Comprehensive Characterization of the FATs Gene Family in Maize: Phylogeny, Expression Patterns, and Regulatory Networks
Abstract
1. Introduction
2. Materials and Methods
2.1. Identification of FATs Family Members in Maize
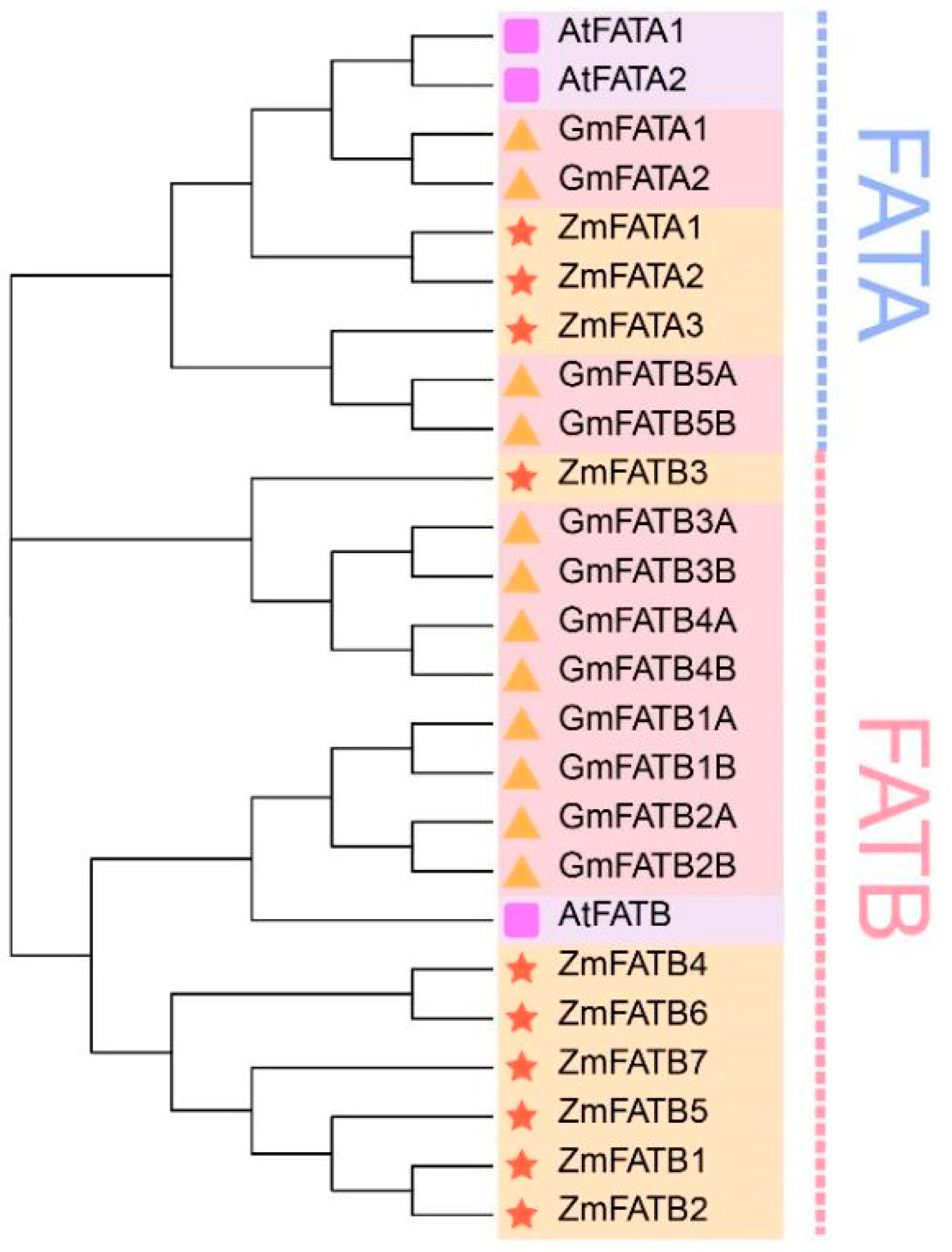
2.2. Phylogenetic Analysis of ZmFATs Family Members
2.3. Gene Structure and Motif Analysis of ZmFATs Family
2.4. Chromosomal Localization and Collinearity Analysis of Maize ZmFAT Genes
2.5. Cis-Elements Analysis
2.6. Functional Annotation Analysis
2.7. Protein Interaction Prediction
2.8. miRNA Interaction Analysis with Maize ZmFATs
2.9. Transcriptome Analysis
2.10. qRT–PCR Analysis
2.11. Statistical Analysis
3. Results
3.1. Identification and Characterization of Maize FAT Family Members
3.2. Phylogenetic Classification of the Maize FATs Family
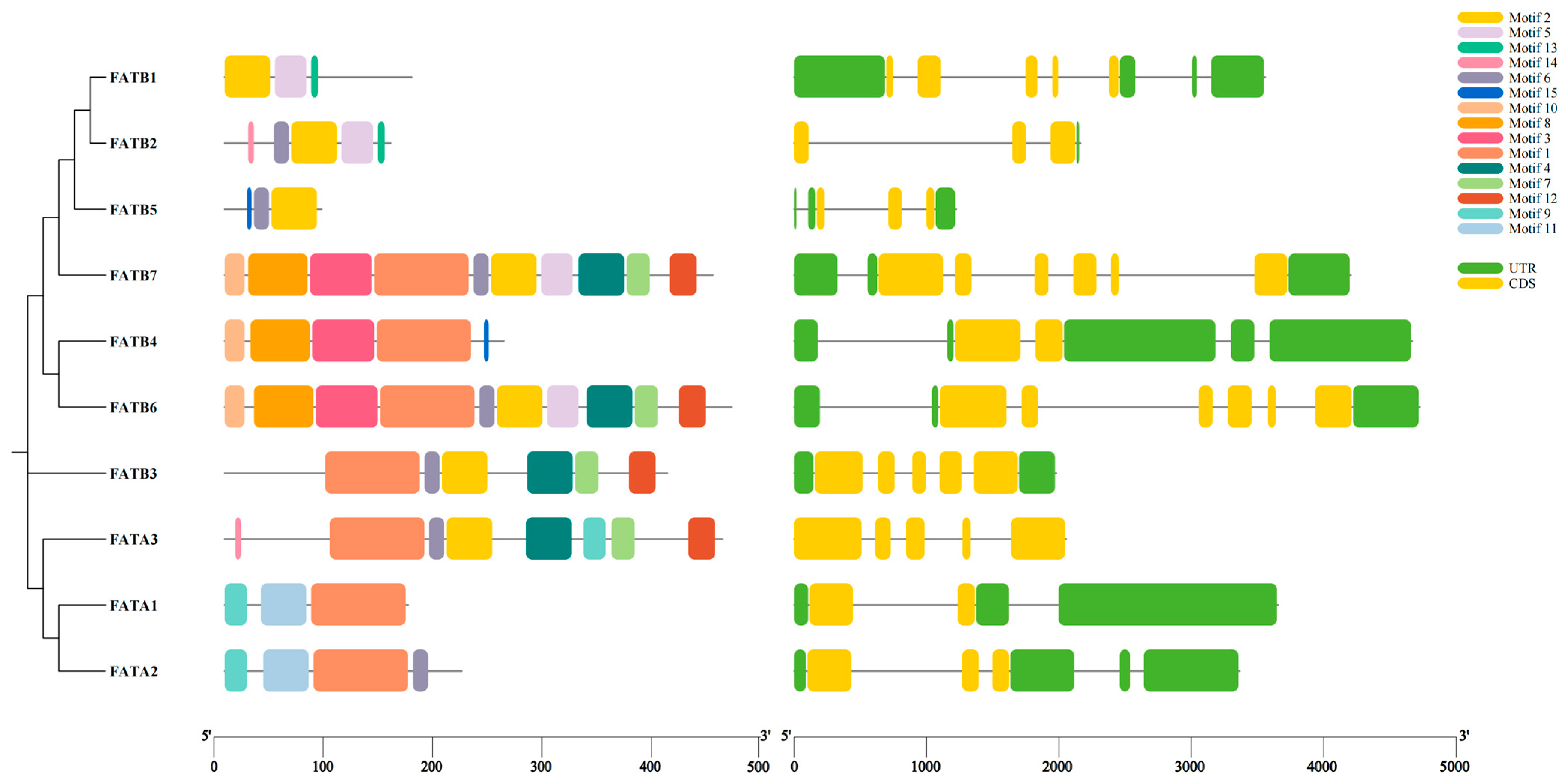
3.3. Analysis of Conserved Motifs and Gene Structure
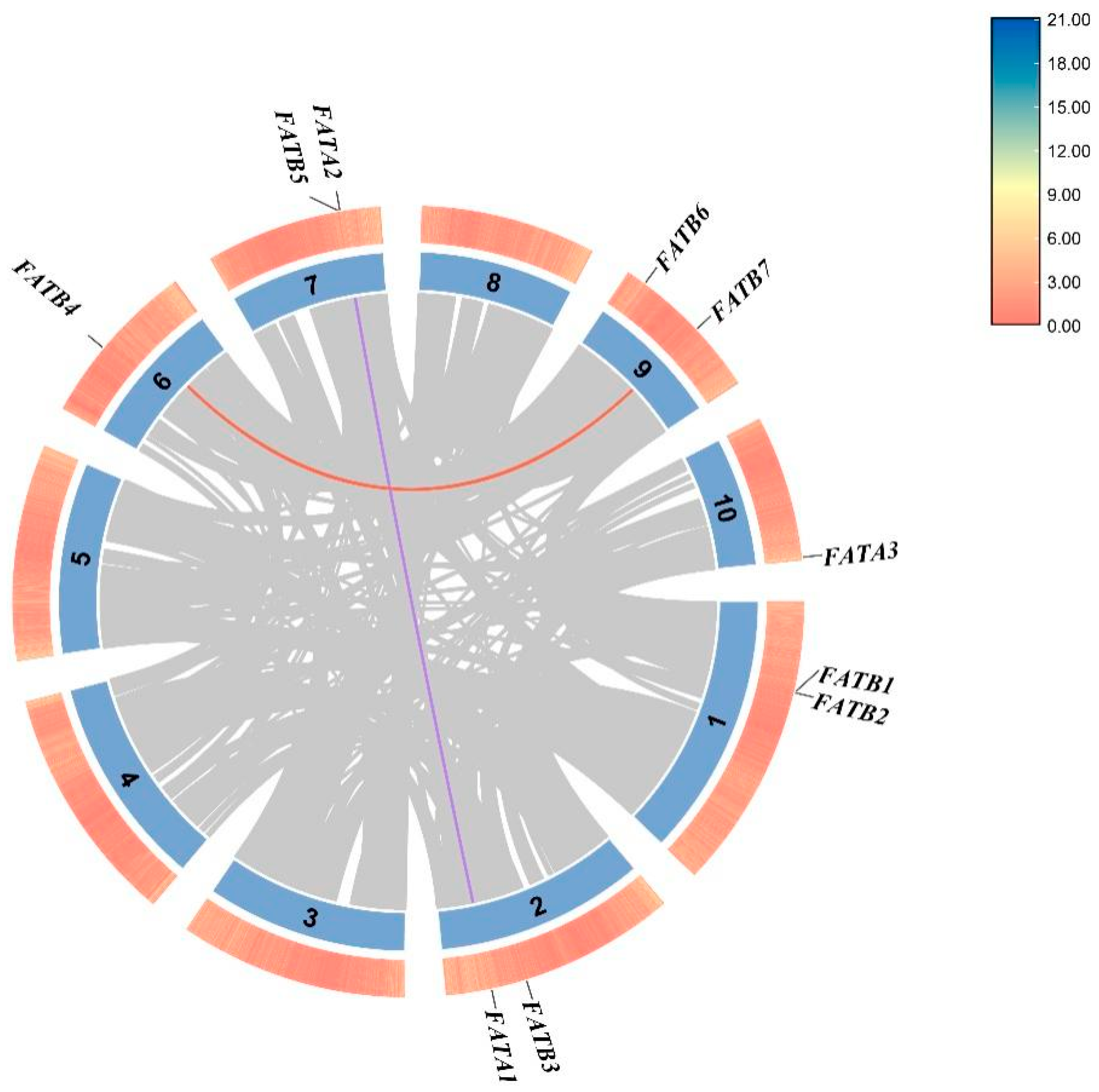
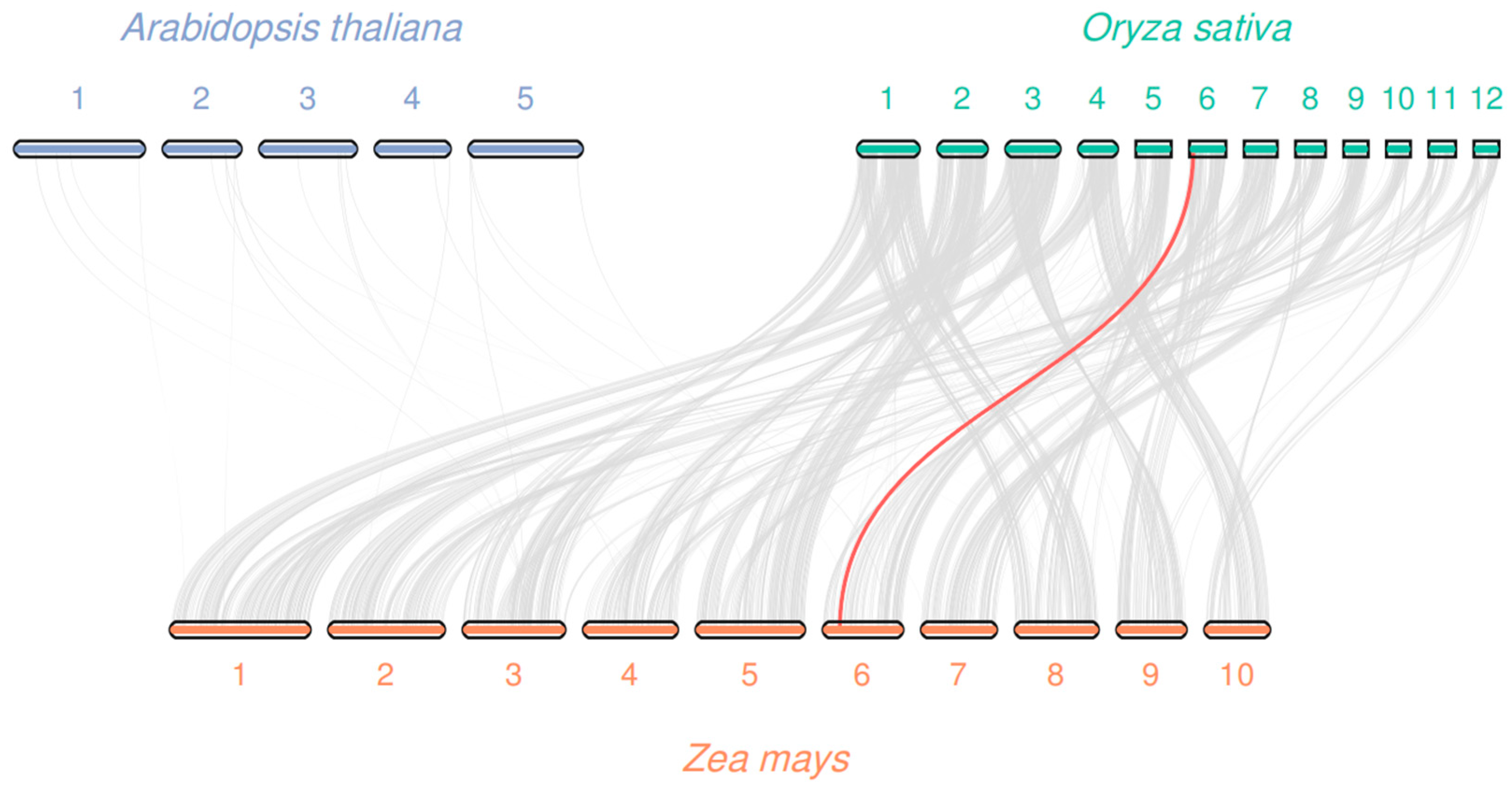
3.4. Chromosomal Localization and Collinearity Analysis
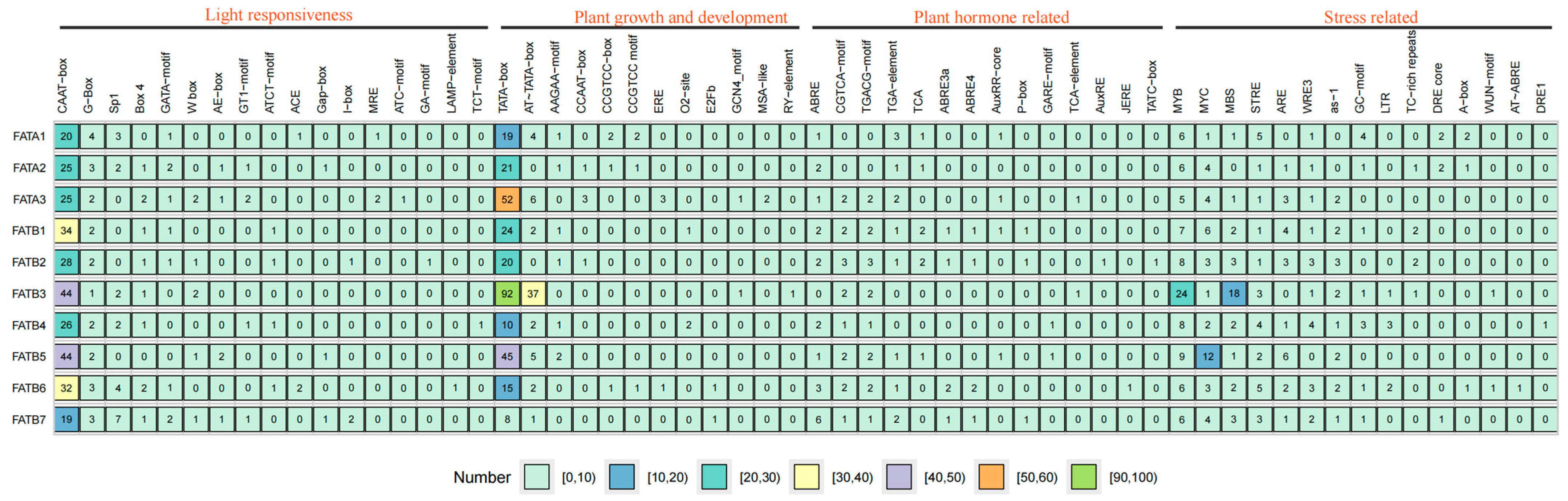
3.5. Cis-Acting Element Analysis
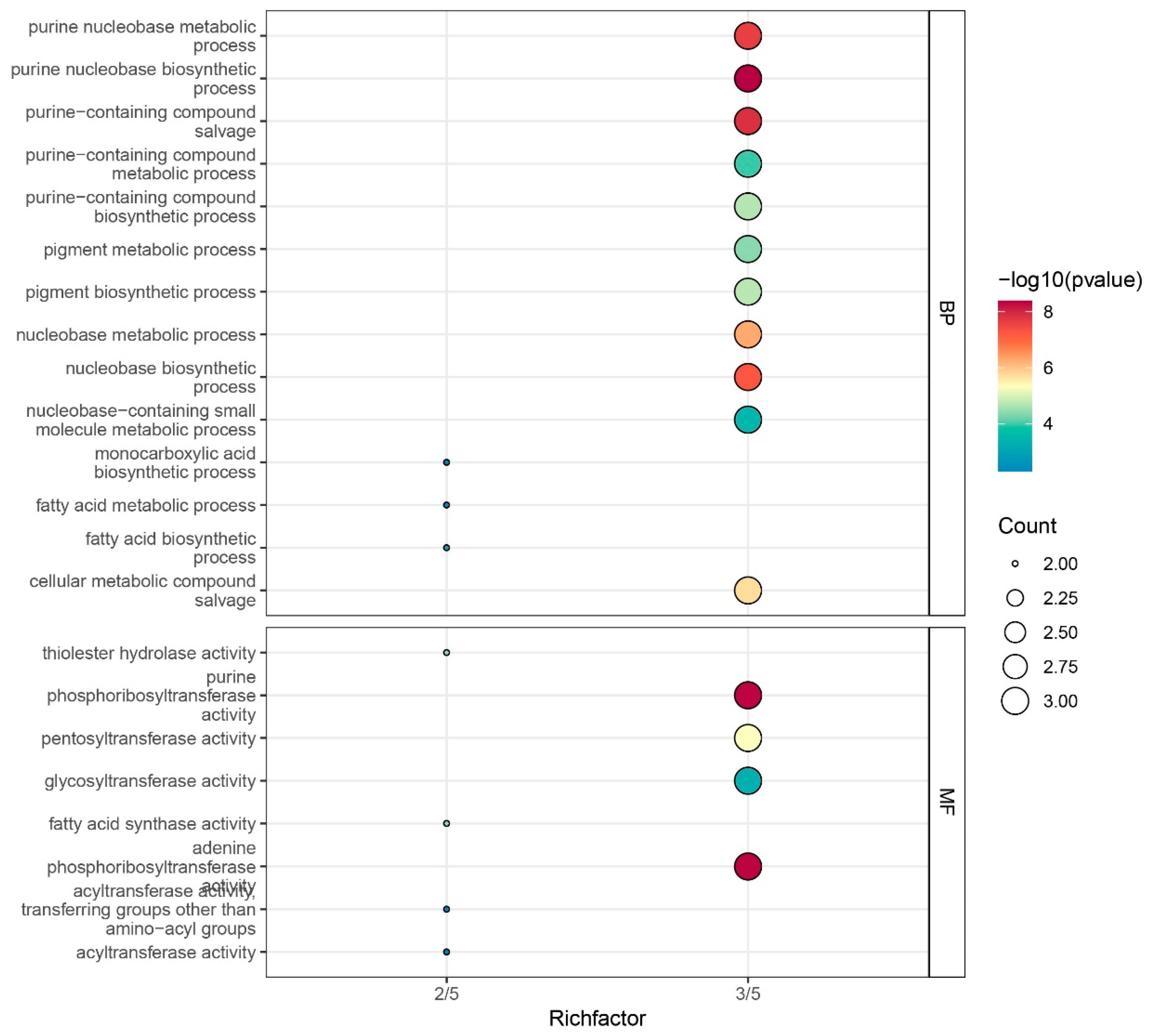
3.6. Functional Annotation Analysis of ZmFAT Genes
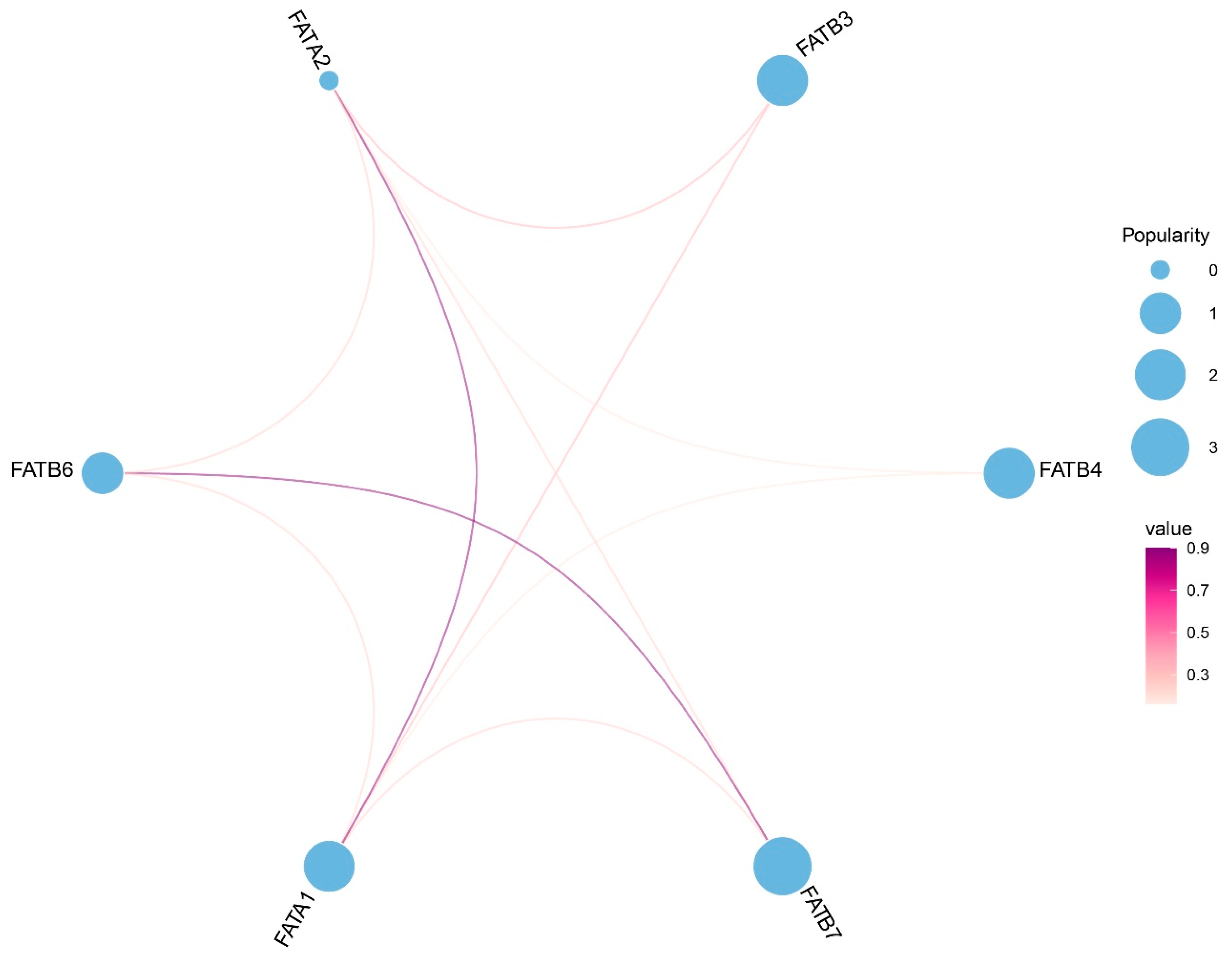
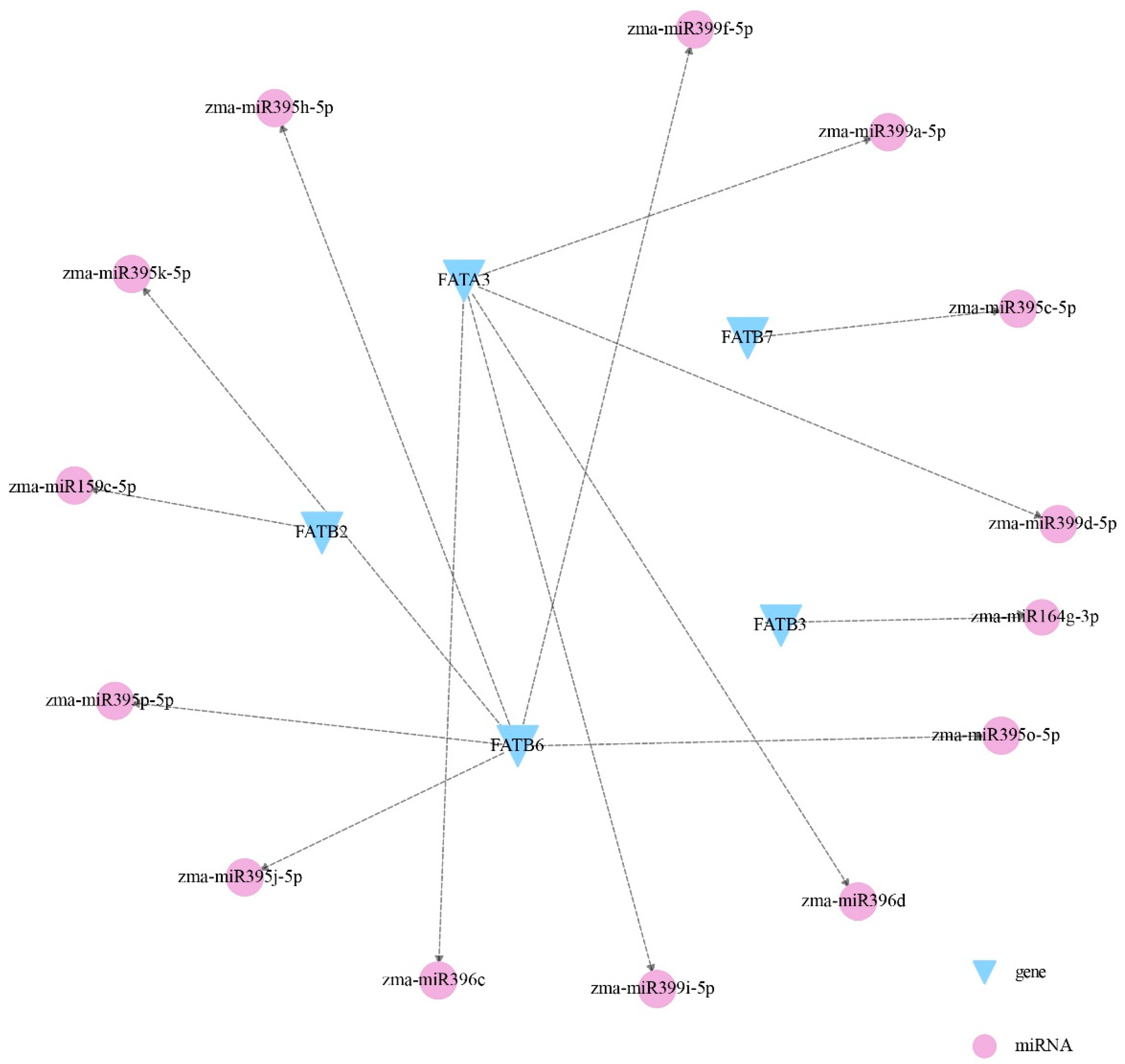
3.7. Interaction Analysis Among ZmFATs and miRNA Regulation Analysis
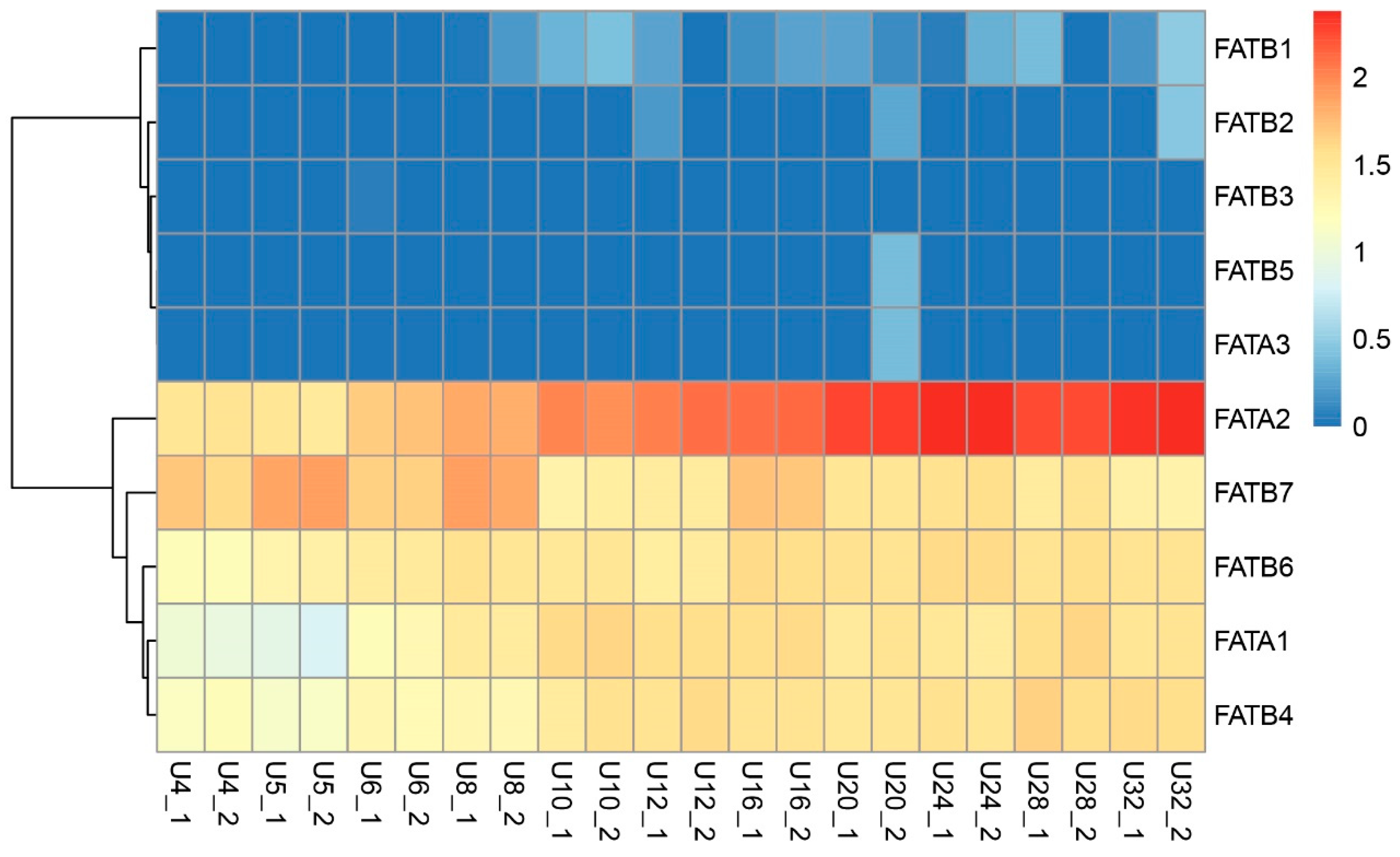
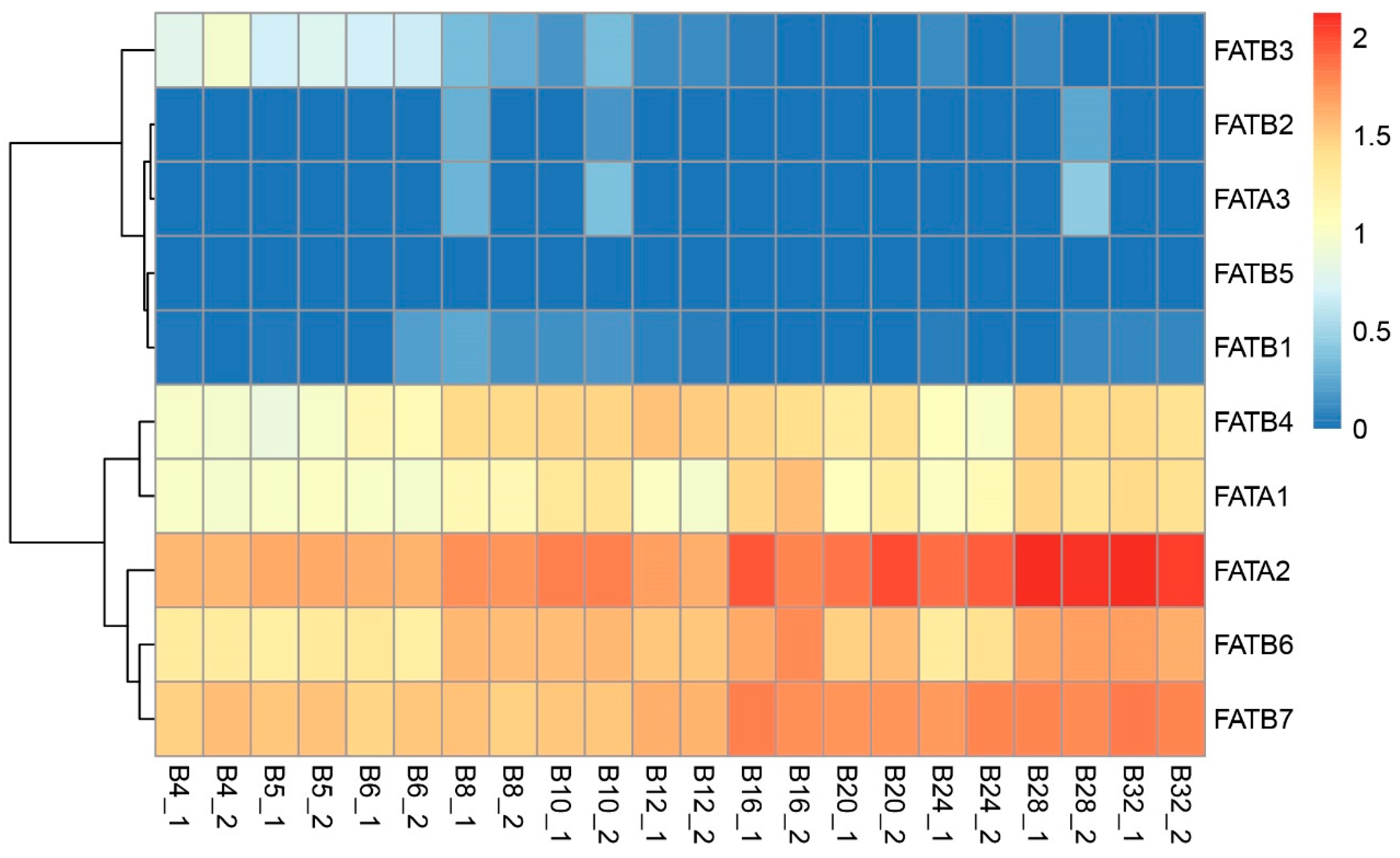
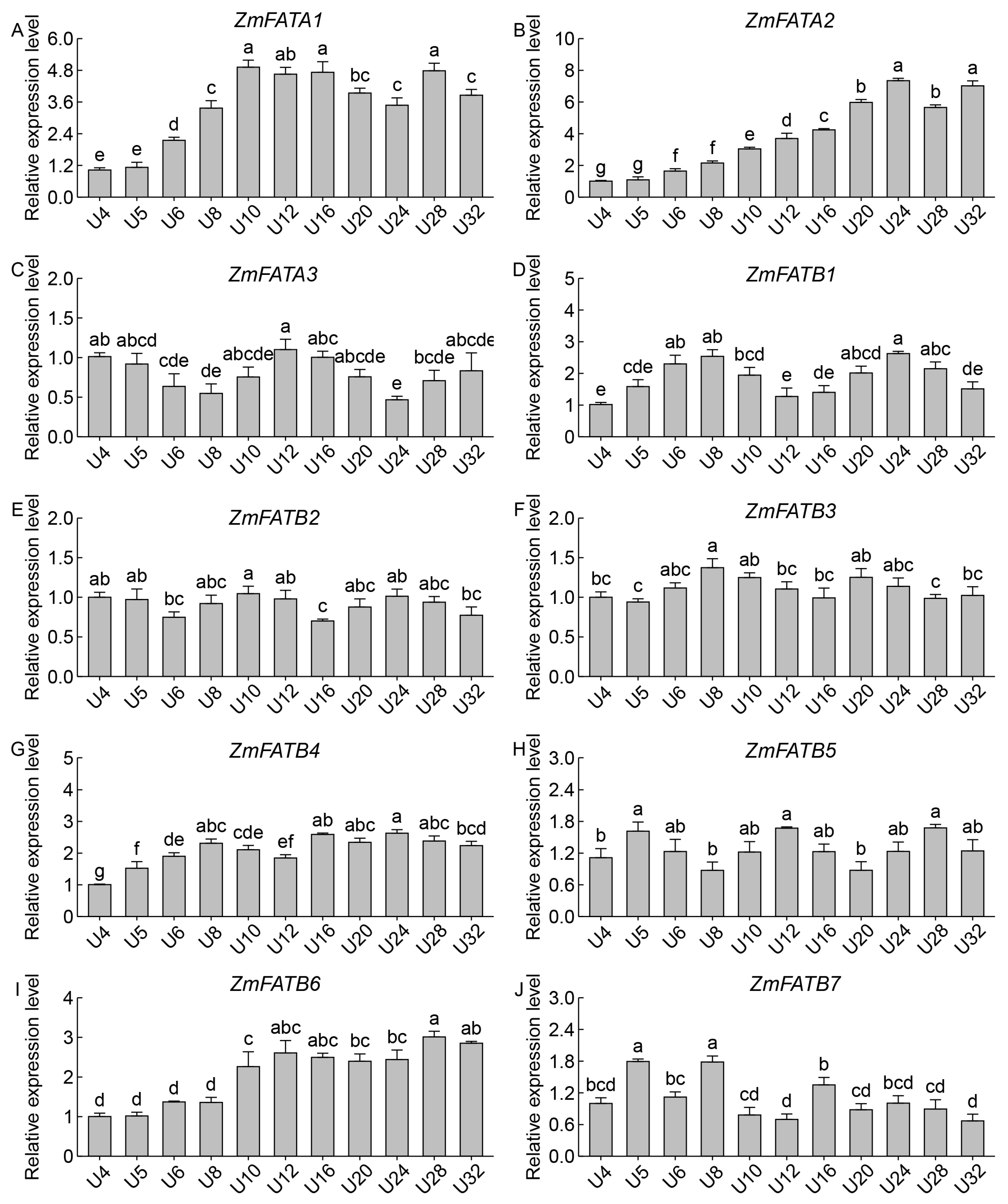
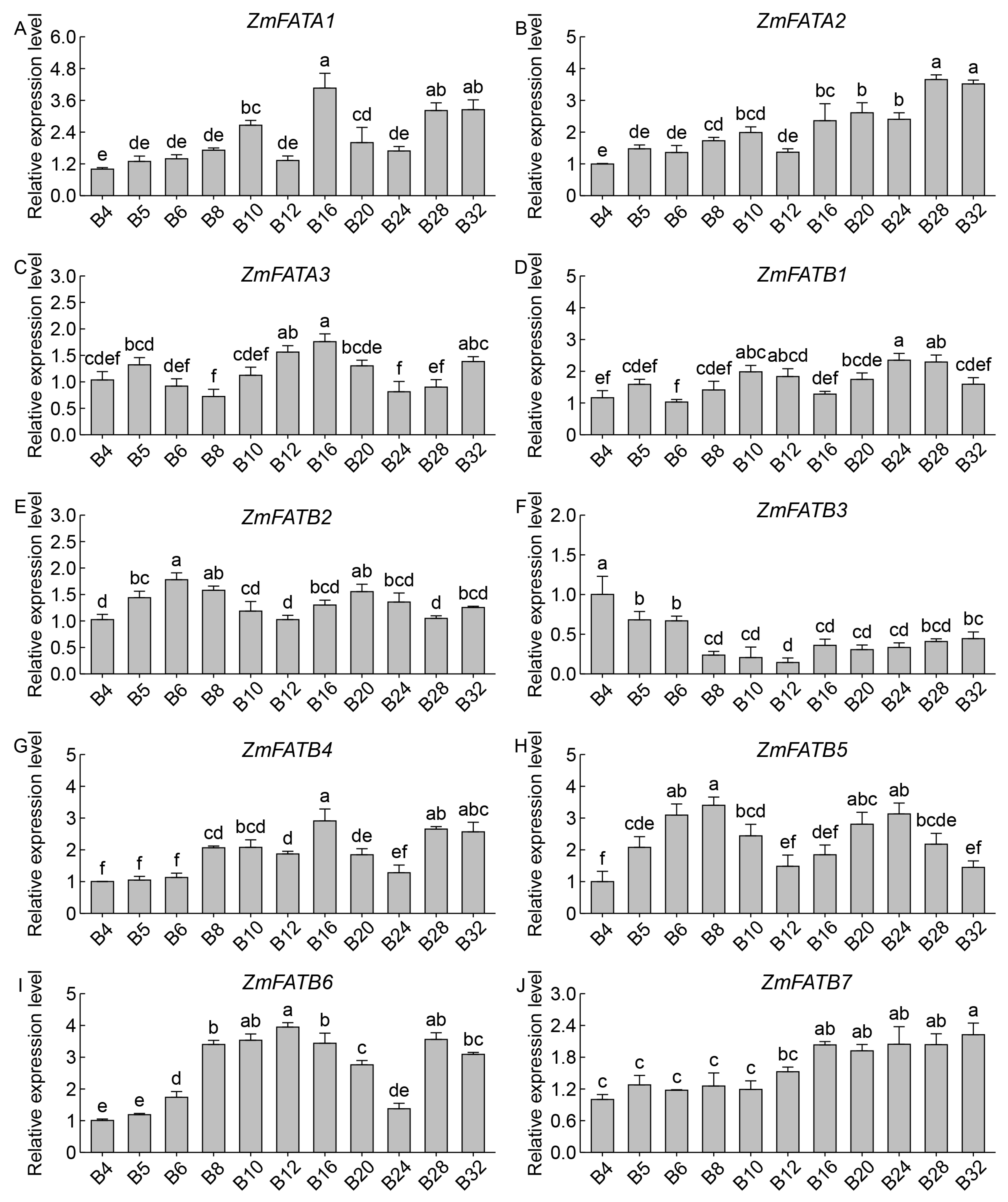
3.8. Analysis of ZmFATs Expression Levels in Maize Seeds at Different Developmental Stages
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Correction Statement
References
- Xu, Y.; Singer, S.D.; Chen, G. Protein interactomes for plant lipid biosynthesis and their biotechnological applications. Plant Biotechnol. J. 2023, 21, 1734–1744. [Google Scholar] [CrossRef]
