Identification of a Novel lncRNA LNC_001186 and Its Effects on CPB2 Toxin-Induced Apoptosis of IPEC-J2 Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal Sample Collection
2.2. IPEC-J2 Cells Culture and Damage Model Establishment
2.3. RNA Isolation and cDNA Synthesis
2.4. Rapid Amplification of cDNA Ends (RACE)
2.5. Cytoplasmic/Nuclear RNA Isolation Experiment
2.6. RNA-FISH
2.7. Bioinformatics Analysis
2.8. Overexpression Vector Construction and siRNA Synthesis
2.9. Cell Transfection
2.10. CCK-8 Assay
2.11. Flow Cytometry
2.12. LDH Activity Test
2.13. RT-qPCR
2.14. Western Blot
2.15. Statistical Analysis
3. Results
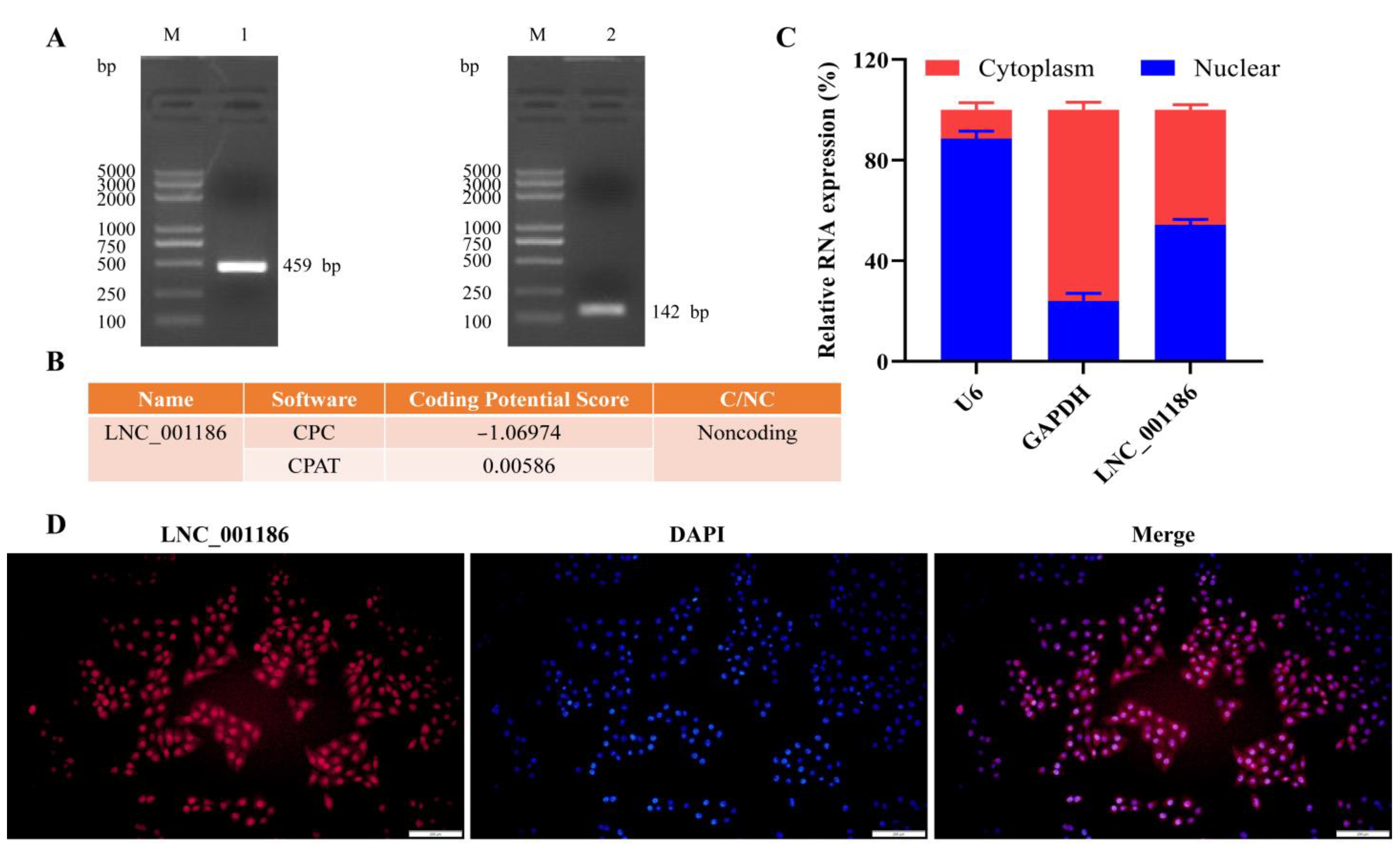
3.1. Pig LNC_001186 Characteristics
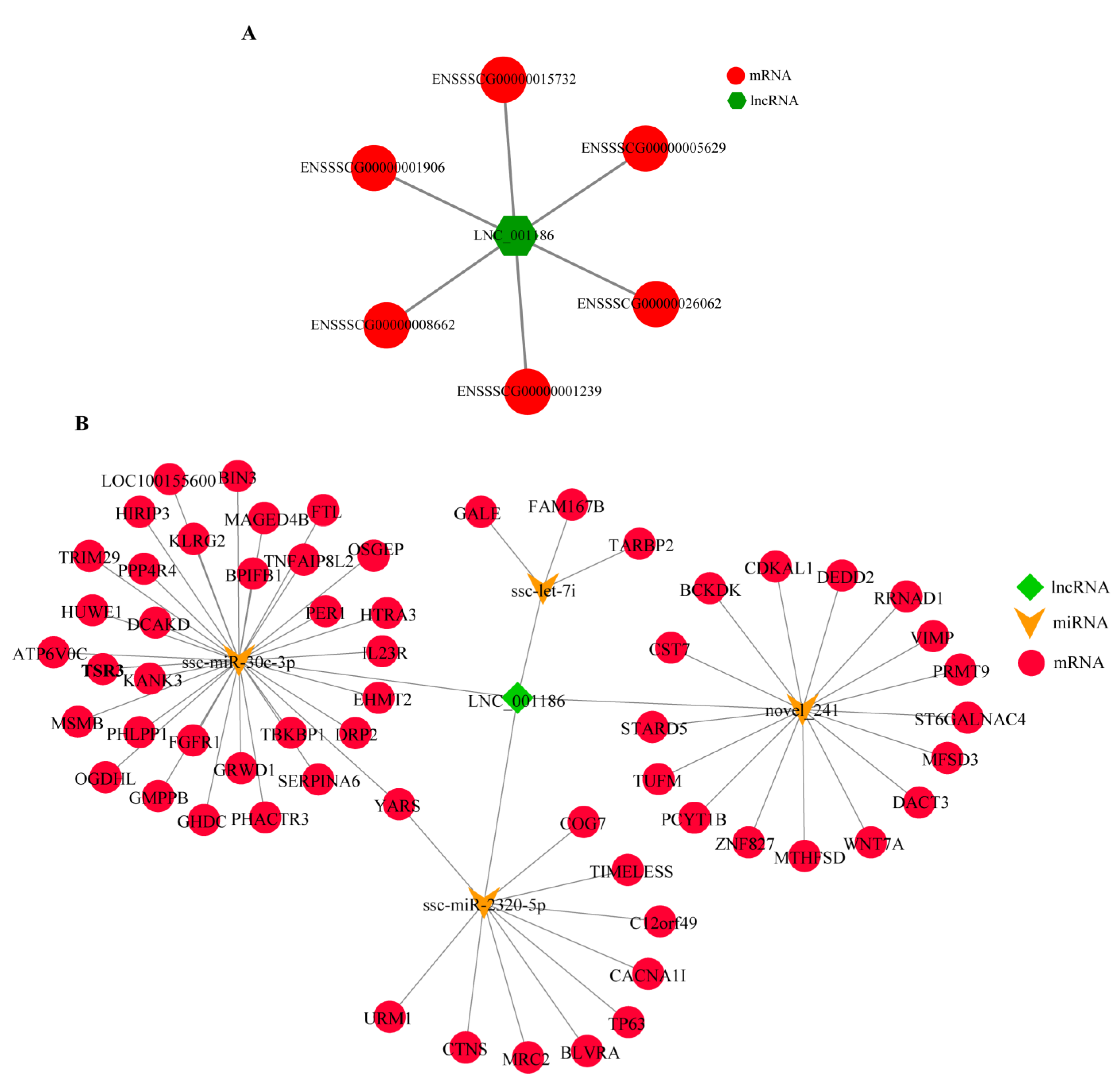
3.2. LNC_001186 Regulatory Network Construction
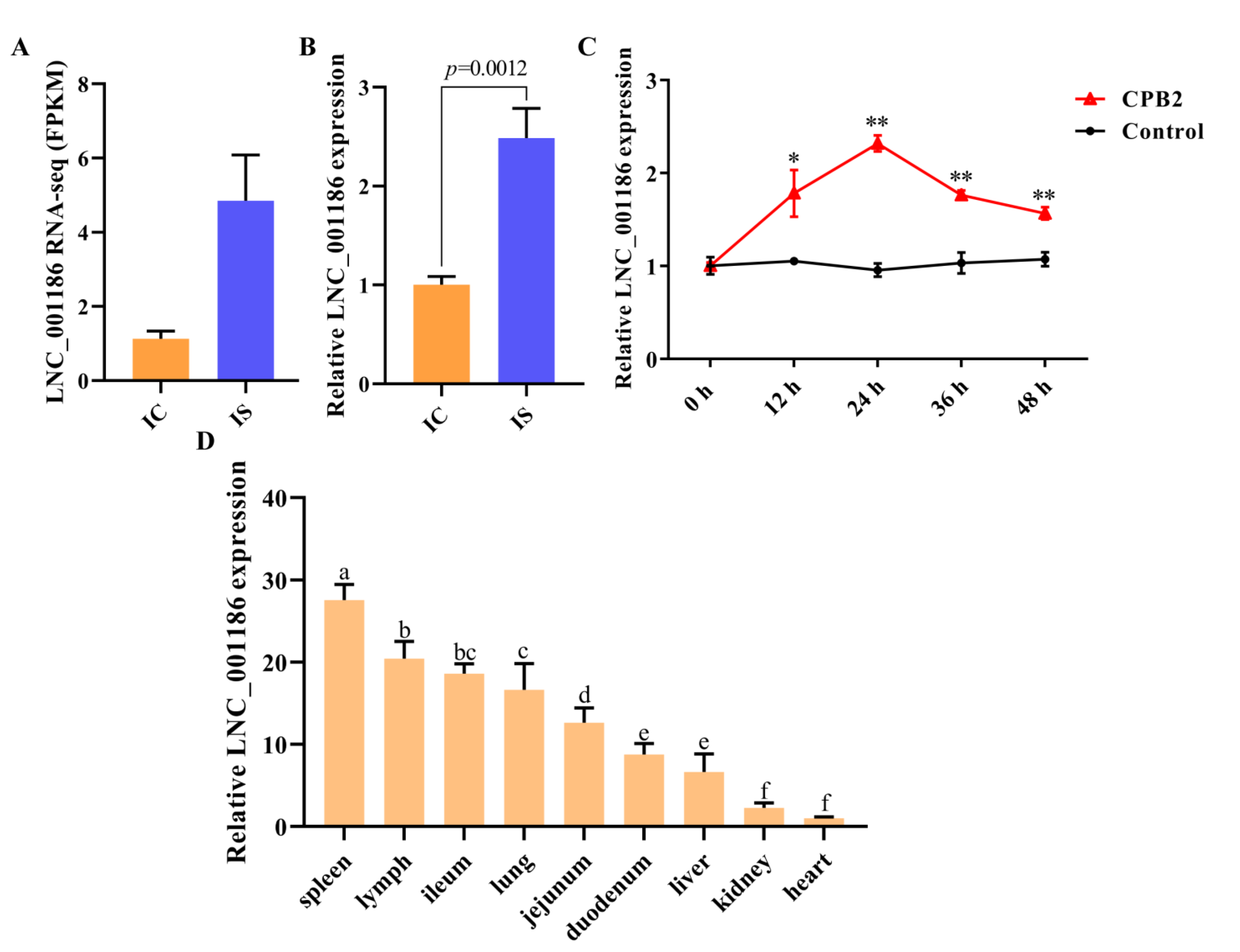
3.3. LNC_001186 Expression Pattern Analysis
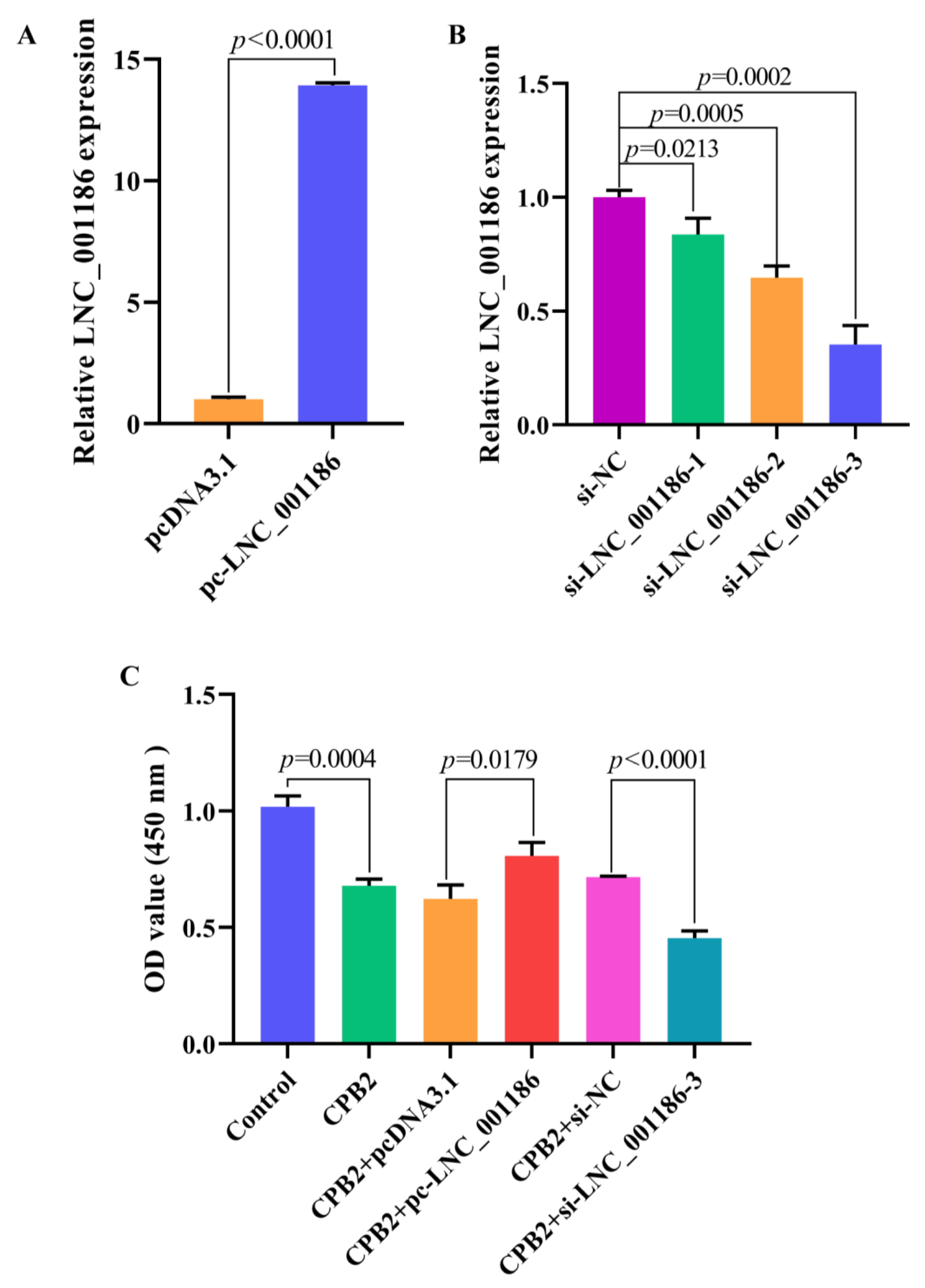
3.4. LNC_001186 Improved CPB2 Toxin-Treated IPEC-J2 Cell Viability
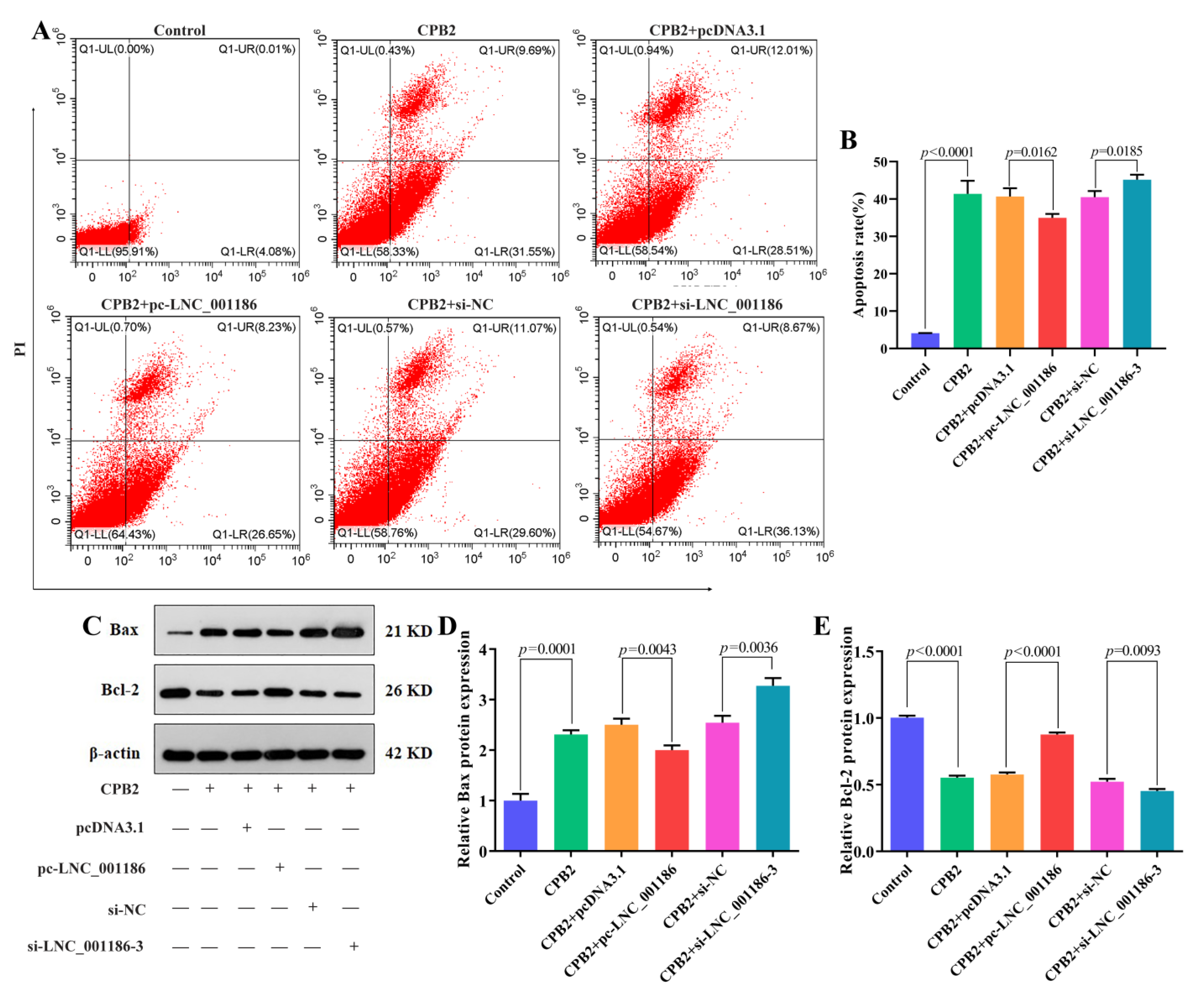
3.5. LNC_001186 Suppressed CPB2 Toxin-Induced IPEC-J2 Cell Apoptosis
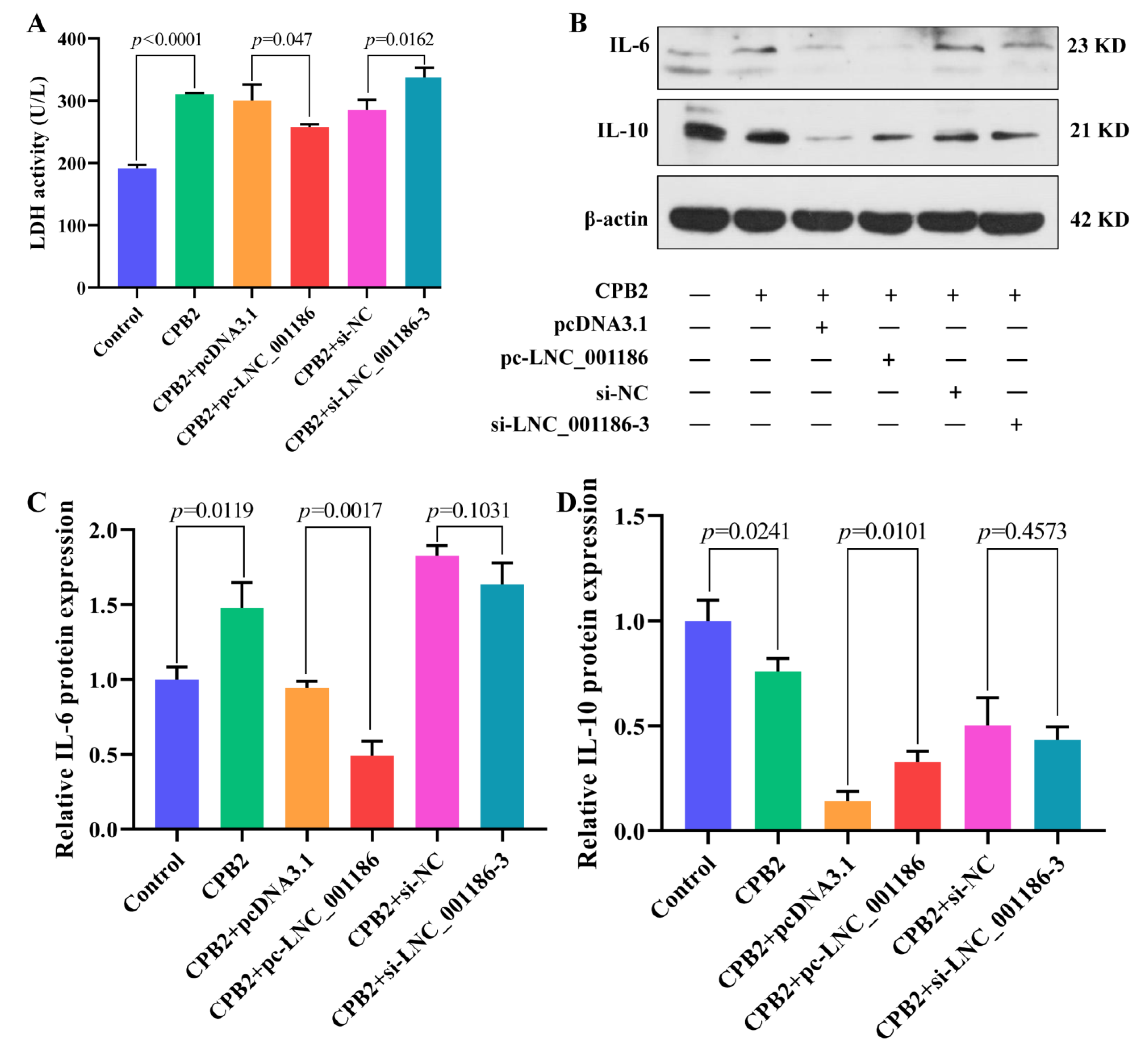
3.6. LNC_001186 Alleviated CPB2 Toxin-Induced IPEC-J2 Cell Cytotoxicity and Inflammation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Huang, X.; Yang, Q.; Yuan, J.; Liu, L.; Sun, W.; Jiang, Y.; Zhao, S.; Zhang, S.; Huang, W.; Gun, S. Effect of Genetic Diversity in Swine Leukocyte Antigen-DRA Gene on Piglet Diarrhea. Genes 2016, 7, 36. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z.; Cai, L.; Huang, X.; Sun, W.; Li, S.; Wang, P.; Yang, Q.; Jiang, T.; Gun, S. Histological and Comparative Transcriptome Analyses Provide Insights into Small Intestine Health in Diarrheal Piglets after Infection with Clostridium Perfringens Type C. Animals 2019, 9, 269. [Google Scholar] [CrossRef] [PubMed]
