Genome-Wide Analysis Reveals Extensive Changes in LncRNAs during Skeletal Muscle Development in Hu Sheep
Abstract
:1. Introduction
2. Materials and Methods
2.1. Animals and Sample Collection
2.2. Library Preparation
2.3. Clustering, Sequencing, and Transcriptome Assembly
2.4. Prediction of Multiple-Exon Long Non-Coding RNA and Protein-Coding RNA
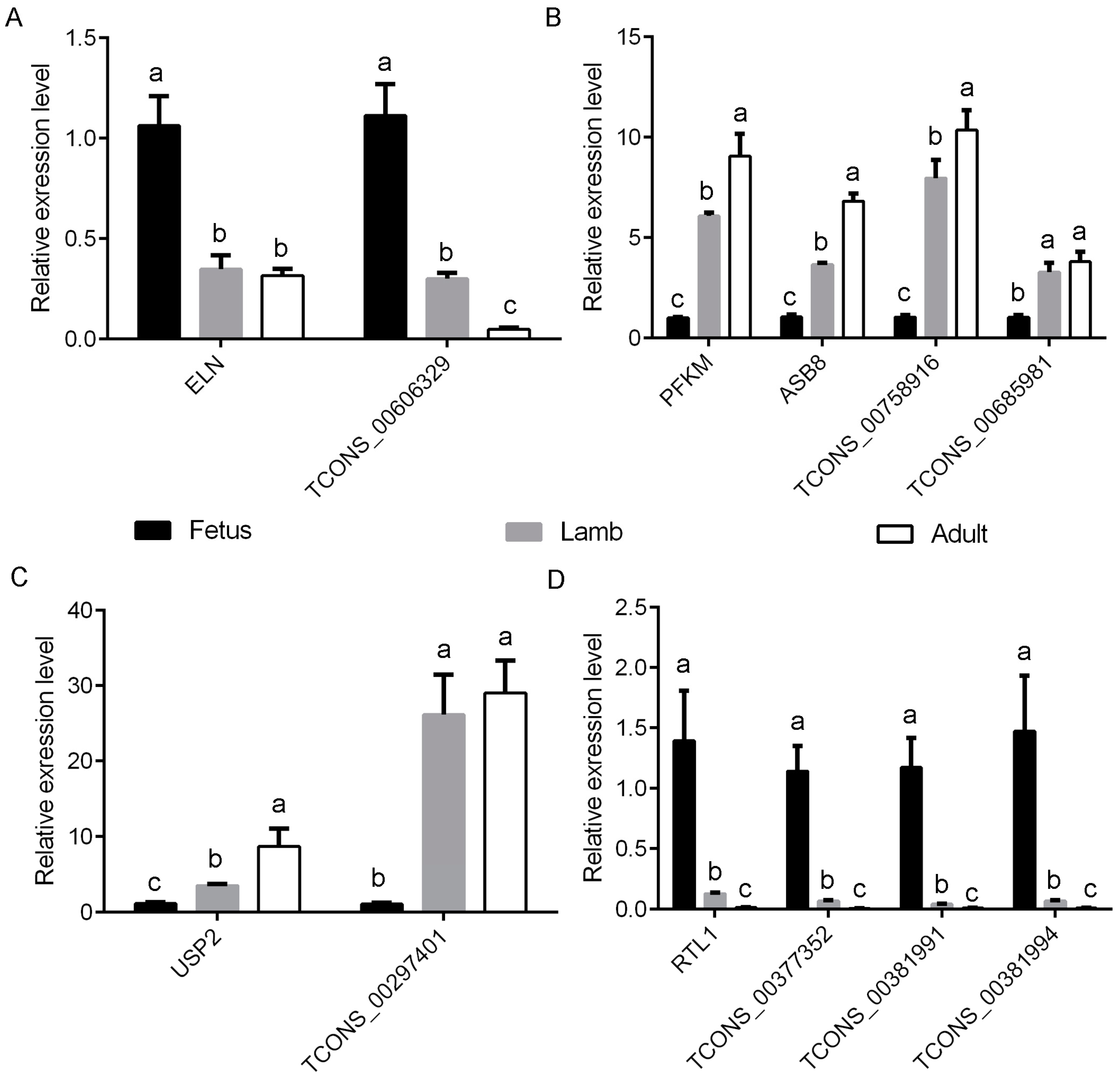
2.5. Differential Expression Analysis and Quantitative Real-Time Polymerase Chain Reaction Validation
2.6. Target Gene Prediction and Functional Annotation Analysis
2.7. Construction and Verification of the LncRNA-Gene Co-Expression Network
2.8. Statistical Analysis
3. Results
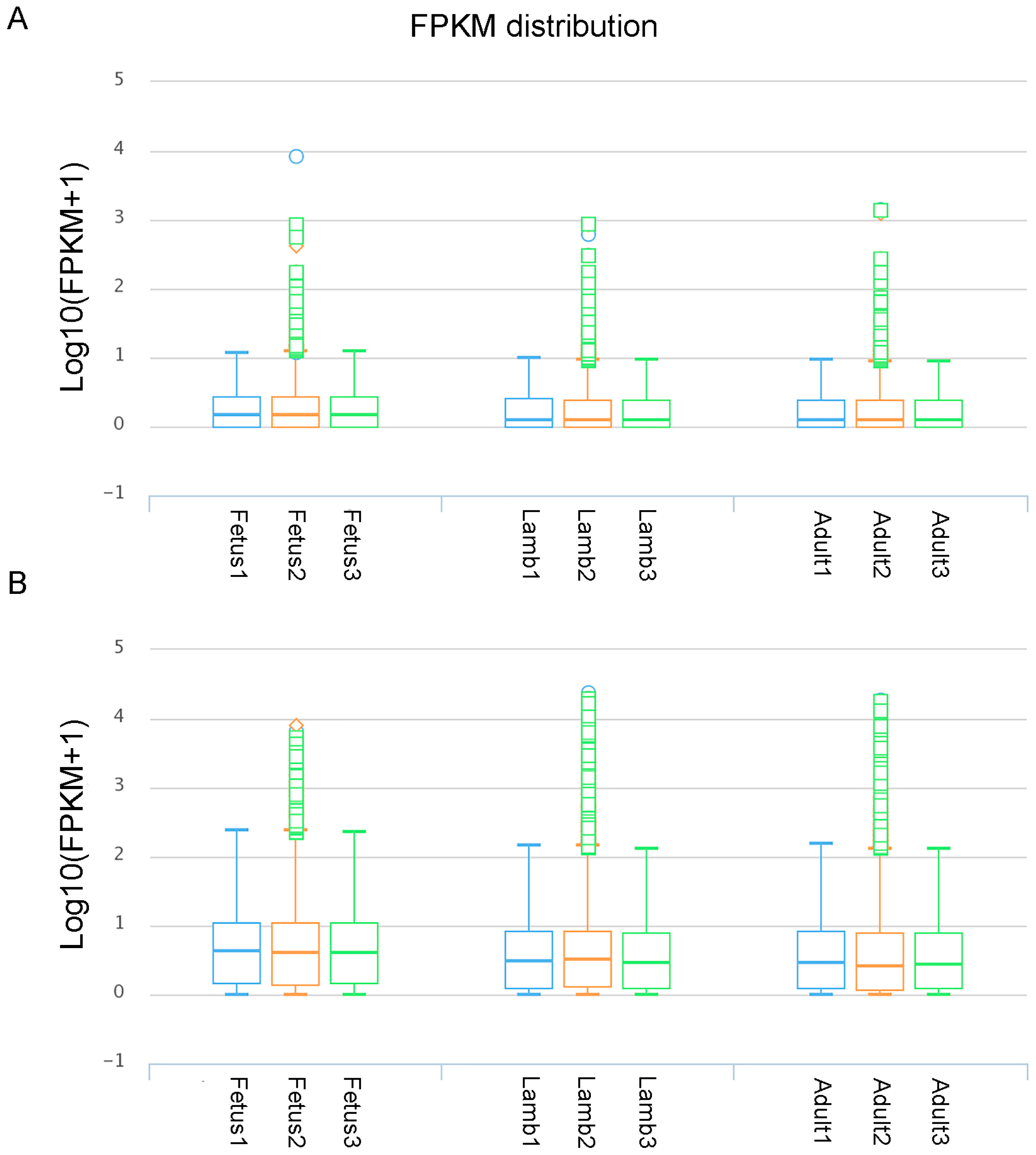
3.1. Read Mapping and Transcript Assembly
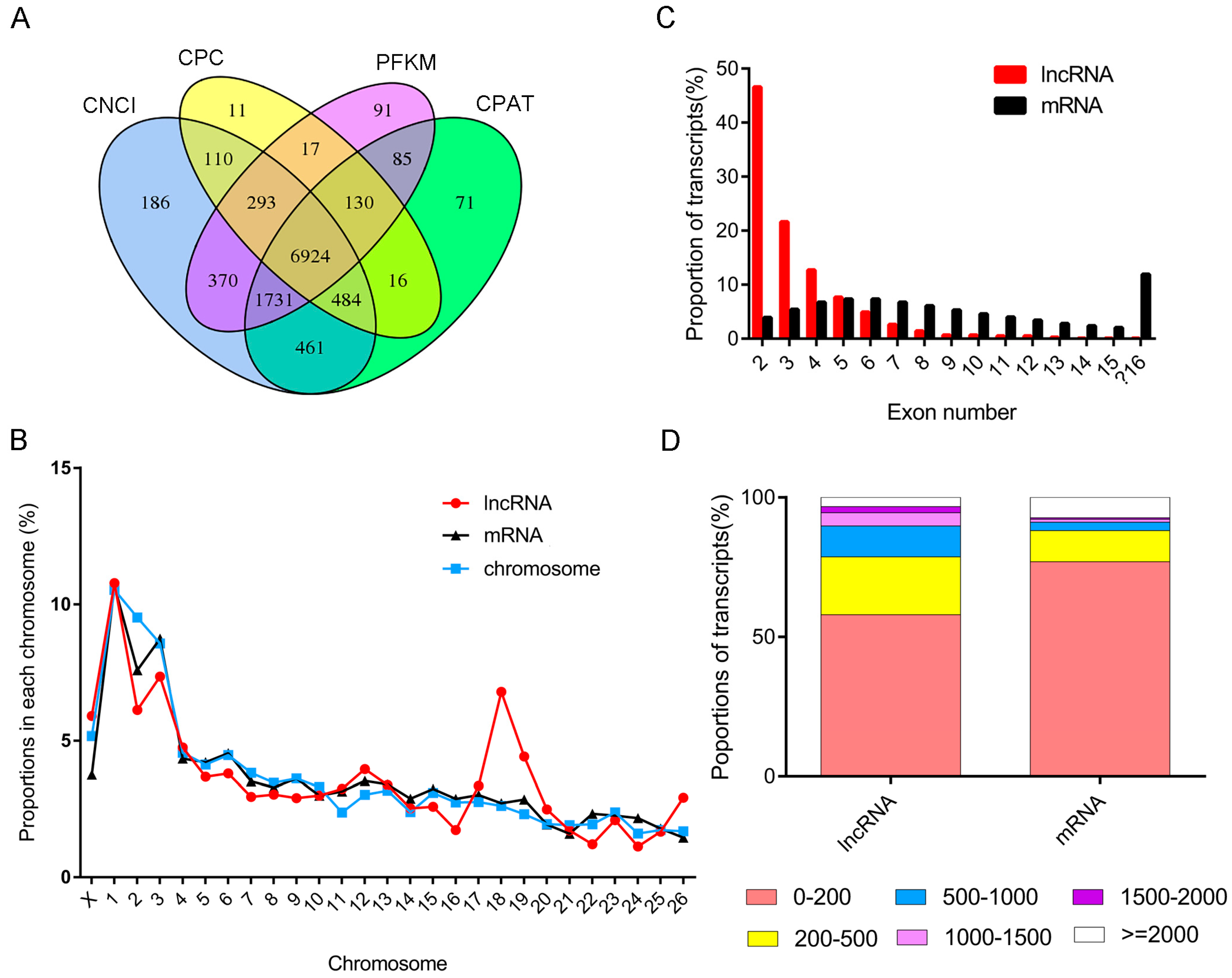
3.2. Identification and Characterization of Long Non-Coding RNA in Sheep Muscle
3.3. Differential Expression Analysis and Target Gene Prediction
3.4. Bioinformatics Analysis of the Target Genes of the Differentially Expressed Long Non-Coding RNAs and Differentially Expressed Genes
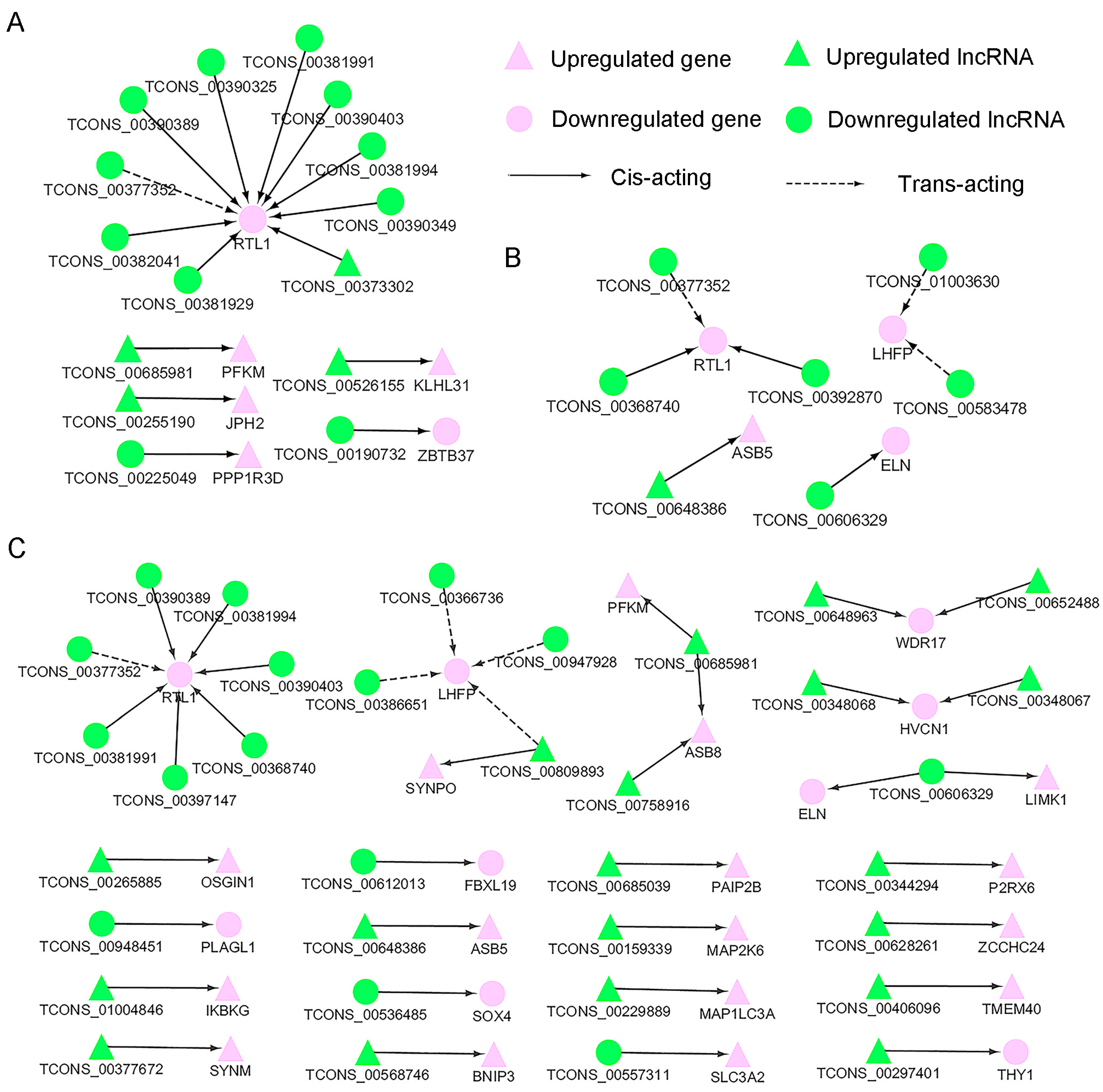
3.5. Screening of Potential Functional Long Non-Coding RNAs Involved in Myofiber Growth
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Edmondson, D.G.; Lyons, G.E.; Martin, J.F.; Olson, E.N. Mef2 gene-expression marks the cardiac and skeletal-muscle lineages during mouse embryogenesis. Development 1994, 120, 1251–1263. [Google Scholar] [PubMed]
- Yokoyama, S.; Asahara, H. The myogenic transcriptional network. Cell. Mol. Life Sci. 2011, 68, 1843–1849. [Google Scholar] [CrossRef] [PubMed]
- Giudice, J.; Loehr, J.A.; Rodney, G.G.; Cooper, T.A. Alternative splicing of four trafficking genes regulates myofiber structure and skeletal muscle physiology. Cell Rep. 2016, 17, 1923–1933. [Google Scholar] [CrossRef] [PubMed]
- Clop, A.; Marcq, F.; Takeda, H.; Pirottin, D.; Tordoir, X.; Bibe, B.; Bouix, J.; Caiment, F.; Elsen, J.M.; Eychenne, F.; et al. A mutation creating a potential illegitimate microrna target site in the myostatin gene affects muscularity in sheep. Nat. Genet. 2006, 38, 813–818. [Google Scholar] [CrossRef] [PubMed]
- Freking, B.A.; Murphy, S.K.; Wylie, A.A.; Rhodes, S.J.; Keele, J.W.; Leymaster, K.A.; Jirtle, R.L.; Smith, T.P. Identification of the single base change causing the callipyge muscle hypertrophy phenotype, the only known example of polar overdominance in mammals. Genome Res. 2002, 12, 1496–1506. [Google Scholar] [CrossRef] [PubMed]
