S.; Dijkstra, B.W.; Colson, C.; van Heuvel, M.; Misset, O. Bacterial lipases. FEMS Microbiol. Rev. 1994, 15, 29–63. [Google Scholar] [CrossRef] [PubMed]
- Brenneis, R.; Baeck, B. Esterification of fatty acids using Candida antarctica lipase A in water-abundant systems. Biotechnol. Lett. 2012, 34, 1459–1463. [Google Scholar] [CrossRef] [PubMed]
- Messaoudi, A.; Belguith, H.; Ghram, I.; Hamida, J.B. LIPABASE: A database for “true” lipase family enzymes. Int. J. Bioinform. Res. Appl. 2011, 7, 390. [Google Scholar] [CrossRef] [PubMed]

| Strain | Morphology | Extracellular Lipase at 48 h * |
|---|---|---|
| A04-5 | Black, concentric rings | + |
| A06-6 | White, radial growth | + |
| B07(+)-1(3)N | White, radial growth | - |
| B08-6 | White, radial growth and Green spores | + |
| B09-4 | White, cottony mycelia | - |
| B09-5 | White, cottony mycelia | ++ |
| B09-8 | White, radial growth | ++ |
| B10-4(b1)Emmb | Black, radial growth | - |
| B10(+)-2(4) | White, radial growth | - |
| B10-4(1)-2(1) | Black, radial growth | - |
| B11-6 | Fuchsia, cottony mycelia | ++ |
| B11-7 | Fuchsia, cottony mycelia | ++ |
| B13-1 | White with green concentric rings | +++ |
| B13-3 | White with green concentric rings | +++ |
| B13-4 | Yellow, compact colony | ++ |
| B14-6 | White, radial growth | ++ |
| B17(+)-4(3) | Fuchsia, cottony mycelia | ++ |
| B19-01-3(3) | White, radial growth | + |
| Protein ID | Superfamily | Domains | Domain Description | Active Site Domain | Substrate Binding Pocket | BLAST Hits (Search Homologues in the First 50 Hits) | PSI-BLAST (Homologues Characterized) |
|---|---|---|---|---|---|---|---|
| 551811 | Abhydrolase family cl21494 | LIP pfam03583 Mal_quin-oxido TIGR01320 | Secretory lipase: related with lipases from Candida albicans. Malate:quinone-oxidoreductase: Membrane-associated enzyme as part of the TCA cycle | Active site Ser-His-Asp/Glu. Nucleophilic attack on a carbonyl carbon atom. | Substrate binding pocket related to pfam03583 | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Metharhizium sp., Fusarium sp.) as hypothetical protein and Predicted lipases | Lipase 1 C. albicans (KGR02689) Query cover 94% Identity 33% |
| 87496 | Abhydrolase family cl21494 | LIP pfam03583 | Secretory lipase: Related with lipases from C. albicans | Active site Ser-His-Asp/Glu. Nucleophilic attack on a carbonyl carbon atom. | Substrate binding pocket related to pfam03583 | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Metharhizium sp., Fusarium sp.) as hypothetical protein and Predicted lipases | Lipase 1 C. albicans (KGR02689) Query cover 93% Identity 32% |
| 526309 | Abhydrolase family cl21494 | LIP pfam03583 | Secretory lipase: Related with lipases from C. albicans | Active site Ser-His-Asp/Glu. Nucleophilic attack on a carbonyl carbon atom. | Substrate binding pocket related to pfam03583 | GH16 protein [Trichoderma guizhouense]. Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Metharhizium sp., Fusarium sp., Pochonia sp., Cordyceps sp., etc.) as hypothetical protein. Secretory lipase. Probable lipase precursor | Secretory lipase 5 C. albicans (ADP21191.1) Query cover 92% Identity 39% |
| 514427 | Abhydrolase family cl21494 | LIP pfam03583 DAP2 | Secretory lipase; Related with lipases from C. albicans Dipeptidyl aminopeptidase/acylaminoacyl peptidase:Amino acid transport and metabolism Acetyl xylan esterase (AXE1). | Active site Ser-His-Asp/Glu. Nucleophilic attack on a carbonyl carbon atom. | Substrate binding pocket related to pfam03583 | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Metharhizium sp., Fusarium sp.) as hypothetical protein and prolyl aminopeptidase (secreted protein). | No characterized protein among the first 500 hits |
