Genome-Wide Identification of the BnaRFS Gene Family and Functional Characterization of BnaRFS6 in Brassica napus
Abstract
1. Introduction
2. Results
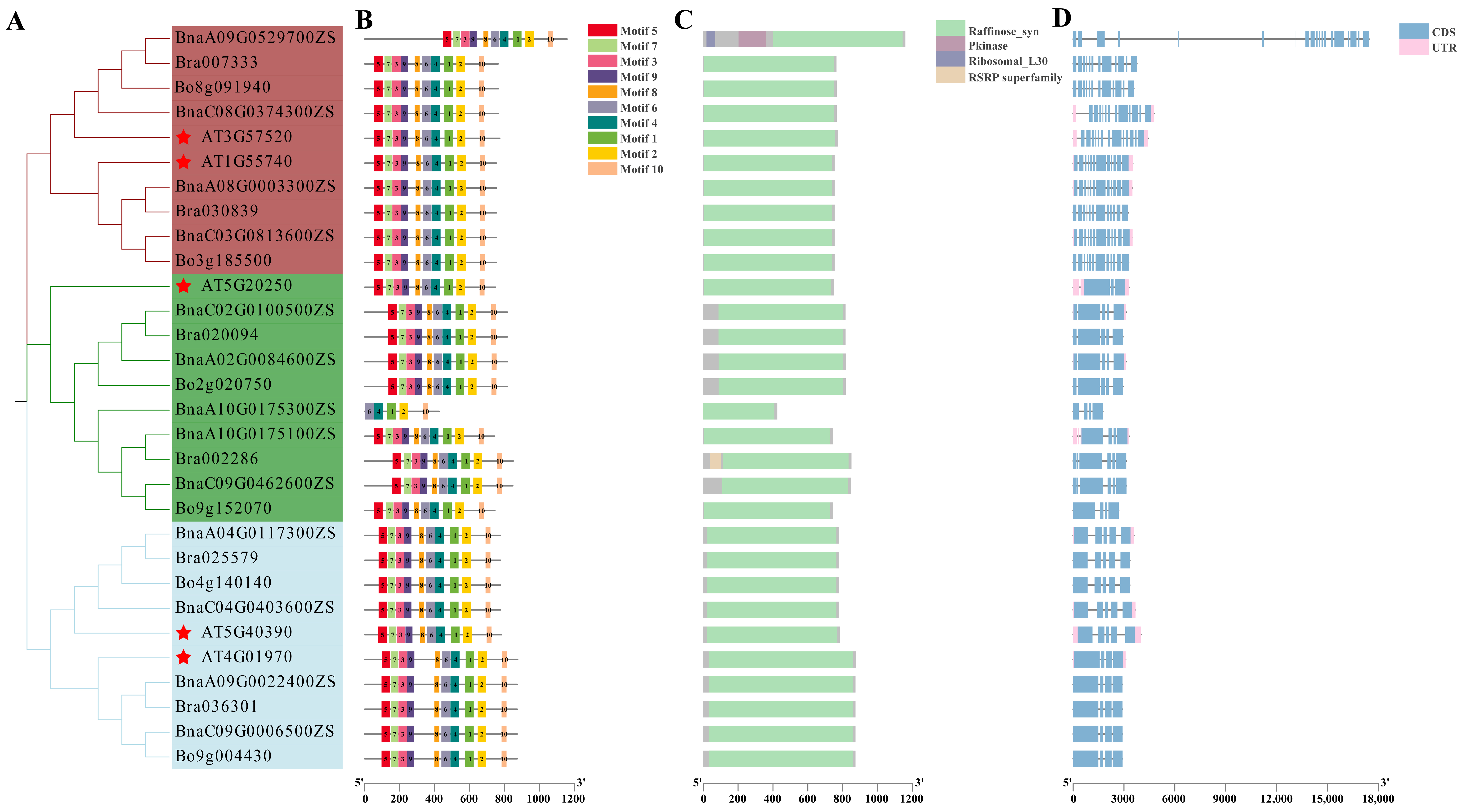
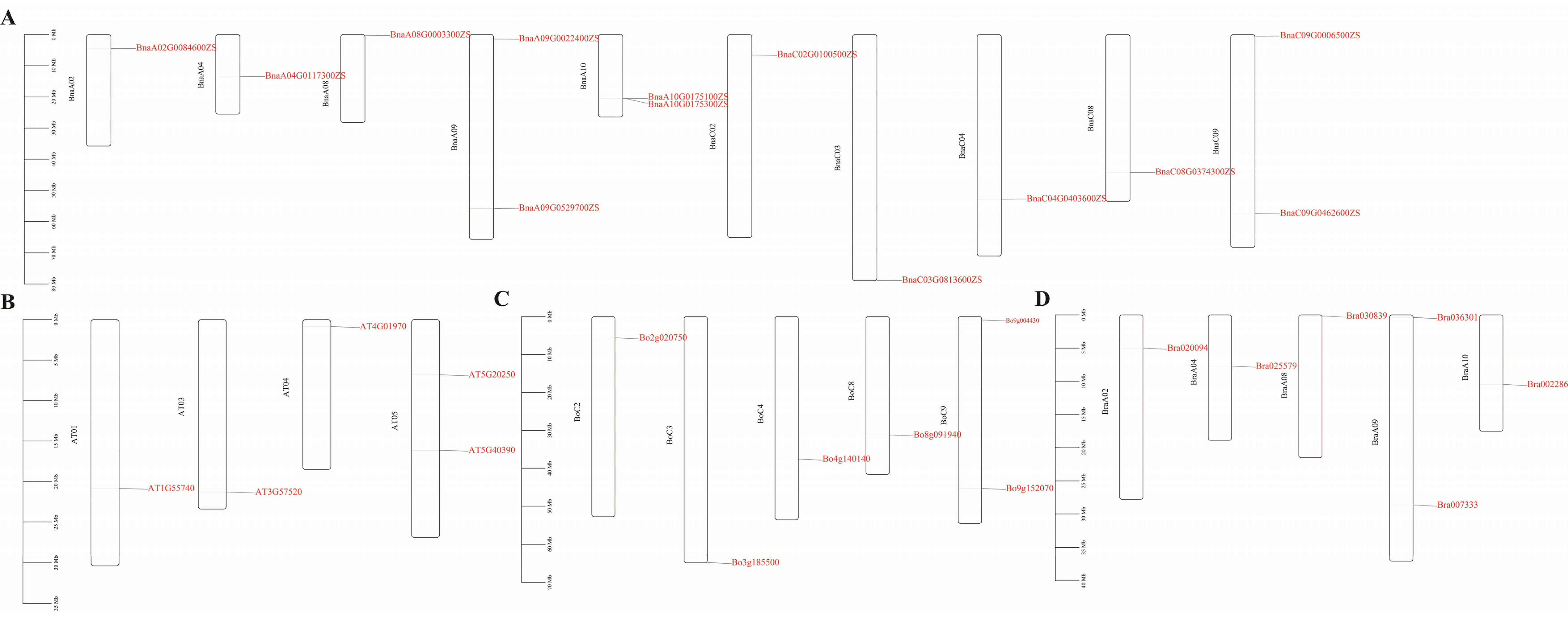
2.1. Bioinformatic Analysis of the RFS Gene Family
2.2. Physicochemical Properties of RFS Proteins in B. napus, B. rapa, and B. oleracea
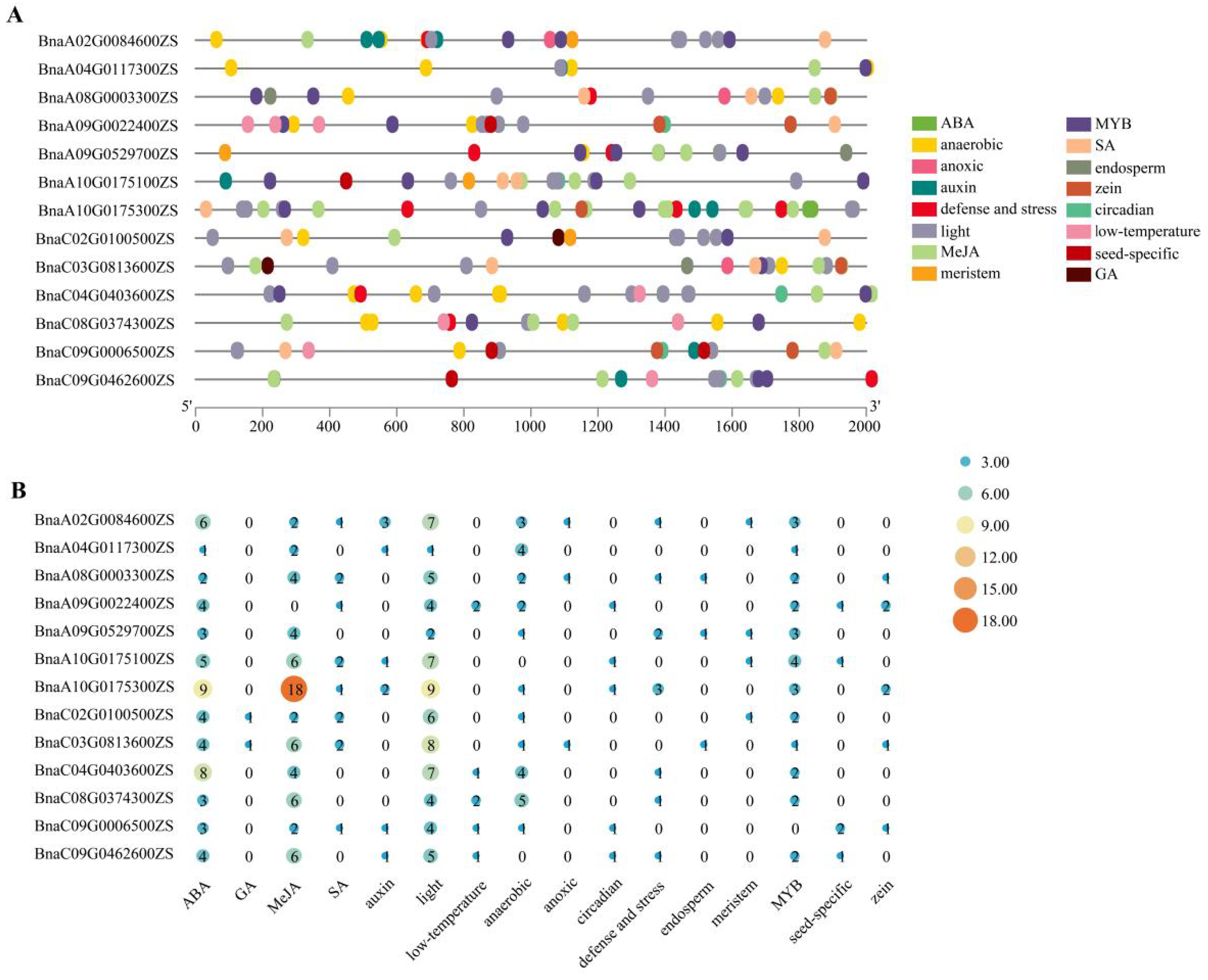
2.3. Cis-Element Analysis of BnaRFS Gene Promoter Regions
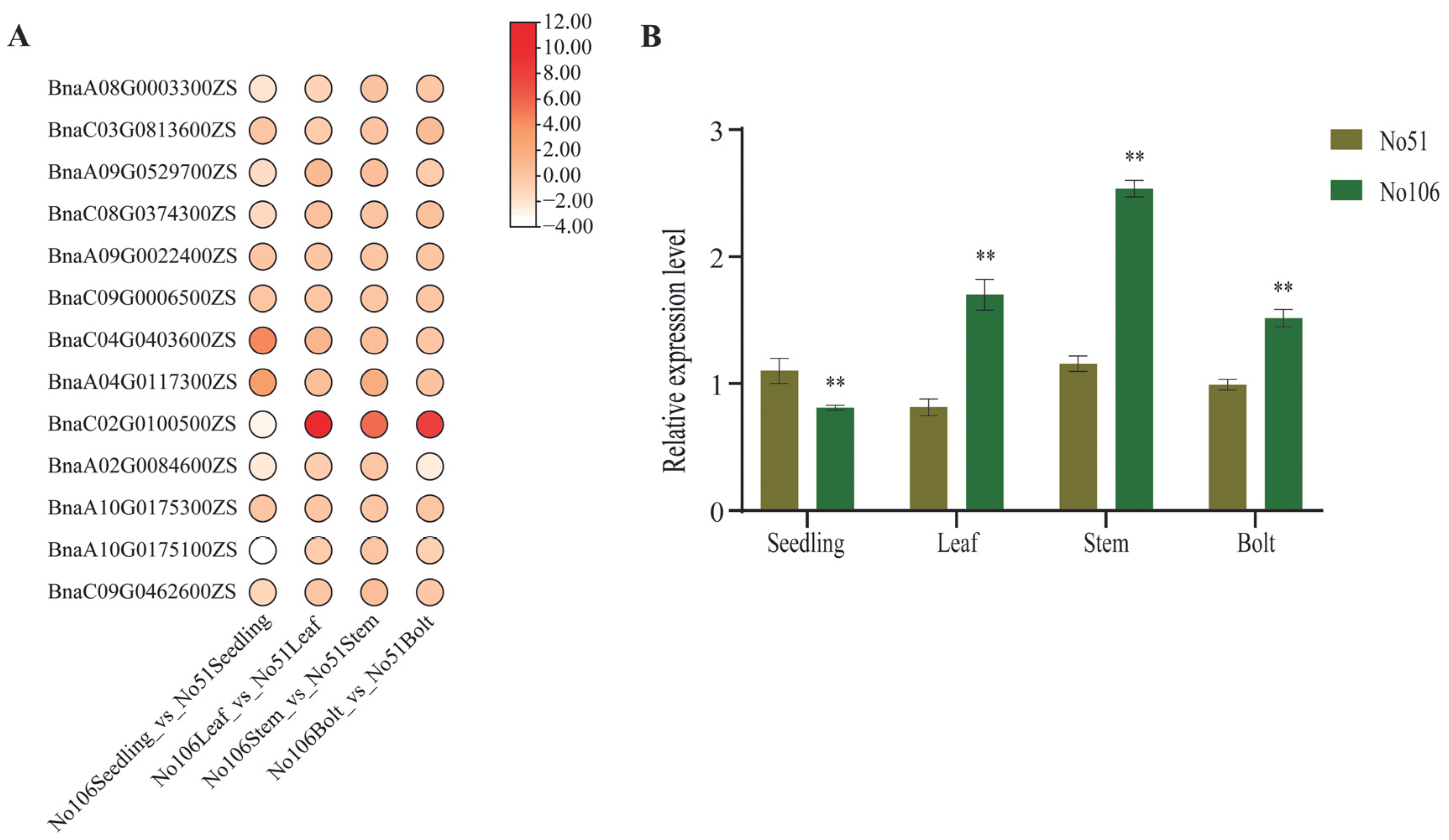
2.4. Expression Analysis of BnaRFS Genes in Different Tissues
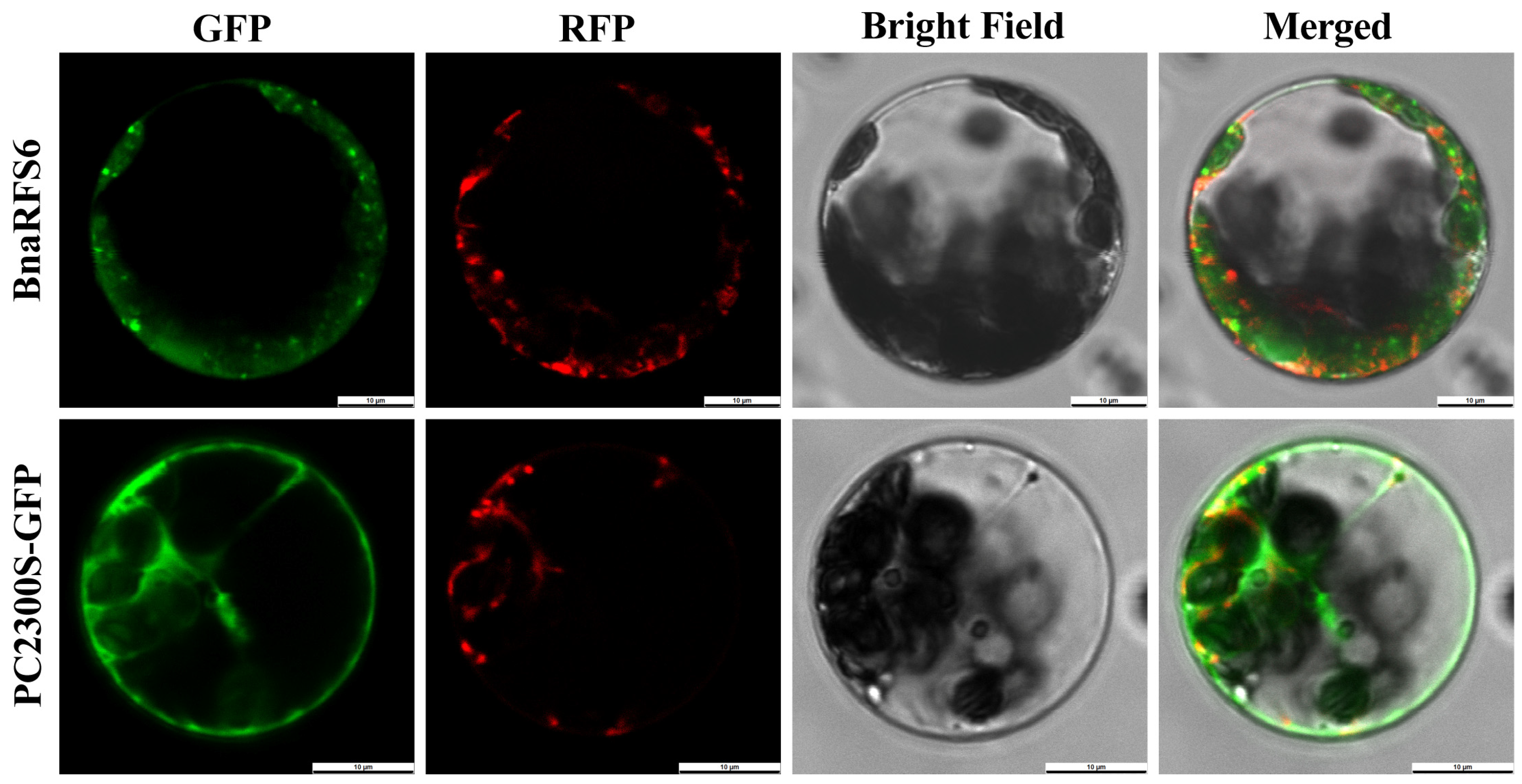
2.5. Cloning and Subcellular Localization of BnaRFS6
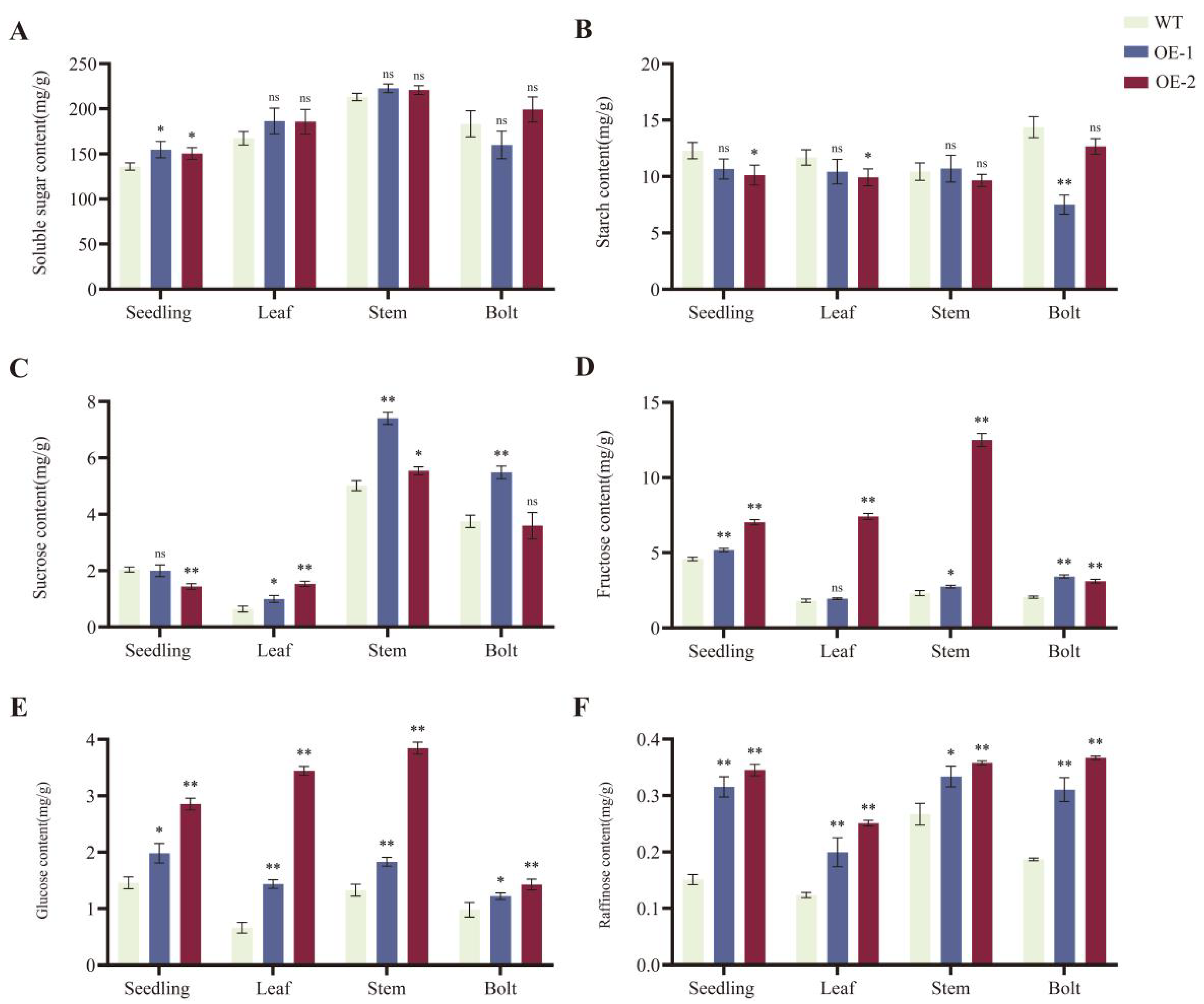
2.6. Overexpression of BnaRFS6 Enhances Soluble Sugar Content in Rapeseed Bolting
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Bioinformatics Analysis
4.3. Gene Cloning and Subcellular Localization
4.4. Construction of Overexpression Vector and Genetic Transformation in B. napus
4.5. Expression Analysis
4.6. Sugar Content Determination
4.7. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Xiao, Z.; Pan, Y.; Wang, C.; Li, X.; Lu, Y.; Tian, Z.; Kuang, L.; Wang, X.; Dun, X.; Wang, H. Multi-Functional Development and Utilization of Rapeseed: Comprehensive Analysis of the Nutritional Value of Rapeseed Sprouts. Foods 2022, 11, 778. [Google Scholar] [CrossRef]
