An Inactive Geminin Mutant That Binds Cdt1
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plasmids
2.2. Protein Expression
2.3. Replication Reactions
2.4. In Vitro Binding Studies
2.5. In Vivo Chromatin Binding Studies
2.6. Isothermal Calorimetry
2.7. HBO1 Acetylation Assays
2.8. Antibodies
3. Results
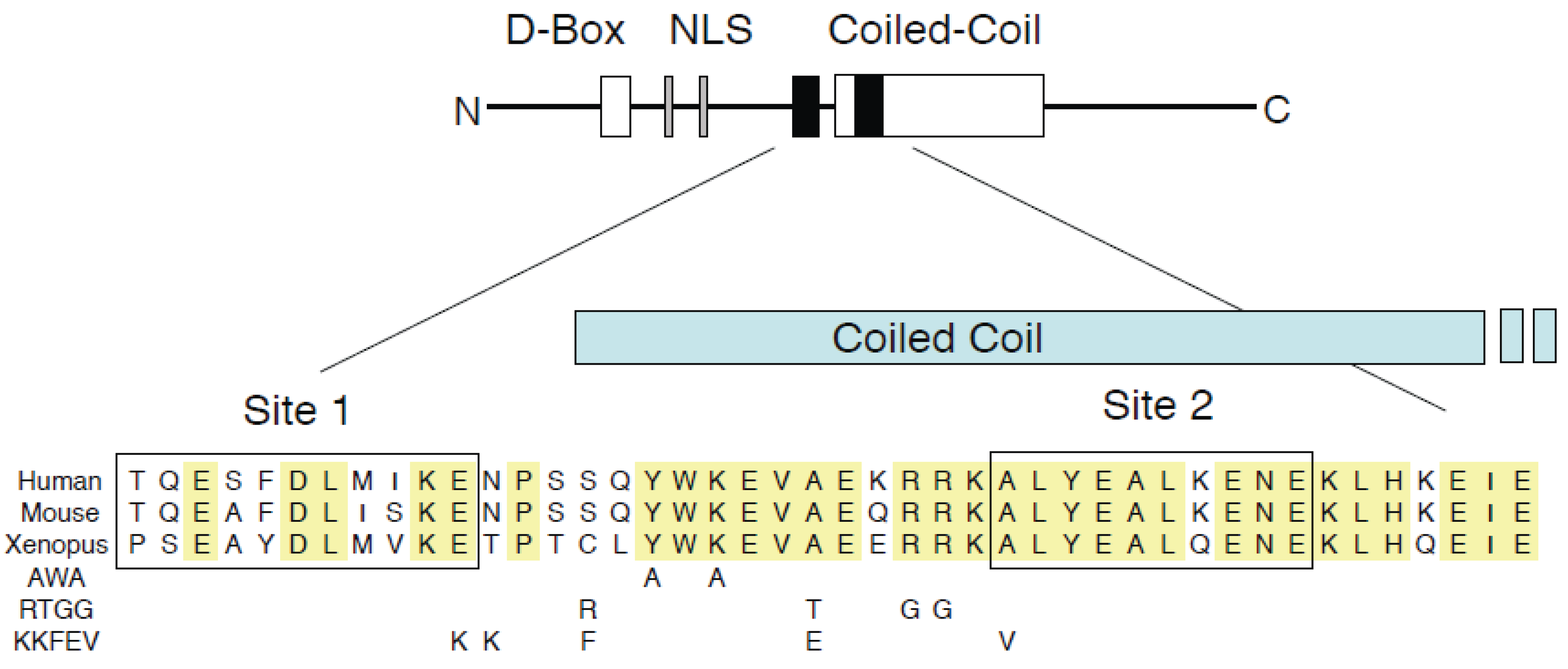
3.1. GemininAWA Binds Cdt1, Yet Fails to Inhibit DNA Replication

3.2. GemininAWA Allows MCM Loading In Vivo
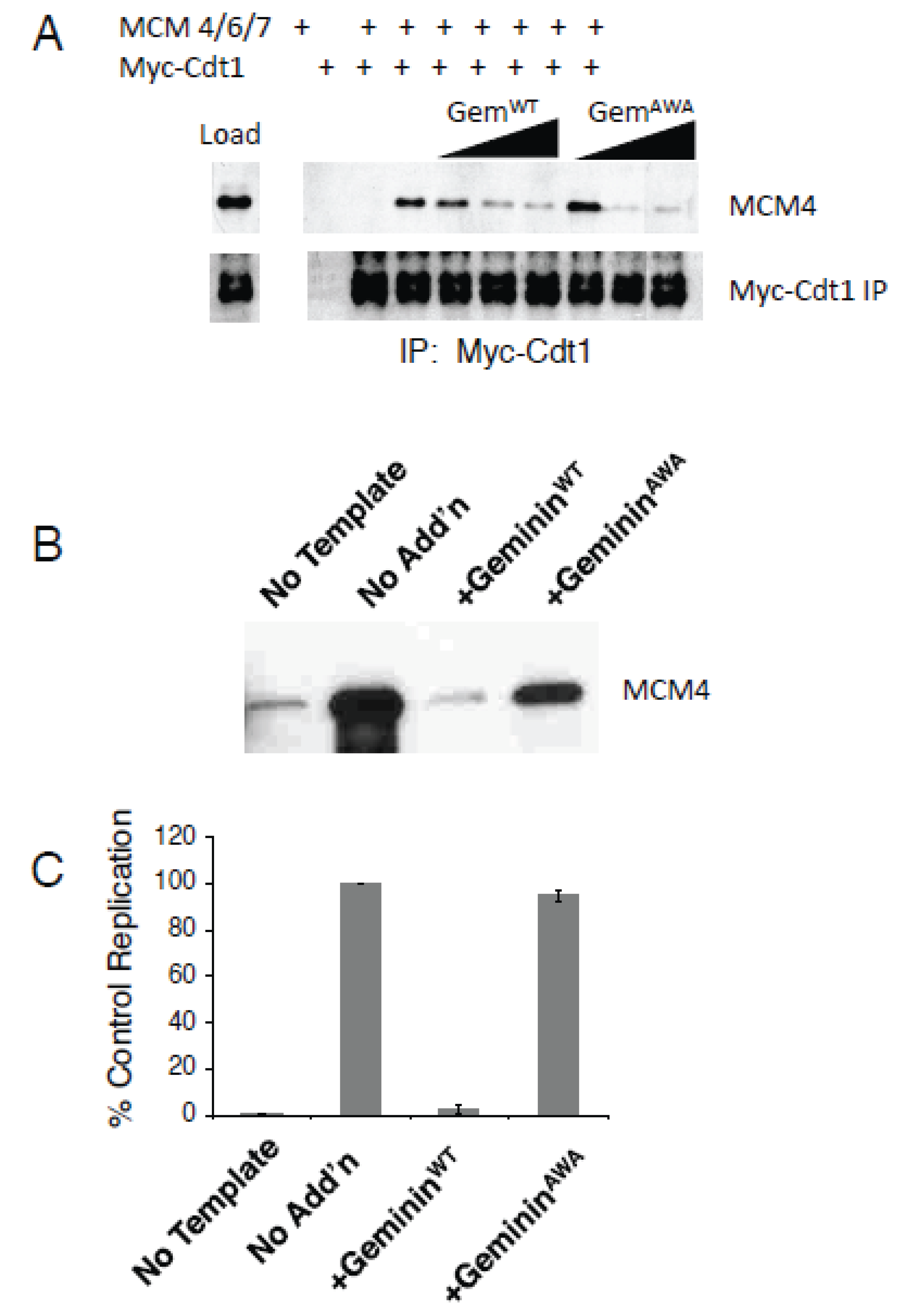
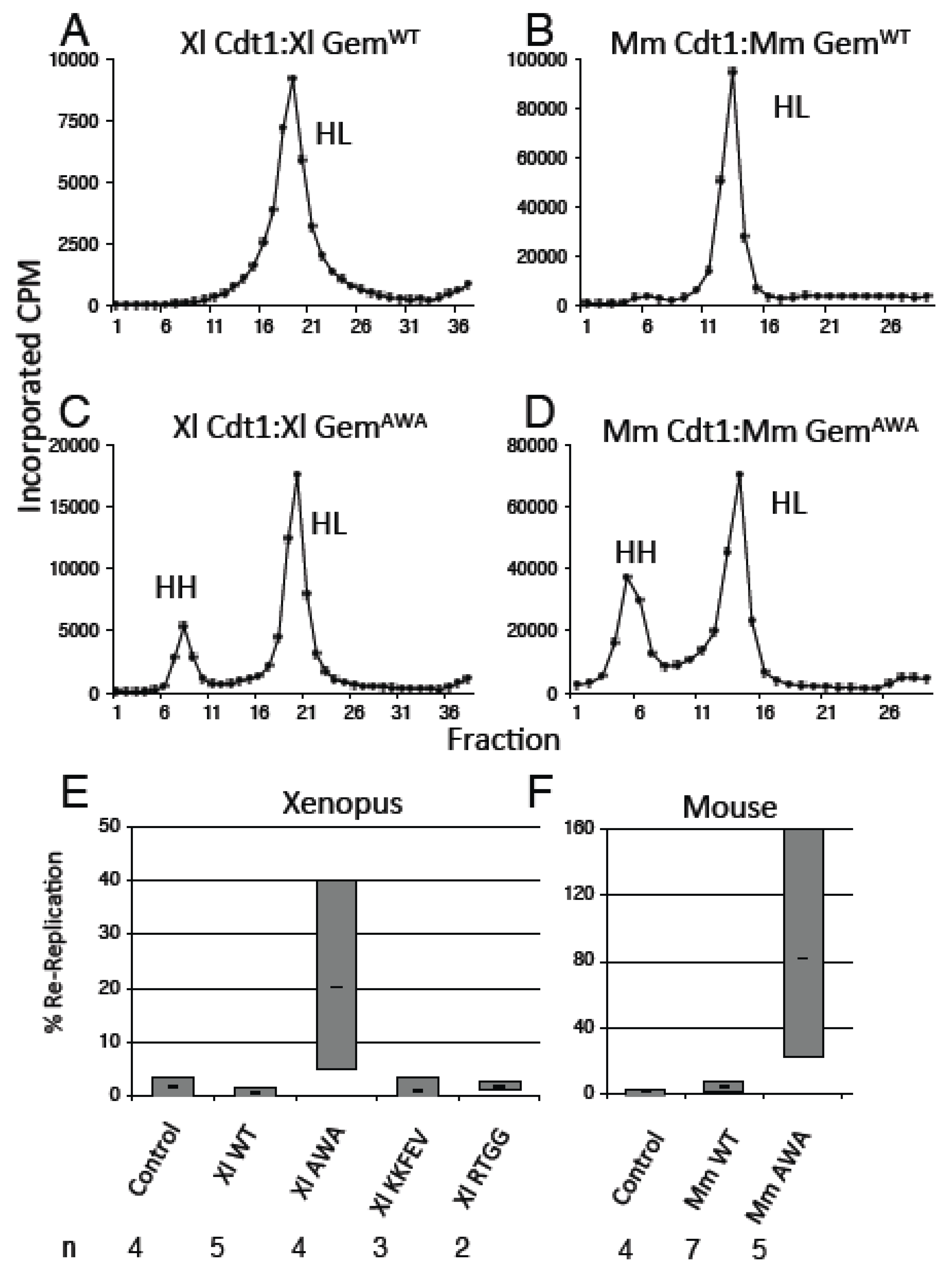
3.3. GemininAWA Does Not Inhibit a Second Round of Replication
3.4. GemininAWA Does Not Inhibit the Histone Acetyltransferase HBO1

4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Arias, E.E.; Walter, J.C. Strength in numbers: Preventing rereplication via multiple mechanisms in eukaryotic cells. Genes Dev. 2007, 21, 497–518. [Google Scholar] [CrossRef] [PubMed]
- Maiorano, D.; Moreau, J.; Mechali, M. XCDT1 is required for the assembly of pre-replicative complexes in Xenopus laevis. Nature 2000, 404, 622–625. [Google Scholar] [CrossRef] [PubMed]
- Arias, E.E.; Walter, J.C. PCNA functions as a molecular platform to trigger Cdt1 destruction and prevent re-replication. Nat. Cell Biol. 2006, 8, 84–90. [Google Scholar] [CrossRef] [PubMed]
- Arias, E.E.; Walter, J.C. Replication-dependent destruction of Cdt1 limits DNA replication to a single round per cell cycle in Xenopus egg extracts. Genes Dev. 2005, 19, 114–126. [Google Scholar] [CrossRef] [PubMed]
- McGarry, T.J.; Kirschner, M.W. Geminin, an inhibitor of DNA replication, is degraded during mitosis. Cell 1998, 93, 1043–1053. [Google Scholar] [CrossRef] [PubMed]