- Jump, D.B. Fatty acid regulation of gene transcription. Crit. Rev. Clin. Lab. Sci. 2004, 41, 41–78. [Google Scholar] [CrossRef] [PubMed]
- Guan, X.; Okazaki, Y.; Zhang, R.; Saito, K.; Nikolau, B.J. Dual-localized enzymatic components constitute the fatty acid synthase systems in mitochondria and plastids. Plant Physiol. 2020, 183, 517–529. [Google Scholar] [CrossRef]
- Ashton, A.R.; Jenkins, C.L.; Whitfeld, P.R. Molecular cloning of two different cDNAs for maize acetyl CoA carboxylase. Plant Mol. Biol. 1994, 24, 35–49. [Google Scholar] [CrossRef]
- Kremer, L.; Nampoothiri, K.M.; Lesjean, S.; Dover, L.G.; Graham, S.; Betts, J.; Brennan, P.J.; Minnikin, D.E.; Locht, C.; Besra, G.S. Biochemical characterization of acyl carrier protein (AcpM) and malonyl-CoA: AcpM transacylase (MtFabD), two major components of Mycobacterium tuberculosis fatty acid synthase II. J. Biol. Chem. 2001, 276, 27967–27974. [Google Scholar] [CrossRef]
- Zhang, S.; Wu, S.; Hou, Q.; Zhao, J.; Fang, C.; An, X.; Wan, X. Fatty acid de novo biosynthesis in plastids: Key enzymes and their critical roles for male reproduction and other processes in plants. Plant Physiol. Biochem. 2024, 210, 108654. [Google Scholar] [CrossRef] [PubMed]
- Aznar-Moreno, J.A.; Sánchez, R.; Gidda, S.K.; Martínez-Force, E.; Moreno-Pérez, A.J.; Venegas Calerón, M.; Garcés, R.; Mullen, R.T.; Salas, J.J. New insights into sunflower (Helianthus annuus L.) FatA and FatB thioesterases, their regulation, structure and distribution. Front. Plant Sci. 2018, 9, 1496. [Google Scholar] [CrossRef]
- Bates, P.D.; Shockey, J. Towards rational control of seed oil composition: Dissecting cellular organization and flux control of lipid metabolism. Plant Physiol. 2025, 197, kiae658. [Google Scholar] [CrossRef]
- Dehesh, K.; Tai, H.; Edwards, P.; Byrne, J.; Jaworski, J.G. Overexpression of 3-ketoacyl-acyl-carrier protein synthase IIIs in plants reduces the rate of lipid synthesis. Plant Physiol. 2001, 125, 1103–1114. [Google Scholar] [CrossRef]