- Collins, F.S.; Lander, E.S.; Rogers, J.; Waterston, R.H.; International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 2004, 431, 931–945. [Google Scholar]
- Cesana, M.; Cacchiarelli, D.; Legnini, I.; Santini, T.; Sthandier, O.; Chinappi, M.; Tramontano, A.; Bozzoni, I. A long noncoding rna controls muscle differentiation by functioning as a competing endogenous RNA. Cell 2011, 147, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhao, Y.; Bao, X.; Zhu, X.; Kwok, Y.K.; Sun, K.; Chen, X.; Huang, Y.; Jauch, R.; Esteban, M.A.; et al. LncRNA dum interacts with dnmts to regulate DPPA2 expression during myogenic differentiation and muscle regeneration. Cell Res. 2015, 25, 335–350. [Google Scholar] [CrossRef] [PubMed]
- Sakakibara, I.; Santolini, M.; Ferry, A.; Hakim, V.; Maire, P. Six homeoproteins and a iinc-rna at the fast myh locus lock fast myofiber terminal phenotype. PLoS Genet. 2014, 10, e1004386. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nelson, B.R.; Makarewich, C.A.; Anderson, D.M.; Winders, B.R.; Troupes, C.D.; Wu, F.; Reese, A.L.; McAnally, J.R.; Chen, X.; Kavalali, E.T.; et al. A peptide encoded by a transcript annotated as long noncoding RNA enhances serca activity in muscle. Science 2016, 351, 271–275. [Google Scholar] [CrossRef] [PubMed]
- Anderson, D.M.; Anderson, K.M.; Chang, C.L.; Makarewich, C.A.; Nelson, B.R.; McAnally, J.R.; Kasaragod, P.; Shelton, J.M.; Liou, J.; Bassel-Duby, R.; et al. A micropeptide encoded by a putative long noncoding RNA regulates muscle performance. Cell 2015, 160, 595–606. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, A.; Pasut, A.; Matsumoto, M.; Yamashita, R.; Fung, J.; Monteleone, E.; Saghatelian, A.; Nakayama, K.I.; Clohessy, J.G.; Pandolfi, P.P. MTORC1 and muscle regeneration are regulated by the linc00961-encoded spar polypeptide. Nature 2017, 541, 228–232. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Zhang, Y.; Li, T.; Ma, Z.; Jia, H.; Chen, Q.; Zhao, Y.; Zhai, L.; Zhong, R.; Li, C.; et al. Long non-coding RNA Linc-RAM enhances myogenic differentiation by interacting with myod. Nat. Commun. 2017, 8, 14016. [Google Scholar] [CrossRef] [PubMed]
- Zhu, M.; Liu, J.; Xiao, J.; Yang, L.; Cai, M.; Shen, H.; Chen, X.; Ma, Y.; Hu, S.; Wang, Z.; et al. Lnc-mg is a long non-coding RNA that promotes myogenesis. Nat. Commun. 2017, 8, 14718. [Google Scholar] [CrossRef] [PubMed]
