Transcriptome Analysis of Paraburkholderia phymatum under Nitrogen Starvation and during Symbiosis with Phaseolus Vulgaris
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains, Media, and Cultivation
2.2. Plant Growth Conditions and Inoculation
2.3. Determination of Symbiotic Properties
2.4. RNA-Sequencing and Data Processing
2.5. Quantitative Reverse Transcription PCR (qRT-PCR) Analysis
2.6. Construction of Paraburkholderia phymatum STM815T Mutant Strains
2.7. Exopolysaccharides Production
2.8. Statistical Analysis
3. Results
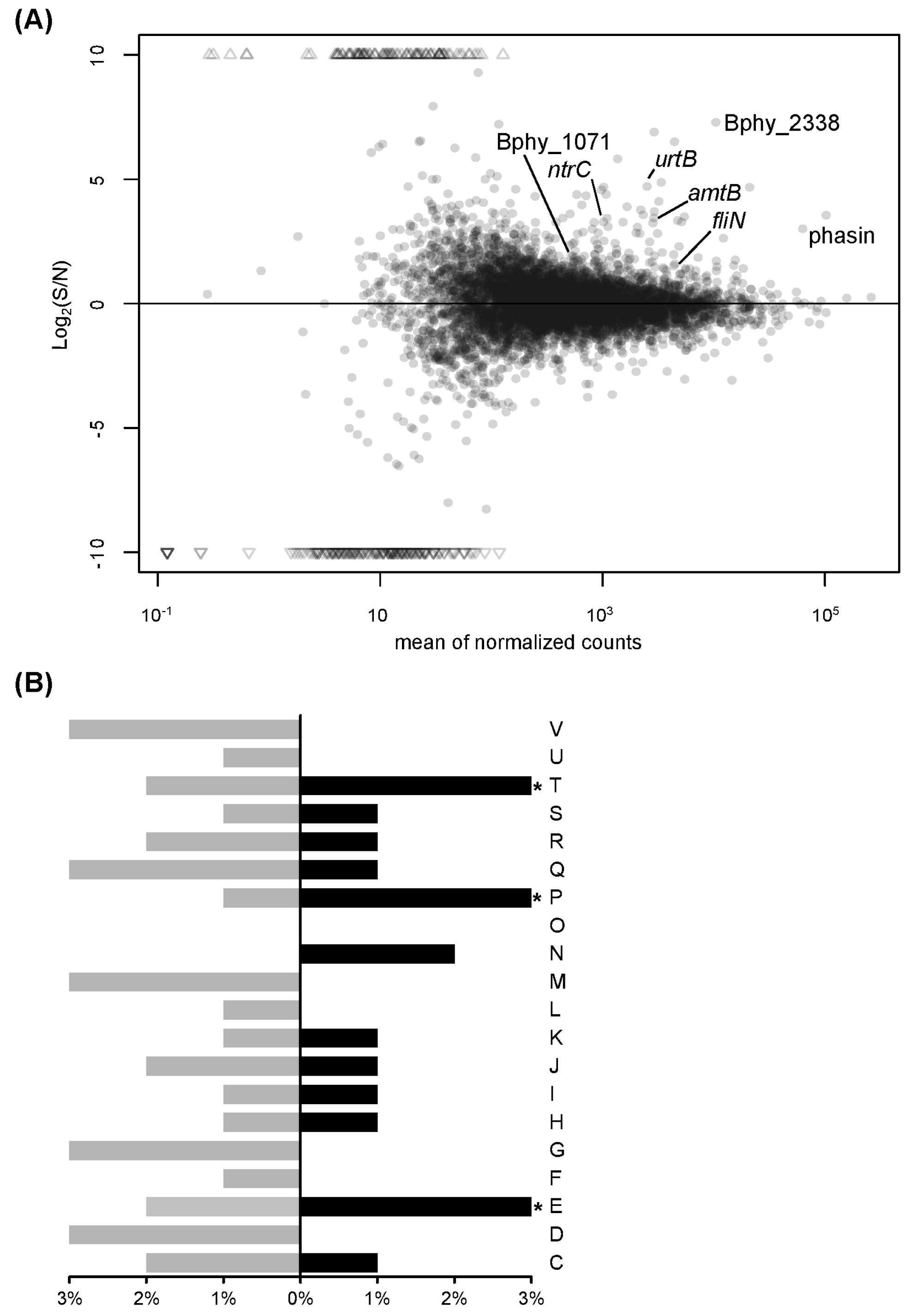
3.1. Transcript Profiling of P. phymatum STM815T in Response to Nitrogen Limitation
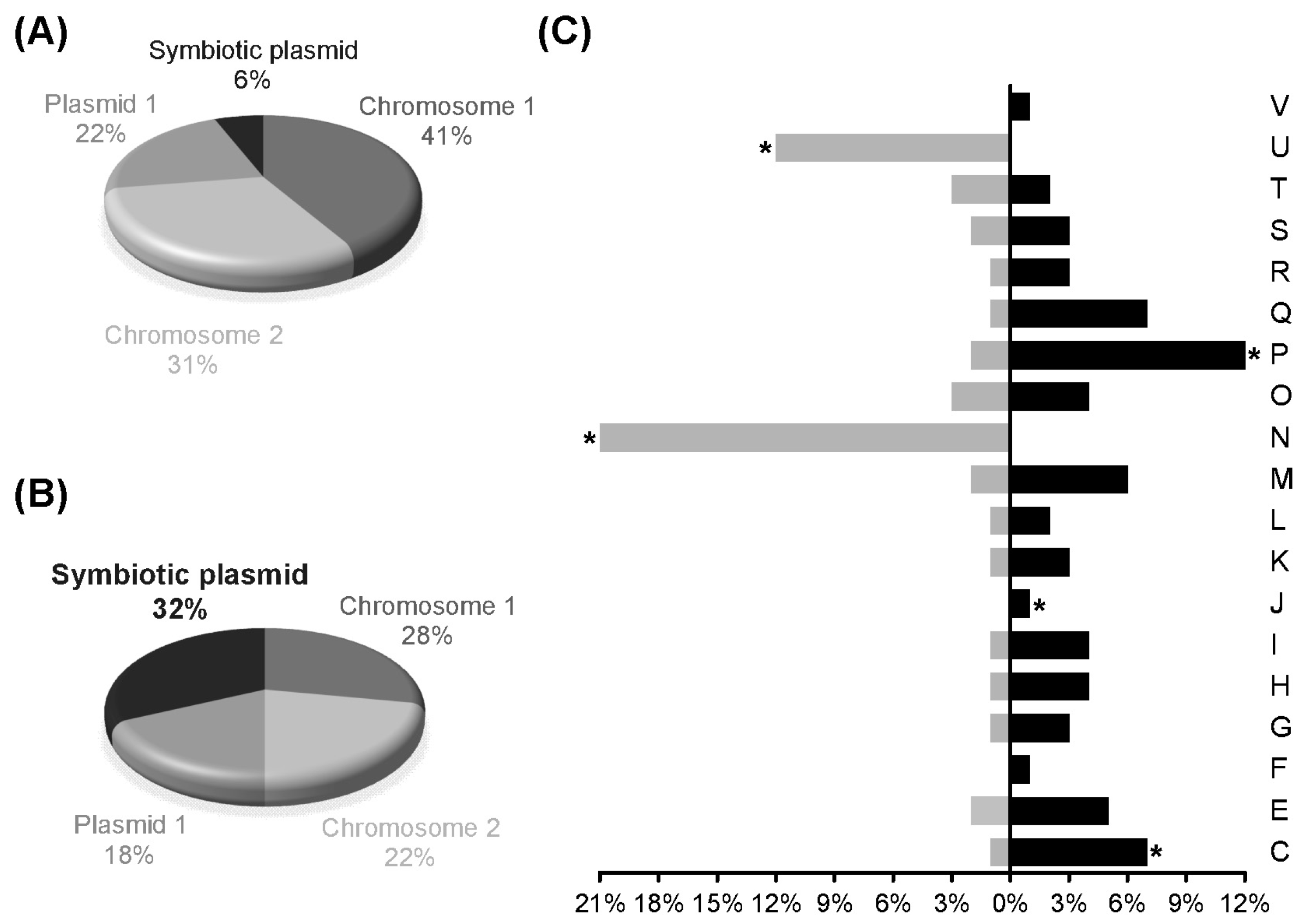
3.2. Transcript Profiling of P. phymatum STM815T during Symbiosis with P. vulgaris
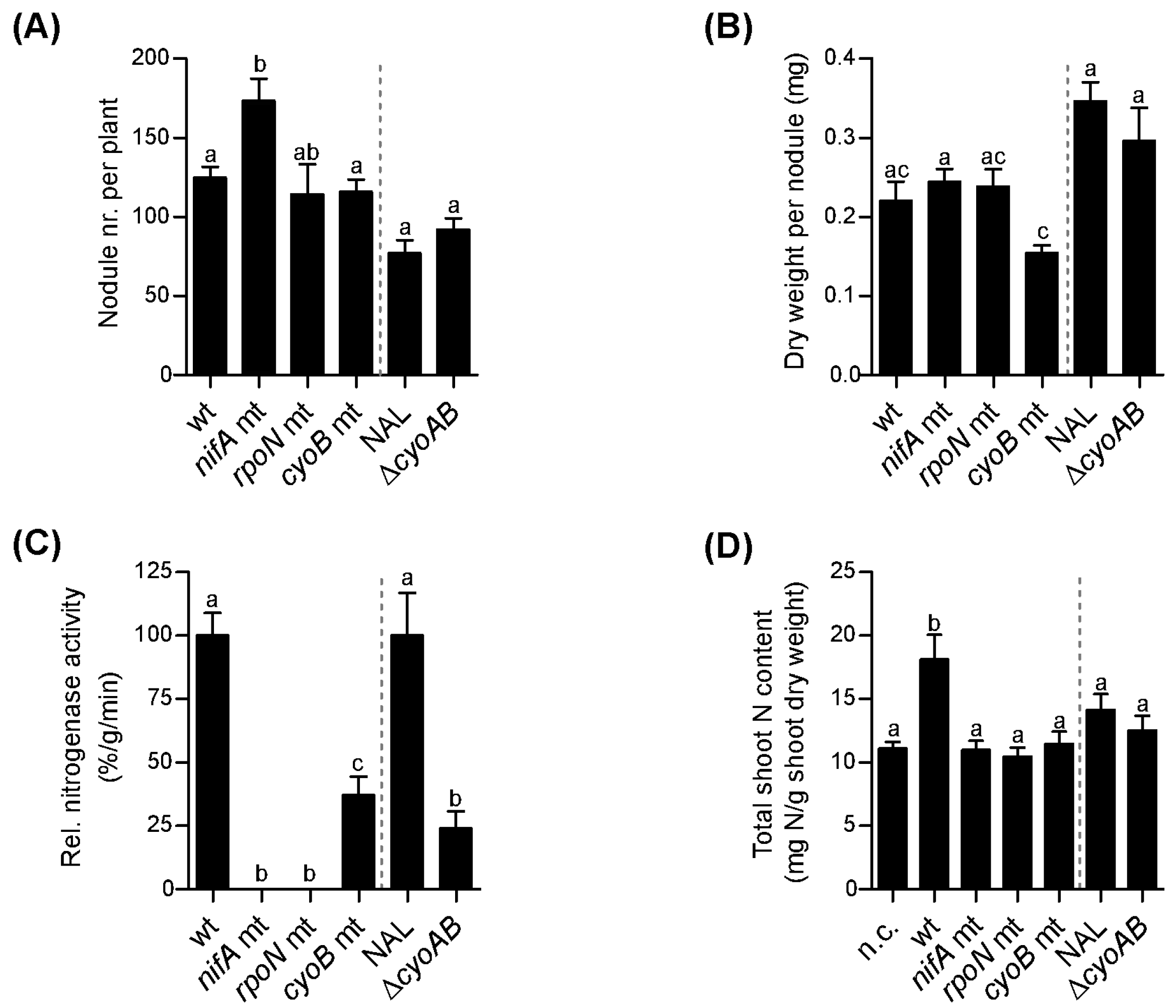
3.3. Role of nifA, rpoN, and cyoB Genes during Symbiosis
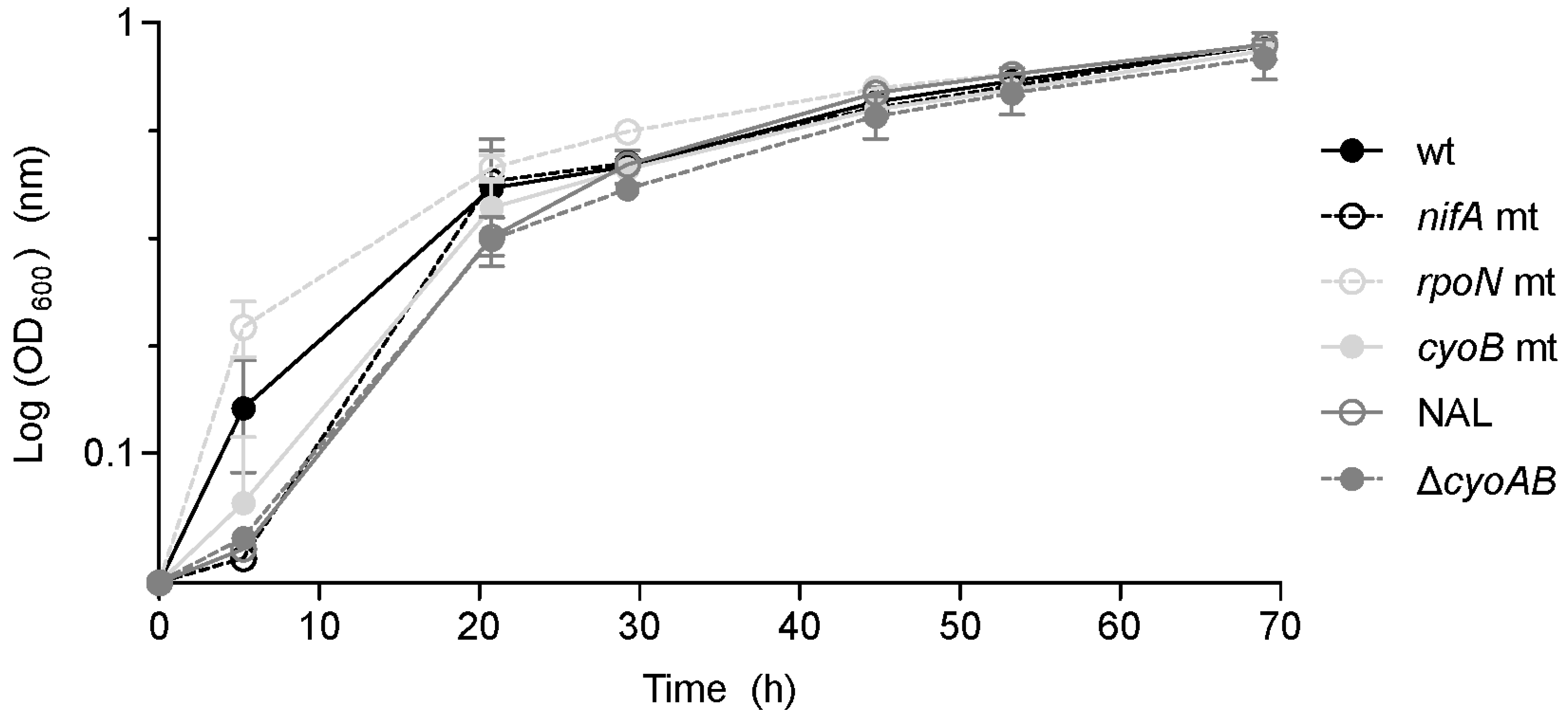
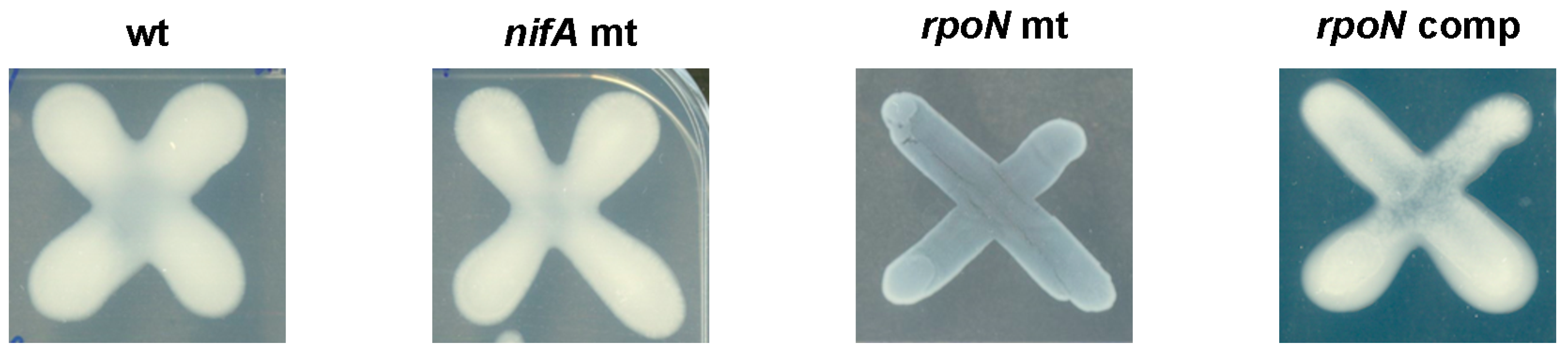
3.4. Role of RpoN in Free-Living Conditions
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Vitousek, P.M.; Hättenschwiler, S.; Olander, L.; Allison, S. Nitrogen and nature. Ambio 2002, 31, 97–101. [Google Scholar] [CrossRef] [PubMed]
- Moulin, L.; Munive, A.; Dreyfus, B.; Boivin-Masson, C. Nodulation of legumes by members of the β-subclass of Proteobacteria. Nature 2001, 411, 948–950. [Google Scholar] [CrossRef] [PubMed]
- Elliott, G.N.; Chen, W.M.; Bontemps, C.; Chou, J.H.; Young, J.P.; Sprent, J.I.; James, E.K. Nodulation of Cyclopia spp. (Leguminosae, Papilionoideae) by Burkholderia tuberum. Ann. Bot. 2007, 100, 1403–1411. [Google Scholar] [CrossRef] [PubMed]
- Elliott, G.N.; Chen, W.M.; Chou, J.H.; Wang, H.C.; Sheu, S.Y.; Perin, L.; Reis, V.M.; Moulin, L.; Simon, M.F.; Bontemps, C.; et al. Burkholderia phymatum is a highly effective nitrogen-fixing symbiont of Mimosa spp. and fixes nitrogen ex planta. New Phytol. 2007, 173, 168–180. [Google Scholar] [CrossRef] [PubMed]
- Mishra, R.P.N.; Tisseyre, P.; Melkonian, R.; Chaintreuil, C.; Miché, L.; Klonowska, A.; Gonzalez, S.; Bena, G.; Laguerre, G.; Moulin, L. Genetic diversity of Mimosa pudica rhizobial symbionts in soils of French Guiana: investigating the origin and diversity of Burkholderia phymatum and other beta-rhizobia. FEMS Microbiol. Ecol. 2012, 79, 487–503. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.M.; Laevens, S.; Lee, T.M.; Coenye, T.; De Vos, P.; Mergeay, M.; Vandamme, P. Ralstonia taiwanensis sp. nov., isolated from root nodules of Mimosa species and sputum of a cystic fibrosis patient. Int. J. Syst. Evol. Microbiol. 2001, 51, 1729–1735. [Google Scholar] [CrossRef] [PubMed]
- Bontemps, C.; Elliott, G.N.; Simon, M.F.; dos Reis, F.B., Jr.; Gross, E.; Lawton, R.C.; Neto, N.E.; de Fatima Loureiro, M.; De Faria, S.M.; Sprent, J.I.; et al. Burkholderia species are ancient symbionts of legumes. Mol. Ecol. 2010, 19, 44–52. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; James, E.K.; Prescott, A.R.; Kierans, M.; Sprent, J.I. Nodulation of Mimosa spp. by the β-proteobacterium Ralstonia taiwanensis. Mol. Plant-Microbe Interact. 2003, 16, 1051–1061. [Google Scholar] [CrossRef] [PubMed]
- Gyaneshwar, P.; Hirsch, A.M.; Moulin, L.; Chen, W.M.; Elliott, G.N.; Bontemps, C.; Estrada-de los Santos, P.; Gross, E.; dos Reis, F.B., Jr.; Sprent, J.I.; et al. Legume-nodulating betaproteobacteria: diversity, host range, and future prospects. Mol. Plant-Microbe Interact. 2011, 24, 1276–1288. [Google Scholar] [CrossRef] [PubMed]
- Angus, A.A.; Lee, A.; Lum, M.R.; Shehayeb, M.; Hessabi, R.; Fujishige, N.A.; Yerrapragada, S.; Kano, S.; Song, N.; Yang, P.; et al. Nodulation and effective nitrogen fixation of Macroptilium atropurpureum (siratro) by Burkholderia tuberum, a nodulating and plant growth promoting beta-proteobacterium, are influenced by environmental factors. Plant Soil 2013, 369, 543–562. [Google Scholar] [CrossRef]