- Higashi, Y.; Saito, K. Lipidomic studies of membrane glycerolipids in plant leaves under heat stress. Prog. Lipid Res. 2019, 75, 100990. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Wang, Y.; Liu, J.; Liu, Y.; Cao, X.; Xue, S. Structural insight into acyl-ACP thioesterase toward substrate specificity design. ACS Chem. Biol. 2017, 12, 2830–2836. [Google Scholar] [CrossRef]
- Bahadır, S.; Abdulla, M.F.; Mostafa, K.; Kavas, M.; Hacıkamiloğlu, S.; Kurt, O.; Yıldırım, K. Exploring the role of FAT genes in Solanaceae species through genome-wide analysis and genome editing. Plant Genome 2024, 17, e20506. [Google Scholar] [CrossRef]
- Salas, J.J.; Ohlrogge, J.B. Characterization of substrate specificity of plant FatA and FatB acyl-ACP thioesterases. Arch. Biochem. Biophys. 2002, 403, 25–34. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Zhang, D.; Lu, M. Cloning and expression analysis of PtFATB gene encoding the acyl-acyl carrier protein thioesterase in Populus tomentosa Carr. J. Genet. Genom. 2007, 34, 267–274. [Google Scholar] [CrossRef]
- Zhou, Z.; Lakhssassi, N.; Knizia, D.; Cullen, M.A.; El Baz, A.; Embaby, M.G.; Liu, S.; Badad, O.; Vuong, T.D.; AbuGhazaleh, A. Genome-wide identification and analysis of soybean acyl-ACP thioesterase gene family reveals the role of GmFAT to improve fatty acid composition in soybean seed. Theor. Appl. Genet. 2021, 134, 3611–3623. [Google Scholar] [CrossRef] [PubMed]
- Peng, Z.; Zhang, H.; Tian, H.; Shan, L.; Zhang, Z.; Ding, H.; Gao, W.; Li, X. The phylogeny and functional characterization of peanut acyl-ACP Thioesterases. J. Plant Growth Regul. 2020, 39, 1381–1392. [Google Scholar] [CrossRef]
- Jones, A.; Davies, H.M.; Voelker, T.A. Palmitoyl-acyl carrier protein (ACP) thioesterase and the evolutionary origin of plant acyl-ACP thioesterases. Plant Cell 1995, 7, 359–371. [Google Scholar]
