Mapping of QTLs for Seed Phorbol Esters, a Toxic Chemical in Jatropha curcas (L.)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials and DNA Extraction
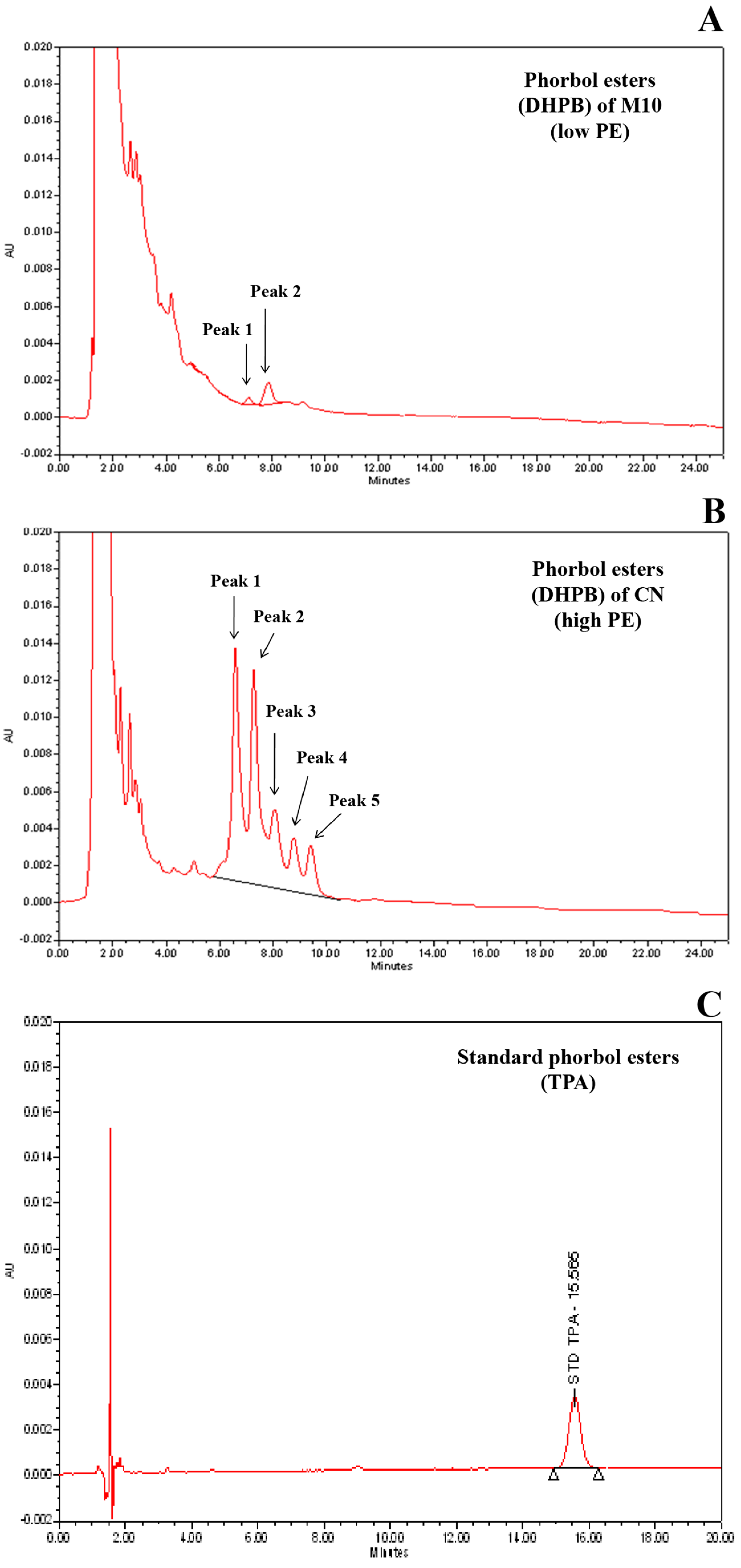
2.2. Quantification of Phorbol Esters Content in Seeds
2.3. SSR Marker Analysis
2.4. Linkage Map Construction and QTL Analysis
2.5. Identification of Physical Locations of Markers Flanking QTLs and Genes Associated with QTL
3. Results
3.1. Variation of Seed PE in Parents and F2 Population
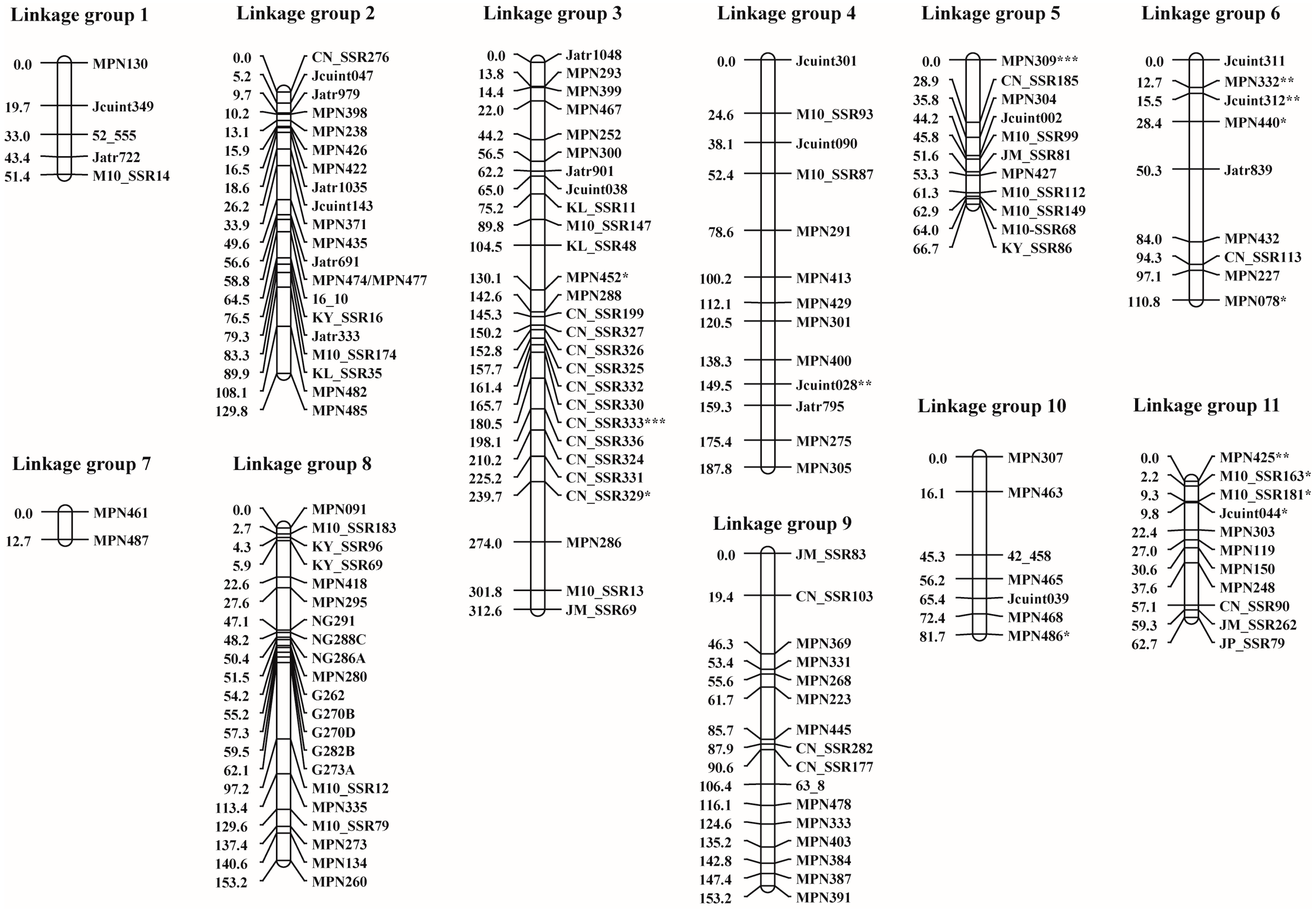
3.2. SSR Polymorphism and Linkage Map
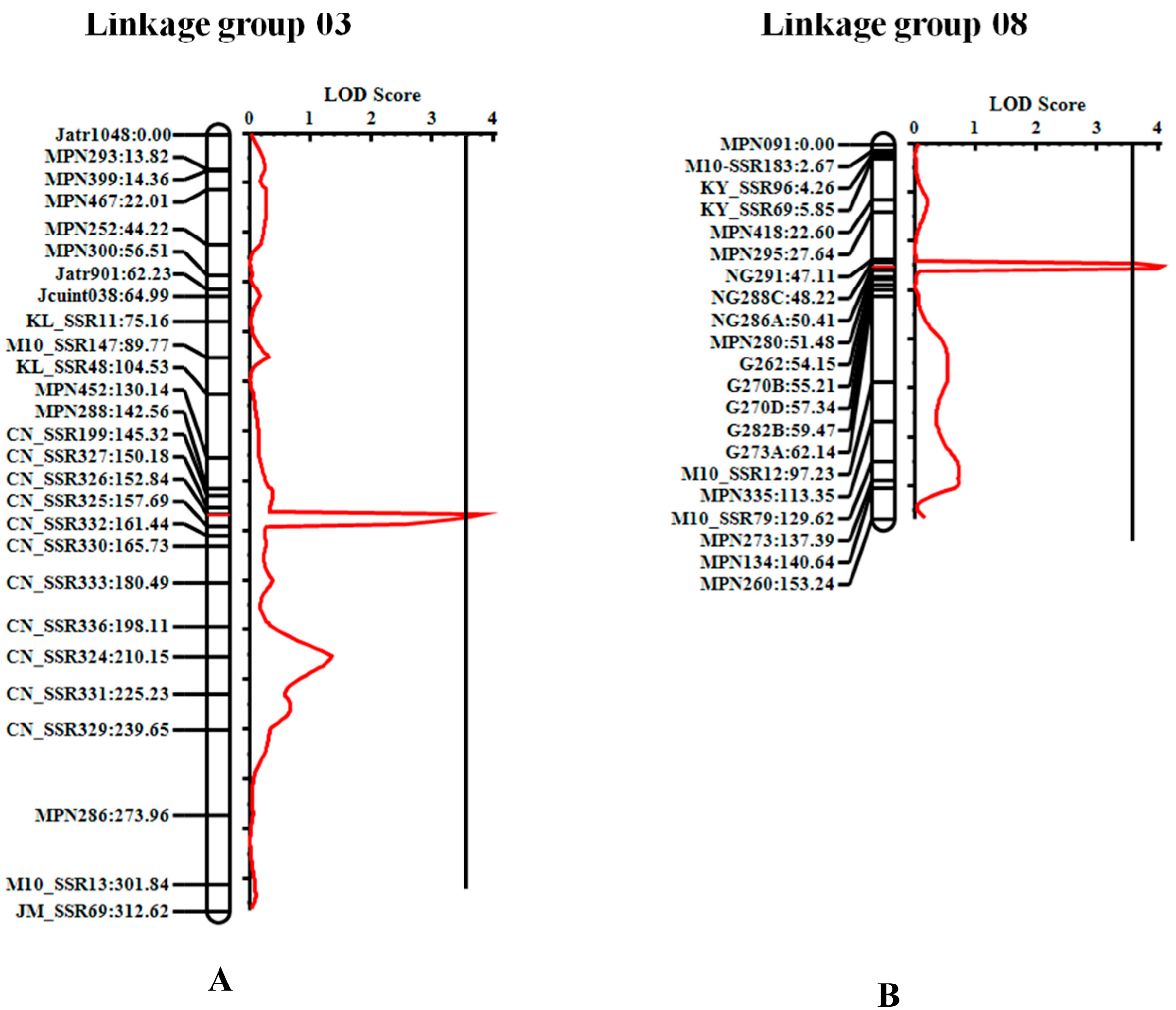
3.3. QTL Controlling Seed PE in Jatropha
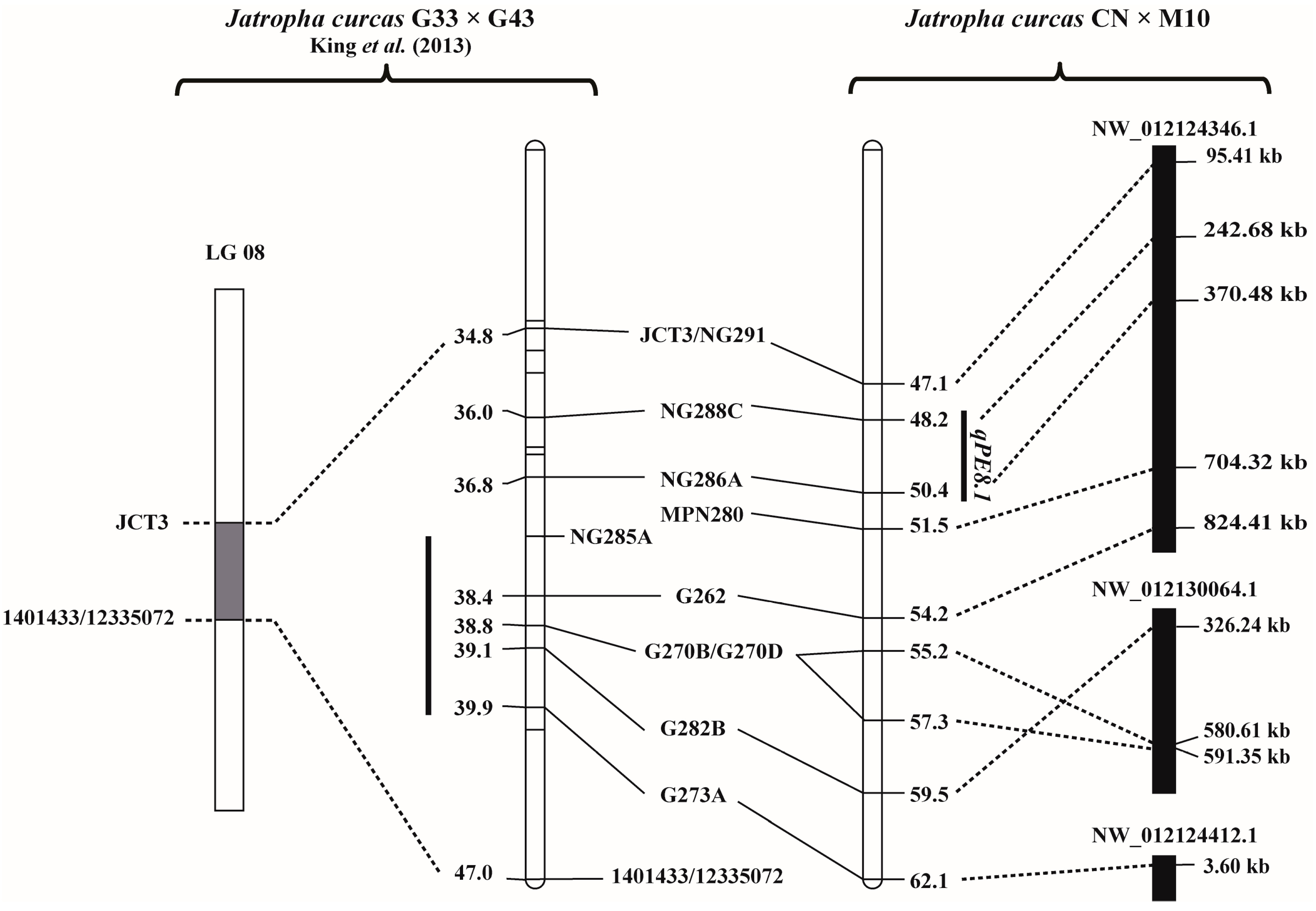
3.4. Physical Location of the PE QTLs and Annotated Genes in the QTL Regions
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gübitz, G.M.; Mittelbach, M.; Trabi, M. Exploitation of the tropical oil seed plant Jatropha curcas L. Bioresour. Technol. 1999, 67, 73–82. [Google Scholar] [CrossRef]
- Openshaw, K. A review of Jatropha curcas: An oil plant of unfulfilled promise. Biomass Bioenergy 2000, 19, 1–15. [Google Scholar] [CrossRef]
