Thymoquinone Potentially Modulates the Expression of Key Onco- and Tumor Suppressor miRNAs in Prostate and Colon Cancer Cell Lines: Insights from PC3 and HCT-15 Cells
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cell Lines
2.2. Cell Viability Assay
2.3. RNA Extraction and cDNA Synthesis
2.4. Computational Target Prediction of Studied Colon and Prostate Cancer miRNAs and Construction of Network Map
2.5. Experimental Validation and Expression Analysis of miRNAs and Target Genes by RT-qPCR
2.6. Statistical Analysis
3. Results
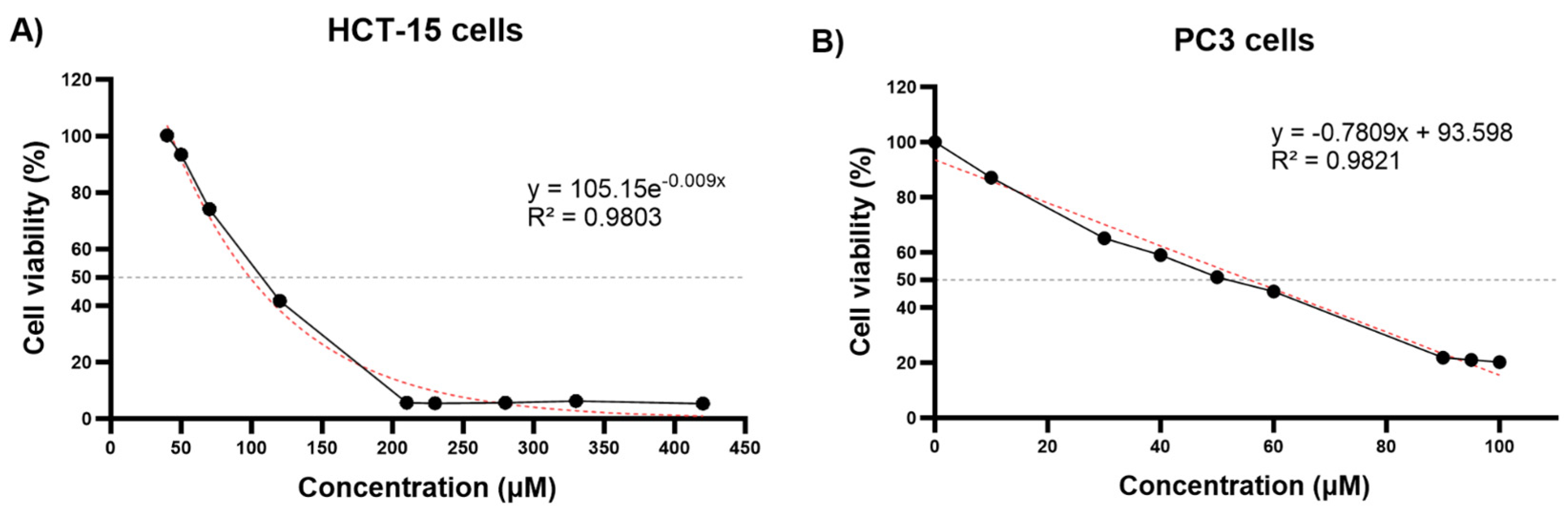
3.1. Cell Viability of HCT-15 and PC3 Cells Following Thymoquinone Treatment and IC50 Value
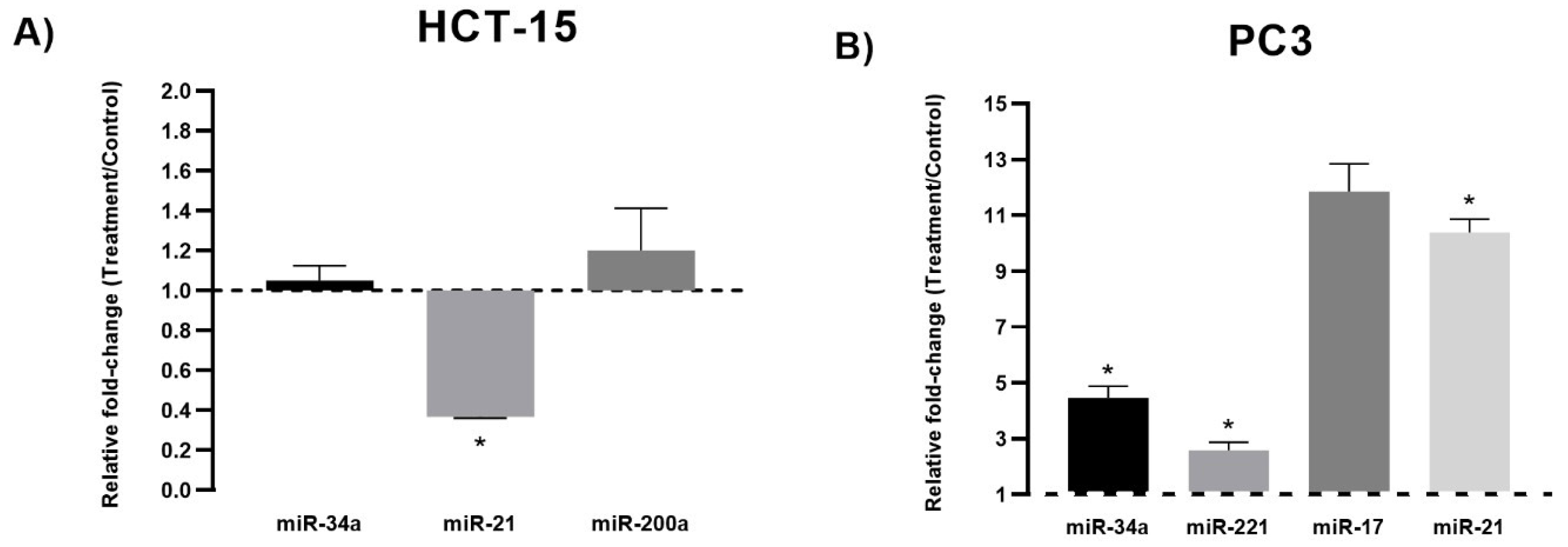
3.2. miRNA Expression Analysis
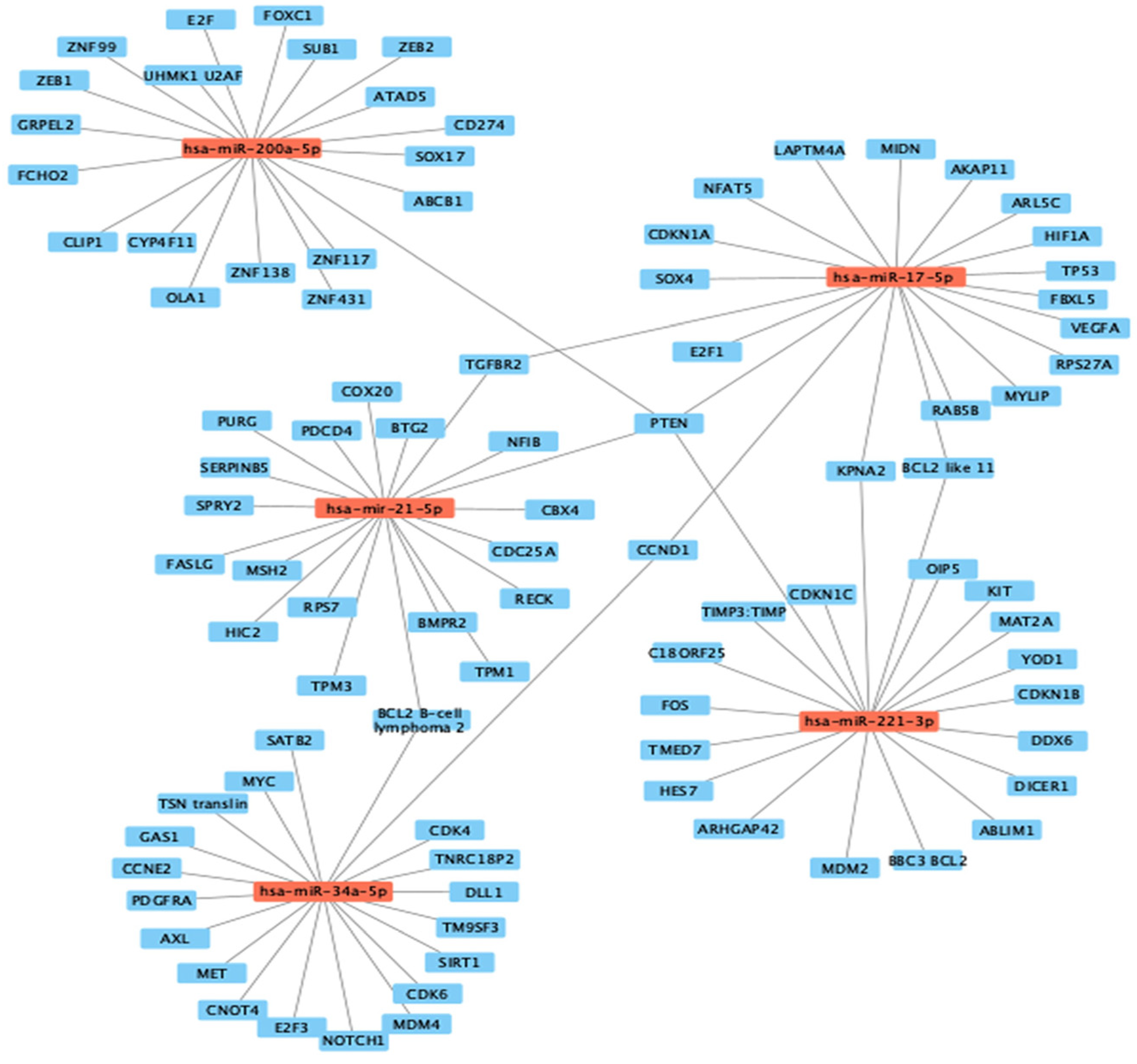
3.3. Computational Target Prediction of Studied Colon and Prostate Cancer miRNAs
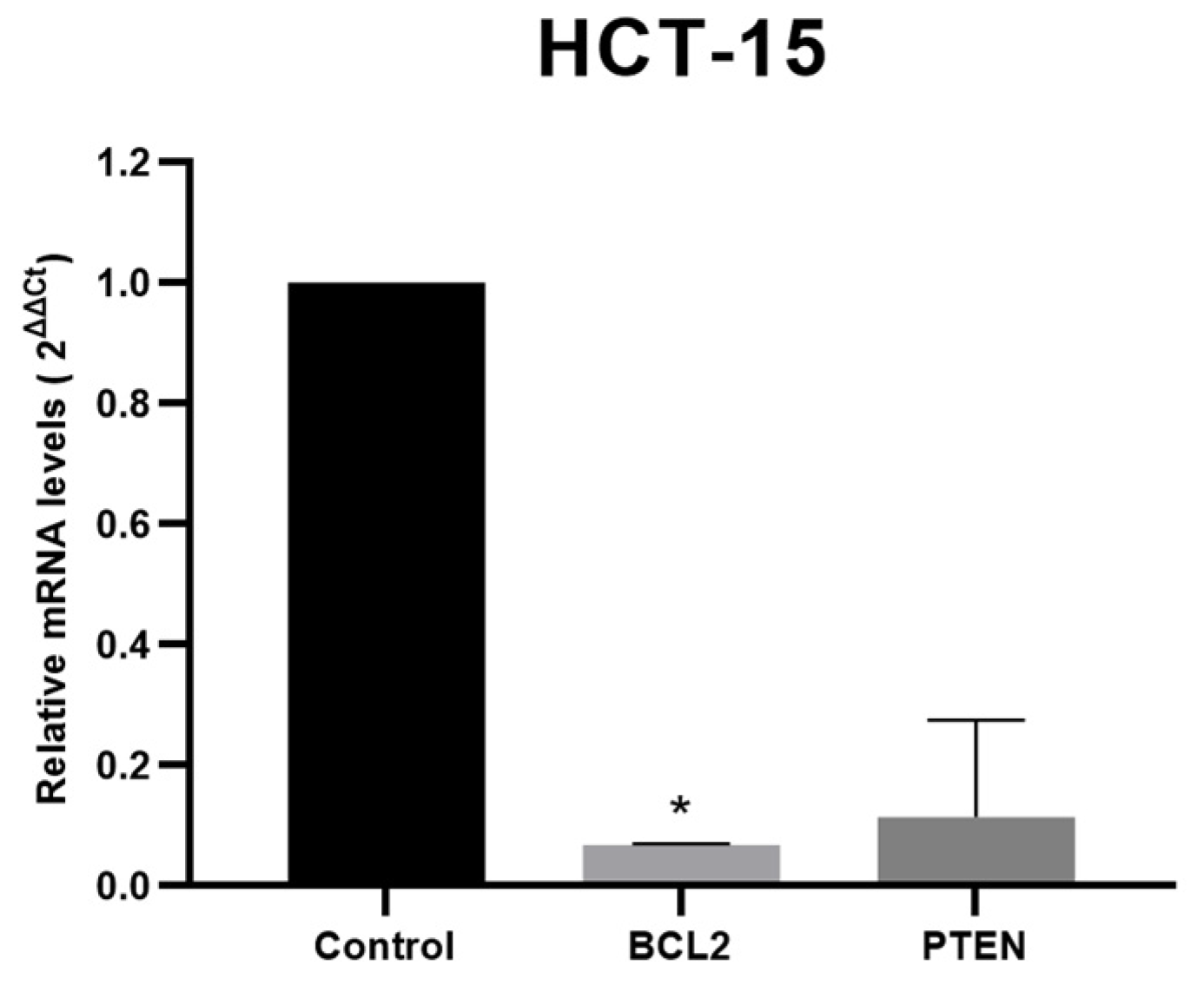
3.4. Target Genes Expression Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chen, S.; Shen, X. Long Noncoding RNAs: Functions and Mechanisms in Colon Cancer. Mol. Cancer 2020, 19, 167. [Google Scholar] [CrossRef] [PubMed]