- Tada, S.; Li, A.; Maiorano, D.; Mechali, M.; Blow, J.J. Repression of origin assembly in metaphase depends on inhibition of RLF-B/Cdt1 by geminin. Nat. Cell Biol. 2001, 3, 107–113. [Google Scholar] [CrossRef] [PubMed]
- Wohlschlegel, J.A.; Dwyer, B.T.; Dhar, S.K.; Cvetic, C.; Walter, J.C.; Dutta, A. Inhibition of eukaryotic DNA replication by geminin binding to Cdt1. Science 2000, 290, 2309–2312. [Google Scholar] [CrossRef] [PubMed]
- Kerns, S.L.; Torke, S.J.; Benjamin, J.M.; McGarry, T.J. Geminin prevents rereplication during xenopus development. J. Biol. Chem. 2007, 282, 5514–5521. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Blow, J.J. Cdt1 downregulation by proteolysis and geminin inhibition prevents DNA re-replication in Xenopus. EMBO J. 2005, 24, 395–404. [Google Scholar] [CrossRef] [PubMed]
- Benjamin, J.M.; Torke, S.J.; Demeler, B.; McGarry, T.J. Geminin has dimerization, Cdt1-binding, and destruction domains that are required for biological activity. J. Biol. Chem. 2004, 279, 45957–45968. [Google Scholar] [CrossRef] [PubMed]
- Yanagi, K.; Mizuno, T.; You, Z.; Hanaoka, F. Mouse geminin inhibits not only Cdt1-MCM6 interactions but also a novel intrinsic Cdt1 DNA binding activity. J. Biol. Chem. 2002, 277, 40871–40880. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Hong, B.; Choi, J.M.; Kim, Y.; Watanabe, S.; Ishimi, Y.; Enomoto, T.; Tada, S.; Cho, Y. Structural basis for inhibition of the replication licensing factor Cdt1 by geminin. Nature 2004, 430, 913–917. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Lu, W.; Santos, R.E.; Frattini, M.G.; Kelly, T.J. Geminin inhibits a late step in the formation of human pre-replicative complexes. J. Biol. Chem. 2014, 289, 30810–30821. [Google Scholar] [CrossRef] [PubMed]