- Beukes, C.W.; Venter, S.N.; Law, I.J.; Phalane, F.L.; Steenkamp, E.T. South African papilionoid legumes are nodulated by diverse Burkholderia with unique nodulation and nitrogen-fixation loci. PLoS ONE 2013, 8, e68406. [Google Scholar] [CrossRef] [PubMed]
- Garau, G.; Yates, R.J.; Deiana, P.; Howieson, J.G. Novel strains of nodulating Burkholderia have a role in nitrogen fixation with papilionoid herbaceous legumes adapted to acid, infertile soils. Soil Biol. Biochem. 2009, 41, 125–134. [Google Scholar] [CrossRef]
- Howieson, J.G.; De Meyer, S.E.; Vivas-Marfisi, A.; Ratnayake, S.; Ardley, J.K.; Yates, R.J. Novel Burkholderia bacteria isolated from Lebeckia ambigua—A perennial suffrutescent legume of the fynbos. Soil Biol. Biochem. 2013, 60, 55–64. [Google Scholar] [CrossRef]
- Lemaire, B.; Chimphango, S.B.M.; Stirton, C.; Rafudeen, S.; Honnay, O.; Smets, E.; Chen, W.M.; Sprent, J.; James, E.K.; Muasya, A.M. Biogeographical patterns of legume-nodulating Burkholderia spp.: From African fynbos to continental scales. Appl. Environ. Microbiol. 2016, 82, 5099–5115. [Google Scholar] [CrossRef] [PubMed]
- Lemaire, B.; Van Cauwenberghe, J.; Verstraete, B.; Chimphango, S.; Stirton, C.; Honnay, O.; Smets, E.; Sprent, J.; James, E.K.; Muasya, A.M. Characterization of the papilionoid-Burkholderia interaction in the Fynbos biome: the diversity and distribution of beta-rhizobia nodulating Podalyria calyptrata (Fabaceae, Podalyrieae). Syst. Appl. Microbiol. 2016, 39, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Talbi, C.; Delgado, M.J.; Girard, L.; Ramírez-Trujillo, A.; Caballero-Mellado, J.; Bedmar, E.J. Burkholderia phymatum strains capable of nodulating Phaseolus vulgaris are present in Moroccan soils. Appl. Environ. Microbiol. 2010, 76, 4587–4591. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.Y.Y.; Ridgway, H.J.; James, T.K.; James, E.K.; Chen, W.M.; Sprent, J.I.; Young, J.P.W.; Andrews, M. Burkholderia sp. induces functional nodules on the South African invasive legume Dipogon lignosus (Phaseoleae) in New Zealand Soils. Microb. Ecol. 2014, 68, 542–555. [Google Scholar] [CrossRef] [PubMed]
- Lemaire, B.; Dlodlo, O.; Chimphango, S.; Stirton, C.; Schrire, B.; Boatwright, J.S.; Honnay, O.; Smets, E.; Sprent, J.; James, E.K.; et al. Symbiotic diversity, specificity and distribution of rhizobia in native legumes of the Core Cape Subregion (South Africa). FEMS Microbiol. Ecol. 2015, 91. [Google Scholar] [CrossRef] [PubMed]
- Beukes, C.W.; Palmer, M.; Manyaka, P.; Chan, W.Y.; Avontuur, J.R.; van Zyl, E.; Huntemann, M.; Clum, A.; Pillay, M.; Palaniappan, K.; et al. Genome data provides high support for generic boundaries in Burkholderia sensu lato. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Sawana, A.; Adeolu, M.; Gupta, R.S. Molecular signatures and phylogenomic analysis of the genus Burkholderia: Proposal for division of this genus into the emended genus Burkholderia containing pathogenic organisms and a new genus Paraburkholderia gen. nov. harboring environmental species. Front. Genet. 2014, 5, 429. [Google Scholar] [CrossRef] [PubMed]
- Vandamme, P.; Goris, J.; Chen, W.-M.; de Vos, P.; Willems, A. Burkholderia tuberum sp. nov. and Burkholderia phymatum sp. nov., nodulate the roots of tropical legumes. Syst. Appl. Microbiol. 2002, 25, 507–512. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.M.; James, E.K.; Coenye, T.; Chou, J.H.; Barrios, E.; de Faria, S.M.; Elliott, G.N.; Sheu, S.Y.; Sprent, J.I.; Vandamme, P. Burkholderia mimosarum sp. nov., isolated from root nodules of Mimosa spp. from Taiwan and South America. Int. J. Syst. Evol. Microbiol. 2006, 56, 1847–1851. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.M.; de Faria, S.M.; James, E.K.; Elliott, G.N.; Lin, K.Y.; Chou, J.H.; Sheu, S.Y.; Cnockaert, M.; Sprent, J.I.; Vandamme, P. Burkholderia nodosa sp. nov., isolated from root nodules on the woody Brazilian legumes Mimosa bimucronata and Mimosa scabrella. Int. J. Syst. Evol. Microbiol. 2007, 57, 1055–1059. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.M.; de Faria, S.M.; Chou, J.H.; James, E.K.; Elliott, G.N.; Sprent, J.I.; Bontemps, C.; Young, J.P.W.; Vandamme, P. Burkholderia sabiae sp. nov., isolated from root nodules of Mimosa caesalpiniifolia. Int. J. Syst. Evol. Microbiol. 2008, 58, 2174–2179. [Google Scholar] [CrossRef] [PubMed]