- Yue, Y.; Guo, T.; Yuan, C.; Liu, J.; Guo, J.; Feng, R.; Niu, C.; Sun, X.; Yang, B. Integrated analysis of the roles of long noncoding RNA and coding RNA expression in sheep (Ovis aries) skin during initiation of secondary hair follicle. PLoS ONE 2016, 11, e0156890. [Google Scholar] [CrossRef] [PubMed]
- Chao, T.L.; Wang, G.Z.; Wang, J.M.; Liu, Z.H.; Ji, Z.B.; Hou, L.; Zhang, C.L. Identification and classification of new transcripts in dorper and small-tailed han sheep skeletal muscle transcriptomes. PLoS ONE 2016, 11, e0159638. [Google Scholar] [CrossRef] [PubMed]
- Ren, H.; Li, L.; Su, H.; Xu, L.; Wei, C.; Zhang, L.; Li, H.; Liu, W.; Du, L. Histological and transcriptome-wide level characteristics of fetal myofiber hyperplasia during the second half of gestation in texel and ujumqin sheep. BMC Genom. 2011, 12, 411. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of rna-seq experiments with tophat and cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed]
- Guttman, M.; Garber, M.; Levin, J.Z.; Donaghey, J.; Robinson, J.; Adiconis, X.; Fan, L.; Koziol, M.J.; Gnirke, A.; Nusbaum, C.; et al. Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincrnas. Nat. Biotechnol. 2010, 28, 503–510. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Zhang, Y.; Ye, Z.Q.; Liu, X.Q.; Zhao, S.Q.; Wei, L.; Gao, G. Cpc: Assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 2007, 35, W345–W349. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Luo, H.; Bu, D.; Zhao, G.; Yu, K.; Zhang, C.; Liu, Y.; Chen, R.; Zhao, Y. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res. 2013, 41, e166. [Google Scholar] [CrossRef] [PubMed]
- Punta, M.; Coggill, P.C.; Eberhardt, R.Y.; Mistry, J.; Tate, J.; Boursnell, C.; Pang, N.; Forslund, K.; Ceric, G.; Clements, J.; et al. The Pfam protein families database. Nucleic Acids Res. 2012, 40, D290–D301. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Park, H.J.; Dasari, S.; Wang, S.; Kocher, J.P.; Li, W. Cpat: Coding-potential assessment tool using an alignment-free logistic regression model. Nucleic Acids Res. 2013, 41, e74. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by rna-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Feng, Z.; Wang, X.; Wang, X.; Zhang, X. Degseq: An R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 2010, 26, 136–138. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Li, M.; Sun, Y.; Cai, H.; Lan, X.; Huang, Y.; Bai, Y.; Qi, X.; Chen, H. The developmental transcriptome sequencing of bovine skeletal muscle reveals a long noncoding RNA, Lncmd, promotes muscle differentiation by sponging mir-125b. Biochim. Biophys. Acta 2016, 1863, 2835–2845. [Google Scholar] [CrossRef] [PubMed]
