Cloning and Characterization of 12 TCP Genes in Medicinal Plant Plantago asiatica via De Novo Transcriptome Assembly
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Growth Conditions
2.2. Transcriptome Library Preparation
2.3. Transcriptome De Novo Assembly
2.4. Gene Annotation and Motif Analysis
2.5. Phylogenetic Tree Analysis
2.6. PaTCPs Amplification and Quantitative PCR
2.7. PaTCPs Subcellular Localization
3. Results
3.1. De Novo Transcriptome Assembly of P. asiatica
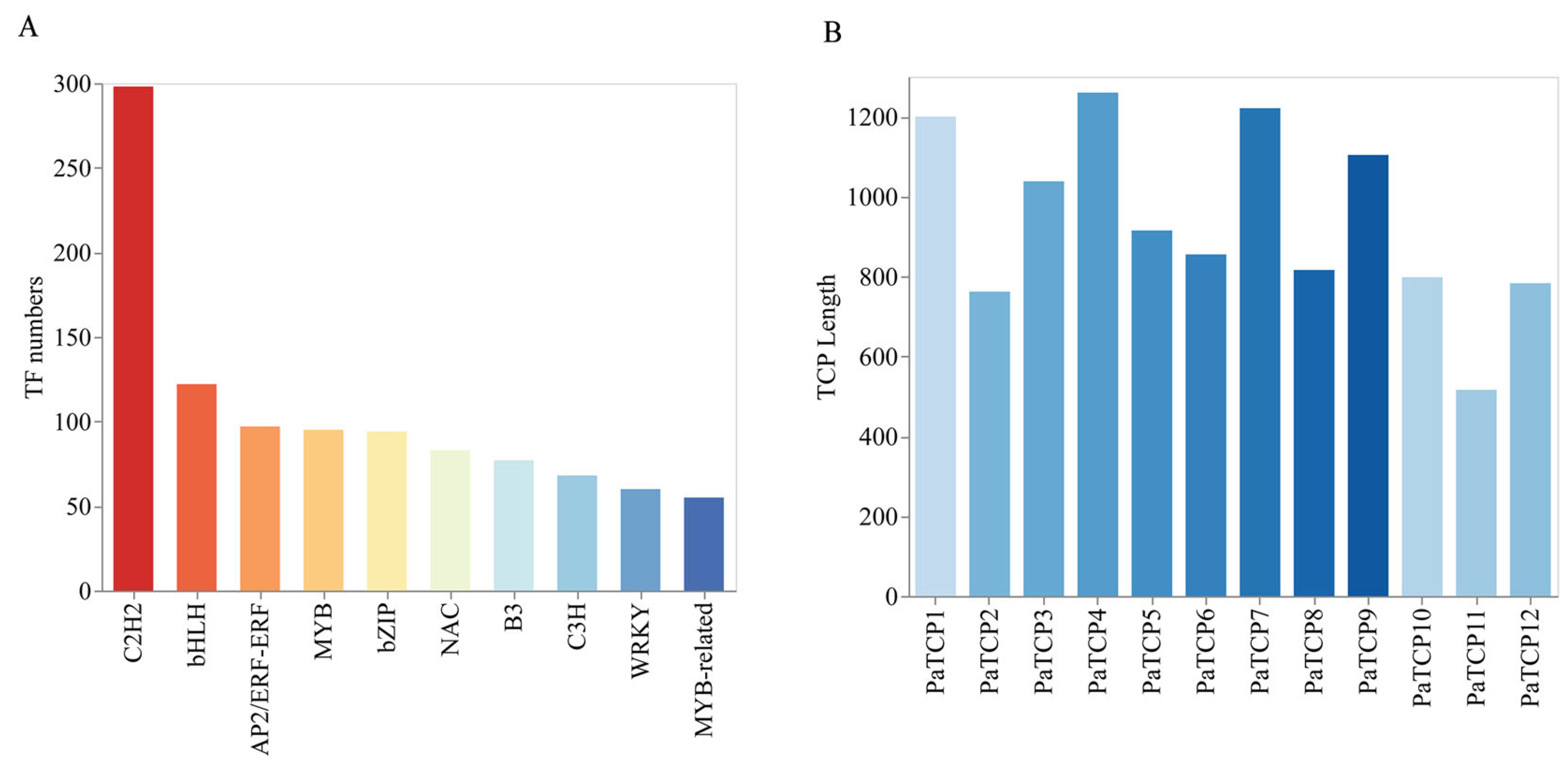
3.2. Unigenes Annotation
3.3. Clone and Sequencing Validation of the 12 TCP TF in P. asiatica
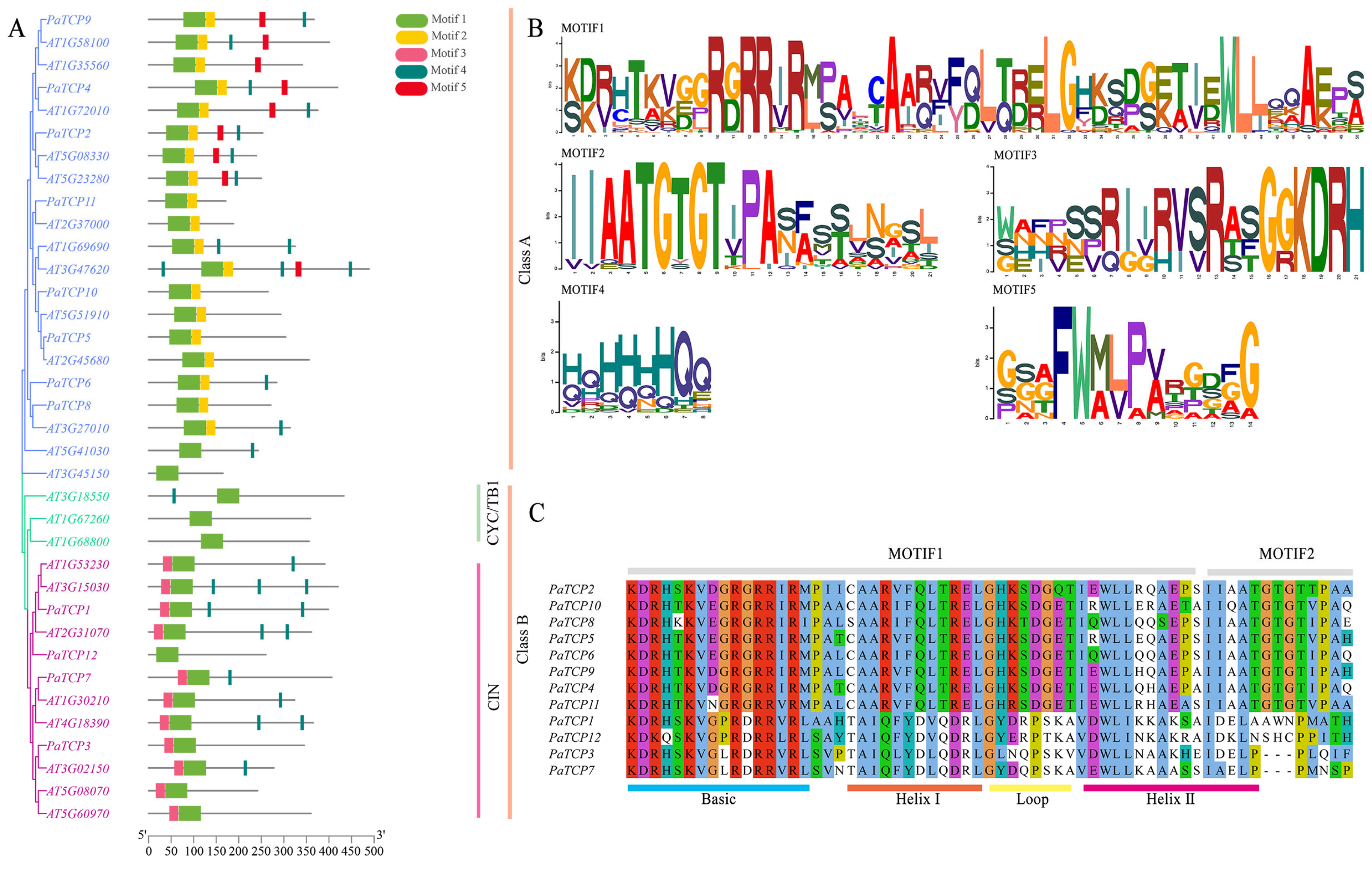
3.4. Phylogenetic Tree and Motif Analysis of PaTCPs
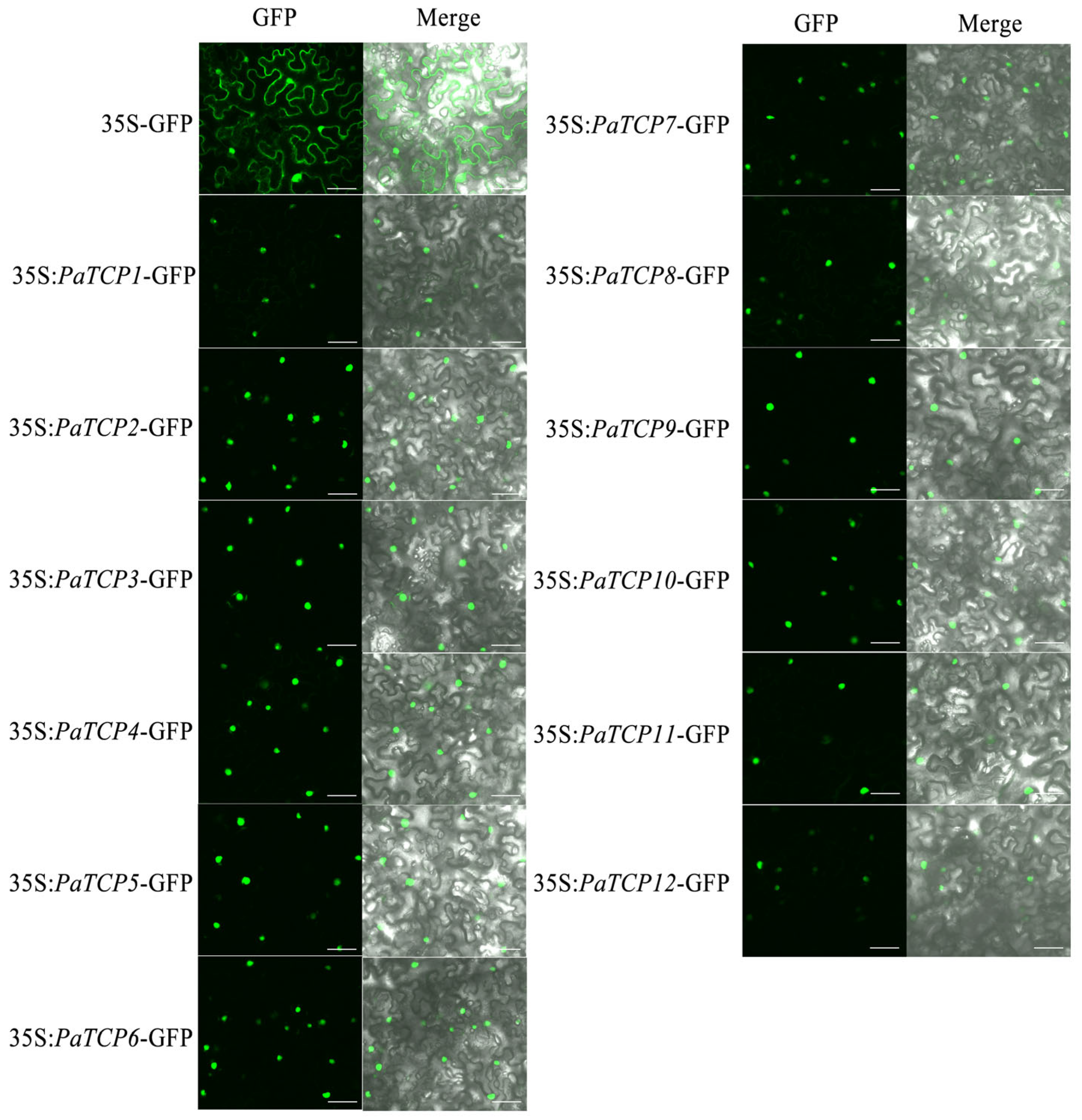
3.5. Subcellular Localization Analysis of the PaTCPs
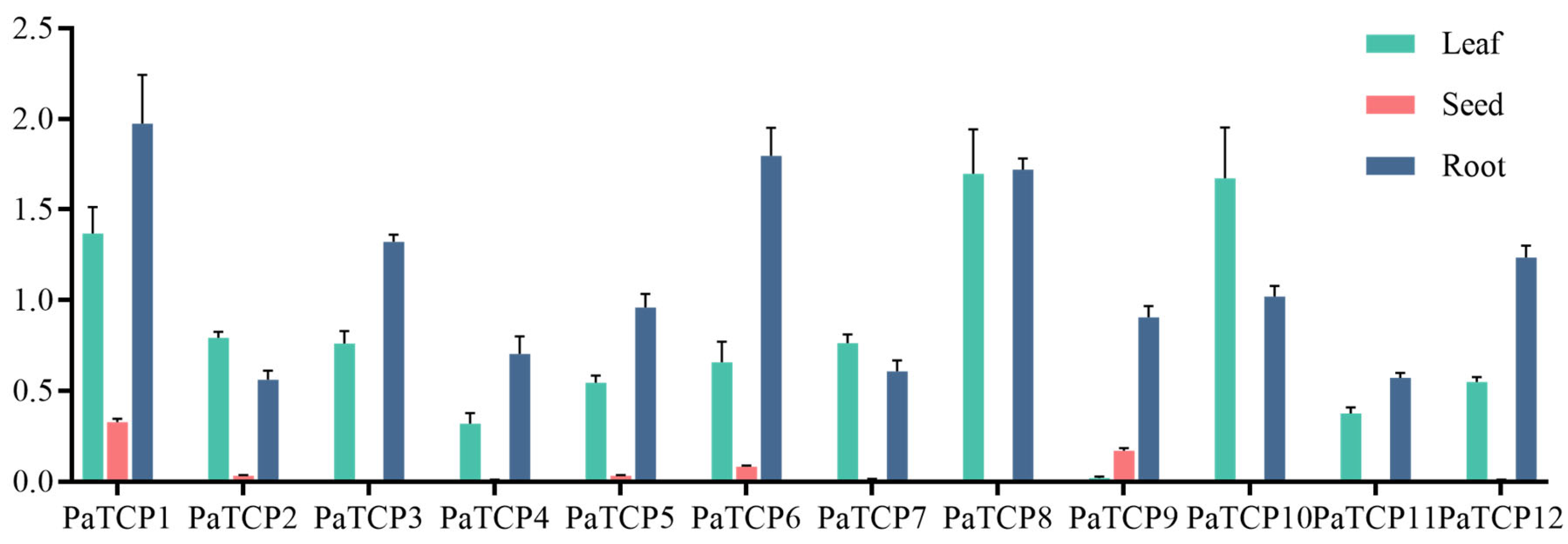
3.6. Tissue-Specific Expression of PaTCPs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Si, H.; Li, R.; Zhang, Q.; Liu, L. Complete Chloroplast Genome of Plantago asiatica and Its Phylogenetic Position in Plantaginaceae. Mitochondrial DNA B Resour. 2022, 7, 819–821. [Google Scholar] [CrossRef]
- Gonçalves, S.; Romano, A. The Medicinal Potential of Plants from the Genus Plantago (Plantaginaceae). Ind. Crops Prod. 2016, 83, 213–226. [Google Scholar] [CrossRef]
- Wen, S.-Y.; Wei, B.-Y.; Ma, J.-Q.; Wang, L.; Chen, Y.-Y. Phytochemicals, Biological Activities, Molecular Mechanisms, and Future Prospects of Plantago asiatica L. J. Agric. Food Chem. 2023, 71, 143–173. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Hou, Q.; Sun, M.; Sun, J.; Zhang, B. Targeted Isolation of Antioxidant Constituents from Plantago asiatica L.and In Vitro Activity Assay. Molecules 2020, 25, 1825. [Google Scholar] [CrossRef] [PubMed]