- Achten, W.M.J.; Nielsen, L.R.; Aerts, R.; Lengkeek, A.G.; Kjaer, E.D.; Trabucco, A.; Hansen, J.K.; Maes, W.H.; Graudal, L.; Akinnifesi, F.K.; et al. Towards domestication of Jatropha curcas. Biofuels 2010, 1, 91–107. [Google Scholar] [CrossRef]
- Haas, H.S.; Sterk, H.; Mittelbach, M. Novel 12-deoxy-16-hydroxy phorbol diesters isolated from the seed oil of Jatropha curcas L. J. Nat. Prod. 2002, 65, 1434–1440. [Google Scholar] [CrossRef] [PubMed]
- Devappa, R.K.; Makkar, H.P.; Becker, K. Jatropha toxicity—A review. J. Toxicol. Environ. Health B Crit. Rev. 2010, 13, 476–507. [Google Scholar] [CrossRef] [PubMed]
- Gomes, K.A.; Almeida, T.C.; Gesteira, A.S.; Lôbo, I.P.; Guimarấes, A.C.R.; Miranda, A.B.; Sluys, M.A.; Cruz, R.S.; Cascardo, J.C.M.; Carels, N. ESTs from seeds to assist the selective breeding of Jatropha curcas L. for oil and active compounds. Genom. Insights 2010, 3, 29–56. [Google Scholar]
- Sato, S.; Hirakawa, H.; Isobe, S.; Fukai, E.; Watanabe, A.; Kato, M.; Kawashima, K.; Minami, C.; Muraki, A.; Nakazaki, N.; et al. Sequence analysis of the genome of an oil-bearing tree, Jatropha curcas L. DNA Res. 2011, 18, 65–76. [Google Scholar] [CrossRef] [PubMed]