- Beltran-Ontiveros, S.A.; Fernandez-Galindo, M.A.; Moreno-Ortiz, J.M.; Contreras-Gutierrez, J.A.; Madueña-Molina, J.; Arambula-Meraz, E.; Leal-Leon, E.; Becerril-Camacho, D.M.; Picos-Cardenas, V.J.; Angulo-Rojo, C.; et al. Incidence, Mortality, and Trends of Prostate Cancer in Mexico from 2000 to 2019: Results from the Global Burden of Disease Study 2019. Cancers 2022, 14, 3184. [Google Scholar] [CrossRef] [PubMed]
- Thorpe, C.S.; Bhangoo, R.S.; Anderson, J.D.; Jason Shen, J.; Vargas, C.E. Prostate Cancer. In Principles and Practice of Particle Therapy; Wiley: New York, NY, USA, 2023; pp. 383–410. [Google Scholar] [CrossRef]
- Rawla, P. Epidemiology of Prostate Cancer. World J. Oncol. 2019, 10, 63. [Google Scholar] [CrossRef]
- Sekhoacha, M.; Riet, K.; Motloung, P.; Gumenku, L.; Adegoke, A.; Mashele, S. Prostate Cancer Review: Genetics, Diagnosis, Treatment Options, and Alternative Approaches. Molecules 2022, 27, 5730. [Google Scholar] [CrossRef] [PubMed]
- Xi, Y.; Xu, P. Global Colorectal Cancer Burden in 2020 and Projections to 2040. Transl. Oncol. 2021, 14, 101174. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Sauer, A.G.; Fedewa, S.A.; Butterly, L.F.; Anderson, J.C.; Cercek, A.; Smith, R.A.; Jemal, A. Colorectal Cancer Statistics, 2020. CA Cancer J. Clin. 2020, 70, 145–164. [Google Scholar] [CrossRef] [PubMed]