- Posthaus, H.; Kittl, S.; Tarek, B.; Bruggisser, J. Clostridium perfringens type C necrotic enteritis in pigs: Diagnosis, pathogenesis, and prevention. J. Vet. Diagn. Investig. 2020, 32, 203–212. [Google Scholar] [CrossRef] [PubMed]
- Petit, L.; Gibert, M.; Popoff, M.R. Clostridium perfringens: Toxinotype and genotype. Trends Microbiol. 1999, 7, 104–110. [Google Scholar] [CrossRef] [PubMed]
- Rood, J.I.; Adams, V.; Lacey, J.; Lyras, D.; McClane, B.A.; Melville, S.B.; Moore, R.J.; Popoff, M.R.; Sarker, M.R.; Songer, J.G.; et al. Expansion of the Clostridium perfringens toxin-based typing scheme. Anaerobe 2018, 53, 5–10. [Google Scholar] [CrossRef]
- Gibert, M.; Jolivet-Reynaud, C.; Popoff, M.R. Beta2 toxin, a novel toxin produced by Clostridium perfringens. Gene 1997, 203, 65–73. [Google Scholar] [CrossRef]
- Uzal, F.A.; Freedman, J.C.; Shrestha, A.; Theoret, J.R.; Garcia, J.; Awad, M.M.; Adams, V.; Moore, R.J.; Rood, J.I.; McClane, B.A. Towards an understanding of the role of Clostridium perfringens toxins in human and animal disease. Future Microbiol. 2014, 9, 361–377. [Google Scholar] [CrossRef]
- Fisher, D.J.; Miyamoto, K.; Harrison, B.; Akimoto, S.; Sarker, M.R.; McClane, B.A. Association of beta2 toxin production with Clostridium perfringens type A human gastrointestinal disease isolates carrying a plasmid enterotoxin gene. Mol. Microbiol. 2005, 56, 747–762. [Google Scholar] [CrossRef]
