Functional Role of the microRNA-200 Family in Breast Morphogenesis and Neoplasia
Abstract
:1. Introduction
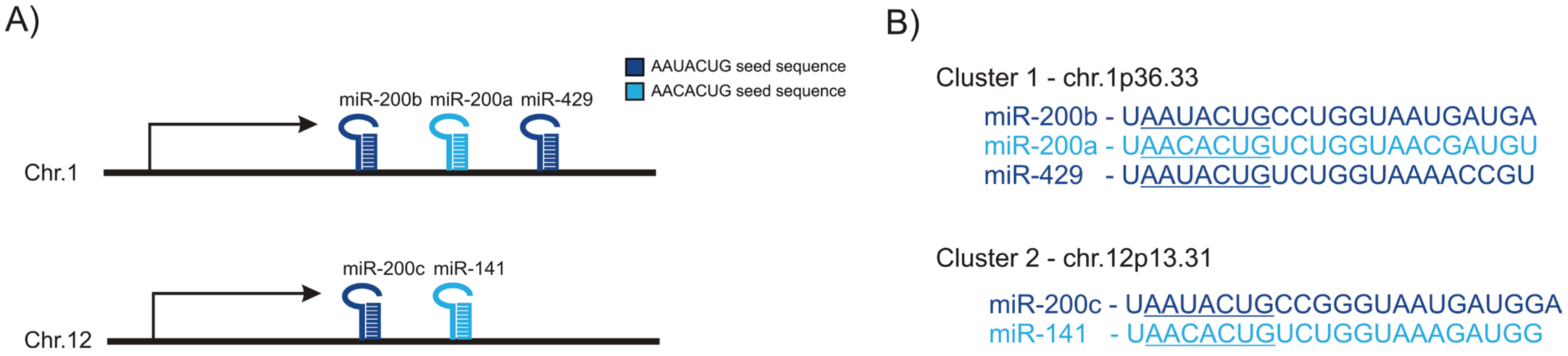
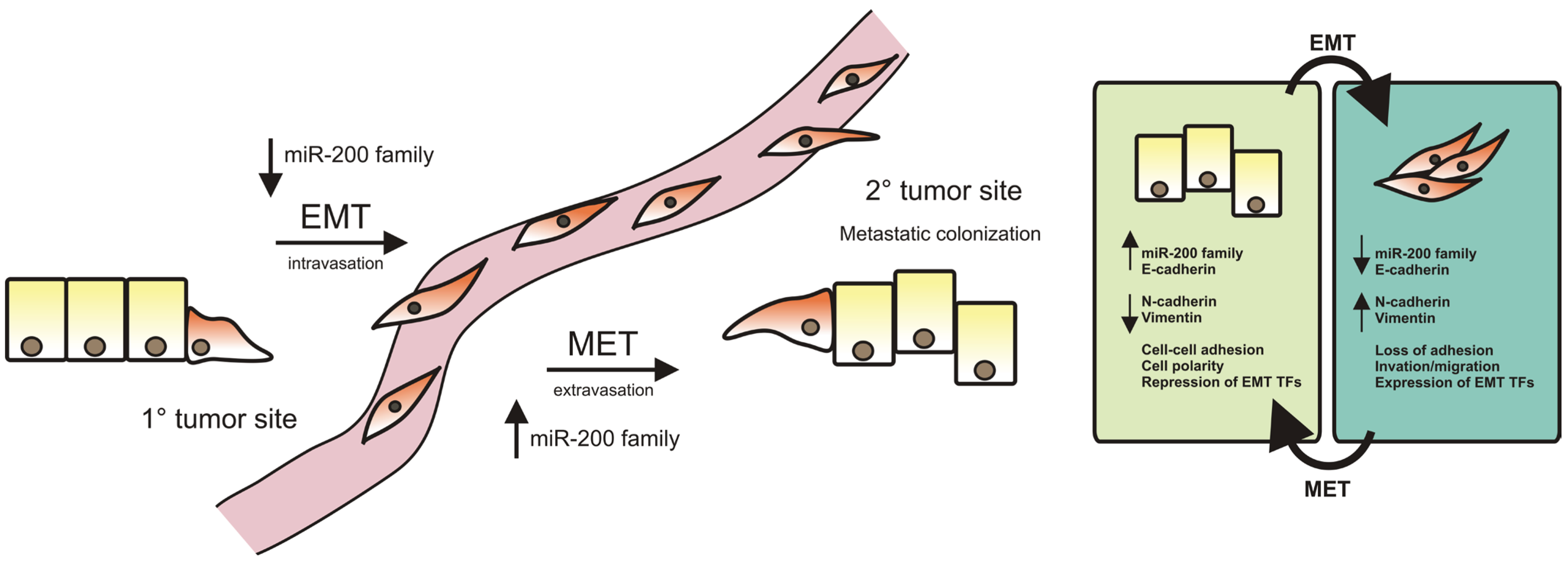
2. miR-200 Family in Epithelial-to-Mesenchymal Transition
3. miR-200 Family in Branching Morphogenesis in the Mammary Gland
4. mir-200 Family and Breast Cancer
5. Future Perspective
6. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Iber, D.; Menshykau, D. The control of branching morphogenesis. Open Biol. 2013, 3. [Google Scholar] [CrossRef]
- Huebner, R.J.; Ewald, A.J. Cellular foundations of mammary tubulogenesis. Semin. Cell Dev. Biol. 2014, 31C, 124–131. [Google Scholar] [CrossRef]
- Affolter, M.; Zeller, R.; Caussinus, E. Tissue remodelling through branching morphogenesis. Nat. Rev. Mol. Cell Biol. 2009, 10, 831–842. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Robinson, G.W.; Karpf, A.B.; Kratochwil, K. Regulation of mammary gland development by tissue interaction. J. Mammary Gland Biol. Neoplasia 1999, 4, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Fata, J.E.; Werb, Z.; Bissell, M.J. Regulation of mammary gland branching morphogenesis by the extracellular matrix and its remodeling enzymes. Breast Cancer Res. 2004, 6, 1–11. [Google Scholar] [PubMed]
- Egeblad, M.; Rasch, M.G.; Weaver, V.M. Dynamic interplay between the collagen scaffold and tumor evolution. Curr. Opin. Cell Biol. 2010, 22, 697–706. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [PubMed]
- Ebert, M.S.; Sharp, P.A. Roles for microRNAs in conferring robustness to biological processes. Cell 2012, 149, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar]
- Griffiths-Jones, S.; Grocock, R.J.; van Dongen, S.; Bateman, A.; Enright, A.J. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 2006, 34, D140–D144. [Google Scholar]
- Lee, Y.; Ahn, C.; Han, J.; Choi, H.; Kim, J.; Yim, J.; Lee, J.; Provost, P.; Rådmark, O.; Kim, S.; et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 2003, 425, 415–419. [Google Scholar]
- Yi, R.; Qin, Y.; Macara, I.G.; Cullen, B.R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 2003, 17, 3011–3016. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Kolb, F.A.; Brondani, V.; Billy, E.; Filipowicz, W. Human Dicer preferentially cleaves dsRNAs at their termini without a requirement for ATP. EMBO J. 2002, 21, 5875–5885. [Google Scholar] [CrossRef] [PubMed]
- Ameres, S.L.; Zamore, P.D. Diversifying microRNA sequence and function. Nat. Rev. Mol. Cell Biol. 2013, 14, 475–488. [Google Scholar] [CrossRef] [PubMed]
- Gu, S.; Jin, L.; Zhang, F.; Sarnow, P.; Kay, M.A. Biological basis for restriction of microRNA targets to the 3' untranslated region in mammalian mRNAs. Nat. Struct. Mol. Biol. 2009, 16, 144–150. [Google Scholar] [CrossRef] [PubMed]