- Ye, C.-L.; Hu, W.-L.; Dai, D.-H. Extraction of Polysaccharides and the Antioxidant Activity from the Seeds of Plantago asiatica L. Int. J. Biol. Macromol. 2011, 49, 466–470. [Google Scholar] [CrossRef]
- Qian, B.; Hu, J.; Dai, L.; Zhou, Y.; Xu, H. Anti-Inflammatory Effect of Plantago asiatica Crude Extract in Rat Gout Arthritis Model. J. Immunotoxicol. 2025, 22, 2453156. [Google Scholar] [CrossRef]
- Astuti, A.D.; Etikawati, N.; Pangastuti, A. Antibacterial Activity of Plantago Major Leaves against Streptococcus Pyogenes ATCC 19615 as a Cause of Tonsilitis. Asian J. Trop. Biotechnol. 2020, 17, 14–21. [Google Scholar] [CrossRef]
- Chiang, L.-C.; Chiang, W.; Chang, M.-Y.; Lin, C.-C. In Vitro Cytotoxic, Antiviral and Immunomodulatory Effects of Plantago Major and Plantago asiatica. Am. J. Chin. Med. 2003, 31, 225–234. [Google Scholar] [CrossRef]
- Zhang, J.; Cui, J.; Gao, J.; Zhang, D.; Lin, D.; Lin, J. Polysaccharides of Plantago asiatica Enhance Antitumor Activity via Regulating Macrophages to M1-like Phenotype. Biomed. Pharmacother. 2023, 159, 114246. [Google Scholar] [CrossRef]
- Lim, Y.; Lee, K.W.; Kim, J.Y.; Kwon, O. A Beverage of Asiatic Plantain Extracts Alleviated Postprandial Oxidative Stress in Overweight Hyperlipidemic Subjects Challenged with a High-Fat Meal: A Preliminary Study. Nutr. Res. 2013, 33, 704–710. [Google Scholar] [CrossRef]