- Bournaud, C.; de Faria, S.M.; Ferreira dos Santos, J.M.; Tisseyre, P.; Silva, M.; Chaintreuil, C.; Gross, E.; James, E.K.; Prin, Y.; Moulin, L. Burkholderia species are the most common and preferred nodulating symbionts of the Piptadenia Group (tribe Mimoseae). PLoS ONE 2013, 8, e63478. [Google Scholar] [CrossRef] [PubMed]
- Sheu, S.Y.; Chou, J.H.; Bontemps, C.; Elliott, G.N.; Gross, E.; dos Reis, F.B., Jr.; Melkonian, R.; Moulin, L.; James, E.K.; Sprent, J.I.; et al. Burkholderia diazotrophica sp. nov., isolated from root nodules of Mimosa spp. Int. J. Syst. Evol. Microbiol. 2013, 63, 435–441. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Aguilar, L.; Salazar-Salazar, C.; Méndez, R.D.; Caballero-Mellado, J.; Hirsch, A.M.; Vásquez-Murrieta, M.S.; Estrada-de Los Santos, P. Burkholderia caballeronis sp. nov., a nitrogen fixing species isolated from tomato (Lycopersicon esculentum) with the ability to effectively nodulate Phaseolus vulgaris. Antonie Van Leeuwenhoek 2013, 104, 1063–1071. [Google Scholar] [CrossRef] [PubMed]
- De Meyer, S.E.; Cnockaert, M.; Ardley, J.K.; Maker, G.; Yates, R.; Howieson, J.G.; Vandamme, P. Burkholderia sprentiae sp. nov., isolated from Lebeckia ambigua root nodules. Int. J. Syst. Evol. Microbiol. 2013, 63, 3950–3957. [Google Scholar] [CrossRef] [PubMed]
- De Meyer, S.E.; Cnockaert, M.; Ardley, J.K.; Trengove, R.D.; Garau, G.; Howieson, J.G.; Vandamme, P. Burkholderia rhynchosiae sp. nov., isolated from Rhynchosia ferulifolia root nodules. Int. J. Syst. Evol. Microbiol. 2013, 63, 3944–3949. [Google Scholar] [CrossRef] [PubMed]
- De Meyer, S.E.; Cnockaert, M.; Ardley, J.K.; Van Wyk, B.E.; Vandamme, P.A.; Howieson, J.G. Burkholderia dilworthii sp. nov., isolated from Lebeckia ambigua root nodules. Int. J. Syst. Evol. Microbiol. 2014, 64, 1090–1095. [Google Scholar] [CrossRef] [PubMed]
- Mavengere, N.R.; Ellis, A.G.; Le Roux, J.J. Burkholderia aspalathi sp. nov., isolated from root nodules of the South African legume Aspalathus abietina Thunb. Int. J. Syst. Evol. Microbiol. 2014, 64, 1906–1912. [Google Scholar] [CrossRef] [PubMed]
- Steenkamp, E.T.; van Zyl, E.; Beukes, C.W.; Avontuur, J.R.; Chan, W.Y.; Palmer, M.; Mthombeni, L.S.; Phalane, F.L.; Sereme, T.K.; Venter, S.N. Burkholderia kirstenboschensis sp. nov. nodulates papilionoid legumes indigenous to South Africa. Syst. Appl. Microbiol. 2015, 38, 545–554. [Google Scholar] [CrossRef] [PubMed]
- Sheu, S.Y.; Chen, M.H.; Liu, W.Y.Y.; Andrews, M.; James, E.K.; Ardley, J.K.; De Meyer, S.E.; James, T.K.; Howieson, J.G.; Coutinho, B.G.; Chen, W.M. Burkholderia dipogonis sp. nov, isolated from root nodules of Dipogon lignosus in New Zealand and Western Australia. Int. J. Syst. Evol. Microbiol. 2015, 65, 4716–4723. [Google Scholar] [CrossRef] [PubMed]
- Bournaud, C.; Moulin, L.; Cnockaert, M.; de Faria, S.; Prin, Y.; Severac, D.; Vandamme, P. Paraburkholderia piptadeniae sp. nov. and Paraburkholderia ribeironis sp. nov., two root-nodulating symbiotic species of Piptadenia gonoacantha in Brazil. Int. J. Syst. Evol. Microbiol. 2017, 67, 432–440. [Google Scholar] [CrossRef] [PubMed]
- Estrada-de los Santos, P.; Rojas-Rojas, F.U.; Tapia-García, E.Y.; Vásquez-Murrieta, M.S.; Hirsch, A.M. To split or not to split: An opinion on dividing the genus Burkholderia. Ann. Microbiol. 2016, 66, 1303–1314. [Google Scholar] [CrossRef]