| 510832 | Alpha/Beta hydrolase fold cl26327 | PLN02872 Abhydro_lipase (pfam04083) Mhpc (COG0596) | Triacylglycerol lipase Partial alpha/beta hydrolase lipase region: Pimeloyl-ACP methyl ester carboxylesterase | - | - | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Metharhizium sp., Fusarium sp., Colletotrichum sp., etc.) as hypothetical protein, triglyceride lipase-cholesterol esterase, alpha/beta-hydrolase, Sterol esterase 2. | Yeh2p Saccharomyces cerevisiae YJM1419 (AJV63174). Query cover 58% Identity 35% |
| 92423 | Alpha/Beta hydrolase fold cl26327 | PLN02872 Abhydro_lipase (pfam04083) MhpC (Cpg0596) | Triacylglycerol lipase Partial alpha/beta-hydrolase lipase region Pimeloyl-ACP methyl ester carboxylesterase | - | - | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Metharhizium sp., Fusarium sp., Colletotrichum sp., etc.) as steryl ester lipase TPL1, lysosomal acid lipase/cholesteryl ester hydrolase, triglyceride lipase-cholesterol esterase, hypothetical protein, alpha/beta hydrolase fold-1 | Tgl1p S. cerevisiae YJM1083 (AJS39941) Query cover 80% Identity 43% |
| 79895 | WD40 Cl25539 | WD40 (cd00200) WD40 (COG2319) WD40 (smart00320) Wd40 (pfam00400) PHA03247 Atrophin-1 (pfam03154) PLN00171 Amelogenin (smart00818) PABP-1234 (TIGR01628) | WD40 domain, found in many eukaryotic proteins with a wide variety of functions. Ancestral coatomer element 1 (ACE1) of COPII with role in vesicular traffic. Atrophin-1 family domain Protein SPA1-related Cell adhesion proteins Polyadenylate binding protein: | - | - | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Metharhizium sp., Fusarium sp., Colletotrichum sp., etc.) as hypothetical protein, Vesicle coat complex COPII, Sec31, transport protein sec31, transporter. | Web1p S. cevevisiae (AAA50367) Query cover 99% Identity 31% |
| 135964 | Abhydrolase family cl21494. | EstA (COG1075) PGAP1 (pfam07819) | Triacylglycerol esterase/lipase PGAP1-like protein | Active site Ser-His-Asp/Glu. Nucleophilic attack on a carbonyl carbon atom. | - | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Hirsutella sp., Stachybotrys sp., Fusarium sp., Ophiocordyceps sp., Colletotrichum sp., etc.) as hypothetical protein, Triacylglycerol lipase, Lipase 2, related to TGL2-triacylglycerol lipase, GPI inositol-deacylase PGAP1-like protein, PGAP1-like protein lipase. | No characterized protein among the first 500 hits |
| 492160 | DRF_GBD Superfamily Cl05720 DRF_FH3 Superfamily Cl05717 FH2 Superfamily cl19758 | Drf_GBD (pfam06371) Drf_FH3 (pfam06367) FH2 (pfam02181) FH2 (smart00498) PHA03307 | Diaphanous GTPase-binding domain; Rho proteins, leading to activation of the Drf protein. Formin Homology 2 Domain Involved in rearrangements of the actin cytoskeleton. Transcriptional regulator ICP4-like | - | - | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Tolypocladium sp., Pochonia sp., Metharhizium sp., Fusarium sp., Colletotrichum sp., etc.) as cytokinesis protein sepA, Rho-GTPase effector BNI1 and related formin, involved in mating, karyogamy and meiosis, hypothetical protein, SepA/Bni1 | PSI-BLAST analysis did not run |