- Gao, X.; Yang, Q.; Huang, X.; Yan, Z.; Zhang, S.; Luo, R.; Wang, P.; Wang, W.; Xie, K.; Jiang, T.; et al. Effects of Clostridium perfringens beta2 toxin on apoptosis, inflammation, and barrier function of intestinal porcine epithelial cells. Microb. Pathog. 2020, 147, 104379. [Google Scholar] [CrossRef]
- Luo, R.; Yang, Q.; Huang, X.; Yan, Z.; Gao, X.; Wang, W.; Xie, K.; Wang, P.; Gun, S. Clostridium perfringens beta2 toxin induced in vitro oxidative damage and its toxic assessment in porcine small intestinal epithelial cell lines. Gene 2020, 759, 144999. [Google Scholar] [CrossRef]
- Ulitsky, I. Interactions between short and long noncoding RNAs. FEBS Lett. 2018, 592, 2874–2883. [Google Scholar] [CrossRef] [PubMed]
- Hansji, H.; Leung, E.Y.; Baguley, B.C.; Finlay, G.J.; Askarian-Amiri, M.E. Keeping abreast with long non-coding RNAs in mammary gland development and breast cancer. Front. Genet. 2014, 5, 379. [Google Scholar] [CrossRef] [PubMed]
- Kazimierczyk, M.; Kasprowicz, M.K.; Kasprzyk, M.E.; Wrzesinski, J. Human Long Noncoding RNA Interactome: Detection, Characterization and Function. Int. J. Mol. Sci. 2020, 21, 1027. [Google Scholar] [CrossRef] [PubMed]
- Blokhin, I.; Khorkova, O.; Hsiao, J.; Wahlestedt, C. Developments in lncRNA drug discovery: Where are we heading? Expert Opin. Drug Discov. 2018, 13, 837–849. [Google Scholar] [CrossRef]
- Bhardwaj, V.; Tan, Y.Q.; Wu, M.M.; Ma, L.; Zhu, T.; Lobie, P.E.; Pandey, V. Long non-coding RNAs in recurrent ovarian cancer: Theranostic perspectives. Cancer Lett. 2021, 502, 97–107. [Google Scholar] [CrossRef]