- Jahangir, M.; Kim, H.; Choi, Y.; Verpoorte, R. Health-Affecting Compounds in Brassicaceae. Compr. Rev. Food Sci. Food Saf. 2010, 8, 31–43. [Google Scholar] [CrossRef]
- Fenwick, G.; Griffiths, N.; Heaney, R. Bitterness in brussels sprouts (Brassica oleracea L. var.gemmifera): The role of glucosinolates and their breakdown products. J. Sci. Food Agric. 1983, 34, 73–80. [Google Scholar] [CrossRef]
- Padilla, G.; Cartea, M.; Velasco, P.; de Haro, A.; Ordás, A. Variation of glucosinolates in vegetable crops of Brassica rapa. Phytochemistry 2007, 68, 536–545. [Google Scholar] [CrossRef]
- Dinehart, M.; Hayes, J.; Bartoshuk, L.; Lanier, S.; Duffy, V. Bitter taste markers explain variability in vegetable sweetness, bitterness, and intake. Physiol. Behav. 2006, 87, 304–313. [Google Scholar] [CrossRef] [PubMed]
- Engel, E.; Baty, C.; Le Corre, D.; Souchon, I.; Martin, N. Flavor-active compounds potentially implicated in cooked cauliflower acceptance. J. Agric. Food Chem. 2002, 50, 6459–6467. [Google Scholar] [CrossRef]