- Dixon, R.; Kahn, D. Genetic regulation of biological nitrogen fixation. Nat. Rev. Microbiol. 2004, 2, 621–631. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, R.A.; Klopprogge, K.; Grabbe, R. Regulation of nitrogen fixation in Klebsiella pneumoniae and Azotobacter vinelandii: NifL, transducing two environmental signals to the nif transcriptional activator NifA. J. Mol. Microbiol. Biotechnol. 2002, 4, 235–242. [Google Scholar] [PubMed]
- Fischer, H.M. Genetic regulation of nitrogen fixation in rhizobia. Microbiol. Rev. 1994, 58, 352–386. [Google Scholar] [PubMed]
- Bobik, C.; Meilhoc, E.; Batut, J. FixJ: A major regulator of the oxygen limitation response and late symbiotic functions of Sinorhizobium meliloti. J. Bacteriol. 2006, 188, 4890–4902. [Google Scholar] [CrossRef] [PubMed]
- Terpolilli, J.J.; Hood, G.A.; Poole, P.S. What determines the efficiency of N2-fixing Rhizobium-legume symbioses? Adv. Microb. Physiol. 2012, 60, 325–389. [Google Scholar] [PubMed]
- MacLean, A.M.; Finan, T.M.; Sadowsky, M.J. Genomes of the symbiotic nitrogen-fixing bacteria of legumes. Plant Physiol. 2007, 144, 615–622. [Google Scholar] [CrossRef] [PubMed]
- Amadou, C.; Pascal, G.; Mangenot, S.; Glew, M.; Bontemps, C.; Capela, D.; Carrere, S.; Cruveiller, S.; Dossat, C.; Lajus, A.; et al. Genome sequence of the beta-rhizobium Cupriavidus taiwanensis and comparative genomics of rhizobia. Genome Res. 2008, 18, 1472–1483. [Google Scholar] [CrossRef] [PubMed]
- Tian, C.F.; Zhou, Y.J.; Zhang, Y.M.; Li, Q.Q.; Zhang, Y.Z.; Li, D.F.; Wang, S.; Wang, J.; Gilbert, L.B.; Li, Y.R.; et al. Comparative genomics of rhizobia nodulating soybean suggests extensive recruitment of lineage-specific genes in adaptations. Proc. Natl. Acad. Sci. USA 2012, 109, 8629–8634. [Google Scholar] [CrossRef] [PubMed]
- De Meyer, S.E.; Briscoe, L.; Martínez-Hidalgo, P.; Agapakis, C.M.; Estrada-de los Santos, P.; Seshadri, R.; Reeve, W.; Weinstock, G.; O’Hara, G.; Howieson, J.G.; et al. Symbiotic Burkholderia species show diverse arrangements of nif/fix and nod genes and lack typical high-affinity cytochrome cbb3 oxidase genes. Mol. Plant-Microbe Interact. 2016, 29, 609–619. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wei, S.; Wang, F.; James, E.K.; Guo, X.; Zagar, C.; Xia, L.G.; Dong, X.; Wang, Y.P. Burkholderia and Cupriavidus spp. are the preferred symbionts of Mimosa spp. in Southern China. FEMS Microbiol. Ecol. 2012, 80, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Gehlot, H.S.; Tak, N.; Kaushik, M.; Mitra, S.; Chen, W.M.; Poweleit, N.; Panwar, D.; Poonar, N.; Parihar, R.; Tak, A.; et al. An invasive Mimosa in India does not adopt the symbionts of its native relatives. Ann. Bot. 2013, 112, 179–196. [Google Scholar] [CrossRef] [PubMed]
- dos Reis, F.B., Jr.; Simon, M.F.; Gross, E.; Boddey, R.M.; Elliott, G.N.; Neto, N.E.; de Fatima Loureiro, M.; de Queiroz, L.P.; Scotti, M.R.; Chen, W.-M.; et al. Nodulation and nitrogen fixation by Mimosa spp. in the Cerrado and Caatinga biomes of Brazil. New Phytol. 2010, 186, 934–946. [Google Scholar] [CrossRef] [PubMed]
- Moulin, L.; Klonowska, A.; Caroline, B.; Booth, K.; Vriezen, J.A.; Melkonian, R.; James, E.K.; Young, J.P.W.; Bena, G.; Hauser, L.; et al. Complete Genome sequence of Burkholderia phymatum STM815T, a broad host range and efficient nitrogen-fixing symbiont of Mimosa species. Stand. Genomic Sci. 2014, 9, 763–774. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.H. Experiments in Molecular Genetics; Cold Spring Harbor Lab Press: Cold Spring Harbor, NY, USA, 1972; pp. 352–355. [Google Scholar]
- Clark, D.J.; Maaløe, M. DNA replication and the division cycle in Escherichia coli. J. Mol. Biol. 1967, 23, 99–112. [Google Scholar] [CrossRef]
- Regensburger, B.; Hennecke, H. RNA polymerase from Rhizobium japonicum. Arch. Microbiol. 1983, 135, 103–109. [Google Scholar] [CrossRef] [PubMed]