- Laosatit, K.; Tanya, P.; Somta, P.; Ruang-Areerate, P.; Sonthirod, C.; Tangphatsornruang, S.; Juntawong, P.; Srinives, P. De novo transcriptome analysis of apical meristem of Jatrophas spp. using 454 pyrosequencing platform, and identification of SNP and EST-SSR markers. Plant Mol. Biol. Rep. 2016, 34, 786–793. [Google Scholar] [CrossRef]
- King, A.J.; Brown, G.D.; Gilday, A.D.; Larson, T.R.; Graham, I.A. Production of bioactive diterpenoids in the Euphorbiaceae depends on evolutionarily conserved gene clusters. Plant Cell 2014, 26, 3286–3298. [Google Scholar] [CrossRef] [PubMed]
- Ng, A. Genes Implicated in Phorbol Ester (PE) Biosynthesis and Interception through Downregulation of Casbene Synthase Genes. Ph.D. Thesis, Nanyang Technological University, Singapore, 2014. [Google Scholar]
- Becker, K.; Makkar, H.P.S. Jatropha curcas: A potential source for tomorrow’s oil and biodiesel. Lipid Technol. 2008, 20, 104–107. [Google Scholar] [CrossRef]
- King, A.; Montes, L.R.; Clarke, J.G.; Affleck, J.; Li, Y.; Witsenboer, H.; Vossen, E.V.; Linde, P.V.; Tripathi, Y.; Tavares, E.; et al. Linkage mapping in the oilseed crop Jatropha curcas L. reveals a locus controlling the biosynthesis of PE which cause seed toxicity. Plant Biotechnol. J. 2013, 11, 986–996. [Google Scholar] [CrossRef] [PubMed]
- Jongcherdchutrakul, A. Inheritance of Phorbol Esters Content in Jatropha Seeds (Jatropha curcas L.). Master’s Thesis, Kasetsart University, Bangkok, Thailand, 2012. [Google Scholar]
- Laosatit, K.; Saensuk, C.; Tanya, P.; Srinives, P. Development and characterization of EST-SSR markers from Jatropha curcas EST database and their transferability across jatropha-related species/genus. Biologia 2013, 68, 41–47. [Google Scholar] [CrossRef]
- Tanya, P.; Dachapak, S.; Tar, M.M.; Srinives, P. New microsatellite markers classifiying nontoxic and toxic Jatropha curcas. J. Genet. 2011, 90, 76–78. [Google Scholar]
- Wang, C.M.; Liu, P.; Yi, C.; Gu, K.; Sun, F.; Li, L.; Lo, L.C.; Liu, X.; Feng, F.; Lin, G.; et al. A first generation microsatellite- and SNP based linkage map of Jatropha. PLoS ONE 2011, 6. [Google Scholar] [CrossRef] [PubMed]