- Milne, R.; Grof, C.; Patrick, J. Mechanisms of phloem unloading: Shaped by cellular pathways, their conductances and sink function. Curr. Opin. Plant Biol. 2018, 43, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Geiger, D. Plant glucose transporter structure and function. Pflug. Arch.-Eur. J. Physiol. 2020, 472, 1111–1128. [Google Scholar] [CrossRef]
- Li, S.; Li, T.; Kim, W.; Kitaoka, M.; Yoshida, S.; Nakajima, M.; Kobayashi, H. Characterization of raffinose synthase from rice (Oryza sativa L. var. Nipponbare). Biotechnol. Lett. 2007, 29, 635–640. [Google Scholar] [CrossRef]
- Gangl, R.; Tenhaken, R. Raffinose Family Oligosaccharides Act As Galactose Stores in Seeds and Are Required for Rapid Germination of Arabidopsis in the Dark. Front. Plant Sci. 2016, 7, 1115. [Google Scholar] [CrossRef] [PubMed]
- Shi, Q.; Han, G.; Liu, Y.; Jiang, J.; Jia, Y.; Li, X. Nutrient composition and quality traits of dried jujube fruits in seven producing areas based on metabolomics analysis. Food Chem. 2022, 385, 132627. [Google Scholar] [CrossRef]
- Zhou, M.; Zhang, Q.; Zhou, M.; Sun, Z.; Zhu, X.; Shao, J.; Tang, Y.; Wu, Y. Genome-wide identification of genes involved in raffinose metabolism in Maize. Glycobiology 2012, 22, 1775–1785. [Google Scholar] [CrossRef]