- Miotto, B.; Struhl, K. HBO1 histone acetylase is a coactivator of the replication licensing factor Cdt1. Genes Dev. 2008, 22, 2633–2638. [Google Scholar] [CrossRef] [PubMed]
- Miotto, B.; Struhl, K. HBO1 histone acetylase activity is essential for DNA replication licensing and inhibited by Geminin. Mol. Cell 2010, 37, 57–66. [Google Scholar] [CrossRef] [PubMed]
- Wong, P.G.; Glozak, M.A.; Cao, T.V.; Vaziri, C.; Seto, E.; Alexandrow, M. Chromatin unfolding by Cdt1 regulates MCM loading via opposing functions of HBO1 and HDAC11-geminin. Cell Cycle 2010, 9, 4351–4363. [Google Scholar] [CrossRef] [PubMed]
- Iizuka, M.; Matsui, T.; Takisawa, H.; Smith, M.M. Regulation of replication licensing by acetyltransferase Hbo1. Mol. Cell Biol. 2006, 26, 1098–1108. [Google Scholar] [CrossRef] [PubMed]
- McGarry, T.J. Measurement of geminin activity in Xenopus egg extracts. Methods Mol. Biol. 2005, 296, 263–278. [Google Scholar] [PubMed]
- You, Z.; Ishimi, Y.; Masai, H.; Hanaoka, F. Roles of MCM7 and MCM4 subunits in the DNA helicase activity of the mouse MCM4/6/7 complex. J. Biol. Chem. 2002, 277, 42471–42479. [Google Scholar] [CrossRef] [PubMed]
- Saxena, S.; Yuan, P.; Dhar, S.K.; Senga, T.; Takeda, D.; Robinson, H.; Kornbluth, S.; Swaminathan, K.; Dutta, A. A dimerized coiled-coil domain and an adjoining part of geminin interact with two sites on Cdt1 for replication inhibition. Mol. Cell 2004, 15, 245–258. [Google Scholar] [CrossRef] [PubMed]