- Meng, L.; Li, H.; Zhang, L.; Wang, J. QTL IciMapping: Integrated software for genetic linkage map construction and quantitative trait locus mapping in bi-parental populations. Crop J. 2015, 3, 265–279. [Google Scholar] [CrossRef]
- Li, H.; Ye, G.; Wang, J. A modified algorithm for the improvement of composite interval mapping. Genetics 2007, 175, 361–374. [Google Scholar] [CrossRef] [PubMed]
- Beavis, W.B. QTL analyses: Power, precision, and accuracy. In Molecular Dissection of Complex Traits; Patterson, A.H., Ed.; CRC Press: Boca Raton, FL, USA, 1998. [Google Scholar]
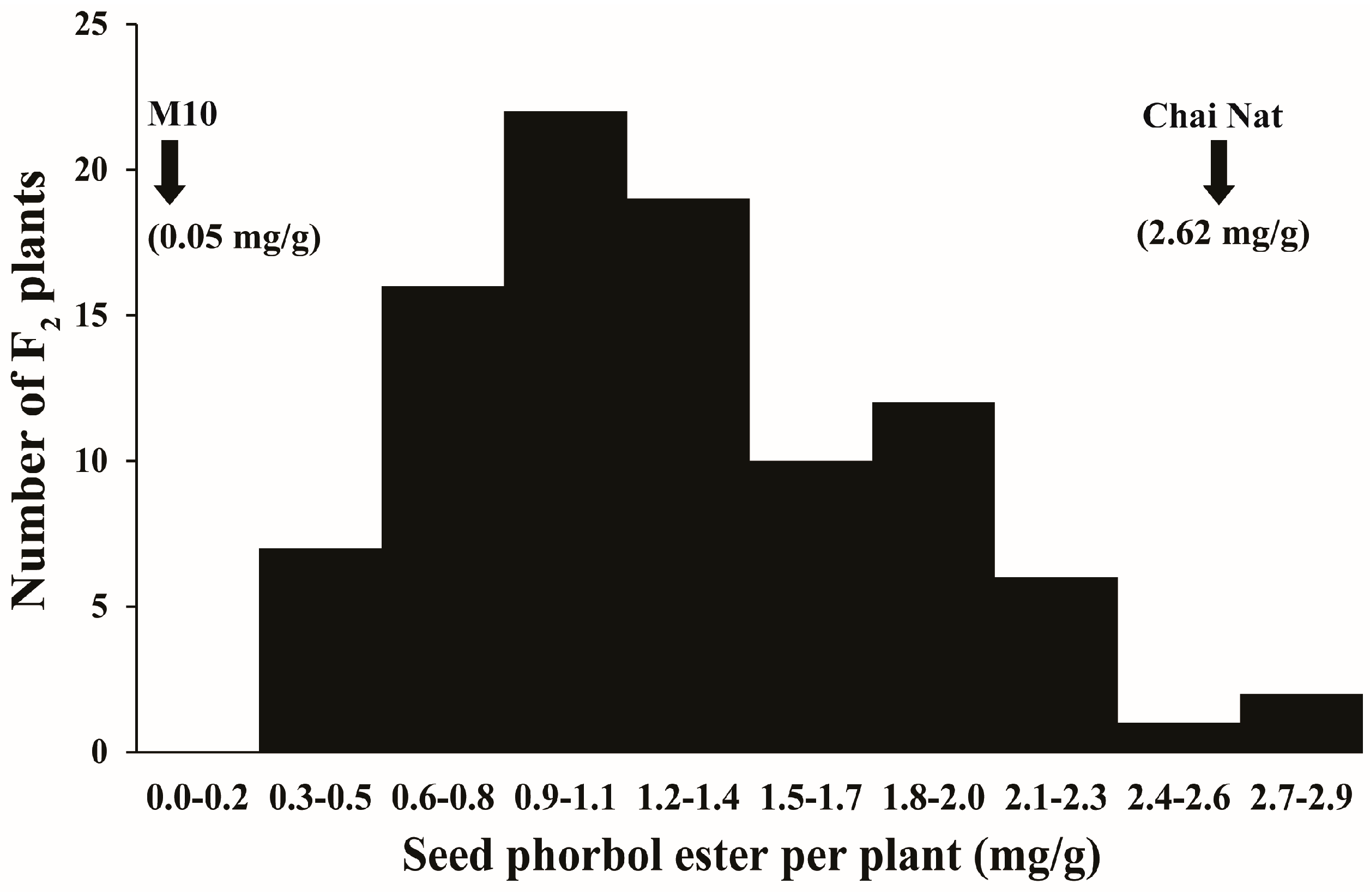


| LG a | Marker Name | R2 (%) b | Position | LOD Score |
|---|---|---|---|---|
| 3 | CN_SSR199 | 11.74 | 145.32 | 2.58 |
| 3 | CN_SSR327 | 12.25 | 150.18 | 2.70 |
| 3 | CN_SSR326 | 15.88 | 152.84 | 3.57 |
| 3 | CN_SSR330 | 11.67 | 165.73 | 2.56 |
| 3 | CN_SSR336 | 12.88 | 198.11 | 2.84 |
| 8 | NG291 | 14.20 | 47.11 | 3.16 |
| 8 | NG288C | 13.06 | 48.22 | 2.89 |
| 8 | NG286A | 16.99 | 50.41 | 3.84 |
| 8 | MPN280 | 16.40 | 51.48 | 3.70 |
| 8 | G262 | 13.55 | 54.15 | 3.00 |
| 8 | G270B | 15.69 | 55.21 | 3.52 |
| 8 | G270D | 15.64 | 57.34 | 3.51 |
| 8 | G282B | 13.94 | 59.47 | 3.10 |