- Weryszko-Chmielewska, E.; Matysik-Woźniak, A.; Sulborska, A.; Rejdak, R. Commercially Important Properties of Plants of the Genus Plantago. Acta Agrobot. 2012, 65, 11–20. [Google Scholar] [CrossRef]
- Alvarez-Buylla, E.R.; Benítez, M.; Dávila, E.B.; Chaos, A.; Espinosa-Soto, C.; Padilla-Longoria, P. Gene Regulatory Network Models for Plant Development. Curr. Opin. Plant Biol. 2007, 10, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Karlebach, G.; Shamir, R. Modelling and Analysis of Gene Regulatory Networks. Nat. Rev. Mol. Cell Biol. 2008, 9, 770–780. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Rajewsky, N. The Evolution of Gene Regulation by Transcription Factors and microRNAs. Nat. Rev. Genet. 2007, 8, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Hajheidari, M.; Huang, S.-S.C. Elucidating the Biology of Transcription Factor-DNA Interaction for Accurate Identification of Cis-Regulatory Elements. Curr. Opin. Plant Biol. 2022, 68, 102232. [Google Scholar] [CrossRef]
- Wittkopp, P.J.; Kalay, G. Cis-Regulatory Elements: Molecular Mechanisms and Evolutionary Processes Underlying Divergence. Nat. Rev. Genet. 2012, 13, 59–69. [Google Scholar] [CrossRef]
- Khan, S.-A.; Li, M.-Z.; Wang, S.-M.; Yin, H.-J. Revisiting the Role of Plant Transcription Factors in the Battle against Abiotic Stress. Int. J. Mol. Sci. 2018, 19, 1634. [Google Scholar] [CrossRef]
- Jain, M. Next-Generation Sequencing Technologies for Gene Expression Profiling in Plants. Brief. Funct. Genom. 2012, 11, 63–70. [Google Scholar] [CrossRef]