- Li, T.; Zhang, Y.; Wang, D.; Liu, Y.; Dirk, L.; Goodman, J.; Downie, A.; Wang, J.; Wang, G.; Zhao, T. Regulation of Seed Vigor by Manipulation of Raffinose Family Oligosaccharides in Maize and Arabidopsis thaliana. Mol. Plant 2017, 10, 1540–1555. [Google Scholar] [CrossRef]
- Jing, Y.; Lang, S.; Wang, D.; Xue, H.; Wang, X. Functional characterization of galactinol synthase and raffinose synthase in desiccation tolerance acquisition in developing Arabidopsis seeds. J. Plant Physiol. 2018, 230, 109–121. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Zhang, Y.; Liu, Y.; Li, X.; Hao, G.; Han, Q.; Dirk, L.; Downie, A.; Ruan, Y.; Wang, J.; et al. Raffinose synthase enhances drought tolerance through raffinose synthesis or galactinol hydrolysis in maize and Arabidopsis plants. J. Biol. Chem. 2020, 295, 8064–8077. [Google Scholar] [CrossRef]
- Peterbauer, T.; Mach, L.; Mucha, J.; Richter, A. Functional expression of a cDNA encoding pea (Pisum sativum L.) raffinose synthase, partial purification of the enzyme from maturing seeds, and steady-state kinetic analysis of raffinose synthesis. Planta 2002, 215, 839–846. [Google Scholar] [CrossRef]
- Sui, X.; Meng, F.; Wang, H.; Wei, Y.; Li, R.; Wang, Z.; Hu, L.; Wang, S.; Zhang, Z. Molecular cloning, characteristics and low temperature response of raffinose synthase gene in Cucumis sativus L. J. Plant Physiol. 2012, 169, 1883–1891. [Google Scholar] [CrossRef]