| 8 | G273A | 15.57 | 62.14 | 3.49 |
| QTL Name | LG a | Marker Interval | Position (cM) b | LOD Score | Confidence Interval (cM) c | PVE (%) d | Additive Effect e | Dominant Effect | [d/a] f |
|---|---|---|---|---|---|---|---|---|---|
| qPE3.1 | 3 | CN_SSR326–CN_SSR325 | 153 | 3.82 | 152.5–156.5 | 14.10 | −0.09 | −0.41 | 4.56 |
| qPE8.1 | 8 | NG288C–NG286A | 50 | 4.06 | 48.5–51.5 | 15.49 | 0.31 | 0.02 | 0.06 |
| qPE3.1 | ||||
| Name | Position | Annotation | GO Term | KEGG |
| LOC105634082 | NW_012124225.1:1070722..1072343 | Uncharacterized protein At4g22758 | - | - |
| qPE8.1 | ||||
| Name | Position | Annotation | GO Term | KEGG |
| LOC105650633 | NW_012130064.1:251091..252362 | Peptidyl-prolylcis-trans isomerase CYP26-2, chloroplastic | GO:0003755 | K03768 |
| LOC105650634 | NW_012130064.1:244937..251195 | Uncharacterized LOC105650634 | GO:0008270 | K03768 |
| LOC105650635 | NW_012130064.1:253456..257226 | Single-stranded DNA-binding protein, mitochondrial | GO:0003697 | K03111 |
| LOC105650637 | NW_012130064.1:259463..262260 | Uncharacterized LOC105650637 | - | - |
| LOC105650638 | NW_012130064.1:262323..266102 | Uncharacterized LOC105650638 | - | - |