| 77338 | Abhydrolase family cl21494. | Lipase_3 (cd00519) Lipase_3 (pfam001764) PLN02310 Lip2 (COG3675) | Lipases (Class 3). Lipase that can hydrolyze long-chain acyl-triglycerides into di- and monoglycerides, glycerol and free fatty acids. | Active site Ser-His-Asp/Glu. Nucleophilic elbow on conserved domain Lipase_3: GHSLG | Active site flap/lid on conserved domain Lipase_3 (11 amino acids). | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Tolypocladium sp., Pochonia sp., Metharhizium sp., Fusarium sp., Colletotrichum sp., etc.) as lipase, hypothetical protein, triacylglycerol lipase, Mono- and diacylglycerol lipase, Putative feruloyl esterase A, Alpha/Beta hydrolase protein. | Triacylglycerol lipase FGL2 Fusarium graminearum (ABW74155) Query cover 89% Identity 62% |
| 514252 | Lipase (class 3). Alpha/beta hydrolase cd00519 | Lipase_3 (pfam01764) Lipase_3 (cd00519) CVT17 (COG5153) PRK11071 | Lipase that can hydrolyze long-chain acyl-triglycerides into di- and monoglycerides, glycerol and free fatty acids. Putative lipase essential for disintegration of autophagic bodies inside the vacuole Esterase YqiA | Active site Ser-His-Asp/Glu. Nucleophilic elbow on conserved domain Lipase_3: GHSLG | Active site flap/lid on conserved domain Lipase_3 (11 amino acids). | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Tolypocladium sp., Pochonia sp., Metharhizium sp., Fusarium sp., Colletotrichum sp., etc.) as autophagy related lipase Atg15, triacylglycerol lipase, hypothetical protein, alpha/beta-hydrolase, related to starvation induced protein PSI-7, autophagy lipase. | Hypothetical protein FGSG_02519 F. graminearum PH-1 (XP_011318453) Query cover 92% Identity 70% |
| 502433 | Predicted triacylglycerol lipase activity, Lipase_3. | Lipase 3 (cd00519) Lipase_3 (pfam01764) PLN02847 AF-4 (pfam05110) | Lipase that can hydrolyze long-chain acyl-triglycerides into di- and monoglycerides, glycerol and free fatty acids. AF-4 Proto-oncoprotein; Nuclear proteins linked to human disease | Active site Ser-His-Asp/Glu. Nucleophilic elbow on conserved domain Lipase_3: GHSLG | Active site flap/lid on conserved domain Lipase_3 (11 amino acids). | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Tolypocladium sp., Pochonia sp., Metharhizium sp., Ophiocordyceps sp., Fusarium sp., Colletotrichum sp., etc.) as esterase/lipase, hypothetical protein, Sn1-specific diacylglycerol lipase beta, Lipase, Lipase, class 3, | No characterized protein among the first 500 hits |
| 78181 | Predicted triacylglycerol lipase activity, Lipase_3. Lipid transport and metabolism. | Lipase_3 (cd00519) Lipase_3 (pfam01764) PLN00413 Lip2 | Lipase that can hydrolyze long-chain acyl-triglycerides into di- and monoglycerides, glycerol and free fatty acids. | Active site Ser-His-Asp/Glu. Nucleophilic elbow on conserved domain Lipase_3: GHSLG | Active site flap/lid on conserved domain Lipase_3 (11 amino acids). | Genome Sequence and Annotation of different fungi (e.g., Trichoderma sp., Purpureocillium sp., Fusarium sp., Tolypocladium sp., etc.) as extracellular lipase-like protein, hypothetical protein, Lipase, probable triacylglycerol lipase precursor, Chain A, Crystal Structure of Lipase | Lipase Fusarium heterosporum (AAB34680) Query cover 98% Identity 52% |
| Protein ID | aa Residues | Molecular Weight (kDa) | SignalP (Secreted) | TMHMM Domains | Putative Cellular Localization | Continue for Next Analysis |
|---|---|---|---|---|---|---|
| 551811 | 379 | 41.185 | No | No | Cytoplasmic | Yes |
| 87496 | 430 | 46.62 | Yes | No | Extracellular | Yes |
| 526309 | 452 | 47.99 | Yes | No | Extracellular | Yes |
| 514427 | 454 | 49.25 | Yes | No | Extracellular | Yes |
| 510832 | 722 | 82.80 | No | No | Cytoplasmic | No |
| 92423 | 552 | 62.361 | No | 1 | Ambiguous location (plasmatic, endoplasmic reticulum, Golgi Apparatus) | Yes |
