Comparative Analysis of the Complete Chloroplast Genome of Four Known Ziziphus Species
Abstract
:1. Introduction
2. Materials and Methods
2.1. Samples
2.2. Genome Sequencing
2.3. Chloroplast Genome Assembly
2.4. Genome Annotation
2.5. Identification of Repeat Sequences and Simple Sequence Repeats
2.6. Genome-Wide Sequence Variations and Gene Selective Pressure Analysis among Four Ziziphus Chloroplast Genomes
2.7. Phylogenetic Analysis
3. Results
3.1. Genome Assembly and Validation
3.2. Genome Structure, Gene Content, and Features
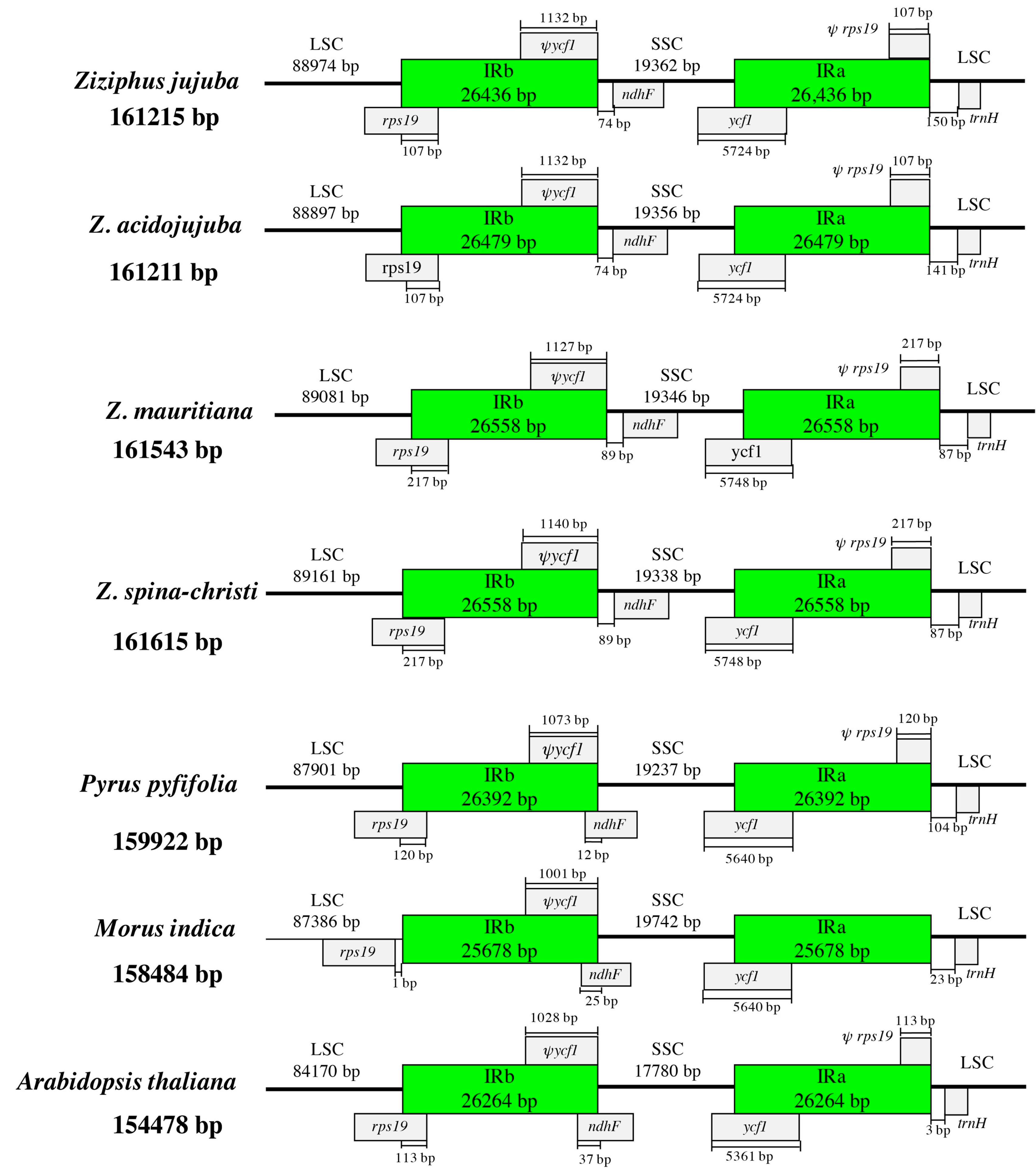
3.3. IR Expansion and Contraction
3.4. Repetitive Sequences
3.5. Single Sequence Repeat Polymorphism
3.6. Global Sequence Variation across Chloroplast Genomes
3.7. Evolutionary Rates
3.8. Phylogenetic Analysis Based on the Protein-Coding Genes
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Islam, M.B.; Simmons, M.P. A thorny dilemma: Testing alternative intrageneric classifications within Ziziphus (Rhamnaceae). Syst. Bot. 2006, 31, 826–842. [Google Scholar] [CrossRef]
- Liu, M.J.; Zhao, Z.H. Germplasm resources and production of jujube in China. Acta Hortic. 2009, 840, 25–32. [Google Scholar] [CrossRef]
- Shengrui, Y. Past, present, and future of jujubes—Chinese Dates in the United States. Hortscience 2013, 48, 672–680. [Google Scholar]
- Dov, P.; Albert, N.; Ali, I.; Debesaye, S.; Djibrilla, I. How domesticated Ziziphus mauritiana (Lam) spread in the Sahel region of Africa and in Ethiopia. Chron. Hort. 2016, 56, 5. [Google Scholar]
- Bal, J.S. Development and production of Indian jujube (Ber) in India. Acta Hortic. 2016, 1116, 15–22. [Google Scholar] [CrossRef]
- Dafni, A.; Levy, S.; Lev, E. The ethnobotany of Christ’s Thorn Jujube (Ziziphus spina-christi) in Israel. J. Ethnobiol. Ethnomed. 2005, 1, 8. [Google Scholar] [CrossRef] [PubMed]
- Saied, A.S.; Gebauer, J.; Hammer, K.; Buerkert, A. Ziziphus spina-christi (L.) Willd.: A multipurpose fruit tree. Genet. Resour. Crop. Evol. 2007, 55, 929–937. [Google Scholar] [CrossRef]