- Croce, C.M.; Calin, G.A. miRNAs, cancer, and stem cell division. Cell 2005, 122, 6–7. [Google Scholar] [CrossRef] [PubMed]
- Arnold, C.P.; Tan, R.; Zhou, B.; Yue, S.-B.; Schaffert, S.; Biggs, J.R.; Doyonnas, R.; Lo, M.-C.; Perry, J.M.; Renault, V.M.; et al. MicroRNA programs in normal and aberrant stem and progenitor cells. Genome Res. 2011, 21, 798–810. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.A.; Lin, H. The microRNA regulation of stem cells. Wiley Interdiscip. Rev. Dev. Biol. 2012, 1, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Leonardo, T.R.; Schultheisz, H.L.; Loring, J.F.; Laurent, L.C. The functions of microRNAs in pluripotency and reprogramming. Nat. Cell Biol. 2012, 14, 1114–1121. [Google Scholar] [CrossRef] [PubMed]
- Farazi, T.A.; Hoell, J.I.; Morozov, P.; Tuschl, T. MicroRNAs in human cancer. Adv. Exp. Med. Biol. 2013, 774, 1–20. [Google Scholar] [PubMed]
- Feng, X.; Wang, Z.; Fillmore, R.; Xi, Y. MiR-200, a new star miRNA in human cancer. Cancer Lett. 2014, 344, 166–173. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.-N.; Chen, S.-Y.; Hwang, S.-M.; Yu, C.-C.; Su, M.-W.; Mai, W.; Wang, H.-W.; Cheng, W.-C.; Schuyler, S.C.; Ma, N.; et al. miR-200c and GATA binding protein 4 regulate human embryonic stem cell renewal and differentiation. Stem Cell Res. 2014, 12, 338–353. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Tetzlaff, M.T.; Cui, R.; Xu, X. miR-200c inhibits melanoma progression and drug resistance through down-regulation of BMI-1. Am. J. Pathol. 2012, 181, 1823–1835. [Google Scholar] [CrossRef] [PubMed]
- Park, S.-M.; Gaur, A.B.; Lengyel, E.; Peter, M.E. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev. 2008, 22, 894–907. [Google Scholar] [CrossRef] [PubMed]
- Shimono, Y.; Zabala, M.; Cho, R.W.; Lobo, N.; Dalerba, P.; Qian, D.; Diehn, M.; Liu, H.; Panula, S.P.; Chiao, E.; et al. Downregulation of miRNA-200c links breast cancer stem cells with normal stem cells. Cell 2009, 138, 592–603. [Google Scholar] [CrossRef] [PubMed]
- Hill, L.; Browne, G.; Tulchinsky, E. ZEB/miR-200 feedback loop: At the crossroads of signal transduction in cancer. Int. J. Cancer J. Int. Cancer 2013, 132, 745–754. [Google Scholar] [CrossRef]
- Micalizzi, D.S.; Farabaugh, S.M.; Ford, H.L. Epithelial-mesenchymal transition in cancer: Parallels between normal development and tumor progression. J. Mammary Gland Biol. Neoplasia 2010, 15, 117–134. [Google Scholar] [CrossRef] [PubMed]
- Guarino, M.; Tosoni, A.; Nebuloni, M. Direct contribution of epithelium to organ fibrosis: Epithelial-mesenchymal transition. Hum. Pathol. 2009, 40, 1365–1376. [Google Scholar] [CrossRef] [PubMed]
- Moustakas, A.; Heldin, C.-H. Signaling networks guiding epithelial-mesenchymal transitions during embryogenesis and cancer progression. Cancer Sci. 2007, 98, 1512–1520. [Google Scholar] [CrossRef] [PubMed]