| 79895 | 1254 | 134.85 | No | No | Mitochondria | No |
| 135964 | 339 | 37.51 | No | No | Endoplasmic reticulum | no |
| 492160 | 1782 | 198.07 | No | No | Nuclear | No |
| 77338 | 404 | 44.5 | Yes | 1 | Extracellular | Yes |
| 514252 | 613 | 65.4 | Yes | No | Extracellular | Yes |
| 502433 | 1059 | 115.6 | No | No | Cytosolic/nuclear | Yes |
| 78181 | 340 | 36.2 | Yes | No | Extracellular | Yes |
| Protein & | Template | Reference Art | Pentapeptide | Lid Domain | Catalytic Triad | Oxyanion |
|---|---|---|---|---|---|---|
| 551811 | 3GUU | Ericsson et al. [78] | GYSGG Gly160-Gly164 | Asn195-Ser287 | Ser162, Asp309, His341 | Asp92 Gly163 |
| 87496 | GYSGG Gly188-Gly192 | Leu223-Glu313 | Ser190, Asp334, His368 | Asp96 Gly191 | ||
| 526309 | GYSGG Gy208-Gly212 | Asn243-Asp332 | Ser210, Asp352, His384 | Asp116 Gly211 | ||
| 514427 | GHSQG Gly226-Gly230 | Ala258-Phe349 | Ser228, Asp373, His405 | Ile146 Gln229 | ||
| 510832 | 1K8Q | Roussel et al. [79] and Selvan et al. [80] | CHSQG Cys431-Gly435 | Phe496-Glu550 | Ser433, Asp634, His667 | Leu349 Gln434 |
| 92423 | GFSQG Gly224-Gly228 | Ile286-Ile321 | Ser226, Asp396, His422 | Leu139 Gln227 | ||
| 79895 # | No structural homologue of lipase was identified | |||||
| 135964 | 4 X6U | Dror et al. [81] | AHSMG Ala-152-Gly156 | Phe189-Pro200 | Ser154, Asp275, His297 | Leu70 Met155 |
| 492160 # | No structural homologue of lipase was identified | |||||
| 77338 | 3O0D | Bordes et al. [82] | GHSLG Gly214-Gly218 | Thr137-Tyr154 | Ser216, Asp282, His343 | Thr137 Leu217 |
| 514252 | GHSLG Gly316-Gly320 | Thr230-Trp262 | Ser318, Asp379, His457 | Thr230 Leu319 | ||
| 502433 * | Complete protein sequence does not model with lipase | |||||
| 78181 | 3NGM | Lou et al. [83] | GHSLG Gly174-Gly178 | Asn115-Phe126 | Ser176, Asp230, His289 | Ser114 Leu177 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Canseco-Pérez, M.A.; Castillo-Avila, G.M.; Chi-Manzanero, B.; Islas-Flores, I.; Apolinar-Hernández, M.M.; Rivera-Muñoz, G.; Gamboa-Angulo, M.; Sanchez-Teyer, F.; Couoh-Uicab, Y.; Canto-Canché, B. Fungal Screening on Olive Oil for Extracellular Triacylglycerol Lipases: Selection of a Trichoderma harzianum Strain and Genome Wide Search for the Genes. Genes 2018, 9, 62. https://doi.org/10.3390/genes9020062
Canseco-Pérez MA, Castillo-Avila GM, Chi-Manzanero B, Islas-Flores I, Apolinar-Hernández MM, Rivera-Muñoz G, Gamboa-Angulo M, Sanchez-Teyer F, Couoh-Uicab Y, Canto-Canché B. Fungal Screening on Olive Oil for Extracellular Triacylglycerol Lipases: Selection of a Trichoderma harzianum Strain and Genome Wide Search for the Genes. Genes. 2018; 9(2):62. https://doi.org/10.3390/genes9020062
Chicago/Turabian StyleCanseco-Pérez, Miguel Angel, Genny Margarita Castillo-Avila, Bartolomé Chi-Manzanero, Ignacio Islas-Flores, Max M. Apolinar-Hernández, Gerardo Rivera-Muñoz, Marcela Gamboa-Angulo, Felipe Sanchez-Teyer, Yeny Couoh-Uicab, and Blondy Canto-Canché. 2018. "Fungal Screening on Olive Oil for Extracellular Triacylglycerol Lipases: Selection of a Trichoderma harzianum Strain and Genome Wide Search for the Genes" Genes 9, no. 2: 62. https://doi.org/10.3390/genes9020062
APA StyleCanseco-Pérez, M. A., Castillo-Avila, G. M., Chi-Manzanero, B., Islas-Flores, I., Apolinar-Hernández, M. M., Rivera-Muñoz, G., Gamboa-Angulo, M., Sanchez-Teyer, F., Couoh-Uicab, Y., & Canto-Canché, B. (2018). Fungal Screening on Olive Oil for Extracellular Triacylglycerol Lipases: Selection of a Trichoderma harzianum Strain and Genome Wide Search for the Genes. Genes, 9(2), 62. https://doi.org/10.3390/genes9020062