- White, J.D.; Vuocolo, T.; McDonagh, M.; Grounds, M.D.; Harper, G.S.; Cockett, N.E.; Tellam, R. Analysis of the callipyge phenotype through skeletal muscle development; association of dlk1 with muscle precursor cells. Differentiation 2008, 76, 283–298. [Google Scholar] [CrossRef] [PubMed]
- Vuocolo, T.; Byrne, K.; White, J.; McWilliam, S.; Reverter, A.; Cockett, N.E.; Tellam, R.L. Identification of a gene network contributing to hypertrophy in callipyge skeletal muscle. Physiol. Genom. 2007, 28, 253–272. [Google Scholar] [CrossRef] [PubMed]
- Butchart, L.C.; Fox, A.; Shavlakadze, T.; Grounds, M.D. The long and short of non-coding RNAs during post-natal growth and differentiation of skeletal muscles: Focus on lncRNA and miRNAs. Differentiation 2016, 92, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Mu, Y.; Ma, L.; Wang, C.; Tang, Z.; Yang, S.; Zhou, R.; Hu, X.; Li, M.H.; Li, K. Systematic identification and characterization of long intergenic non-coding RNAs in fetal porcine skeletal muscle development. Sci. Rep. 2015, 5, 8957. [Google Scholar] [CrossRef] [PubMed]
- Zhan, S.; Dong, Y.; Zhao, W.; Guo, J.; Zhong, T.; Wang, L.; Li, L.; Zhang, H. Genome-wide identification and characterization of long non-coding rnas in developmental skeletal muscle of fetal goat. BMC Genom. 2016, 17, 666. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Bai, M.; Xiang, L.; Zhang, G.; Ma, W.; Jiang, H. Comparative transcriptome profiling of longissimus muscle tissues from qianhua mutton merino and small tail han sheep. Sci. Rep. 2016, 6, 33586. [Google Scholar] [CrossRef] [PubMed]
- Adiconis, X.; Borges-Rivera, D.; Satija, R.; DeLuca, D.S.; Busby, M.A.; Berlin, A.M.; Sivachenko, A.; Thompson, D.A.; Wysoker, A.; Fennell, T.; et al. Comparative analysis of RNA sequencing methods for degraded or low-input samples. Nat. Methods 2013, 10, 623–629. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- White, N.M.; Cabanski, C.R.; Silva-Fisher, J.M.; Dang, H.X.; Govindan, R.; Maher, C.A. Transcriptome sequencing reveals altered long intergenic non-coding RNAs in lung cancer. Genome Biol. 2014, 15, 429. [Google Scholar] [CrossRef] [PubMed]