- Peinado, H.; Olmeda, D.; Cano, A. Snail, Zeb and bHLH factors in tumour progression: An alliance against the epithelial phenotype? Nat. Rev. Cancer 2007, 7, 415–428. [Google Scholar] [CrossRef]
- Gregory, P.A.; Bert, A.G.; Paterson, E.L.; Barry, S.C.; Tsykin, A.; Farshid, G.; Vadas, M.A.; Khew-Goodall, Y.; Goodall, G.J. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat. Cell Biol. 2008, 10, 593–601. [Google Scholar] [CrossRef] [PubMed]
- Korpal, M.; Lee, E.S.; Hu, G.; Kang, Y. The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J. Biol. Chem. 2008, 283, 14910–14914. [Google Scholar] [CrossRef] [PubMed]
- Burk, U.; Schubert, J.; Wellner, U.; Schmalhofer, O.; Vincan, E.; Spaderna, S.; Brabletz, T. A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep. 2008, 9, 582–589. [Google Scholar] [CrossRef] [PubMed]
- Bracken, C.P.; Gregory, P.A.; Kolesnikoff, N.; Bert, A.G.; Wang, J.; Shannon, M.F.; Goodall, G.J. A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res. 2008, 68, 7846–7854. [Google Scholar] [CrossRef] [PubMed]
- Vrba, L.; Garbe, J.C.; Stampfer, M.R.; Futscher, B.W. Epigenetic regulation of normal human mammary cell type-specific miRNAs. Genome Res. 2011, 21, 2026–2037. [Google Scholar] [CrossRef] [PubMed]
- Wiklund, E.D.; Bramsen, J.B.; Hulf, T.; Dyrskjøt, L.; Ramanathan, R.; Hansen, T.B.; Villadsen, S.B.; Gao, S.; Ostenfeld, M.S.; Borre, M.; et al. Coordinated epigenetic repression of the miR-200 family and miR-205 in invasive bladder cancer. Int. J. Cancer J. Int. Cancer 2011, 128, 1327–1334. [Google Scholar] [CrossRef]
- Neves, R.; Scheel, C.; Weinhold, S.; Honisch, E.; Iwaniuk, K.M.; Trompeter, H.-I.; Niederacher, D.; Wernet, P.; Santourlidis, S.; Uhrberg, M. Role of DNA methylation in miR-200c/141 cluster silencing in invasive breast cancer cells. BMC Res. Notes 2010, 3, e219. [Google Scholar] [CrossRef]
- Davalos, V.; Moutinho, C.; Villanueva, A.; Boque, R.; Silva, P.; Carneiro, F.; Esteller, M. Dynamic epigenetic regulation of the microRNA-200 family mediates epithelial and mesenchymal transitions in human tumorigenesis. Oncogene 2012, 31, 2062–2074. [Google Scholar] [CrossRef] [PubMed]
- Lim, Y.-Y.; Wright, J.A.; Attema, J.L.; Gregory, P.A.; Bert, A.G.; Smith, E.; Thomas, D.; Lopez, A.F.; Drew, P.A.; Khew-Goodall, Y.; et al. Epigenetic modulation of the miR-200 family is associated with transition to a breast cancer stem-cell-like state. J. Cell Sci. 2013, 126, 2256–2266. [Google Scholar] [CrossRef] [PubMed]
- Pozzi, A.; Zent, R. Extracellular matrix receptors in branched organs. Curr. Opin. Cell Biol. 2011, 23, 547–553. [Google Scholar] [CrossRef] [PubMed]