- Pessi, G.; Braunwalder, R.; Grunau, A.; Omasits, U.; Ahrens, C.H.; Eberl, L. Response of Burkholderia cenocepacia H111 to micro-oxia. PLoS ONE 2013, 8, e72939. [Google Scholar] [CrossRef] [PubMed]
- Talbi, C.; Argandoña, M.; Salvador, M.; Alché, J.D.; Vargas, C.; Bedmar, E.J.; Delgado, M.J. Burkholderia phymatum improves salt tolerance of symbiotic nitrogen fixation in Phaseolus vulgaris. Plant Soil 2012, 367, 673–685. [Google Scholar] [CrossRef]
- Hahn, M.; Hennecke, H. Localized Mutagenesis in Rhizobium japonicum. Mol. Gen. Genet. 1984, 193, 46–52. [Google Scholar] [CrossRef]
- Göttfert, M.; Hitz, S.; Hennecke, H. Identification of nodS and nodU, two inducible genes inserted between the Bradyrhizobium japonicum nodYABC and nodIJ genes. Mol. Plant-Microbe Interact. 1990, 3, 308–316. [Google Scholar] [CrossRef] [PubMed]
- Koch, M.; Delmotte, N.; Rehrauer, H.; Vorholt, J.A.; Pessi, G.; Hennecke, H. Rhizobial adaptation to hosts, a new facet in the legume root-nodule symbiosis. Mol. Plant-Microbe Interact. 2010, 23, 784–790. [Google Scholar] [CrossRef] [PubMed]
- Lardi, M.; Bolzan de Campos, S.; Purtschert, G.; Eberl, L.; Pessi, G. Competition experiments for legume infection identify Burkholderia phymatum as a highly competitive β-rhizobium. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Pessi, G.; Ahrens, C.H.; Rehrauer, H.; Lindemann, A.; Hauser, F.; Fischer, H.-M.; Hennecke, H. Genome-wide transcript analysis of Bradyrhizobium japonicum bacteroids in soybean root nodules. Mol. Plant-Microbe Interact. 2007, 20, 1353–1363. [Google Scholar] [CrossRef] [PubMed]
- Lardi, M.; Aguilar, C.; Pedrioli, A.; Omasits, U.; Suppiger, A.; Cárcamo-Oyarce, G.; Schmid, N.; Ahrens, C.H.; Eberl, L.; Pessi, G. σ54-dependent response to nitrogen limitation and virulence in Burkholderia cenocepacia H111. Appl. Environ. Microbiol. 2015, 81, 12. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef] [PubMed]
- Powell, S.; Szklarczyk, D.; Trachana, K.; Roth, A.; Kuhn, M.; Muller, J.; Arnold, R.; Rattei, T.; Letunic, I.; Doerks, T.; et al. eggNOG v3.0: Orthologous groups covering 1133 organisms at 41 different taxonomic ranges. Nucleic Acids Res. 2012, 40. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Beringer, J.E. R Factor Transfer in Rhizobium leguminosarum. J. Gen. Microbiol. 1974, 84, 188–198. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Lardi, M.; Pedrioli, A.; Eberl, L.; Pessi, G. NtrC-dependent control of exopolysaccharide synthesis and motility in Burkholderia cenocepacia H111. PLoS ONE 2017, 12, e0180362. [Google Scholar] [CrossRef] [PubMed]
- Lardi, M.; Pessi, G. Nitrogen Sources Utilization of P. phymatum Using Biolog PM3 Plates; University of Zurich: Zurich, Switzerland, 2017. [Google Scholar]
- Reitzer, L. Nitrogen assimilation and global regulation in Escherichia coli. Annu. Rev. Microbiol. 2003, 57, 155–176. [Google Scholar] [CrossRef] [PubMed]
- Huergo, L.F.; Chandra, G.; Merrick, M. PII signal transduction proteins: Nitrogen regulation and beyond. FEMS Microbiol. Rev. 2013, 37, 251–283. [Google Scholar] [CrossRef] [PubMed]
- Lardi, M.; Pessi, G. Differential Transcript Expression by P. phymarum Grown under Micro-oxic and Aerobic Conditions; University of Zurich: Zurich, Switzerland, 2017. [Google Scholar]
- Yoshida, K.; Takemoto, Y.; Sotsuka, T.; Tanaka, K.; Takenaka, S. PhaP phasins play a principal role in poly-β-hydroxybutyrate accumulation in free-living Bradyrhizobium japonicum. BMC Microbiol. 2013, 13, 290. [Google Scholar] [CrossRef] [PubMed]
- Labes, M.; Rastogi, V.; Watson, R.; Finan, T.M. Symbiotic nitrogen fixation by a nifA deletion mutant of Rhizobium meliloti: The role of an unusual ntrC allele. J. Bacteriol. 1993, 175, 2662–2673. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhao, K.; Liu, M.; Burgess, R.R. Promoter and regulon analysis of nitrogen assimilation factor, σ54, reveal alternative strategy for E. coli MG1655 flagellar biosynthesis. Nucleic Acids Res. 2009, 38, 1273–1283. [Google Scholar] [CrossRef] [PubMed]