- Nam, J.-W.; Yeon, J.; Jeong, J.; Cho, E.; Kim, H.B.; Hur, Y.; Lee, K.-R.; Yi, H. Overexpression of acyl-ACP thioesterases, CpFatB4 and CpFatB5, induce distinct gene expression reprogramming in developing seeds of Brassica napus. Int. J. Mol. Sci. 2019, 20, 3334. [Google Scholar] [CrossRef]
- Lin, N.; Ai, T.; Gao, J.; Fan, L.; Wang, S.; Chen, F. Identification of novel acyl-ACP thioesterase gene ClFATB1 from Cinnamomum longepaniculatum. Biochemistry 2013, 78, 1298–1303. [Google Scholar] [CrossRef]
- Aznar-Moreno, J.A.; Venegas-Calerón, M.; Martínez-Force, E.; Garcés, R.; Salas, J.J. Acyl carrier proteins from sunflower (Helianthus annuus L.) seeds and their influence on FatA and FatB acyl-ACP thioesterase activities. Planta 2016, 244, 479–490. [Google Scholar] [CrossRef]
- Liu, B.; Sun, Y.; Wang, X.; Xue, J.; Wang, J.; Jia, X.; Li, R. Identification and functional characterization of acyl-ACP thioesterases B (GhFatBs) responsible for palmitic acid accumulation in cotton seeds. Int. J. Mol. Sci. 2022, 23, 12805. [Google Scholar] [CrossRef] [PubMed]
- Dormann, P.; Voelker, T.A.; Ohlrogge, J.B. Accumulation of palmitate in Arabidopsis mediated by the acyl-acyl carrier protein thioesterase FATB1. Plant Physiol. 2000, 123, 637–644. [Google Scholar] [CrossRef]
- Han, H.; Wu, W.; Hou, H.; Zhang, M.; Guo, A.; Zhou, Y.; Liu, J.; Li, K.; Bai, S.; Li, B. Function analysis of transcription factor OSR1 regulating osmotic stress resistance in maize. Biochem. Biophys. Res. Commun. 2024, 714, 149956. [Google Scholar] [CrossRef]
- Stamenković, O.S.; Kostić, M.D.; Tasić, M.B.; Djalović, I.G.; Mitrović, P.M.; Biberdžić, M.O.; Veljković, V.B. Kinetic, thermodynamic and optimization study of the corn germ oil extraction process. Food Bioprod. Process. 2020, 120, 91–103. [Google Scholar] [CrossRef]
- Maki, K.; Dicklin, M.; Cassens, M.; Bell, M.; Bunczek, M.; Eren, F. Predictors of Cholesterol Lowering with Corn Oil Consumption: Results from a Pooled Analysis of Randomized, Free-living Feeding Trials (P08-112-19). Curr. Dev. Nutr. 2019, 3, nzz044.P008-112-019. [Google Scholar] [CrossRef]