- El Zarif, T.; Yibirin, M.; De Oliveira-Gomes, D.; Machaalani, M.; Nawfal, R.; Bittar, G.; Bahmad, H.F.; Bitar, N. Overcoming Therapy Resistance in Colon Cancer by Drug Repurposing. Cancers 2022, 14, 2105. [Google Scholar] [CrossRef]
- Hossain, M.S.; Karuniawati, H.; Jairoun, A.A.; Urbi, Z.; Ooi, D.J.; John, A.; Lim, Y.C.; Kaderi Kibria, K.M.; Mohiuddin, A.K.M.; Ming, L.C.; et al. Colorectal Cancer: A Review of Carcinogenesis, Global Epidemiology, Current Challenges, Risk Factors, Preventive and Treatment Strategies. Cancers 2022, 14, 1732. [Google Scholar] [CrossRef]
- Cao, W.; Chen, H.D.; Yu, Y.W.; Li, N.; Chen, W.Q. Changing Profiles of Cancer Burden Worldwide and in China: A Secondary Analysis of the Global Cancer Statistics 2020. Chin. Med. J. 2021, 134, 783–791. [Google Scholar] [CrossRef]
- Sawicki, T.; Ruszkowska, M.; Danielewicz, A.; Niedźwiedzka, E.; Arłukowicz, T.; Przybyłowicz, K.E. A Review of Colorectal Cancer in Terms of Epidemiology, Risk Factors, Development, Symptoms and Diagnosis. Cancers 2021, 13, 2025. [Google Scholar] [CrossRef]
- Cáncer. Available online: https://www.who.int/es/news-room/fact-sheets/detail/cancer (accessed on 12 August 2023).
- Cancer Treatments|Cancer Survivors|CDC. Available online: https://www.cdc.gov/cancer/survivors/patients/treatments.htm (accessed on 12 August 2023).