- Leroy, P.; Mostov, K.E. Slug is required for cell survival during partial epithelial-mesenchymal transition of HGF-induced tubulogenesis. Mol. Biol. Cell 2007, 18, 1943–1952. [Google Scholar] [CrossRef] [PubMed]
- Ewald, A.J.; Brenot, A.; Duong, M.; Chan, B.S.; Werb, Z. Collective epithelial migration and cell rearrangements drive mammary branching morphogenesis. Dev. Cell 2008, 14, 570–581. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Mani, S.A.; Donaher, J.L.; Ramaswamy, S.; Itzykson, R.A.; Come, C.; Savagner, P.; Gitelman, I.; Richardson, A.; Weinberg, R.A. Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell 2004, 117, 927–939. [Google Scholar] [CrossRef] [PubMed]
- Guarino, M. Epithelial-mesenchymal transition and tumour invasion. Int. J. Biochem. Cell Biol. 2007, 39, 2153–2160. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Song, F.; Chen, X.; Li, Y.; Fan, J.; Wu, X. Bmi-1 regulates epithelial-to-mesenchymal transition to promote migration and invasion of breast cancer cells. Int. J. Clin. Exp. Pathol. 2014, 7, 3057–3064. [Google Scholar] [PubMed]
- Lo, A.T.; Mori, H.; Mott, J.; Bissell, M.J. Constructing three-dimensional models to study mammary gland branching morphogenesis and functional differentiation. J. Mammary Gland Biol. Neoplasia 2012, 17, 103–110. [Google Scholar] [CrossRef] [PubMed]
- Bockmeyer, C.L.; Christgen, M.; Müller, M.; Fischer, S.; Ahrens, P.; Länger, F.; Kreipe, H.; Lehmann, U. MicroRNA profiles of healthy basal and luminal mammary epithelial cells are distinct and reflected in different breast cancer subtypes. Breast Cancer Res. Treat. 2011, 130, 735–745. [Google Scholar] [CrossRef] [PubMed]
- Péchoux, C.; Gudjonsson, T.; Ronnov-Jessen, L.; Bissell, M.J.; Petersen, O.W. Human mammary luminal epithelial cells contain progenitors to myoepithelial cells. Dev. Biol. 1999, 206, 88–99. [Google Scholar]
- Gudjonsson, T.; Adriance, M.C.; Sternlicht, M.D.; Petersen, O.W.; Bissell, M.J. Myoepithelial cells: Their origin and function in breast morphogenesis and neoplasia. J. Mammary Gland Biol. Neoplasia 2005, 10, 261–272. [Google Scholar] [CrossRef] [PubMed]
- Villadsen, R.; Fridriksdottir, A.J.; Rønnov-Jessen, L.; Gudjonsson, T.; Rank, F.; LaBarge, M.A.; Bissell, M.J.; Petersen, O.W. Evidence for a stem cell hierarchy in the adult human breast. J. Cell Biol. 2007, 177, 87–101. [Google Scholar] [CrossRef] [PubMed]
- Song, S.J.; Poliseno, L.; Song, M.S.; Ala, U.; Webster, K.; Ng, C.; Beringer, G.; Brikbak, N.J.; Yuan, X.; Cantley, L.C.; et al. MicroRNA-antagonism regulates breast cancer stemness and metastasis via TET-family-dependent chromatin remodeling. Cell 2013, 154, 311–324. [Google Scholar] [CrossRef] [PubMed]
- Avril-Sassen, S.; Goldstein, L.D.; Stingl, J.; Blenkiron, C.; le Quesne, J.; Spiteri, I.; Karagavriilidou, K.; Watson, C.J.; Tavaré, S.; Miska, E.A.; et al. Characterisation of microRNA expression in post-natal mouse mammary gland development. BMC Genomics 2009, 10, e548. [Google Scholar]