- Wang, J.-K.; Li, Y.; Zhao, X.-L.; Liu, Y.-B.; Tan, J.; Xing, Y.-Y.; Adi, D.; Wang, Y.-T.; Fu, Z.-Y.; Ma, Y.-T. Ablation of plasma prekallikrein decreases low-density lipoprotein cholesterol by stabilizing low-density lipoprotein receptor and protects against atherosclerosis. Circulation 2022, 145, 675–687. [Google Scholar] [CrossRef] [PubMed]
- Marchler-Bauer, A.; Derbyshire, M.K.; Gonzales, N.R.; Lu, S.; Chitsaz, F.; Geer, L.Y.; Geer, R.C.; He, J.; Gwadz, M.; Hurwitz, D.I. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 2015, 43, D222–D226. [Google Scholar] [CrossRef]
- Lei, B.; Song, M.; Li, X.; Dang, X.; Qin, R.; Zhu, S.; An, X.; Liu, Q.; Yao, X.; Nie, Y. SMART V1.0: A database for small molecules with functional implications in plants. Interdiscip. Sci. Comput. Life Sci. 2022, 14, 279–283. [Google Scholar] [CrossRef] [PubMed]
- Duvaud, S.; Gabella, C.; Lisacek, F.; Stockinger, H.; Ioannidis, V.; Durinx, C. Expasy, the Swiss Bioinformatics Resource Portal, as designed by its users. Nucleic Acids Res. 2021, 49, W216–W227. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Wang, X.; Sun, Y.; Joseph, P.V.; Paterson, A.H. Detection of colinear blocks and synteny and evolutionary analyses based on utilization of MCScanX. Nat. Protoc. 2024, 19, 2206–2229. [Google Scholar] [CrossRef]
- Rombauts, S.; Déhais, P.; Van Montagu, M.; Rouzé, P. PlantCARE, a plant cis-acting regulatory element database. Nucleic Acids Res. 1999, 27, 295–296. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Nastou, K.; Koutrouli, M.; Kirsch, R.; Mehryary, F.; Hachilif, R.; Hu, D.; Peluso, M.E.; Huang, Q.; Fang, T. The STRING database in 2025: Protein networks with directionality of regulation. Nucleic Acids Res. 2025, 53, D730–D737. [Google Scholar] [CrossRef]