- Debela, D.T.; Muzazu, S.G.Y.; Heraro, K.D.; Ndalama, M.T.; Mesele, B.W.; Haile, D.C.; Kitui, S.K.; Manyazewal, T. New Approaches and Procedures for Cancer Treatment: Current Perspectives. SAGE Open Med. 2021, 9. [Google Scholar] [CrossRef] [PubMed]
- Ballout, F.; Monzer, A.; Fatfat, M.; El Ouweini, H.; Jaffa, M.A.; Abdel-Samad, R.; Darwiche, N.; Abou-Kheir, W.; Gali-Muhtasib, H. Thymoquinone Induces Apoptosis and DNA Damage in 5-Fluorouracil-Resistant Colorectal Cancer Stem/Progenitor Cells. Oncotarget 2020, 11, 2959–2972. [Google Scholar] [CrossRef] [PubMed]
- Salehi, B.; Fokou, P.V.T.; Yamthe, L.R.T.; Tali, B.T.; Adetunji, C.O.; Rahavian, A.; Mudau, F.N.; Martorell, M.; Setzer, W.N.; Rodrigues, C.F.; et al. Phytochemicals in Prostate Cancer: From Bioactive Molecules to Upcoming Therapeutic Agents. Nutrients 2019, 11, 1483. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Hazra, J.; Pal, K.; Nelson, V.K.; Pal, M. Prostate Cancer: Therapeutic Prospect with Herbal Medicine. Curr. Res. Pharmacol. Drug Discov. 2021, 2, 100034. [Google Scholar] [CrossRef]
- Homayoonfal, M.; Asemi, Z.; Yousefi, B. Targeting MicroRNAs with Thymoquinone: A New Approach for Cancer Therapy. Cell. Mol. Biol. Lett. 2021, 26, 43. [Google Scholar] [CrossRef]
- Khan, M.A.; Tania, M.; Fu, J. Epigenetic Role of Thymoquinone: Impact on Cellular Mechanism and Cancer Therapeutics. Drug Discov. Today 2019, 24, 2315–2322. [Google Scholar] [CrossRef]
- Gomathinayagam, R.; Ha, J.H.; Jayaraman, M.; Song, Y.S.; Isidoro, C.; Dhanasekaran, D.N. Chemopreventive and Anticancer Effects of Thymoquinone: Cellular and Molecular Targets. J. Cancer Prev. 2020, 25, 136. [Google Scholar] [CrossRef]
- Singh, S.K.; Apata, T.; Gordetsky, J.B.; Singh, R. Docetaxel Combined with Thymoquinone Induces Apoptosis in Prostate Cancer Cells via Inhibition of the PI3K/AKT Signaling Pathway. Cancers 2019, 11, 1390. [Google Scholar] [CrossRef]
- Karimian, A.; Majidinia, M.; Moliani, A.; Alemi, F.; Asemi, Z.; Yousefi, B. The Modulatory Effects of Two Bioflavonoids, Quercetin and Thymoquinone on the Expression Levels of DNA Damage and Repair Genes in Human Breast, Lung and Prostate Cancer Cell Lines. Pathol. Res. Pr. 2022, 240, 154143. [Google Scholar] [CrossRef]
- Almajali, B.; Al-Jamal, H.A.N.; Taib, W.R.W.; Ismail, I.; Johan, M.F.; Doolaanea, A.A.; Ibrahim, W.N. Thymoquinone, as a Novel Therapeutic Candidate of Cancers. Pharmaceuticals 2021, 14, 369. [Google Scholar] [CrossRef]
- Ruiz-Manriquez, L.M.; Estrada-Meza, C.; Benavides-Aguilar, J.A.; Ledesma-Pacheco, S.J.; Torres-Copado, A.; Serrano-Cano, F.I.; Bandyopadhyay, A.; Pathak, S.; Chakraborty, S.; Srivastava, A.; et al. Phytochemicals Mediated Modulation of MicroRNAs and Long Non-Coding RNAs in Cancer Prevention and Therapy. Phytother. Res. 2022, 36, 705–729. [Google Scholar] [CrossRef]
- Zhu, J.; Xu, Y.; Liu, S.; Qiao, L.; Sun, J.; Zhao, Q. MicroRNAs Associated with Colon Cancer: New Potential Prognostic Markers and Targets for Therapy. Front. Bioeng. Biotechnol. 2020, 8, 523746. [Google Scholar] [CrossRef]
- Bhardwaj, V.; Mandal, A.K.A. Next-Generation Sequencing Reveals the Role of Epigallocatechin-3-Gallate in Regulating Putative Novel and Known MicroRNAs Which Target the MAPK Pathway in Non-Small-Cell Lung Cancer A549 Cells. Molecules 2019, 24, 368. [Google Scholar] [CrossRef]
- Lv, Y.; Duanmu, J.; Fu, X.; Li, T.; Jiang, Q. Identifying a New MicroRNA Signature as a Prognostic Biomarker in Colon Cancer. PLoS ONE 2020, 15, e0228575. [Google Scholar] [CrossRef] [PubMed]
- Cozar, J.M.; Robles-Fernandez, I.; Rodriguez-Martinez, A.; Puche-Sanz, I.; Vazquez-Alonso, F.; Lorente, J.A.; Martinez-Gonzalez, L.J.; Alvarez-Cubero, M.J. The Role of MiRNAs as Biomarkers in Prostate Cancer. Mutat. Res./Rev. Mutat. Res. 2019, 781, 165–174. [Google Scholar] [CrossRef]
- Cochetti, G.; Rossi de Vermandois, J.A.; Maulà, V.; Giulietti, M.; Cecati, M.; Del Zingaro, M.; Cagnani, R.; Suvieri, C.; Paladini, A.; Mearini, E. Role of MiRNAs in Prostate Cancer: Do We Really Know Everything? Urol. Oncol. Semin. Orig. Investig. 2020, 38, 623–635. [Google Scholar] [CrossRef] [PubMed]
- Menon, A.; Abd-Aziz, N.; Khalid, K.; Poh, C.L.; Naidu, R. MiRNA: A Promising Therapeutic Target in Cancer. Int. J. Mol. Sci. 2022, 23, 11502. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Liu, Z. The Emerging Role of MicroRNAs in Breast Cancer. J. Oncol. 2020, 2020, 9160905. [Google Scholar] [CrossRef]
- Sharma, N.; Baruah, M.M. The MicroRNA Signatures: Aberrantly Expressed MiRNAs in Prostate Cancer. Clin. Transl. Oncol. 2018, 21, 126–144. [Google Scholar] [CrossRef]
- Zhang, L.; Liao, Y.; Tang, L. MicroRNA-34 Family: A Potential Tumor Suppressor and Therapeutic Candidate in Cancer. J. Exp. Clin. Cancer Res. 2019, 38, 53. [Google Scholar] [CrossRef]