- van Heeswijk, W.C.; Westerhoff, H.V.; Boogerd, F.C. Nitrogen assimilation in Escherichia coli: Putting molecular data into a systems perspective. Microbiol. Mol. Biol. Rev. 2013, 77, 628–695. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.I.; Donati, A.J.; Hahn, D.; Tisa, L.S.; Chang, W.S. Alteration of the exopolysaccharide production and the transcriptional profile of free-living Frankia strain CcI3 under nitrogen-fixing conditions. Appl. Microbiol. Biotechnol. 2013, 97, 10499–10509. [Google Scholar] [CrossRef] [PubMed]
- Williams, A.; Wilkinson, A.; Krehenbrink, M.; Russo, D.M.; Zorreguieta, A.; Downie, J.A. Glucomannan-mediated attachment of Rhizobium leguminosarum to pea root hairs is required for competitive nodule infection. J. Bacteriol. 2008, 190, 4706–4715. [Google Scholar] [CrossRef] [PubMed]
- Downie, J.A. The roles of extracellular proteins, polysaccharides and signals in the interactions of rhizobia with legume roots. FEMS Microbiol. Rev. 2010, 34, 150–170. [Google Scholar] [CrossRef] [PubMed]
- Scharf, B.E.; Hynes, M.F.; Alexandre, G.M. Chemotaxis signaling systems in model beneficial plant–bacteria associations. Plant Mol. Biol. 2016, 90, 549–559. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.Y.; Arsène, F.; Elmerich, C. Characterization of the ntrBC genes of Azospirillam brasilense Sp7: Their involvement in the regulation of nitrogenase synthesis and activity. Mol. Gen. Genet. 1993, 240, 188–196. [Google Scholar] [CrossRef] [PubMed]
- Hakoyama, T.; Niimi, K.; Watanabe, H.; Tabata, R.; Matsubara, J.; Sato, S.; Nakamura, Y.; Tabata, S.; Jichun, L.; Matsumoto, T.; et al. Host plant genome overcomes the lack of a bacterial gene for symbiotic nitrogen fixation. Nature 2009, 462, 514–517. [Google Scholar] [CrossRef] [PubMed]
- McLean, P.A.; Dixon, R.A. Requirement of nifV gene for production of wild-type nitrogenase enzyme in Klebsiella pneumoniae. Nature 1981, 292, 655–656. [Google Scholar] [CrossRef] [PubMed]
- Mora, Y.; Diaz, R.; Vargas-Lagunas, C.; Peralta, H.; Guerrero, G.; Aguilar, A.; Encarnacion, S.; Girard, L.; Mora, J. Nitrogen-fixing rhizobial strains isolated from common bean seeds: Phylogeny, physiology, and genome analysis. Appl. Environ. Microbiol. 2014, 80, 5644–5654. [Google Scholar] [CrossRef] [PubMed]
- Yamaura, M.; Uchiumi, T.; Higashi, S.; Abe, M.; Kucho, K.I. Identification by suppression subtractive hybridization of Frankia genes induced under nitrogen-fixing conditions. Appl. Environ. Microbiol. 2010, 76, 1692–1694. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.M.; Prell, J.; James, E.K.; Sheu, D.S.; Sheu, S.Y. Biosynthesis of branched-chain amino acids is essential for effective symbioses between betarhizobia and Mimosa pudica. Microbiology 2012, 158, 1758–1766. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.M.; Prell, J.; James, E.K.; Sheu, D.S.; Sheu, S.Y. Effect of phosphoglycerate mutase and fructose 1,6-bisphosphatase deficiency on symbiotic Burkholderia phymatum. Microbiology 2012, 158, 1127–1136. [Google Scholar] [CrossRef] [PubMed]
- Green, L.S.; Karr, D.B.; Emerich, D.W. Isocitrate dehydrogenase and glyoxylate cycle enzyme activities in Bradyrhizobium japonicum under various growth conditions. Arch. Microbiol. 1998, 169, 445–451. [Google Scholar] [CrossRef] [PubMed]
- Delmotte, N.; Ahrens, C.H.; Knief, C.; Qeli, E.; Koch, M.; Fischer, H.M.; Vorholt, J.A.; Hennecke, H.; Pessi, G. An integrated proteomics and transcriptomics reference data set provides new insights into the Bradyrhizobium japonicum bacteroid metabolism in soybean root nodules. Proteomics 2010, 10, 1391–1400. [Google Scholar] [CrossRef] [PubMed]
- Gourion, B.; Delmotte, N.; Bonaldi, K.; Nouwen, N.; Vorholt, J.A.; Giraud, E. Bacterial RuBisCO is required for efficient Bradyrhizobium/Aeschynomene symbiosis. PLoS ONE 2011, 6, e21900. [Google Scholar] [CrossRef] [PubMed]