- Singhal, A.; Cao, S.; Churas, C.; Pratt, D.; Fortunato, S.; Zheng, F.; Ideker, T. Multiscale community detection in Cytoscape. PLoS Comput. Biol. 2020, 16, e1008239. [Google Scholar] [CrossRef] [PubMed]
- Farid, B.; Saddique, M.A.B.; Tahir, M.H.N.; Ikram, R.M.; Ali, Z.; Akbar, W. Expression divergence of BAG gene family in maize under heat stress. BMC Plant Biol. 2025, 25, 16. [Google Scholar] [CrossRef] [PubMed]
- Qian, B.; Wang, Q.; Zhang, C.; Guo, J.; Yu, Z.; Han, J.; Xia, H.; Zhao, R.; Yin, Y. Exploring the Roles of TALE Gene Family in Maize Drought Stress Responses. Agronomy 2024, 14, 1267. [Google Scholar] [CrossRef]
- Hall, J.A.; Van Saun, R.J.; Tornquist, S.J.; Gradin, J.L.; Pearson, E.G.; Wander, R.C. Effect of type of dietary polyunsaturated fatty acid supplement (corn oil or fish oil) on immune responses in healthy horses. J. Vet. Intern. Med. 2004, 18, 880–886. [Google Scholar] [CrossRef]
- Apgar, J.L.; Shively, C.A.; Tarka Jr, S.M. Digestibility of cocoa butter and corn oil and their influence on fatty acid distribution in rats. J. Nutr. 1987, 117, 660–665. [Google Scholar] [CrossRef]
- Tan, K.W.M.; Lee, Y.K. Expression of the heterologous Dunaliella tertiolecta fatty acyl-ACP thioesterase leads to increased lipid production in Chlamydomonas reinhardtii. J. Biotechnol. 2017, 247, 60–67. [Google Scholar] [CrossRef]
- Wang, H.; Shi, J.; Guo, W.; Sun, X.; Niu, S.; Chen, L.; Liu, S.; Ma, L. The identification and expression analysis of walnut Acyl-ACP thioesterases. Front. Genet. 2024, 15, 1409159. [Google Scholar] [CrossRef]
- Li, Y.; Xian, X.; Guo, L.; Zhang, J.; Gan, C.; Wang, Z.; Li, H.; Li, X.; Yuan, X.; Zhang, N. CsbZIP50 binds to the G-box/ABRE motif in CsRD29A promoter to enhance drought tolerance in cucumber. Environ. Exp. Bot. 2022, 199, 104884. [Google Scholar] [CrossRef]
- Wang, Y.; Xu, H.; Liu, W.; Wang, N.; Qu, C.; Jiang, S.; Fang, H.; Zhang, Z.; Chen, X. Methyl jasmonate enhances apple’cold tolerance through the JAZ–MYC2 pathway. Plant Cell Tissue Organ Cult. (PCTOC) 2019, 136, 75–84. [Google Scholar] [CrossRef]