- Wang, Y.; Cao, Y.; Qin, G. Multifaceted Roles of TCP Transcription Factors in Fate Determination. New Phytol. 2025, 245, 95–101. [Google Scholar] [CrossRef]
- Zhou, H.; Hwarari, D.; Ma, H.; Xu, H.; Yang, L.; Luo, Y. Genomic Survey of TCP Transcription Factors in Plants: Phylogenomics, Evolution and Their Biology. Front. Genet. 2022, 13, 1060546. [Google Scholar] [CrossRef]
- Cubas, P.; Lauter, N.; Doebley, J.; Coen, E. The TCP Domain: A Motif Found in Proteins Regulating Plant Growth and Development. Plant J. 1999, 18, 215–222. [Google Scholar] [CrossRef]
- Nicolas, M.; Cubas, P. TCP Factors: New Kids on the Signaling Block. Curr. Opin. Plant Biol. 2016, 33, 33–41. [Google Scholar] [CrossRef] [PubMed]
- Li, S. The Arabidopsis Thaliana TCP Transcription Factors: A Broadening Horizon beyond Development. Plant Signal Behav. 2015, 10, e1044192. [Google Scholar] [CrossRef] [PubMed]
- Ling, L.; Zhang, W.; An, Y.; Du, B.; Wang, D.; Guo, C. Genome-Wide Analysis of the TCP Transcription Factor Genes in Five Legume Genomes and Their Response to Salt and Drought Stresses. Funct. Integr. Genom. 2020, 20, 537–550. [Google Scholar] [CrossRef]
- Xu, X.; Zhang, X.; Fan, Y.; Zhou, H.; Pu, X. Genome-Wide Identification and Expression Analysis of the TCP Transcription Factor Family and Its Response to Abiotic Stress in Rapeseed (Brassica napus L.). 3 Biotech 2025, 15, 119. [Google Scholar] [CrossRef]
- Martín-Trillo, M.; Cubas, P. TCP Genes: A Family Snapshot Ten Years Later. Trends Plant Sci. 2010, 15, 31–39. [Google Scholar] [CrossRef]