- De Marco, V.; Gillespie, P.J.; Li, A.; Karantzelis, N.; Christodoulou, E.; Klompmaker, R.; van Gerwen, S.; Fish, A.; Petoukhov, M.V.; Iliou, M.S.; et al. Quaternary structure of the human Cdt1-Geminin complex regulates DNA replication licensing. Proc. Natl. Acad. Sci. USA 2009, 106, 19807–19812. [Google Scholar]
- Leno, G.H.; Laskey, R.A. DNA replication in cell-free extracts from Xenopus laevis. Methods Cell Biol. 1991, 36, 561–579. [Google Scholar] [PubMed]
- Blow, J.J.; Laskey, R.A. Initiation of DNA replication in nuclei and purified DNA by a cell-free extract of Xenopus eggs. Cell 1986, 47, 577–587. [Google Scholar] [CrossRef] [PubMed]
- Lutzmann, M.; Maiorano, D.; Mechali, M. A Cdt1-geminin complex licenses chromatin for DNA replication and prevents rereplication during S phase in Xenopus. EMBO J. 2006, 25, 5764–5774. [Google Scholar] [CrossRef] [PubMed]
- Murray, A.W.; Kirschner, M.W. Cyclin synthesis drives the early embryonic cell cycle. Nature 1989, 339, 275–280. [Google Scholar] [CrossRef] [PubMed]
- Burke, T.W.; Cook, J.G.; Asano, M.; Nevins, J.R. Replication factors MCM2 and ORC1 interact with the histone acetyltransferase HBO1. J. Biol. Chem. 2001, 276, 15397–15408. [Google Scholar] [CrossRef] [PubMed]
- Iizuka, M.; Stillman, B. Histone acetyltransferase HBO1 interacts with the ORC1 subunit of the human initiator protein. J. Biol. Chem. 1999, 274, 23027–23034. [Google Scholar] [CrossRef] [PubMed]
- Ait-Si-Ali, S.; Ramirez, S.; Robin, P.; Trouche, D.; Harel-Bellan, A. A rapid and sensitive assay for histone acetyl-transferase activity. Nucleic Acids Res. 1998, 26, 3869–3870. [Google Scholar] [CrossRef] [PubMed]