- Le Guillou, S.; Marthey, S.; Laloë, D.; Laubier, J.; Mobuchon, L.; Leroux, C.; le Provost, F. Characterisation and comparison of lactating mouse and bovine mammary gland miRNomes. PLoS ONE 2014, 9, e91938. [Google Scholar]
- Galio, L.; Droineau, S.; Yeboah, P.; Boudiaf, H.; Bouet, S.; Truchet, S.; Devinoy, E. MicroRNA in the ovine mammary gland during early pregnancy: Spatial and temporal expression of miR-21, miR-205, and miR-200. Physiol. Genomics 2013, 45, 151–161. [Google Scholar] [CrossRef] [PubMed]
- Rebustini, I.T.; Hayashi, T.; Reynolds, A.D.; Dillard, M.L.; Carpenter, E.M.; Hoffman, M.P. miR-200c regulates FGFR-dependent epithelial proliferation via Vldlr during submandibular gland branching morphogenesis. Dev. Camb. Engl. 2012, 139, 191–202. [Google Scholar]
- Patel, V.; Hajarnis, S.; Williams, D.; Hunter, R.; Huynh, D.; Igarashi, P. MicroRNAs regulate renal tubule maturation through modulation of Pkd1. J. Am. Soc. Nephrol. JASN 2012, 23, 1941–1948. [Google Scholar] [CrossRef]
- Gudjonsson, T.; Villadsen, R.; Nielsen, H.L.; Rønnov-Jessen, L.; Bissell, M.J.; Petersen, O.W. Isolation, immortalization, and characterization of a human breast epithelial cell line with stem cell properties. Genes Dev. 2002, 16, 693–706. [Google Scholar] [CrossRef] [PubMed]
- Sigurdsson, V.; Hilmarsdottir, B.; Sigmundsdottir, H.; Fridriksdottir, A.J.R.; Ringnér, M.; Villadsen, R.; Borg, A.; Agnarsson, B.A.; Petersen, O.W.; Magnusson, M.K.; et al. Endothelial induced EMT in breast epithelial cells with stem cell properties. PLoS ONE 2011, 6, e23833. [Google Scholar]
- Bergthorsson, J.T.; Magnusson, M.K.; Gudjonsson, T. Endothelial-rich microenvironment supports growth and branching morphogenesis of prostate epithelial cells. Prostate 2013, 73, 884–896. [Google Scholar] [CrossRef] [PubMed]
- Franzdóttir, S.R.; Axelsson, I.T.; Arason, A.J.; Baldursson, O.; Gudjonsson, T.; Magnusson, M.K. Airway branching morphogenesis in three dimensional culture. Respir. Res. 2010, 11, 162. [Google Scholar]
- Olson, P.; Lu, J.; Zhang, H.; Shai, A.; Chun, M.G.; Wang, Y.; Libutti, S.K.; Nakakura, E.K.; Golub, T.R.; Hanahan, D. MicroRNA dynamics in the stages of tumorigenesis correlate with hallmark capabilities of cancer. Genes Dev. 2009, 23, 2152–2165. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wang, J. MicroRNA-mediated breast cancer metastasis: From primary site to distant organs. Oncogene 2012, 31, 2499–2511. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Pan, X.; Cobb, G.P.; Anderson, T.A. microRNAs as oncogenes and tumor suppressors. Dev. Biol. 2007, 302, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Bendoraite, A.; Knouf, E.C.; Garg, K.S.; Parkin, R.K.; Kroh, E.M.; O’Briant, K.C.; Ventura, A.P.; Godwin, A.K.; Karlan, B.Y.; Drescher, C.W.; et al. Regulation of miR-200 family microRNAs and ZEB transcription factors in ovarian cancer: Evidence supporting a mesothelial-to-epithelial transition. Gynecol. Oncol. 