- Sancak, Y.; Peterson, T.R.; Shaul, Y.D.; Lindquist, R.A.; Thoreen, C.C.; Bar-Peled, L.; Sabatini, D.M. The RAG GTPases bind raptor and mediate amino acid signaling to mTORC1. Science 2008, 320, 1496–1501. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Kong, X.; Li, F.; Tan, B.; Li, Y.; Duan, Y.; Yin, Y.; He, J.; Hu, C.; Blachier, F.; et al. Co-dependence of genotype and dietary protein intake to affect expression on amino acid/peptide transporters in porcine skeletal muscle. Amino Acids 2016, 48, 75–90. [Google Scholar] [CrossRef] [PubMed]
- Jorgensen, L.H.; Blain, A.; Greally, E.; Laval, S.H.; Blamire, A.M.; Davison, B.J.; Brinkmeier, H.; MacGowan, G.A.; Schroder, H.D.; Bushby, K.; et al. Long-term blocking of calcium channels in mdx mice results in differential effects on heart and skeletal muscle. Am. J. Pathol. 2011, 178, 273–283. [Google Scholar] [CrossRef] [PubMed]
- Farini, A.; Sitzia, C.; Cassinelli, L.; Colleoni, F.; Parolini, D.; Giovanella, U.; Maciotta, S.; Colombo, A.; Meregalli, M.; Torrente, Y. Inositol 1,4,5-trisphosphate (ip3)-dependent Ca2+ signaling mediates delayed myogenesis in duchenne muscular dystrophy fetal muscle. Development 2016, 143, 658–669. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.; Couturier, A.; Kubens, J.F.; Most, E.; Mooren, F.C.; Kruger, K.; Ringseis, R.; Eder, K. Niacin supplementation induces type II to type I muscle fiber transition in skeletal muscle of sheep. Acta Vet. Scand. 2013, 55, 85. [Google Scholar] [CrossRef] [PubMed]
- Schiaffino, S.; Dyar, K.A.; Ciciliot, S.; Blaauw, B.; Sandri, M. Mechanisms regulating skeletal muscle growth and atrophy. FEBS J. 2013, 280, 4294–4314. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.K.; Li, J.; Liu, J.; Guo, B.; Leung, A.; Zhang, G.; Zhang, B.T. Icaritin requires phosphatidylinositol 3 kinase (PI3K)/akt signaling to counteract skeletal muscle atrophy following mechanical unloading. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Ectors, F.; Davis, E.E.; Pirottin, D.; Cheng, H.; Farnir, F.; Hadfield, T.; Cockett, N.; Charlier, C.; Georges, M.; et al. Ectopic expression of retrotransposon-derived PEG11/RTL1 contributes to the callipyge muscular hypertrophy. PLoS ONE 2015, 10, e0140594. [Google Scholar] [CrossRef] [PubMed]