- Wu, Z.; Fan, H.; Jin, J.; Gao, S.; Huang, R.; Wu, S.; Bao, W. Insight into mechanisms of pig lncRNA FUT3-AS1 regulating E. coli F18-bacterial diarrhea. PLoS Pathog. 2022, 18, e1010584. [Google Scholar] [CrossRef]
- Gao, X.; Sun, X.; Yao, X.; Wang, Y.; Li, Y.; Jiang, X.; Han, Y.; Zhong, L.; Wang, L.; Song, H.; et al. Downregulation of the Long Noncoding RNA IALNCR Targeting MAPK8/JNK1 Promotes Apoptosis and Antagonizes Bovine Viral Diarrhea Virus Replication in Host Cells. J. Virol. 2022, 96, e0111322. [Google Scholar] [CrossRef]
- Tanuj, G.N.; Khan, O.; Malla, W.A.; Rajak, K.K.; Chandrashekar, S.; Kumar, A.; Dhara, S.K.; Gupta, P.K.; Mishra, B.P.; Dutt, T.; et al. Integrated analysis of long-noncoding RNA and circular RNA expression in Peste-des-Petits-Ruminants Virus (PPRV) infected marmoset B lymphocyte (B95a) cells. Microb. Pathog. 2022, 170, 105702. [Google Scholar] [CrossRef]
- Zhang, J.; Gan, L.; Sun, P.; Wang, J.; Li, D.; Cao, Y.; Fu, Y.; Li, P.; Bai, X.; Li, K.; et al. The long non-coding RNA LNC_000397 negatively regulates PRRSV replication through induction of interferon-stimulated genes. Virol. J. 2022, 19, 40. [Google Scholar] [CrossRef]
- Huang, X.; Sun, W.; Yan, Z.; Shi, H.; Yang, Q.; Wang, P.; Li, S.; Liu, L.; Zhao, S.; Gun, S. Integrative Analyses of Long Non-coding RNA and mRNA Involved in Piglet Ileum Immune Response to Clostridium perfringens Type C Infection. Front. Cell. Infect. Microbiol. 2019, 9, 130. [Google Scholar] [CrossRef]