| LOC105650639 | NW_012130064.1:267598..271421 | Zinc finger BED domain-containing protein DAYSLEEPER | GO:0003677, GO:0046983, GO:0003676 | K03680 |
| LOC105650640 | NW_012130064.1:273262..278864 | Translation initiation factor eIF-2B subunit delta | GO:0044237 | K03680 |
| LOC105650641 | NW_012130064.1:288075..288870 | Oleosin 1-like | GO:0016021, GO:0012511 | - |
| LOC105650642 | NW_012130064.1:291937..292732 | Uncharacterized LOC105650642 | - | K13111 |
| LOC105650712 | NW_012130064.1:279771..287264 | Pseudogene | - | - |
| LOC105650643 | NW_012130064.1:297597..314028 | Suppressor of mec-8 and unc-52 protein homolog 1 | GO:0005515 | K13111 |
| LOC105650713 | NW_012130064.1:317012..317392 | Inactive protein FON2 SPARE1-like | - | - |
| LOC105650644 | NW_012130064.1:324023..347807 | Transcription initiation factor TFIID subunit 2 | GO:0008237, GO:0008270, GO:0005488 | K03128 |
| LOC105650714 | NW_012130064.1:367045..368556 | Transcription factor CYCLOIDEA | - | K08735 |
| LOC105650645 | NW_012130064.1:376579..386656 | DNA mismatch repair protein MSH2 | GO:0005524, GO:0030983, GO:0006298 | K08735 |