2010, 116, 117–125. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Xu, Y.; Ding, L.; Yao, H.; Yu, H.; Zhou, T.; Si, J. Down-regulation of miR-141 in gastric cancer and its involvement in cell growth. J. Gastroenterol. 2009, 44, 556–561. [Google Scholar] [CrossRef] [PubMed]
- Gravgaard, K.H.; Lyng, M.B.; Laenkholm, A.-V.; Søkilde, R.; Nielsen, B.S.; Litman, T.; Ditzel, H.J. The miRNA-200 family and miRNA-9 exhibit differential expression in primary versus corresponding metastatic tissue in breast cancer. Breast Cancer Res. Treat. 2012, 134, 207–217. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.-G.; Zhu, W.-Y.; Huang, Y.-Y.; Ma, L.-N.; Zhou, S.-Q.; Wang, Y.-K.; Zeng, F.; Zhou, J.-H.; Zhang, Y.-K. High expression of serum miR-21 and tumor miR-200c associated with poor prognosis in patients with lung cancer. Med. Oncol. Northwood Lond. Engl. 2012, 29, 618–626. [Google Scholar] [CrossRef]
- Snowdon, J.; Zhang, X.; Childs, T.; Tron, V.A.; Feilotter, H. The microRNA-200 family is upregulated in endometrial carcinoma. PLoS ONE 2011, 6, e22828. [Google Scholar] [CrossRef] [PubMed]
- Perou, C.M.; Sørlie, T.; Eisen, M.B.; van de Rijn, M.; Jeffrey, S.S.; Rees, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A.; et al. Molecular portraits of human breast tumours. Nature 2000, 406, 747–752. [Google Scholar] [CrossRef] [PubMed]
- Sorlie, T.; Tibshirani, R.; Parker, J.; Hastie, T.; Marron, J.S.; Nobel, A.; Deng, S.; Johnsen, H.; Pesich, R.; Geisler, S.; et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc. Natl. Acad. Sci. USA 2003, 100, 8418–8423. [Google Scholar] [CrossRef] [PubMed]
- Prat, A.; Parker, J.S.; Karginova, O.; Fan, C.; Livasy, C.; Herschkowitz, J.I.; He, X.; Perou, C.M. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 2010, 12, R68. [Google Scholar] [CrossRef] [PubMed]
- Sarrió, D.; Rodriguez-Pinilla, S.M.; Hardisson, D.; Cano, A.; Moreno-Bueno, G.; Palacios, J. Epithelial-mesenchymal transition in breast cancer relates to the basal-like phenotype. Cancer Res. 2008, 68, 989–997. [Google Scholar]
- Castilla, M.Á.; Díaz-Martín, J.; Sarrió, D.; Romero-Pérez, L.; López-García, M.Á.; Vieites, B.; Biscuola, M.; Ramiro-Fuentes, S.; Isacke, C.M.; Palacios, J. MicroRNA-200 family modulation in distinct breast cancer phenotypes. PLoS ONE 2012, 7, e47709. [Google Scholar]
- Cascione, L.; Gasparini, P.; Lovat, F.; Carasi, S.; Pulvirenti, A.; Ferro, A.; Alder, H.; He, G.; Vecchione, A.; Croce, C.M.; et al. Integrated microRNA and mRNA signatures associated with survival in triple negative breast cancer. PLoS ONE 2013, 8, e55910. [Google Scholar] [CrossRef] [PubMed]