- Huang, X.Y.; Sun, W.Y.; Yan, Z.Q.; Shi, H.R.; Yang, Q.L.; Wang, P.F.; Li, S.G.; Liu, L.X.; Zhao, S.G.; Gun, S.B. Novel Insights reveal Anti-microbial Gene Regulation of Piglet Intestine Immune in response to Clostridium perfringens Infection. Sci. Rep. 2019, 9, 1963. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Hu, J.; Liu, S.; Chen, B.; Xiao, M.; Li, Y.; Liao, Y.; Rai, K.R.; Zhao, Z.; Ouyang, J.; et al. RDUR, a lncRNA, Promotes Innate Antiviral Responses and Provides Feedback Control of NF-κB Activation. Front. Immunol. 2021, 12, 672165. [Google Scholar] [CrossRef]
- Yang, Y.; Li, M.; Ma, Y.; Ye, W.; Si, Y.; Zheng, X.; Liu, H.; Cheng, L.; Zhang, L.; Zhang, H.; et al. LncRNA NEAT1 Potentiates SREBP2 Activity to Promote Inflammatory Macrophage Activation and Limit Hantaan Virus Propagation. Front. Microbiol. 2022, 13, 849020. [Google Scholar] [CrossRef] [PubMed]
- Pawar, K.; Hanisch, C.; Palma Vera, S.E.; Einspanier, R.; Sharbati, S. Down regulated lncRNA MEG3 eliminates mycobacteria in macrophages via autophagy. Sci. Rep. 2016, 6, 19416. [Google Scholar] [CrossRef]
- Luo, F.; Wen, Y.; Zhao, L.; Su, S.; Lei, W.; Chen, L.; Chen, C.; Huang, Q.; Li, Z. LncRNA ZEB1-AS1/miR-1224-5p/MAP4K4 axis regulates mitochondria-mediated HeLa cell apoptosis in persistent Chlamydia trachomatis infection. Virulence 2022, 13, 444–457. [Google Scholar] [CrossRef]
- Batista, P.J.; Chang, H.Y. Long noncoding RNAs: Cellular address codes in development and disease. Cell 2013, 152, 1298–1307. [Google Scholar] [CrossRef]
- Wen, X.; Gao, L.; Guo, X.; Li, X.; Huang, X.; Wang, Y.; Xu, H.; He, R.; Jia, C.; Liang, F. lncSLdb: A resource for long non-coding RNA subcellular localization. Database 2018, 2018, 1–6. [Google Scholar] [CrossRef]