- Cook, J.G.; Chasse, D.A.; Nevins, J.R. The regulated association of Cdt1 with minichromosome maintenance proteins and Cdc6 in mammalian cells. J. Biol. Chem. 2004, 279, 9625–9633. [Google Scholar] [CrossRef] [PubMed]
- Gillespie, P.J.; Li, A.; Blow, J.J. Reconstitution of licensed replication origins on Xenopus sperm nuclei using purified proteins. BMC Biochem. 2001, 2, 15. [Google Scholar] [CrossRef] [PubMed]
- Powell, S.K.; MacAlpine, H.K.; Prinz, J.A.; Li, Y.; Belsky, J.A.; MacAlpine, D.M. Dynamic loading and redistribution of the MCM2-7 helicase complex through the cell cycle. EMBO J. 2015, 34, 531–543. [Google Scholar] [CrossRef] [PubMed]
- Symeonidou, I.E.; Kotsantis, P.; Roukos, V.; Rapsomaniki, M.A.; Grecco, H.E.; Bastiaens, P.; Taraviras, S.; Lygerou, Z. Multi-step loading of human minichromosome maintenance proteins in live human cells. J. Biol. Chem. 2013, 288, 35852–35867. [Google Scholar] [CrossRef] [PubMed]
- Bleichert, F.; Botchan, M.R.; Berger, J.M. Crystal structure of the eukaryotic origin recognition complex. Nature 2015, 519, 321–326. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Suchyta, M.; Miotto, B.; McGarry, T.J. An Inactive Geminin Mutant That Binds Cdt1. Genes 2015, 6, 252-266. https://doi.org/10.3390/genes6020252
Suchyta M, Miotto B, McGarry TJ. An Inactive Geminin Mutant That Binds Cdt1. Genes. 2015; 6(2):252-266. https://doi.org/10.3390/genes6020252
Chicago/Turabian StyleSuchyta, Marissa, Benoit Miotto, and Thomas J. McGarry. 2015. "An Inactive Geminin Mutant That Binds Cdt1" Genes 6, no. 2: 252-266. https://doi.org/10.3390/genes6020252
APA StyleSuchyta, M., Miotto, B., & McGarry, T. J. (2015). An Inactive Geminin Mutant That Binds Cdt1. Genes, 6(2), 252-266. https://doi.org/10.3390/genes6020252