- Bidwell, C.A.; Kramer, L.N.; Perkins, A.C.; Hadfield, T.S.; Moody, D.E.; Cockett, N.E. Expression of PEG11 and PEG11as transcripts in normal and callipyge sheep. BMC Biol. 2004, 2. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hitachi, K.; Tsuchida, K. Myostatin-deficiency in mice increases global gene expression at the Dlk1-Dio3 locus in the skeletal muscle. Oncotarget 2017, 8, 5943–5953. [Google Scholar] [CrossRef] [PubMed]
- Fleming-Waddell, J.N.; Olbricht, G.R.; Taxis, T.M.; White, J.D.; Vuocolo, T.; Craig, B.A.; Tellam, R.L.; Neary, M.K.; Cockett, N.E.; Bidwell, C.A. Effect of Dlk1 and Rtl1 but not Meg3 or Meg8 on muscle gene expression in callipyge lambs. PLoS ONE 2009, 4, e7399. [Google Scholar] [CrossRef] [PubMed]
- Mo, C.F.; Wu, F.C.; Tai, K.Y.; Chang, W.C.; Chang, K.W.; Kuo, H.C.; Ho, H.N.; Chen, H.F.; Lin, S.P. Loss of non-coding rna expression from the Dlk1-Dio3 imprinted locus correlates with reduced neural differentiation potential in human embryonic stem cell lines. Stem Cell Res. Ther. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Davis, E.; Caiment, F.; Tordoir, X.; Cavaille, J.; Ferguson-Smith, A.; Cockett, N.; Georges, M.; Charlier, C. Rnai-mediated allelic trans-interaction at the imprinted Rtl1/Peg11 locus. Curr. Biol. 2005, 15, 743–749. [Google Scholar] [CrossRef] [PubMed]
- Clark, A.L.; Naya, F.J. MicroRNAs in the myocyte enhancer factor 2 (MEF2)-regulated GTL2-DIO3 noncoding RNA locus promote cardiomyocyte proliferation by targeting the transcriptional coactivator cited2. J. Biol. Chem. 2015, 290, 23162–23172. [Google Scholar] [CrossRef] [PubMed]
- Sabater-Molina, M.; Navarro, M.; Saez, E.G.M.; Garrido, I.; Pascual-Figal, D.; Carrillo, J.G.; Blanes, J.R.G. Mutation in JPH2 cause dilated cardiomyopathy. Clin. Genet. 2016, 90, 468–469. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; VanDusen, N.J.; Zhang, L.; Gu, W.; Sethi, I.; Guatimosim, S.; Ma, Q.; Jardin, B.D.; Ai, Y.; Zhang, D.; et al. Analysis of cardiac myocyte maturation using CASAAV, a platform for rapid dissection of cardiac myocyte gene function in vivo. Circ. Res. 2017, 120, 1874–1888. [Google Scholar] [CrossRef] [PubMed]
- Petit, M.M.R.; Schoenmakers, E.F.P.M.; Huysmans, C.; Geurts, J.M.W.; Mandahl, N.; Van de Ven, W.J.M. LHFP, a novel translocation partner gene of HMGIC in a lipoma, is a member of a new family of LHFP-like genes. Genomics 1999, 57, 438–441. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, A.; Wyczalkowska-Tomasik, A.; Zendzian-Piotrowska, M.; Czarkowska-Paczek, B. Training differentially regulates elastin level and proteolysis in skeletal and heart muscles and aorta in healthy rats. Biol. Open 2016, 5, 556–562. [Google Scholar] [CrossRef] [PubMed]