- Latos, P.A.; Pauler, F.M.; Koerner, M.V.; Şenergin, H.B.; Hudson, Q.J.; Stocsits, R.R.; Allhoff, W.; Stricker, S.H.; Klement, R.M.; Warczok, K.E.; et al. Airn transcriptional overlap, but not its lncRNA products, induces imprinted Igf2r silencing. Science 2012, 338, 1469–1472. [Google Scholar] [CrossRef]
- Atala, A. Re: Integrative Analyses Reveal a Long Noncoding RNA-Mediated Sponge Regulatory Network in Prostate Cancer. J. Urol. 2017, 197 Pt 1, 701. [Google Scholar] [CrossRef]
- Zhang, H.; Zou, X.; Liu, F. Silencing TTTY15 mitigates hypoxia-induced mitochondrial energy metabolism dysfunction and cardiomyocytes apoptosis via TTTY15/let-7i-5p and TLR3/NF-κB pathways. Cell. Signal. 2020, 76, 109779. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.J.; Liu, R.; Han, X.J.; Qiu, C.L.; Dong, G.L.; Liu, Z.Q.; Liu, L.H.; Luo, Y.; Jiang, L.P. Knockdown of the long non-coding RNA MALAT1 ameliorates TNF-α-mediated endothelial cell pyroptosis via the miR-30c-5p/Cx43 axis. Mol. Med. Rep. 2023, 27, 90. [Google Scholar] [CrossRef] [PubMed]
- Pickard, M.R.; Williams, G.T. The hormone response element mimic sequence of GAS5 lncRNA is sufficient to induce apoptosis in breast cancer cells. Oncotarget 2016, 7, 10104–10116. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Cheng, Q.; Xia, M.; Huang, X.; He, X.; Liao, J. Hypoxia-Induced lncRNA-NEAT1 Sustains the Growth of Hepatocellular Carcinoma via Regulation of miR-199a-3p/UCK2. Front. Oncol. 2020, 10, 998. [Google Scholar] [CrossRef]
- Bai, J.; Li, H.; Chen, X.; Chen, L.; Hu, Y.; Liu, L.; Zhao, Y.; Zuo, W.; Zhang, B.; Yin, C. LncRNA-AC009948.5 promotes invasion and metastasis of lung adenocarcinoma by binding to miR-186-5p. Front. Oncol. 2022, 12, 949951. [Google Scholar] [CrossRef]