- Egert, A.; Keller, F.; Peters, S. Abiotic stress-induced accumulation of raffinose in Arabidopsis leaves is mediated by a single raffinose synthase (RS5, At5g40390). BMC Plant Biol. 2013, 13, 218. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Meng, D.; Li, J.; Bao, Y.; Yu, P.; Dou, G.; Guo, J.; Tang, C.; Lv, J.; Wang, X.; et al. Analysis of the Rice Raffinose Synthase (OsRS) Gene Family and Haplotype Diversity. Int. J. Mol. Sci. 2024, 11, 9815. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Yang, Y.; Wang, T.; Wang, Y.; Zhang, X.; Min, D.; Zhang, X. Analysis of Raffinose Synthase Gene Family in Bread Wheat and Identification of Drought Resistance and Salt Tolerance Function of TaRS15-3B. Int. J. Mol. Sci. 2023, 24, 11185. [Google Scholar] [CrossRef]
- Cui, R.; Wang, X.; Malik, W.; Lu, X.; Chen, X.; Wang, D.; Wang, J.; Wang, S.; Chen, C.; Guo, L.; et al. Genome-wide identification and expression analysis of Raffinose synthetase family in cotton. BMC Bioinform. 2021, 22, 356. [Google Scholar] [CrossRef]
- Sweetlove, L.; Beard, K.; Nunes-Nesi, A.; Fernie, A.; Ratcliffe, R. Not just a circle: Flux modes in the plant TCA cycle. Trends Plant Sci. 2010, 15, 462–470. [Google Scholar] [CrossRef]