- Soma, F.; Takahashi, F.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Cellular phosphorylation signaling and gene expression in drought stress responses: ABA-dependent and ABA-independent regulatory systems. Plants 2021, 10, 756. [Google Scholar] [CrossRef]
- Li, Y.; Hu, Z.; Dong, Y.; Xie, Z. Overexpression of the cotton trihelix transcription factor GhGT23 in Arabidopsis mediates salt and drought stress tolerance by binding to GT and MYB promoter elements in stress-related genes. Front. Plant Sci. 2023, 14, 1144650. [Google Scholar] [CrossRef]
- Li, Z.; Liu, W.; Chen, Q.; Zhang, S.; Mei, Z.; Yu, L.; Wang, C.; Mao, Z.; Chen, Z.; Chen, X.; et al. Mdm-miR858 targets MdMYB9 and MdMYBPA1 to participate anthocyanin biosynthesis in red-fleshed apple. Plant J. 2023, 113, 1295–1309. [Google Scholar] [CrossRef] [PubMed]
- He, M.; Qin, C.-X.; Wang, X.; Ding, N.-Z. Plant unsaturated fatty acids: Biosynthesis and regulation. Front. Plant Sci. 2020, 11, 390. [Google Scholar] [CrossRef] [PubMed]
- Thelen, J.J.; Ohlrogge, J.B. Metabolic engineering of fatty acid biosynthesis in plants. Metab. Eng. 2002, 4, 12–21. [Google Scholar] [CrossRef]
- Wang, Y.; Shen, Y.; Dong, W.; Cai, X.; Yang, J.; Chen, Y.; Jia, B.; Sun, M.; Sun, X. PHD17 acts as a target of miR1320 to negatively control cold tolerance via JA-activated signaling in rice. Crop J. 2024, 12, 1447–1458. [Google Scholar] [CrossRef]
- Megha, S.; Basu, U.; Kav, N.N. Regulation of low temperature stress in plants by microRNAs. Plant Cell Environ. 2018, 41, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Shen, L.; Ren, D.; Hu, J.; Chen, G.; Zhu, L.; Gao, Z.; Zhang, G.; Guo, L.; Zeng, D. Characterization, expression, and interaction analyses of OsMORF gene family in rice. Genes 2019, 10, 694. [Google Scholar] [CrossRef] [PubMed]

| Protein ID | Name | aa | CDS | MW | pI | GRAVY |