- Van Wilpe, S.; Koornstra, R.; Den Brok, M.; De Groot, J.W.; Blank, C.; De Vries, J.; Gerritsen, W.; Mehra, N. Lactate dehydrogenase: A marker of diminished antitumor immunity. Oncoimmunology 2020, 9, 1731942. [Google Scholar] [CrossRef]
- Zou, M.; Zhai, Y.; Mei, X.; Wei, X. Lactate dehydrogenase and the severity of adenoviral pneumonia in children: A meta-analysis. Front. Pediatr. 2022, 10, 1059728. [Google Scholar] [CrossRef]
- Hong, J.; Xiao, X.; Li, T.; Wang, H.; Hua, Q. Long non-coding RNA NONHSAT217600.1 is involved in the regulation of neodymium oxide-induced cytotoxicity in 16HBE cells. Mol. Cell. Toxicol. 2023. [Google Scholar] [CrossRef]
- Castellanos-Rubio, A.; Fernandez-Jimenez, N.; Kratchmarov, R.; Luo, X.; Bhagat, G.; Green, P.H.; Schneider, R.; Kiledjian, M.; Bilbao, J.R.; Ghosh, S. A long noncoding RNA associated with susceptibility to celiac disease. Science 2016, 352, 91–95. [Google Scholar] [CrossRef]
- Manevski, M.; Devadoss, D.; Long, C.; Singh, S.P.; Nasser, M.W.; Borchert, G.M.; Nair, M.N.; Rahman, I.; Sopori, M.; Chand, H.S. Increased Expression of LASI lncRNA Regulates the Cigarette Smoke and COPD Associated Airway Inflammation and Mucous Cell Hyperplasia. Front. Immunol. 2022, 13, 803362. [Google Scholar] [CrossRef]
- Xie, B.; Qiao, M.; Xuan, J. lncRNA MEG3 Downregulation Relieves Intracerebral Hemorrhage by Inhibiting Oxidative Stress and Inflammation in an miR-181b-Dependent Manner. Med. Sci. Monit. 2021, 27, e929435. [Google Scholar] [CrossRef] [PubMed]
| Primers | Sequences (5′ to 3′) |