- Van den Ende, W. Multifunctional fructans and raffinose family oligosaccharides. Front. Plant Sci. 2013, 4, 247. [Google Scholar] [CrossRef] [PubMed]
- Leggewie, G.; Kolbe, A.; Lemoine, R.; Roessner, U.; Lytovchenko, A.; Zuther, E.; Kehr, J.; Frommer, W.; Riesmeier, J.; Willmitzer, L.; et al. Overexpression of the sucrose transporter SoSUT1 in potato results in alterations in leaf carbon partitioning and in tuber metabolism but has little impact on tuber morphology. Planta 2003, 217, 158–167. [Google Scholar] [CrossRef]
- Wu, F.; Shen, S.; Lee, L.; Lee, S.; Chan, M.; Lin, C. Tape-Arabidopsis Sandwich—A simpler Arabidopsis protoplast isolation method. Plant Methods 2009, 5, 16. [Google Scholar] [CrossRef]
- Pfaffl, M. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, 2003–2007. [Google Scholar] [CrossRef]
- Roe, J. The determination of sugar in blood and spinal fluid with anthrone reagent. J. Biol. Chem. 1955, 212, 335–343. [Google Scholar] [CrossRef] [PubMed]
- Filip, M.; Vlassa, M.; Coman, V.; Halmagyi, A. Simultaneous determination of glucose, fructose, sucrose and sorbitol in the leaf and fruit peel of different apple cultivars by the HPLC-RI optimized method. Food Chem. 2015, 199, 653–659. [Google Scholar] [CrossRef]