- Liu, M.J.; Zhao, J.; Cai, Q.L.; Liu, G.C.; Wang, J.R.; Zhao, Z.H.; Liu, P.; Dai, L.; Yan, G.; Wang, W.J.; et al. The complex jujube genome provides insights into fruit tree biology. Nat. Commun. 2014, 5, 5315. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhang, C.; Zhao, X.; Fei, Z.; Wan, K.; Zhang, Z.; Pang, X.; Yin, X.; Bai, Y.; Sun, X.; et al. The Jujube genome provides insights into genome evolution and the domestication of sweetness/acidity taste in fruit trees. PLoS Genet. 2016, 12, e1006433. [Google Scholar] [CrossRef] [PubMed]
- Kearse, M.; Moir, R.; Wilson, A.A.; Stoneshavas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Ripma, L.A.; Simpson, M.G.; Hasenstab-Lehman, K. Geneious! Simplified genome skimming methods for phylogenetic systematic studies: A case study in Oreocarya (Boraginaceae). Appl. Plant Sci. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Schattner, P.; Brooks, A.N.; Lowe, T.M. The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res. 2005, 33, W686–W689. [Google Scholar] [CrossRef] [PubMed]
- Lohse, M.; Drechsel, O.; Bock, R. Organellar genome draw (OGDRAW): A tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr. Genet. 2007, 52, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Kurtz, S.; Choudhuri, J.V.; Ohlebusch, E.; Schleiermacher, C.; Stoye, J.; Giegerich, R. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 2001, 29, 4633–4642. [Google Scholar] [CrossRef] [PubMed]
- Martins, W.S.; Lucas, D.C.S.; de Souza Neves, K.F.; Bertioli, D.J. WebSat—A web software for microsatellite marker development. Bioinformation 2009, 3, 282. [Google Scholar] [CrossRef] [PubMed]
- Mayor, C.; Brudno, M.; Schwartz, J.; Poliakov, A.; Rubin, E.; Frazer, K.; Pachter, L.; Dubchak, I. VISTA: Visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 2000, 16, 1046–1047. [Google Scholar] [CrossRef] [PubMed]
- Li, W.H.; Wu, C.I.; Luo, C.C. A new method for estimating synonymous and nonsynonymous rates of nucleotide substitution considering the relative likelihood of nucleotide and codon changes. Mol. Biol. Evol. 1985, 2, 150–174. [Google Scholar] [PubMed]