- Cantalapiedra, C.P.; Hernández-Plaza, A.; Letunic, I.; Bork, P.; Huerta-Cepas, J. eggNOG-Mapper v2: Functional Annotation, Orthology Assignments, and Domain Prediction at the Metagenomic Scale. Mol. Biol. Evol. 2021, 38, 5825–5829. [Google Scholar] [CrossRef]
- Huerta-Cepas, J.; Szklarczyk, D.; Heller, D.; Hernández-Plaza, A.; Forslund, S.K.; Cook, H.; Mende, D.R.; Letunic, I.; Rattei, T.; Jensen, L.J.; et al. eggNOG 5.0: A Hierarchical, Functionally and Phylogenetically Annotated Orthology Resource Based on 5090 Organisms and 2502 Viruses. Nucleic Acids Res. 2019, 47, D309–D314. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Gene Ontology Consortium. The Gene Ontology (GO) Database and Informatics Resource. Nucleic Acids Res. 2004, 32, D258–D261. [Google Scholar] [CrossRef]
- Guo, A.-Y.; Chen, X.; Gao, G.; Zhang, H.; Zhu, Q.-H.; Liu, X.-C.; Zhong, Y.-F.; Gu, X.; He, K.; Luo, J. PlantTFDB: A Comprehensive Plant Transcription Factor Database. Nucleic Acids Res. 2008, 36, D966–D969. [Google Scholar] [CrossRef]
- Chen, C.; Wu, Y.; Li, J.; Wang, X.; Zeng, Z.; Xu, J.; Liu, Y.; Feng, J.; Chen, H.; He, Y.; et al. TBtools-II: A “One for All, All for One” Bioinformatics Platform for Biological Big-Data Mining. Mol. Plant 2023, 16, 1733–1742. [Google Scholar] [CrossRef]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef] [PubMed]
- de Moraes Russo, C.A.; Selvatti, A.P. Bootstrap and Rogue Identification Tests for Phylogenetic Analyses. Mol. Biol. Evol. 2018, 35, 2327–2333. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Yu, H.; Lin, S.; Gao, Y. Molecular Characterization of FT and FD Homologs from Eriobotrya Deflexa Nakai Forma Koshunensis. Front. Plant Sci. 2016, 7, 8. [Google Scholar] [CrossRef]
