Computational Biology Helps Understand How Polyploid Giant Cancer Cells Drive Tumor Success
Abstract
1. Introduction
2. Polyploidy in Cancer
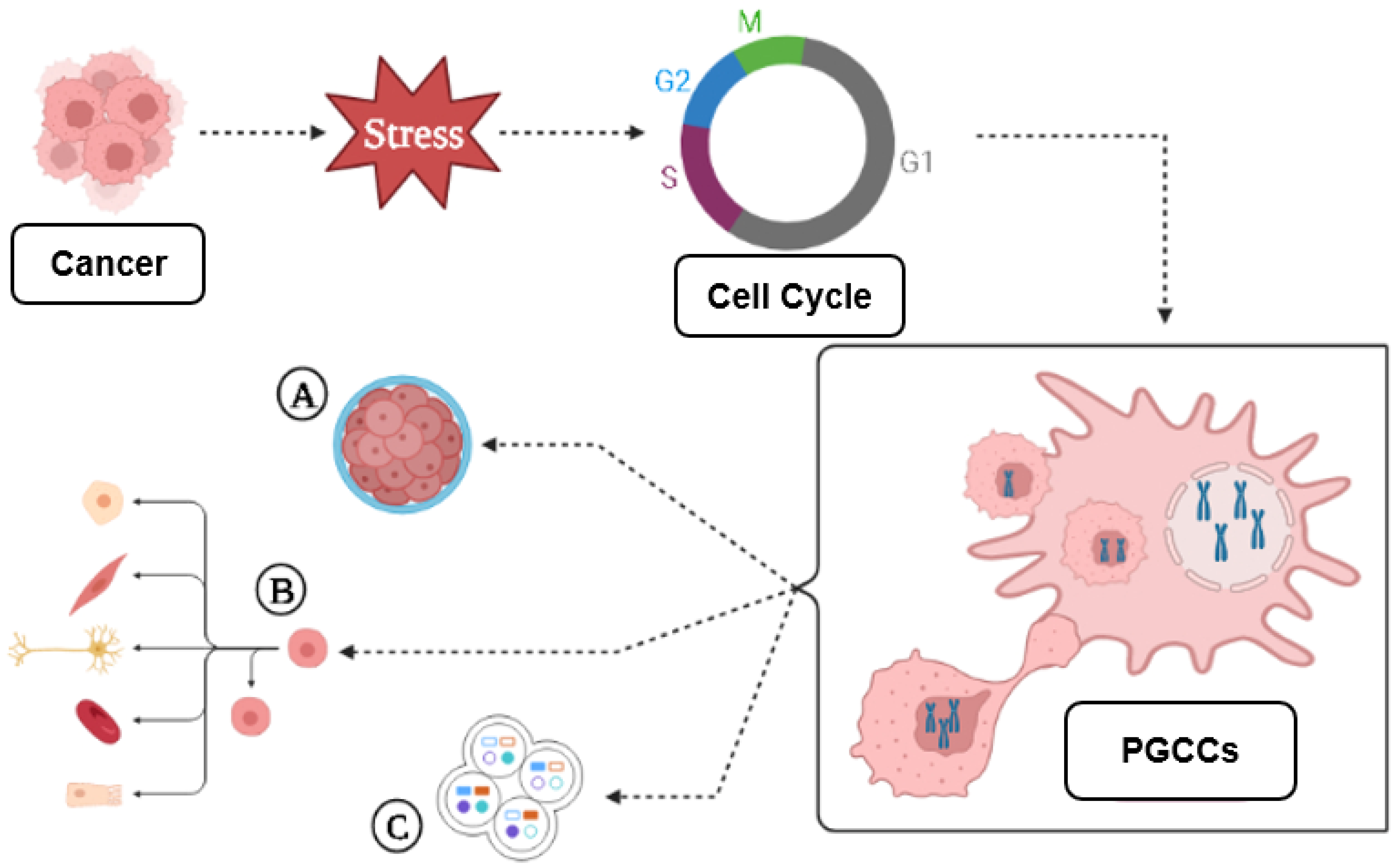
3. A Brief Introduction to PGCCs