- Li, W.H. Unbiased estimation of the rates of synonymous and nonsynonymous substitution. J. Mol. Evol. 1993, 36, 96–99. [Google Scholar] [CrossRef] [PubMed]
- Pamilo, P.; Bianchi, N. Evolution of the Zfx and Zfy genes: Rates and interdependence between the genes. Mol. Biol. Evol. 1993, 10, 271–281. [Google Scholar] [PubMed]
- Yang, Z. PAML 4: Phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 2007, 24, 1586–1591. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Swofford, D.L. PAUP*: Phylogenetic Analysis Using Parsimony, Version 4.0 b10; Sinauer Associates: Sunderland, MA, USA, 2003. [Google Scholar] [CrossRef]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [PubMed]
- Jansen, R.K.; Cai, Z.; Raubeson, L.A.; Daniell, H.; Depamphilis, C.W.; Leebens-Mack, J.; Muller, K.F.; Guisinger-Bellian, M.; Haberle, R.C.; Hansen, A.K.; et al. Analysis of 81 genes from 64 plastid genomes resolves relationships in angiosperms and identifies genome-scale evolutionary patterns. Proc. Natl. Acad. Sci. USA 2007, 104, 19369–19374. [Google Scholar] [CrossRef] [PubMed]
- Asatryan, A.; Tel-Zur, N. Intraspecific and interspecific crossability in three Ziziphus species (Rhamnaceae). Genet. Resour. Crop. Evol. 2013, 61, 215–233. [Google Scholar] [CrossRef]
- Terakami, S.; Matsumura, Y.; Kurita, K.; Kanamori, H.; Katayose, Y.; Yamamoto, T.; Katayama, H. Complete sequence of the chloroplast genome from pear (Pyrus pyrifolia): Genome structure and comparative analysis. Tree Genet. Genomes 2012, 8, 841–854. [Google Scholar] [CrossRef]
- Lu, R.S.; Li, P.; Qiu, Y.X. The complete chloroplast genomes of three Cardiocrinum (Liliaceae) Species: Comparative genomic and phylogenetic analyses. Front. Plant Sci. 2016, 7, 2054. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Yang, X.; Zhang, C.; Yin, X.; Liu, S.; Li, X. Development of chloroplast microsatellite markers and analysis of chloroplast diversity in Chinese Jujube (Ziziphus jujuba Mill.) and Wild Jujube (Ziziphus acidojujuba Mill.). PLoS ONE 2015, 10, e0134519. [Google Scholar] [CrossRef] [PubMed]