- Volinia, S.; Galasso, M.; Sana, M.E.; Wise, T.F.; Palatini, J.; Huebner, K.; Croce, C.M. Breast cancer signatures for invasiveness and prognosis defined by deep sequencing of microRNA. Proc. Natl. Acad. Sci. USA 2012, 109, 3024–3029. [Google Scholar] [CrossRef] [PubMed]
- Dykxhoorn, D.M.; Wu, Y.; Xie, H.; Yu, F.; Lal, A.; Petrocca, F.; Martinvalet, D.; Song, E.; Lim, B.; Lieberman, J. miR-200 enhances mouse breast cancer cell colonization to form distant metastases. PLoS ONE 2009, 4, e7181. [Google Scholar] [CrossRef] [PubMed]
- Korpal, M.; Ell, B.J.; Buffa, F.M.; Ibrahim, T.; Blanco, M.A.; Celià-Terrassa, T.; Mercatali, L.; Khan, Z.; Goodarzi, H.; Hua, Y.; et al. Direct targeting of Sec23a by miR-200s influences cancer cell secretome and promotes metastatic colonization. Nat. Med. 2011, 17, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Roslan, S.; Johnstone, C.N.; Wright, J.A.; Bracken, C.P.; Anderson, M.; Bert, A.G.; Selth, L.A.; Anderson, R.L.; Goodall, G.J.; et al. MiR-200 can repress breast cancer metastasis through ZEB1-independent but moesin-dependent pathways. Oncogene 2014, 33, 4077–4088. [Google Scholar] [CrossRef] [PubMed]
- Banyard, J.; Chung, I.; Wilson, A.M.; Vetter, G.; le Béchec, A.; Bielenberg, D.R.; Zetter, B.R. Regulation of epithelial plasticity by miR-424 and miR-200 in a new prostate cancer metastasis model. Sci. Rep. 2013, 3. [Google Scholar] [CrossRef]
- Andre, F.; Job, B.; Dessen, P.; Tordai, A.; Michiels, S.; Liedtke, C.; Richon, C.; Yan, K.; Wang, B.; Vassal, G.; et al. Molecular characterization of breast cancer with high-resolution oligonucleotide comparative genomic hybridization array. Clin. Cancer Res. 2009, 15, 441–451. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.-H.; Chen, C.-L.; More, S.V.; Hsiao, P.-W.; Hung, W.-C.; Li, W.-S. The Tetraindole SK228 Reverses the Epithelial-to-Mesenchymal Transition of Breast Cancer Cells by Up-Regulating Members of the miR-200 Family. PLoS ONE 2014, 9, e101088. [Google Scholar] [CrossRef] [PubMed]
- Ling, H.; Fabbri, M.; Calin, G.A. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat. Rev. Drug Discov. 2013, 12, 847–865. [Google Scholar] [CrossRef] [PubMed]
- Pecot, C.V.; Rupaimoole, R.; Yang, D.; Akbani, R.; Ivan, C.; Lu, C.; Wu, S.; Han, H.-D.; Shah, M.Y.; Rodriguez-Aguayo, C.; et al. Tumour angiogenesis regulation by the miR-200 family. Nat. Commun. 2013, 4. [Google Scholar] [CrossRef]
- Howe, E.N.; Cochrane, D.R.; Cittelly, D.M.; Richer, J.K. miR-200c targets a NF-κB up-regulated TrkB/NTF3 autocrine signaling loop to enhance anoikis sensitivity in triple negative breast cancer. PLoS ONE 2012, 7, e49987. [Google Scholar] [CrossRef] [PubMed]