- Sparkes, I.A.; Runions, J.; Kearns, A.; Hawes, C. Rapid, Transient Expression of Fluorescent Fusion Proteins in Tobacco Plants and Generation of Stably Transformed Plants. Nat. Protoc. 2006, 1, 2019–2025. [Google Scholar] [CrossRef]
- Haas, B.J.; Papanicolaou, A.; Yassour, M.; Grabherr, M.; Blood, P.D.; Bowden, J.; Couger, M.B.; Eccles, D.; Li, B.; Lieber, M.; et al. De Novo Transcript Sequence Reconstruction from RNA-Seq Using the Trinity Platform for Reference Generation and Analysis. Nat. Protoc. 2013, 8, 1494–1512. [Google Scholar] [CrossRef]
- Cubas, P.; Vincent, C.; Coen, E. An Epigenetic Mutation Responsible for Natural Variation in Floral Symmetry. Nature 1999, 401, 157–161. [Google Scholar] [CrossRef]
- Liu, M.-M.; Wang, M.-M.; Yang, J.; Wen, J.; Guo, P.-C.; Wu, Y.-W.; Ke, Y.-Z.; Li, P.-F.; Li, J.-N.; Du, H. Evolutionary and Comparative Expression Analyses of TCP Transcription Factor Gene Family in Land Plants. Int. J. Mol. Sci. 2019, 20, 3591. [Google Scholar] [CrossRef]
- Luo, D.; Carpenter, R.; Copsey, L.; Vincent, C.; Clark, J.; Coen, E. Control of Organ Asymmetry in Flowers of Antirrhinum. Cell 1999, 99, 367–376. [Google Scholar] [CrossRef]
- Su, S.; Xiao, W.; Guo, W.; Yao, X.; Xiao, J.; Ye, Z.; Wang, N.; Jiao, K.; Lei, M.; Peng, Q.; et al. The CYCLOIDEA–RADIALIS Module Regulates Petal Shape and Pigmentation, Leading to Bilateral Corolla Symmetry in Torenia fournieri (Linderniaceae). New Phytol. 2017, 215, 1582–1593. [Google Scholar] [CrossRef] [PubMed]