- Neubig, K.M.; Whitten, W.M.; Carlsward, B.S.; Blanco, M.A.; Endara, L.; Williams, N.H.; Moore, M. Phylogenetic utility of ycf1 in orchids: A plastid gene more variable than matK. Plant Syst. Evol. 2009, 277, 75–84. [Google Scholar] [CrossRef]
- Sarkinen, T.; George, M. Predicting plastid marker variation: Can complete plastid genomes from closely related species help? PLoS ONE 2013, 8, e82266. [Google Scholar] [CrossRef] [PubMed]
- Neubig, K.M.; Abbott, J.R. Primer development for the plastid region ycf1 in Annonaceae and other magnoliids. Am. J. Bot. 2010, 97, e52–e55. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Shi, C.; Gao, L.Z. Plastid genome sequence of a wild woody oil species, Prinsepia utilis, provides insights into evolutionary and mutational patterns of rosaceae chloroplast genomes. PLoS ONE 2013, 8, e73946. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Shi, C.; Liu, Y.; Mao, S.-Y.; Gao, L.-Z. Thirteen Camellia chloroplast genome sequences determined by high-throughput sequencing: Genome structure and phylogenetic relationships. BMC Evol. Biol. 2014, 14, 151. [Google Scholar] [CrossRef] [PubMed]
- Zheng, B.; Halperin, T.; Hruskova-Heidingsfeldova, O.; Adam, Z.; Clarke, A.K. Characterization of chloroplast Clp proteins in Arabidopsis: Localization, tissue specificity and stress responses. Physiol. Plant. 2002, 114, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Kode, V.; Mudd, E.A.; Iamtham, S.; Day, A. The tobacco plastid accD gene is essential and is required for leaf development. Plant J. 2005, 44, 237–244. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.J.; Lee, H.L. Complete chloroplast genome sequences from Korean ginseng (Panax schinseng Nees) and comparative analysis of sequence evolution among 17 vascular plants. DNA Res. 2004, 11, 247–261. [Google Scholar] [CrossRef] [PubMed]
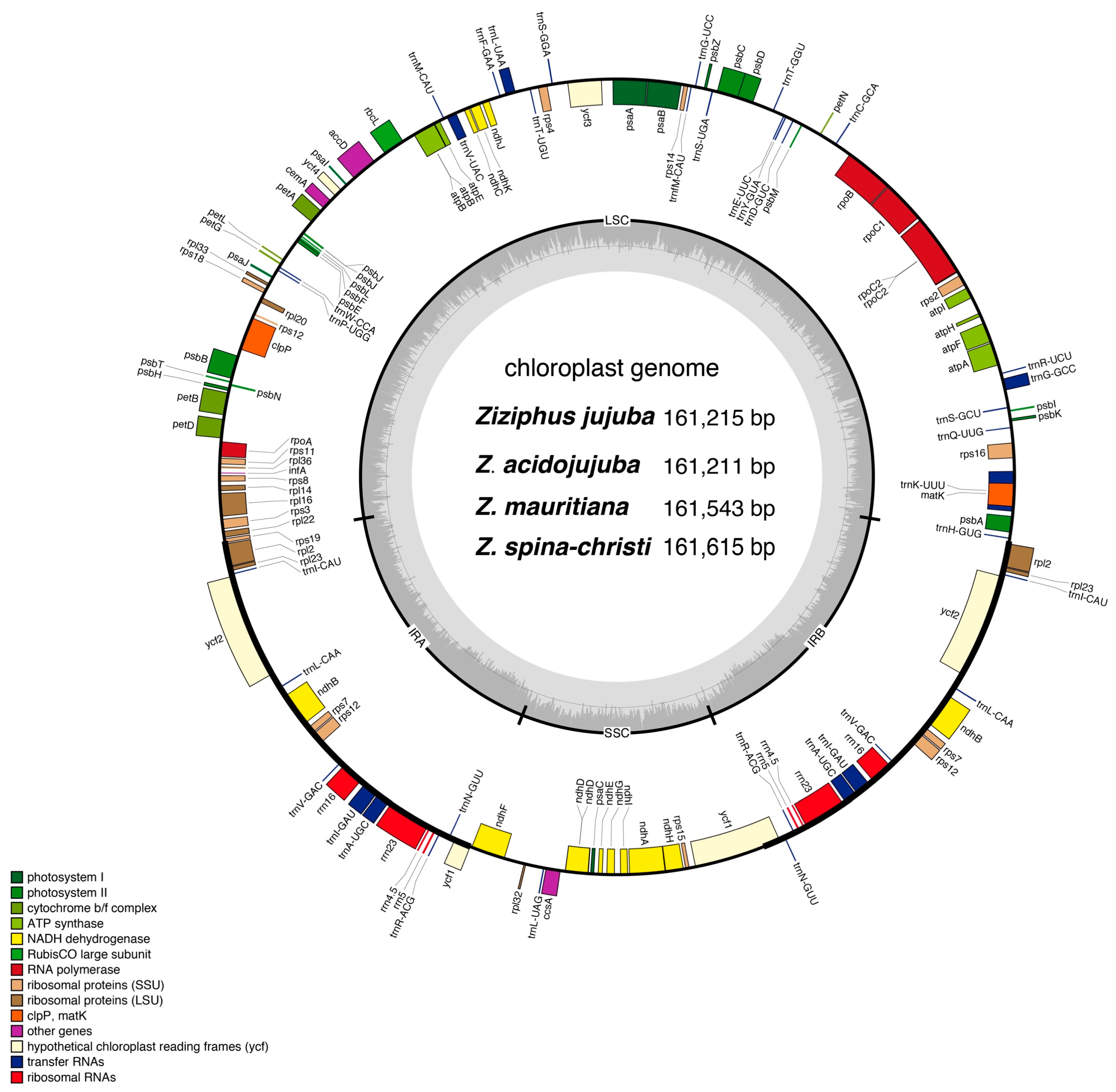



| Category | Group | Genes |
|---|---|---|
| Self-replication | Large subunit of ribosome (LSU) | rpl2(×2)*, rpl14, rpl16*, rpl20, rpl22, rpl23(×2), rpl32, rpl33, rpl36 |
| Small subunit of ribosome (SSU) | rps2, rps3, rps4, rps7(×2), rps8, rps11, rps12*(×2, part), rps14, rps15, rps16*, rps18, rps19 | |