- Humphries, B.; Wang, Z.; Oom, A.L.; Fisher, T.; Tan, D.; Cui, Y.; Jiang, Y.; Yang, C. MicroRNA-200b targets protein kinase Cα and suppresses triple negative breast cancer metastasis. Carcinogenesis 2014. [Google Scholar] [CrossRef]
- Tam, W.L.; Lu, H.; Buikhuisen, J.; Soh, B.S.; Lim, E.; Reinhardt, F.; Wu, Z.J.; Krall, J.A.; Bierie, B.; Guo, W.; et al. Protein kinase C α is a central signaling node and therapeutic target for breast cancer stem cells. Cancer Cell 2013, 24, 347–364. [Google Scholar] [CrossRef] [PubMed]
- Renwick, N.; Cekan, P.; Masry, P.A.; McGeary, S.E.; Miller, J.B.; Hafner, M.; Li, Z.; Mihailovic, A.; Morozov, P.; Brown, M.; et al. Multicolor microRNA FISH effectively differentiates tumor types. J. Clin. Investig. 2013, 123, 2694–2702. [Google Scholar] [PubMed]
- Weaver, V.M.; Petersen, O.W.; Wang, F.; Larabell, C.A.; Briand, P.; Damsky, C.; Bissell, M.J. Reversion of the malignant phenotype of human breast cells in three-dimensional culture and in vivo by integrin blocking antibodies. J. Cell Biol. 1997, 137, 231–245. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Weaver, V.M.; Petersen, O.W.; Larabell, C.A.; Dedhar, S.; Briand, P.; Lupu, R.; Bissell, M.J. Reciprocal interactions between beta1-integrin and epidermal growth factor receptor in three-dimensional basement membrane breast cultures: A different perspective in epithelial biology. Proc. Natl. Acad. Sci. USA 1998, 95, 14821–14826. [Google Scholar] [CrossRef] [PubMed]
- Truong, H.H.; Xiong, J.; Ghotra, V.P.S.; Nirmala, E.; Haazen, L.; le Dévédec, S.E.; Balcioğlu, H.E.; He, S.; Snaar-Jagalska, B.E.; Vreugdenhil, E.; et al. β1 integrin inhibition elicits a prometastatic switch through the TGFβ-miR-200-ZEB network in E-cadherin-positive triple-negative breast cancer. Sci. Signal. 2014, 7, ra15. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Hilmarsdottir, B.; Briem, E.; Bergthorsson, J.T.; Magnusson, M.K.; Gudjonsson, T. Functional Role of the microRNA-200 Family in Breast Morphogenesis and Neoplasia. Genes 2014, 5, 804-820. https://doi.org/10.3390/genes5030804
Hilmarsdottir B, Briem E, Bergthorsson JT, Magnusson MK, Gudjonsson T. Functional Role of the microRNA-200 Family in Breast Morphogenesis and Neoplasia. Genes. 2014; 5(3):804-820. https://doi.org/10.3390/genes5030804
Chicago/Turabian StyleHilmarsdottir, Bylgja, Eirikur Briem, Jon Thor Bergthorsson, Magnus Karl Magnusson, and Thorarinn Gudjonsson. 2014. "Functional Role of the microRNA-200 Family in Breast Morphogenesis and Neoplasia" Genes 5, no. 3: 804-820. https://doi.org/10.3390/genes5030804
APA StyleHilmarsdottir, B., Briem, E., Bergthorsson, J. T., Magnusson, M. K., & Gudjonsson, T. (2014). Functional Role of the microRNA-200 Family in Breast Morphogenesis and Neoplasia. Genes, 5(3), 804-820. https://doi.org/10.3390/genes5030804