- Gemayel, R.; Chavali, S.; Pougach, K.; Legendre, M.; Zhu, B.; Boeynaems, S.; van der Zande, E.; Gevaert, K.; Rousseau, F.; Schymkowitz, J.; et al. Variable Glutamine-Rich Repeats Modulate Transcription Factor Activity. Mol. Cell 2015, 59, 615–627. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Li, J.; Wen, X.; Zhang, Q.; Dai, S. Genome-Wide Identification of the TCP Gene Family in Chrysanthemum Lavandulifolium and Its Homologs Expression Patterns during Flower Development in Different Chrysanthemum Species. Front. Plant Sci. 2023, 14, 1276123. [Google Scholar] [CrossRef] [PubMed]
- Shad, M.A.; Wu, S.; Rao, M.J.; Luo, X.; Huang, X.; Wu, Y.; Zhou, Y.; Wang, L.; Ma, C.; Hu, L. Evolution and Functional Dynamics of TCP Transcription Factor Gene Family in Passion Fruit (Passiflora edulis). Plants 2024, 13, 2568. [Google Scholar] [CrossRef]
- Yu, Z.; Tian, C.; Guan, Y.; He, J.; Wang, Z.; Wang, L.; Lin, S.; Guan, Z.; Fang, W.; Chen, S.; et al. Expression Analysis of TCP Transcription Factor Family in Autopolyploids of Chrysanthemum Nankingense. Front. Plant Sci. 2022, 13, 860956. [Google Scholar] [CrossRef]
- Liu, C.; Lv, T.; Shen, Y.; Liu, T.; Liu, M.; Hu, J.; Liu, S.; Jiang, Y.; Zhang, M.; Zhao, M.; et al. Genome-Wide Identification and Integrated Analysis of TCP Genes Controlling Ginsenoside Biosynthesis in Panax ginseng. BMC Plant Biol. 2024, 24, 47. [Google Scholar] [CrossRef]

| Item | Value |
|---|---|
| Total number of filtered reads | 70,762,701 |
| Total length of filtered reads (bp) | 47,612,229 |
| Total numbers of unigenes | 59,728 |
| GC percentage (%) | 45.75 |