|---|---|---|---|---|---|---|
| Zm00001d006080 | ZmFATA1 | 157 | 471 | 16,956.17 | 8.68 | −0.17 |
| Zm00001d021046 | ZmFATA2 | 203 | 609 | 22,119.05 | 8.24 | −0.14 |
| Zm00001d026569 | ZmFATA3 | 426 | 1278 | 47,666.77 | 6.37 | −0.34 |
| Zm00001d029901 | ZmFATB1 | 160 | 480 | 17,915.04 | 5.4 | −0.54 |
| Zm00001d029902 | ZmFATB2 | 142 | 426 | 16,029.88 | 8.64 | −0.75 |
| Zm00001d005148 | ZmFATB3 | 379 | 1137 | 42,303.49 | 9.19 | −0.44 |
| Zm00001d036137 | ZmFATB4 | 239 | 717 | 25,933.46 | 8.56 | −0.08 |
| Zm00001d021004 | ZmFATB5 | 83 | 249 | 9737.42 | 10.29 | −0.37 |
| Zm00001d045387 | ZmFATB6 | 434 | 1302 | 47,759.83 | 9.01 | −0.39 |
| Zm00001d046454 | ZmFATB7 | 418 | 1254 | 46,396.3 | 8.25 | −0.36 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.; Hou, S.; Sun, Y.; Li, S.; Sun, M.; Guo, B.; Wang, L.; Cai, Q.; Li, X.; Li, S.; et al. Comprehensive Characterization of the FATs Gene Family in Maize: Phylogeny, Expression Patterns, and Regulatory Networks. Genes 2025, 16, 1035. https://doi.org/10.3390/genes16091035
Li Y, Hou S, Sun Y, Li S, Sun M, Guo B, Wang L, Cai Q, Li X, Li S, et al. Comprehensive Characterization of the FATs Gene Family in Maize: Phylogeny, Expression Patterns, and Regulatory Networks. Genes. 2025; 16(9):1035. https://doi.org/10.3390/genes16091035
Chicago/Turabian StyleLi, Yunlong, Shuai Hou, Yan Sun, Shujun Li, Minghao Sun, Baitao Guo, Luyao Wang, Quan Cai, Xin Li, Sinan Li, and et al. 2025. "Comprehensive Characterization of the FATs Gene Family in Maize: Phylogeny, Expression Patterns, and Regulatory Networks" Genes 16, no. 9: 1035. https://doi.org/10.3390/genes16091035
APA StyleLi, Y., Hou, S., Sun, Y., Li, S., Sun, M., Guo, B., Wang, L., Cai, Q., Li, X., Li, S., & Zhang, J. (2025). Comprehensive Characterization of the FATs Gene Family in Maize: Phylogeny, Expression Patterns, and Regulatory Networks. Genes, 16(9), 1035. https://doi.org/10.3390/genes16091035