- D’Andrea, P.; Civita, D.; Cok, M.; Ulloa Severino, L.; Vita, F.; Scaini, D.; Casalis, L.; Lorenzon, P.; Donati, I.; Bandiera, A. Myoblast adhesion, proliferation and differentiation on human elastin-like polypeptide (HELP) hydrogels. J. Appl. Biomater. Funct. Mater. 2017, 15, e43–e53. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- D’Andrea, P.; Scaini, D.; Severino, L.U.; Borelli, V.; Passamonti, S.; Lorenzon, P.; Bandiera, A. In vitro myogenesis induced by human recombinant elastin-like proteins. Biomaterials 2015, 67, 240–253. [Google Scholar] [CrossRef] [PubMed]





| Sample | Fetus | Lamb | Adult |
|---|---|---|---|
| Total Mapped Reads | 65,578,070 | 65,591,958 | 71,241,551 |
| Uniq Mapped Reads | 60,846,465(92.76%) | 60,372,363(92.06%) | 66,521,969(93.43%) |
| Multiple Mapped Reads | 4,731,605(7.24%) | 5,219,595(7.94%) | 4,719,582(6.57%) |
| Reads Map to ‘+’ | 34,126,308(52.04%) | 34,153,600(52.07%) | 36,918,137(51.82%) |
| Reads Map to ‘−’ | 31,451,762(47.96%) | 31,438,358(47.93%) | 34,323,414(48.18%) |
| Reads Map to exonic region | 45.1% | 53.8% | 53.7% |
| Reads Map to intronic region | 25.4% | 19.6% | 19.2% |
| Reads Map to intergenic region | 29.5% | 26.6% | 27.1% |
| #DEG Set (Control/Experiment) | Total Annotated | Nr | COG | GO | KEGG | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Targets | DEGs | Targets | DEGs | Targets | DEGs | Targets | DEGs | Targets | DEGs | |
| Fetus/Adult | 150 | 3564 | 150 | 3564 | 57 | 1353 | 137 | 3317 | 83 | 2353 |
| Fetus/Lamb | 56 | 1480 | 56 | 1480 | 21 | 596 | 52 | 1376 | 37 | 972 |
| Lamb/Adult | 33 | 1536 | 33 | 1536 | 9 | 510 | 32 | 1436 | 20 | 997 |
| KEGG Pathway (KEGG Orthology (ko) ID) | Fetus/Lamb | Lamb/Adult | Fetus/Adult | All Unigenes | ||||
|---|---|---|---|---|---|---|---|---|
| DEGs Number | % | DEGs Number | % | DEGs Number | % | Gene Number | % | |
| 1. Oxidative phosphorylation (ko00190) | 74 | 12.67 | 0 | 0.00 | 72 | 5.26 | 164 | 2.11 |
| 2. Carbon metabolism (ko01200) | 40 | 6.85 | 0 | 0.00 | 51 | 3.72 | 112 | 1.44 |
| 3. Cardiac muscle contraction (ko04260) | 32 | 5.48 | 0 | 0.00 | 37 | 2.70 | 91 | 1.17 |
| 4. PI3K-Akt signaling pathway (ko04151) | 0 | 0.00 | 50 | 8.55 | 0 | 0.00 | 374 | 4.81 |
| 5. Adrenergic signaling in cardiomyocytes (ko04261) | 0 | 0.00 | 0 | 0.00 | 48 | 3.50 | 159 | 2.05 |