- Krajewska, J.B.; Fichna, J.; Mosińska, P. One Step Ahead: MiRNA-34 in Colon Cancer-Future Diagnostic and Therapeutic Tool? Crit. Rev. Oncol. Hematol. 2018, 132, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Hermeking, H. MiR-34a and MiR-34b/c Suppress Intestinal Tumorigenesis. Cancer Res. 2017, 77, 2746–2758. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; Yang, M.; Chen, Y.; Chen, W.; Wang, W. MiR-34a Induces Immunosuppression in Colorectal Carcinoma through Modulating a SIRT1/NF-ΚB/B7-H3/TNF-α Axis. Cancer Immunol. Immunother. 2021, 70, 2247–2259. [Google Scholar] [CrossRef] [PubMed]
- Carter, J.V.; O’Brien, S.J.; Burton, J.F.; Oxford, B.G.; Stephen, V.; Hallion, J.; Bishop, C.; Galbraith, N.J.; Eichenberger, M.R.; Sarojini, H.; et al. The MicroRNA-200 Family Acts as an Oncogene in Colorectal Cancer by Inhibiting the Tumor Suppressor RASSF2. Oncol. Lett. 2019, 18, 3994. [Google Scholar] [CrossRef]
- Inoue, J.; Inazawa, J. Cancer-Associated MiRNAs and Their Therapeutic Potential. J. Hum. Genet. 2021, 66, 937–945. [Google Scholar] [CrossRef]
- Liu, T.; Liu, D.; Guan, S.; Dong, M. Diagnostic Role of Circulating MiR-21 in Colorectal Cancer: A Update Meta-Analysis. Ann. Med. 2021, 53, 87. [Google Scholar] [CrossRef]
- Ourô, S.; Mourato, C.; Ferreira, M.P.; Albergaria, D.; Cardador, A.; Castro, R.E.; Maio, R.; Rodrigues, C.M.P. Evaluation of Tissue and Circulating MiR-21 as Potential Biomarker of Response to Chemoradiotherapy in Rectal Cancer. Pharmaceuticals 2020, 13, 246. [Google Scholar] [CrossRef]
- Ourô, S.; Mourato, C.; Velho, S.; Cardador, A.; Ferreira, M.P.; Albergaria, D.; Castro, R.E.; Maio, R.; Rodrigues, C.M.P. Potential of MiR-21 to Predict Incomplete Response to Chemoradiotherapy in Rectal Adenocarcinoma. Front. Oncol. 2020, 10, 577653. [Google Scholar] [CrossRef]
- Zhao, W.; Ning, L.; Wang, L.; Ouyang, T.; Qi, L.; Yang, R.; Wu, Y. MiR-21 Inhibition Reverses Doxorubicin-Resistance and Inhibits PC3 Human Prostate Cancer Cells Proliferation. Andrologia 2021, 53, e14016. [Google Scholar] [CrossRef]
- Kiener, M.; Chen, L.; Krebs, M.; Grosjean, J.; Klima, I.; Kalogirou, C.; Riedmiller, H.; Kneitz, B.; Thalmann, G.N.; Snaar-Jagalska, E.; et al. MiR-221-5p Regulates Proliferation and Migration in Human Prostate Cancer Cells and Reduces Tumor Growth in vivo. BMC Cancer 2019, 19, 627. [Google Scholar] [CrossRef]
- Kurul, N.O.; Ates, F.; Yilmaz, I.; Narli, G.; Yesildal, C.; Senkul, T. The Association of Let-7c, MiR-21, MiR-145, MiR-182, and MiR-221 with Clinicopathologic Parameters of Prostate Cancer in Patients Diagnosed with Low-Risk Disease. Prostate 2019, 79, 1125–1132. [Google Scholar] [CrossRef]
- Dyson, G.; Farran, B.; Bolton, S.; Craig, D.B.; Dombkowski, A.; Beebe-Dimmer, J.L.; Powell, I.J.; Podgorski, I.; Heilbrun, L.K.; Bock, C.H. The Extrema of Circulating MiR-17 Are Identified as Biomarkers for Aggressive Prostate Cancer. Am. J. Cancer Res. 2018, 8, 2088. [Google Scholar] [PubMed]
- Ottman, R.; Levy, J.; Grizzle, W.E.; Chakrabarti, R. The Other Face of MiR-17-92a Cluster, Exhibiting Tumor Suppressor Effects in Prostate Cancer. Oncotarget 2016, 7, 73739. [Google Scholar] [CrossRef] [PubMed]
- HCT-15–CCL-225|ATCC. Available online: https://www.atcc.org/products/ccl-225 (accessed on 12 August 2023).
- HCT-15 Xenograft Model–Altogen Labs. Available online: https://altogenlabs.com/xenograft-models/colon-cancer-xenograft/hct-15-xenograft-model/ (accessed on 12 August 2023).
- HCT-15 91030712. Available online: https://www.sigmaaldrich.com/MX/es/product/sigma/cb_91030712 (accessed on 12 August 2023).
- Hamdan, N.T.; Jwad, B.A.A.A.A.; Jasim, S.A. Synergistic Anticancer Effects of Phycocyanin and Citrullus Colocynthis Extract against WiDr, HCT-15 and HCT-116 Colon Cancer Cell Lines. Gene Rep. 2021, 22, 100972. [Google Scholar] [CrossRef]
- Lian, G.; Li, F.; Yin, Y.; Chen, L.; Yang, J. Herbal Extract of Artemisia Vulgaris (Mugwort) Induces Antitumor Effects in HCT-15 Human Colon Cancer Cells via Autophagy Induction, Cell Migration Suppression and Loss of Mitochondrial Membrane Potential. J. Buon 2018, 23, 73–78. [Google Scholar]
- PC-3 Cells|Adenocarcinoma|CLS. Available online: https://cls.shop/PC-3/300312 (accessed on 12 August 2023).
- Kamalidehghan, B.; Ghafouri-Fard, S.; Motevaseli, E.; Ahmadipour, F. Inhibition of Human Prostate Cancer (PC-3) Cells and Targeting of PC-3-Derived Prostate Cancer Stem Cells with Koenimbin, a Natural Dietary Compound from Murraya koenigii (L) Spreng. Drug Des. Dev. Ther. 2018, 12, 1119–1133. [Google Scholar] [CrossRef]
- Ghasemi, M.; Turnbull, T.; Sebastian, S.; Kempson, I. The Mtt Assay: Utility, Limitations, Pitfalls, and Interpretation in Bulk and Single-Cell Analysis. Int. J. Mol. Sci. 2021, 22, 12827. [Google Scholar] [CrossRef]
- MiRTarBase: The Experimentally Validated MicroRNA-Target Interactions Database. Available online: https://mirtarbase.cuhk.edu.cn/~miRTarBase/miRTarBase_2022/php/index.php (accessed on 12 August 2023).