| N50 value of unigenes (bp) | 1056 |
| N90 value of unigenes (bp) | 369 |
| Mean length of unigenes (bp) | 797.20 |
| Median length of unigenes (bp) | 537.00 |
| Database | Number of Annotated Unigenes | Percentage |
|---|---|---|
| NR | 50,130 | 83.93% |
| COG | 38,632 | 64.68% |
| Pfam | 37,606 | 62.96% |
| Uniprot | 36,934 | 61.84% |
| KEGG | 22,673 | 37.96% |
| GO | 20,990 | 35.14% |
| TF | 1798 | 3.01% |
| Total | 50,684 | 84.86% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lv, X.; Zhang, L.; Hu, Y.; Jing, T.; Liang, Q.; Zhang, Z.; Huang, M.; Yang, H. Cloning and Characterization of 12 TCP Genes in Medicinal Plant Plantago asiatica via De Novo Transcriptome Assembly. Genes 2025, 16, 1021. https://doi.org/10.3390/genes16091021
Lv X, Zhang L, Hu Y, Jing T, Liang Q, Zhang Z, Huang M, Yang H. Cloning and Characterization of 12 TCP Genes in Medicinal Plant Plantago asiatica via De Novo Transcriptome Assembly. Genes. 2025; 16(9):1021. https://doi.org/10.3390/genes16091021
Chicago/Turabian StyleLv, Xingbin, Ling Zhang, Yufang Hu, Tingting Jing, Qi Liang, Zhiyi Zhang, Mingkun Huang, and Hua Yang. 2025. "Cloning and Characterization of 12 TCP Genes in Medicinal Plant Plantago asiatica via De Novo Transcriptome Assembly" Genes 16, no. 9: 1021. https://doi.org/10.3390/genes16091021
APA StyleLv, X., Zhang, L., Hu, Y., Jing, T., Liang, Q., Zhang, Z., Huang, M., & Yang, H. (2025). Cloning and Characterization of 12 TCP Genes in Medicinal Plant Plantago asiatica via De Novo Transcriptome Assembly. Genes, 16(9), 1021. https://doi.org/10.3390/genes16091021