| 6. Citrate cycle (TCA cycle) (ko00020) | 22 | 3.77 | 0 | 0.00 | 22 | 1.61 | 34 | 0.44 |
| 7. Fatty acid metabolism (ko01212) | 16 | 2.74 | 0 | 0.00 | 21 | 1.53 | 51 | 0.66 |
| 8. Pyruvate metabolism (ko00620) | 16 | 2.74 | 0 | 0.00 | 20 | 1.46 | 42 | 0.54 |
| 9. Focal adhesion (ko04510) | 0 | 0.00 | 33 | 5.64 | 0 | 0.00 | 224 | 2.88 |
| 10. Cell cycle (ko04110) | 0 | 0.00 | 27 | 4.62 | 0 | 0.00 | 148 | 1.90 |
| 11. Propanoate metabolism (ko00640) | 10 | 1.71 | 0 | 0.00 | 15 | 1.09 | 27 | 0.35 |
| 12. Proteasome (ko03050) | 0 | 0.00 | 0 | 0.00 | 22 | 1.61 | 47 | 0.60 |
| 13. Biosynthesis of amino acids (ko01230) | 21 | 3.60 | 0 | 0.00 | 0 | 0.00 | 80 | 1.03 |
| 14. Glycolysis/Gluconeogenesis (ko00010) | 19 | 3.25 | 0 | 0.00 | 0 | 0.00 | 70 | 0.90 |
| 15. PPAR signaling pathway (ko03320) | 19 | 3.25 | 0 | 0.00 | 0 | 0.00 | 70 | 0.90 |
| 16. Fatty acid degradation(ko00071) | 14 | 2.40 | 0 | 0.00 | 0 | 0.00 | 44 | 0.57 |
| 17. Alanine, aspartate and glutamate metabolism (ko00250) | 13 | 2.23 | 0 | 0.00 | 0 | 0.00 | 40 | 0.51 |
| 18. 2-Oxocarboxylic acid metabolism (ko01210) | 11 | 1.88 | 0 | 0.00 | 0 | 0.00 | 18 | 0.23 |
| 19. Nicotinate and nicotinamide metabolism (ko00760) | 10 | 1.71 | 0 | 0.00 | 0 | 0.00 | 29 | 0.37 |
| 20. Steroid biosynthesis (ko00100) | 0 | 0.00 | 8 | 1.37 | 0 | 0.00 | 22 | 0.28 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ren, C.; Deng, M.; Fan, Y.; Yang, H.; Zhang, G.; Feng, X.; Li, F.; Wang, D.; Wang, F.; Zhang, Y. Genome-Wide Analysis Reveals Extensive Changes in LncRNAs during Skeletal Muscle Development in Hu Sheep. Genes 2017, 8, 191. https://doi.org/10.3390/genes8080191
Ren C, Deng M, Fan Y, Yang H, Zhang G, Feng X, Li F, Wang D, Wang F, Zhang Y. Genome-Wide Analysis Reveals Extensive Changes in LncRNAs during Skeletal Muscle Development in Hu Sheep. Genes. 2017; 8(8):191. https://doi.org/10.3390/genes8080191
Chicago/Turabian StyleRen, Caifang, Mingtian Deng, Yixuan Fan, Hua Yang, Guomin Zhang, Xu Feng, Fengzhe Li, Dan Wang, Feng Wang, and Yanli Zhang. 2017. "Genome-Wide Analysis Reveals Extensive Changes in LncRNAs during Skeletal Muscle Development in Hu Sheep" Genes 8, no. 8: 191. https://doi.org/10.3390/genes8080191
APA StyleRen, C., Deng, M., Fan, Y., Yang, H., Zhang, G., Feng, X., Li, F., Wang, D., Wang, F., & Zhang, Y. (2017). Genome-Wide Analysis Reveals Extensive Changes in LncRNAs during Skeletal Muscle Development in Hu Sheep. Genes, 8(8), 191. https://doi.org/10.3390/genes8080191