- Huang, H.Y.; Lin, Y.C.D.; Cui, S.; Huang, Y.; Tang, Y.; Xu, J.; Bao, J.; Li, Y.; Wen, J.; Zuo, H.; et al. MiRTarBase Update 2022: An Informative Resource for Experimentally Validated MiRNA–Target Interactions. Nucleic Acids Res. 2022, 50, D222–D230. [Google Scholar] [CrossRef]
- Ko, B.; Raamsdonk, J.M. Van RNA Sequencing of Pooled Samples Effectively Identifies Differentially Expressed Genes. Biology 2023, 12, 812. [Google Scholar] [CrossRef]
- Martins, M.Q.; Fortunato, A.S.; Rodrigues, W.P.; Partelli, F.L.; Campostrini, E.; Lidon, F.C.; Damatta, F.M.; Ramalho, J.C.; Ribeiro-Barros, A.I. Selection and Validation of Reference Genes for Accurate RT-QPCR Data Normalization in Coffea Spp. under a Climate Changes Context of Interacting Elevated [CO2] and Temperature. Front. Plant Sci. 2017, 8, 307. [Google Scholar] [CrossRef]
- Al-Hail, H.; Mirza, F.; Al Hashemi, A.; Ahmad, M.N.; Iqbal, M.; Tang, P.; Hasan, M.R. Evaluation of Automated Molecular Tests for the Detection of SARS-CoV-2 in Pooled Nasopharyngeal and Saliva Specimens. J. Clin. Lab. Anal. 2021, 35, e23876. [Google Scholar] [CrossRef] [PubMed]
- Saffari-Chaleshtori, J.; Heidari-Sureshjani, E.; Moradi, F.; Heidarian, E. The Effects of Thymoquinone on Viability, and Anti-Apoptotic Factors (BCL-XL, BCL-2, MCL-1) in Prostate Cancer (PC3) Cells: An In Vitro and Computer-Simulated Environment Study. Adv. Pharm. Bull. 2019, 9, 490. [Google Scholar] [CrossRef] [PubMed]
- Ünal, T.D.; Hamurcu, Z.; Delibaşı, N.; Çınar, V.; Güler, A.; Gökçe, S.; Nurdinov, N.; Ozpolat, B. Thymoquinone Inhibits Proliferation and Migration of MDA-MB-231 Triple Negative Breast Cancer Cells by Suppressing Autophagy, Beclin-1 and LC3. Anticancer. Agents Med. Chem. 2020, 21, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Almajali, B.; Al-Jamal, H.A.N.; Taib, W.R.W.; Ismail, I.; Johan, M.F.; Doolaanea, A.A.; Ibrahim, W.N.; Tajudin, S.A. Thymoquinone Suppresses Cell Proliferation and Enhances Apoptosis of HL60 Leukemia Cells through Re-Expression of JAK/STAT Negative Regulators. Asian Pac. J. Cancer Prev. 2021, 22, 879. [Google Scholar] [CrossRef]
- Zhou, X.; Wang, F.; Wu, H.; Chen, X.; Zhang, Y.; Lin, J.; Cai, Y.; Xiang, J.; He, N.; Hu, Z.; et al. Thymoquinone Suppresses the Proliferation, Migration and Invasiveness through Regulating ROS, Autophagic Flux and MiR-877-5p in Human Bladder Carcinoma Cells. Int. J. Biol. Sci. 2021, 17, 3456. [Google Scholar] [CrossRef] [PubMed]
- Özkoç, M.; Mutlu Altundag, E. Antiproliferative Effect of Thymoquinone on Human Colon Cancer Cells: Is It Dependent on Glycolytic Pathway? Acibadem Univ. Saglik. Bilim. Derg. 2023, 14, 103–107. [Google Scholar] [CrossRef]
- Tokay, E. Determination of Cytotoxic Effect and Expression Analyses of Apoptotic and Autophagic Related Genes in Thymoquinone-Treated Colon Cancer Cells. Sak. Univ. J. Sci. 2020, 24, 189–196. [Google Scholar] [CrossRef]
- Sayed, S.R.E.; Cristante, J.; Guyon, L.; Denis, J.; Chabre, O.; Cherradi, N. MicroRNA Therapeutics in Cancer: Current Advances and Challenges. Cancers 2021, 13, 2680. [Google Scholar] [CrossRef]
- Li, S.; Wei, X.; He, J.; Cao, Q.; Du, D.; Zhan, X.; Zeng, Y.; Yuan, S.; Sun, L. The Comprehensive Landscape of MiR-34a in Cancer Research. Cancer Metastasis Rev. 2021, 40, 925–948. [Google Scholar] [CrossRef]
- Slabáková, E.; Culig, Z.; Remšík, J.; Souček, K. Alternative Mechanisms of MiR-34a Regulation in Cancer. Cell Death Dis. 2017, 8, e3100. [Google Scholar] [CrossRef]
- Cancer Cell Culture Basics Handbook. Available online: http://assets.thermofisher.com/TFS-Assets/BID/Handbooks/cancer-cell-culture-basics-handbook.pdf (accessed on 1 November 2022).