| DNA dependent RNA polymerase | rpoA, rpoB, rpoC1*, rpoC2 | |
| Ribosomal RNA | rrn4.5(×2), rrn5(×2), rrn16(×2), rrn23(×2) | |
| Transfer RNAs (tRNA) | trnA-UGC(×2)*, trnC-GCA, trnD-GUC, trnE-UUC, trnF-GAA, trnG-GCC*, trnG-UCC, trnH-GUG, trnI-CAU(×2), trnI-GAU(×2)*, trnK-UUU*, trnL-CAA(×2), trnL-UAA*, trnL-UAG, trnM-CAU, trnfM-CAU, trnN-GUU(×2), trnP-UGG, trnQ-UUG, trnR-ACG(×2), trnR-UCU, trnS-GCU, trnS-UGA, trnS-GGA, trnT-UGU, trnT-GGU, trnV-GAC(×2), trnV-UAC*, trnW-CCA, trnY-GUA | |
| Photosynthesis | Photosystem I | psaA, psaB, psaC, psaI, psaJ, ycf3**, ycf4 |
| Photosystem II | psbA, psbB, psbC, psbD, psbE, psbF, psbH, psbI, psbJ, psbK, psbL, psbM, psbN, psbZ, psbT | |
| NADH dehydrogenase | ndhA*, ndhB(×2)*, ndhC, ndhD, ndhE, ndhF, ndhG, ndhH, ndhI, ndhJ, ndhK | |
| Cytochrome b/f complex | petA, petB*, petD*, petG, petL, petN | |
| Subunits of ATP synthase | atpA, atpB, atpE, atpF*, atpH, atpI, | |
| Large subunit of rubisco | rbcL | |
| Other genes | Translational initiation factor | infA |
| ATP-dependent protease subunit p gene | clpP** | |
| Maturase | matK | |
| Envelop membrane protein | cemA | |
| Unknown function | Subunit of acetyl-CoA-carboxylase | accD |
| c-type cytochrome synthesis gene | ccsA | |
| Hypothetical chloroplast reading frames | ycf1(×2), ycf2(×2) |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, J.; Chen, R.; Li, X. Comparative Analysis of the Complete Chloroplast Genome of Four Known Ziziphus Species. Genes 2017, 8, 340. https://doi.org/10.3390/genes8120340
Huang J, Chen R, Li X. Comparative Analysis of the Complete Chloroplast Genome of Four Known Ziziphus Species. Genes. 2017; 8(12):340. https://doi.org/10.3390/genes8120340
Chicago/Turabian StyleHuang, Jian, Ruihong Chen, and Xingang Li. 2017. "Comparative Analysis of the Complete Chloroplast Genome of Four Known Ziziphus Species" Genes 8, no. 12: 340. https://doi.org/10.3390/genes8120340
APA StyleHuang, J., Chen, R., & Li, X. (2017). Comparative Analysis of the Complete Chloroplast Genome of Four Known Ziziphus Species. Genes, 8(12), 340. https://doi.org/10.3390/genes8120340