- Lardi, M.; Pessi, G. Nitrogenase Activity of Bean Nodules Induced by P. phymatum nifA Mutant Strain Measured 28 Days Post Infection; University of Zurich: Zurich, Switzerland, 2017. [Google Scholar]
- Hauser, F.; Pessi, G.; Friberg, M.; Weber, C.; Rusca, N.; Lindemann, A.; Fischer, H.M.; Hennecke, H. Dissection of the Bradyrhizobium japonicum NifA+σ54 regulon, and identification of a ferredoxin gene (fdxN) for symbiotic nitrogen fixation. Mol. Genet. Genomics 2007, 278, 255–271. [Google Scholar] [CrossRef] [PubMed]
- Lardi, M.; Murset, V.; Fischer, H.M.; Mesa, S.; Ahrens, C.H.; Zamboni, N.; Pessi, G. Metabolomic profiling of Bradyrhizobium diazoefficiens-induced root nodules reveals both host plant-specific and developmental signatures. Int. J. Mol. Sci. 2016, 17, 815. [Google Scholar] [CrossRef] [PubMed]
| Locus ID 1 | Description 1 | Gene Name | FC (S vs. N) 2 | FC (S vs. N) 3 |
|---|---|---|---|---|
| Bphy_0257 | Ammonium transporter | amtB | 19.7 ± 4.6 | 10.9 |
| Bphy_0326 | RNA polymerase factor σ54 | rpoN | 1.4 ± 0.1 | 0.7 |
| Bphy_1479 | Nitrogen metabolism transcriptional regulator | ntrC | 8.8 ± 1.2 | 9.7 |
| Bphy_1481 | Glutamine synthetase, type I | glnA | 4.0 ± 0.8 | 1.9 |
| Locus ID 1 | Description 1 | Gene Name | FC (Bacteroids vs. Free-Living) 2 | FC (MO vs. O) 3 |
|---|---|---|---|---|
| Bphy_0326 | RNA polymerase factor σ54 | rpoN | 2.1 ± 0.5 | 1.6 |
| Bphy_1479 | Nitrogen metabolism transcriptional regulator | ntrC | 6.6 ± 1.6 | 0.9 |
| Bphy_3648 | Cytochrome o ubiquinol oxidase, subunit I | cyoB | 201.8 ± 44.0 | 68.3 |
| Bphy_7728 | Transcriptional regulator | nifA | 19.1 ± 2.4 | 4.2 |
| Bphy_7753 | Nitrogenase reductase | nifH | 898.4 ± 174.2 | 567.0 |
| Locus ID 1 | Description 1 | Gene Name | Log2FC (Bacteroids vs. Free-Living) 2 |
|---|---|---|---|
| Energy production and conversion | |||
| Bphy_1368 | isocitrate lyase | 3.1 | |
| Bphy_1649 | alkanesulfonate monooxygenase | 6.0 | |
| Bphy_1848 | 2-oxoacid dehydrogenase subunit E1 | 2.9 | |
| Bphy_2272 | FAD linked oxidase domain-containing protein | 2.9 | |
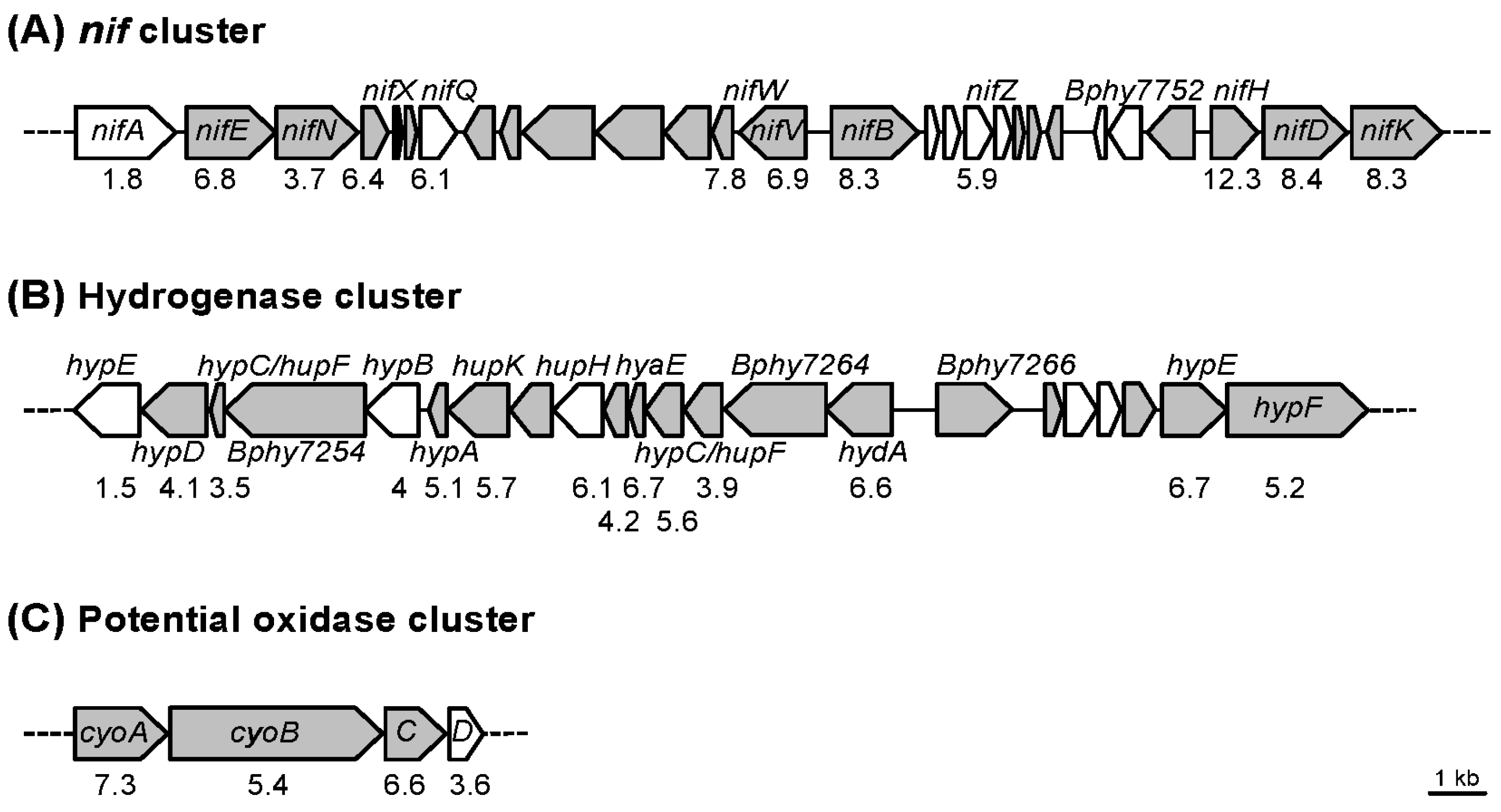
| Bphy_3647 | cytochrome o ubiquinol oxidase, subunit III | cyoC | 6.6 |
| Bphy_3648 | cytochrome o ubiquinol oxidase, subunit I | cyoB | 5.4 |
| Bphy_3649 | ubiquinol oxidase, subunit II | cyoA | 7.3 |
| Bphy_3685 | phosphate acetyltransferase | 2.0 | |
| Bphy_4116 | rubrerythrin | 1.9 | |
| Bphy_4949 | aldehyde dehydrogenase | 2.6 | |
| Bphy_5235 | alkanesulfonate monooxygenase | 6.2 | |
| Bphy_5817 | putative flavodoxin | 3.6 | |
| Bphy_6055 | hypothetical protein | INF | |
| Bphy_6505 | formylmethanofuran dehydrogenase subunit A | 1.9 | |
| Bphy_6506 | formylmethanofuran-tetrahydromethanopterin formyltransferase | 3.9 | |
| Bphy_6671 | 2Fe-2S iron-sulfur cluster binding domain-containing protein | INF | |
| Bphy_6672 | carbon-monoxide dehydrogenase (acceptor) | 3.7 | |
| Bphy_6673 | aldehyde oxidase and xanthine dehydrogenase molybdopterin binding | 4.5 | |
| Bphy_7231 | cytochrome ce class I | 2.8 | |
| Bphy_7232 | xenobiotic (desulfurization)monooxygenase subunit A | 5.3 | |
| Bphy_7262 | hydrogenase expression/formation protein | 5.6 | |
| Bphy_7263 | Ni/Fe-hydrogenase, b-type cytochrome subunit | 3.9 | |
| Bphy_7264 | nickel-dependent hydrogenase large subunit | 6.4 | |
| Bphy_7265 | hydrogenase (NiFe) small subunit | hydA | 6.6 |
| Bphy_7406 | aldehyde dehydrogenase | 7.3 | |
| Bphy_7729 | nitrogenase MoFe cofactor biosynthesis protein | nifE | 6.8 |
| Bphy_7730 | nitrogenase molybdenum-cofactor biosynthesis protein | nifN | 3.7 |
| Bphy_7733 | ferredoxin III, nif-specific | 6.3 | |
| Bphy_7737 | electron-transferring-flavoprotein dehydrogenase | fixC | 6.8 |
| Bphy_7738 | electron transfer flavoprotein α/β-subunit | fixB | 7.1 |
| Bphy_7739 | electron transfer flavoprotein α/β-subunit | fixA | 7.8 |