| Arabidopsis | Homologous Gene | a.a | MW (kDa) | pI | Instability Index | GRAVY |
|---|---|---|---|---|---|---|
| AT1G55740 RFS1 | BnaA08G0003300ZS | 754 | 83.42 | 5.94 | 31.12 | −0.194 |
| BnaC03G0813600ZS | 754 | 83.41 | 6.01 | 32.22 | −0.189 | |
| Bra030839 | 754 | 83.49 | 5.93 | 30.43 | −0.199 | |
| Bo3g185500 | 754 | 83.50 | 6.04 | 31.73 | −0.204 | |
| AT3G57520 RFS2 | BnaA09G0529700ZS | 1159 | 129.20 | 6.8 | 39.04 | −0.259 |
| BnaC08G0374300ZS | 765 | 84.03 | 5.75 | 36.29 | −0.185 | |
| Bra007333 | 764 | 84.03 | 5.68 | 35.05 | −0.189 | |
| Bo8g091940 | 765 | 84.09 | 5.78 | 35.69 | −0.187 | |
| AT4G01970 RFS4 | BnaA09G0022400ZS | 873 | 97.71 | 5.12 | 40.79 | −0.236 |
| BnaC09G0006500ZS | 873 | 97.67 | 5.14 | 40.09 | −0.231 | |
| Bra036301 | 873 | 97.72 | 5.14 | 41.19 | −0.239 | |
| Bo9g004430 | 873 | 97.67 | 5.14 | 40.09 | −0.231 | |
| AT5G40390 RFS5 | BnaC04G0403600ZS | 777 | 85.87 | 4.94 | 33.26 | −0.114 |
| BnaA04G0117300ZS | 777 | 85.81 | 4.96 | 34.21 | −0.12 | |
| Bra025579 | 777 | 85.81 | 4.96 | 34.21 | −0.12 | |
| Bo4g140140 | 778 | 85.92 | 4.98 | 33.78 | −0.089 | |
| AT5G20250 RFS6 | BnaC02G0100500ZS | 816 | 90.45 | 6.25 | 38.26 | −0.233 |
| BnaA02G0084600ZS | 818 | 90.50 | 6.13 | 39.05 | −0.239 | |
| BnaA10G0175300ZS | 425 | 46.66 | 6.06 | 37.6 | −0.159 | |