| LOC105650646 | NW_012130064.1:392247..402394 | Uncharacterized LOC105650646 | GO:0005515, GO:0008270 | - |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Amkul, K.; Laosatit, K.; Somta, P.; Shim, S.; Lee, S.-H.; Tanya, P.; Srinives, P. Mapping of QTLs for Seed Phorbol Esters, a Toxic Chemical in Jatropha curcas (L.). Genes 2017, 8, 205. https://doi.org/10.3390/genes8080205
Amkul K, Laosatit K, Somta P, Shim S, Lee S-H, Tanya P, Srinives P. Mapping of QTLs for Seed Phorbol Esters, a Toxic Chemical in Jatropha curcas (L.). Genes. 2017; 8(8):205. https://doi.org/10.3390/genes8080205
Chicago/Turabian StyleAmkul, Kitiya, Kularb Laosatit, Prakit Somta, Sangrea Shim, Suk-Ha Lee, Patcharin Tanya, and Peerasak Srinives. 2017. "Mapping of QTLs for Seed Phorbol Esters, a Toxic Chemical in Jatropha curcas (L.)" Genes 8, no. 8: 205. https://doi.org/10.3390/genes8080205
APA StyleAmkul, K., Laosatit, K., Somta, P., Shim, S., Lee, S.-H., Tanya, P., & Srinives, P. (2017). Mapping of QTLs for Seed Phorbol Esters, a Toxic Chemical in Jatropha curcas (L.). Genes, 8(8), 205. https://doi.org/10.3390/genes8080205