- Alaufi, O.M.; Noorwali, A.; Zahran, F.; Al-Abd, A.M.; Al-Attas, S. Cytotoxicity of Thymoquinone Alone or in Combination with Cisplatin (CDDP) against Oral Squamous Cell Carcinoma in vitro. Sci. Rep. 2017, 7, 13131. [Google Scholar] [CrossRef]
- Baran, B.; Ozupek, N.-M.; Calibasi-Kocal, G.; Basbinar, Y. MicroRNAs (MiRNAs) in Colorectal Cancer. Oncog. Carcinog. 2018. [Google Scholar] [CrossRef]
- Tian, J.; Cui, P.; Li, Y.; Yao, X.; Wu, X.; Wang, Z.; Li, C. LINC02418 Promotes Colon Cancer Progression by Suppressing Apoptosis via Interaction with MiR-34b-5p/BCL2 Axis. Cancer Cell Int. 2020, 20, 460. [Google Scholar] [CrossRef]
- Ge, X.; Gao, J.; Sun, Q.W.; Wang, C.X.; Deng, W.; Mao, G.Y.; Li, H.Q.; Guo, S.S.; Cheng, J.; Wu, Y.N.; et al. MiR-34a Inhibits the Proliferation, Migration, and Invasion of Oral Squamous Cell Carcinoma by Directly Targeting SATB2. J. Cell. Physiol. 2020, 235, 4856–4864. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.C.; Xu, H.; Wang, X.; Wang, T.; Wu, J. MiR-34a Increases Cisplatin Sensitivity of Osteosarcoma Cells in vitro through up-Regulation of c-Myc and Bim Signal. Cancer Biomark. 2018, 21, 135–144. [Google Scholar] [CrossRef] [PubMed]
- Shaban, N.Z.; Ibrahim, N.K.; Saada, H.N.; El-Rashidy, F.H.; Shaaban, H.M.; Farrag, M.A.; ElDebaiky, K.; Kodous, A.S. MiR-34a and MiR-21 as Biomarkers in Evaluating the Response of Chemo-Radiotherapy in Egyptian Breast Cancer Patients. J. Radiat. Res. Appl. Sci. 2022, 15, 285–292. [Google Scholar] [CrossRef]
- Lim, D.; Cho, J.G.; Yun, E.; Lee, A.; Ryu, H.Y.; Lee, Y.J.; Yoon, S.; Chang, W.; Lee, M.S.; Kwon, B.S.; et al. MicroRNA 34a–AXL Axis Regulates Vasculogenic Mimicry Formation in Breast Cancer Cells. Genes 2021, 12, 9. [Google Scholar] [CrossRef]
- Yin, X.; Liu, C.; Liu, Q.F. Regulation Effect of MiR-34a Expression on Radiosensitivity of Lung Adenocarcinoma Cells by Targeting Bcl-2 and CDK4/6 Signaling Pathways Regulation Effect of MiR-34a Expression on Radiosensi-Tivity of Lung Adenocarcinoma Cells by Tar-Geting Bcl-2 and CDK4/6 Signaling Pathways. J. Cancer Ther. 2022, 13, 187–198. [Google Scholar] [CrossRef]
- Di, Z.; Di, M.; Fu, W.; Tang, Q.; Liu, Y.; Lei, P.; Gu, X.; Liu, T.; Sun, M. Integrated Analysis Identifies a Nine-MicroRNA Signature Biomarker for Diagnosis and Prognosis in Colorectal Cancer. Front. Genet. 2020, 11, 503476. [Google Scholar] [CrossRef]
- Zhang, L.; Cai, Q.Y.; Liu, J.; Peng, J.; Chen, Y.Q.; Sferra, T.J.; Lin, J.M. Ursolic Acid Suppresses the Invasive Potential of Colorectal Cancer Cells by Regulating the TGF-Β1/ZEB1/MiR-200c Signaling Pathway. Oncol. Lett. 2019, 18, 3274. [Google Scholar] [CrossRef]
- Klicka, K.; Grzywa, T.M.; Mielniczuk, A.; Klinke, A.; Włodarski, P.K. The Role of MiR-200 Family in the Regulation of Hallmarks of Cancer. Front. Oncol. 2022, 12, 965231. [Google Scholar] [CrossRef] [PubMed]
- Di Martino, M.T.; Arbitrio, M.; Caracciolo, D.; Cordua, A.; Cuomo, O.; Grillone, K.; Riillo, C.; Caridà, G.; Scionti, F.; Labanca, C.; et al. MiR-221/222 as Biomarkers and Targets for Therapeutic Intervention on Cancer and Other Diseases: A Systematic Review. Mol. Ther. Nucleic Acids 2022, 27, 1191–1224. [Google Scholar] [CrossRef] [PubMed]
- Ni, L.; Xu, J.; Zhao, F.; Dai, X.; Tao, J.; Pan, J.; Shi, A.; Shen, Z.; Su, C.; Zhang, Y. MiR-221-3p-Mediated Downregulation of MDM2 Reverses the Paclitaxel Resistance of Non-Small Cell Lung Cancer in Vitro and in Vivo. Eur. J. Pharmacol. 2021, 899, 174054. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Xu, Y.; Wang, J.; Zhao, S.; Li, J.; Huang, X.; Xu, H.; Zhang, X.; Suo, S.; Lv, Y.; et al. MiR-221-3p Delivered by BMMSC-Derived Microvesicles Promotes the Development of Acute Myelocytic Leukemia. Front. Bioeng. Biotechnol. 2020, 8, 521892. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.H.; Zhao, Z.X.; Dai, J.; Geng, D.C.; Xu, Y.Z. MicroRNA-221 Regulates Osteosarcoma Cell Proliferation, Apoptosis, Migration, and Invasion by Targeting CDKN1B/P27. J. Cell. Biochem. 2019, 120, 4665–4674. [Google Scholar] [CrossRef]
- Jiang, J.; Chang, W.; Fu, Y.; Gao, Y.; Zhao, C.; Zhang, X.; Zhang, S. SAV1, Regulated by MicroRNA-21, Suppresses Tumor Growth in Colorectal Cancer. Biochem. Cell Biol. 2019, 97, 91–99. [Google Scholar] [CrossRef]
- Hill, M.; Tran, N. Global MiRNA to MiRNA Interactions: Impacts for MiR-21. Trends Cell Biol. 2021, 31, 3–5. [Google Scholar] [CrossRef]
- Najjary, S.; Mohammadzadeh, R.; Mokhtarzadeh, A.; Mohammadi, A.; Kojabad, A.B.; Baradaran, B. Role of MiR-21 as an Authentic Oncogene in Mediating Drug Resistance in Breast Cancer. Gene 2020, 738, 144453. [Google Scholar] [CrossRef]