| BnaA10G0175100ZS | 744 | 82.25 | 5.51 | 37.28 | −0.263 | |
| BnaC09G0462600ZS | 848 | 94.13 | 6.5 | 42.8 | −0.312 | |
| Bra020094 | 816 | 90.26 | 6.19 | 38.05 | −0.234 | |
| Bra002286 | 850 | 94.44 | 6.23 | 43.55 | −0.339 | |
| Bo2g020750 | 817 | 90.31 | 6.08 | 38.56 | −0.253 | |
| Bo9g152070 | 745 | 82.38 | 5.8 | 39.35 | −0.252 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhou, B.; Guan, C.; Guan, M. Genome-Wide Identification of the BnaRFS Gene Family and Functional Characterization of BnaRFS6 in Brassica napus. Genes 2025, 16, 1032. https://doi.org/10.3390/genes16091032
Zhou B, Guan C, Guan M. Genome-Wide Identification of the BnaRFS Gene Family and Functional Characterization of BnaRFS6 in Brassica napus. Genes. 2025; 16(9):1032. https://doi.org/10.3390/genes16091032
Chicago/Turabian StyleZhou, Bingqian, Chunyun Guan, and Mei Guan. 2025. "Genome-Wide Identification of the BnaRFS Gene Family and Functional Characterization of BnaRFS6 in Brassica napus" Genes 16, no. 9: 1032. https://doi.org/10.3390/genes16091032
APA StyleZhou, B., Guan, C., & Guan, M. (2025). Genome-Wide Identification of the BnaRFS Gene Family and Functional Characterization of BnaRFS6 in Brassica napus. Genes, 16(9), 1032. https://doi.org/10.3390/genes16091032