|---|---|
| 5′ RACE | Reverse1: CTTCATGAACCTGTGGCAAGTGCCTGAAA |
| 5′ RACE | Reverse2: GGAGGGCATGGCATGCATTGCAAAG |
| 3′ RACE | Forward1: CCCATGGCCACGCCAGAGCCTTAA |
| 3′ RACE | Forward2: GGGATGGAACCTGTGTCCTCCTGGATA |
| Primers | Sequences (5′ to 3′) | Accession No. |
|---|---|---|
| LNC_001186 | Forward: TCTGCCATCTCATCTATTTCGC | / |
| Reverse: GTGGCAAGTGCCTGAAAGAC | ||
| U6 | Forward: GGAACGATACAGAGAAGATTAGC | NC_000015 |
| Reverse: TGGAACGCTTCACGAATTTGCG | ||
| GAPDH | Forward: AGTATGATTCCACCCACGGC | NM_001206359.1 |
| Reverse: TACGTAGCACCAGCATCACC |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xie, K.; Yang, Q.; Yan, Z.; Huang, X.; Wang, P.; Gao, X.; Gun, S. Identification of a Novel lncRNA LNC_001186 and Its Effects on CPB2 Toxin-Induced Apoptosis of IPEC-J2 Cells. Genes 2023, 14, 1047. https://doi.org/10.3390/genes14051047
Xie K, Yang Q, Yan Z, Huang X, Wang P, Gao X, Gun S. Identification of a Novel lncRNA LNC_001186 and Its Effects on CPB2 Toxin-Induced Apoptosis of IPEC-J2 Cells. Genes. 2023; 14(5):1047. https://doi.org/10.3390/genes14051047
Chicago/Turabian StyleXie, Kaihui, Qiaoli Yang, Zunqiang Yan, Xiaoyu Huang, Pengfei Wang, Xiaoli Gao, and Shuangbao Gun. 2023. "Identification of a Novel lncRNA LNC_001186 and Its Effects on CPB2 Toxin-Induced Apoptosis of IPEC-J2 Cells" Genes 14, no. 5: 1047. https://doi.org/10.3390/genes14051047
APA StyleXie, K., Yang, Q., Yan, Z., Huang, X., Wang, P., Gao, X., & Gun, S. (2023). Identification of a Novel lncRNA LNC_001186 and Its Effects on CPB2 Toxin-Induced Apoptosis of IPEC-J2 Cells. Genes, 14(5), 1047. https://doi.org/10.3390/genes14051047