- Jiang, R.; Chen, X.; Ge, S.; Wang, Q.; Liu, Y.; Chen, H.; Xu, J.; Wu, J. MiR-21-5p Induces Pyroptosis in Colorectal Cancer via TGFBI. Front. Oncol. 2021, 10, 610545. [Google Scholar] [CrossRef]
- Liang, G.; Zhu, Y.; Ali, D.J.; Tian, T.; Xu, H.; Si, K.; Sun, B.; Chen, B.; Xiao, Z. Engineered Exosomes for Targeted Co-Delivery of MiR-21 Inhibitor and Chemotherapeutics to Reverse Drug Resistance in Colon Cancer. J. Nanobiotechnol. 2020, 18, 10. [Google Scholar] [CrossRef]
- Ni, H.; Han, Y.; Jin, X. Celastrol Inhibits Colon Cancer Cell Proliferation by Downregulating MiR-21 and PI3K/AKT/GSK-3β Pathway. Int. J. Clin. Exp. Patho 2019, 12, 808–816. [Google Scholar]
- Gomes, S.; Baltazar, F.; Silva, E.; Preto, A. Microbiota-Derived Short-Chain Fatty Acids: New Road in Colorectal Cancer Therapy. Pharmaceutics 2022, 14, 2359. [Google Scholar] [CrossRef] [PubMed]
- Heshmati, M.; Hasani-, N.; Abad, R.; Pashapour, S. Evaluating the Effects of Silymarin on Expressing SBDSP1 and CASC11 Genes in HCT116 Colon Cancer Cells. J. Kermanshah Univ. Med. Sci. 2022, 26. [Google Scholar] [CrossRef]
- Salim, L.Z.A.; Mohan, S.; Othman, R.; Abdelwahab, S.I.; Kamalidehghan, B.; Sheikh, B.Y.; Ibrahim, M.Y. Thymoquinone Induces Mitochondria-Mediated Apoptosis in Acute Lymphoblastic Leukaemia in Vitro. Molecules 2013, 18, 11219–11240. [Google Scholar] [CrossRef] [PubMed]
- Bautista-Sánchez, D.; Arriaga-Canon, C.; Pedroza-Torres, A.; De La Rosa-Velázquez, I.A.; González-Barrios, R.; Contreras-Espinosa, L.; Montiel-Manríquez, R.; Castro-Hernández, C.; Fragoso-Ontiveros, V.; Álvarez-Gómez, R.M.; et al. The Promising Role of MiR-21 as a Cancer Biomarker and Its Importance in RNA-Based Therapeutics. Mol. Ther. Nucleic Acids 2020, 20, 409–420. [Google Scholar] [CrossRef]
- Shakeri, A.; Ghanbari, M.; Tasbandi, A.; Sahebkar, A. Regulation of MicroRNA-21 Expression by Natural Products in Cancer. Phytother. Res. 2021, 35, 3732–3746. [Google Scholar] [CrossRef] [PubMed]
- Shen, Z.; Xu, X.; Lv, L.; Dai, H.; Chen, J.; Chen, B. MiR-21 Overexpression Promotes Esophageal Squamous Cell Carcinoma Invasion and Migration by Repressing Tropomyosin 1. Gastroenterol. Res. Pr. 2020, 2020, 6478653. [Google Scholar] [CrossRef]
- Bica-Pop, C.; Cojocneanu-Petric, R.; Magdo, L.; Raduly, L.; Gulei, D.; Berindan-Neagoe, I. Overview upon MiR-21 in Lung Cancer: Focus on NSCLC. Cell. Mol. Life Sci. 2018, 75, 3539–3551. [Google Scholar] [CrossRef]
- Huang, H.; Li, X.; Zhang, X.; Li, Z.; Han, D.; Gao, W.; Liu, L.; Peng, C.; Zhu, H.; Yu, X. DSCR9/MiR-21-5p Axis Inhibits Pancreatic Cancer Proliferation and Resistance to Gemcitabine via BTG2 Signaling: DSCR9/MiR-21-5p/BTG2 Regulates Pancreatic Cancer. Acta Biochim. Biophys. Sin. 2022, 54, 1775. [Google Scholar] [CrossRef]
- Zhao, Q.; Chen, S.; Zhu, Z.; Yu, L.; Ren, Y.; Jiang, M.; Weng, J.; Li, B. MiR-21 Promotes EGF-Induced Pancreatic Cancer Cell Proliferation by Targeting Spry2. Cell Death Dis. 2018, 9, 1157. [Google Scholar] [CrossRef]
- Folini, M.; Gandellini, P.; Longoni, N.; Profumo, V.; Callari, M.; Pennati, M.; Colecchia, M.; Supino, R.; Veneroni, S.; Salvioni, R.; et al. MiR-21: An Oncomir on Strike in Prostate Cancer. Mol. Cancer 2010, 9, 12. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Osorio-Pérez, S.M.; Estrada-Meza, C.; Ruiz-Manriquez, L.M.; Arvizu-Espinosa, M.G.; Srivastava, A.; Sharma, A.; Paul, S. Thymoquinone Potentially Modulates the Expression of Key Onco- and Tumor Suppressor miRNAs in Prostate and Colon Cancer Cell Lines: Insights from PC3 and HCT-15 Cells. Genes 2023, 14, 1730. https://doi.org/10.3390/genes14091730
Osorio-Pérez SM, Estrada-Meza C, Ruiz-Manriquez LM, Arvizu-Espinosa MG, Srivastava A, Sharma A, Paul S. Thymoquinone Potentially Modulates the Expression of Key Onco- and Tumor Suppressor miRNAs in Prostate and Colon Cancer Cell Lines: Insights from PC3 and HCT-15 Cells. Genes. 2023; 14(9):1730. https://doi.org/10.3390/genes14091730
Chicago/Turabian StyleOsorio-Pérez, Sofía Madeline, Carolina Estrada-Meza, Luis M. Ruiz-Manriquez, María Goretti Arvizu-Espinosa, Aashish Srivastava, Ashutosh Sharma, and Sujay Paul. 2023. "Thymoquinone Potentially Modulates the Expression of Key Onco- and Tumor Suppressor miRNAs in Prostate and Colon Cancer Cell Lines: Insights from PC3 and HCT-15 Cells" Genes 14, no. 9: 1730. https://doi.org/10.3390/genes14091730
APA StyleOsorio-Pérez, S. M., Estrada-Meza, C., Ruiz-Manriquez, L. M., Arvizu-Espinosa, M. G., Srivastava, A., Sharma, A., & Paul, S. (2023). Thymoquinone Potentially Modulates the Expression of Key Onco- and Tumor Suppressor miRNAs in Prostate and Colon Cancer Cell Lines: Insights from PC3 and HCT-15 Cells. Genes, 14(9), 1730. https://doi.org/10.3390/genes14091730