| Bphy_7754 | nitrogenase molybdenum-iron protein α chain | nifD | 8.4 |
| Bphy_7755 | nitrogenase molybdenum-iron protein β chain | nifK | 8.3 |
| Bphy_7804 | electron transfer flavoprotein α/β-subunit | 3.9 | |
| Inorganic ion transport and metabolism | |||
| Bphy_0882 | phosphate ABC transporter, periplasmic protein | 4.3 | |
| Bphy_0883 | phosphate transporter permease subunit | pstC | 2.5 |
| Bphy_0885 | phosphate transporter ATP-binding protein | 2.6 | |
| Bphy_1627 | sulfate ABC transporter inner membrane subunit | cysW | 2.2 |
| Bphy_1629 | sulfate ABC transporter, periplasmic protein | 5.2 | |
| Bphy_1647 | ABC transporter-like protein | 5.9 | |
| Bphy_1648 | binding-protein-dependent transport systems | 3.3 | |
| Bphy_2231 | sulfate adenylyltransferase large subunit | 2.3 | |
| Bphy_2521 | catalase | 4.9 | |
| Bphy_3120 | phosphate ABC transporter, periplasmic protein | 4.2 | |
| Bphy_3602 | ABC transporter related | 4.0 | |
| Bphy_3603 | nitrate/sulfonate/bicarbonate ABC transporter, periplasmic protein | 2.6 | |
| Bphy_3854 | phosphate transporter | 2.7 | |
| Bphy_4233 | Rieske (2Fe-2S) domain-containing protein | 2.9 | |
| Bphy_4622 | phosphonate ABC transporter binding protein | 6.7 | |
| Bphy_5040 | lipoprotein | 7.8 | |
| Bphy_5065 | 2-aminoethylphosphonate ABC transporter, 2-aminoethylphosphonate binding protein | 2.0 | |
| Bphy_5226 | aliphatic sulfonate ABC transporter, periplasmic protein | 4.0 | |
| Bphy_5227 | substrate-binding region of ABC-type glycine betaine transport system | 9.0 | |
| Bphy_5229 | aliphatic sulfonate ABC transporter, periplasmic protein | 5.8 | |
| Bphy_5232 | rhodanese domain-containing protein | 5.6 | |
| Bphy_6080 | taurine ABC transporter, periplasmic binding protein | 7.5 | |
| Bphy_6081 | ABC transporter related | 4.2 | |
| Bphy_6550 | metallophosphoesterase | 3.5 | |
| Bphy_7233 | ABC transporter related | 3.7 | |
| Bphy_7234 | binding-protein-dependent transport systems | 4.6 | |
| Bphy_7235 | binding-protein-dependent transport systems | 4.8 | |
| Bphy_7236 | ABC sulfate ester transporter, periplasmic protein | 4.3 | |
| Bphy_7645 | binding-protein-dependent transport systems | 3.2 | |
| Bphy_7646 | binding-protein-dependent transport systems | 4.4 | |
| Bphy_7647 | ABC transporter related | 4.9 | |
| Bphy_7753 | nitrogenase reductase | nifH | 12.3 |
| Bphy_7808 | nitrogenase reductase | nifH | INF |
| Translation, ribosomal structure and biogenesis | |||
| Bphy_2864 | GCN5-like N-acetyltransferase | 4.2 | |
| Nitrogen Source (s) | Utilization of Nitrogen | |||
|---|---|---|---|---|
| wt | nifA mt | rpoN mt | rpoN complemented | |
| 30 mM NH4Cl | + | + | + | + |
| 0.5 mM NH4Cl | + | + | + | ± |
| 30 mM NO3− | + | + | - | + |
| 15 mM CH4N2O | + | + | - | + |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lardi, M.; Liu, Y.; Purtschert, G.; Bolzan de Campos, S.; Pessi, G. Transcriptome Analysis of Paraburkholderia phymatum under Nitrogen Starvation and during Symbiosis with Phaseolus Vulgaris. Genes 2017, 8, 389. https://doi.org/10.3390/genes8120389
Lardi M, Liu Y, Purtschert G, Bolzan de Campos S, Pessi G. Transcriptome Analysis of Paraburkholderia phymatum under Nitrogen Starvation and during Symbiosis with Phaseolus Vulgaris. Genes. 2017; 8(12):389. https://doi.org/10.3390/genes8120389
Chicago/Turabian StyleLardi, Martina, Yilei Liu, Gabriela Purtschert, Samanta Bolzan de Campos, and Gabriella Pessi. 2017. "Transcriptome Analysis of Paraburkholderia phymatum under Nitrogen Starvation and during Symbiosis with Phaseolus Vulgaris" Genes 8, no. 12: 389. https://doi.org/10.3390/genes8120389
APA StyleLardi, M., Liu, Y., Purtschert, G., Bolzan de Campos, S., & Pessi, G. (2017). Transcriptome Analysis of Paraburkholderia phymatum under Nitrogen Starvation and during Symbiosis with Phaseolus Vulgaris. Genes, 8(12), 389. https://doi.org/10.3390/genes8120389