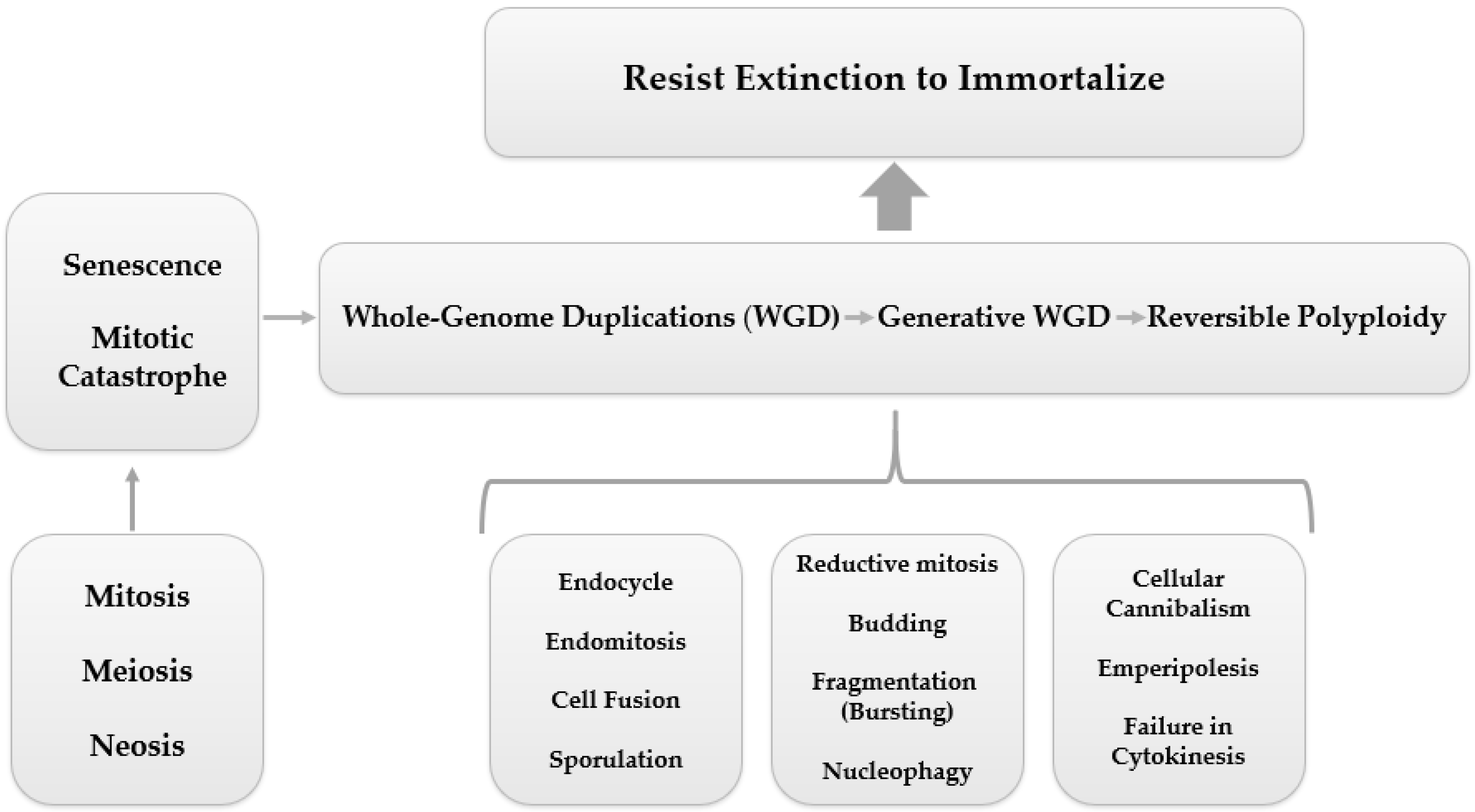
3.1. PGCC’s Giant Cell Cycle
Giant Cell Cycle Possible Outcomes and Fates
3.2. PGCCs Functionalities: Plasticity, Metabolism and Resistance to Therapy
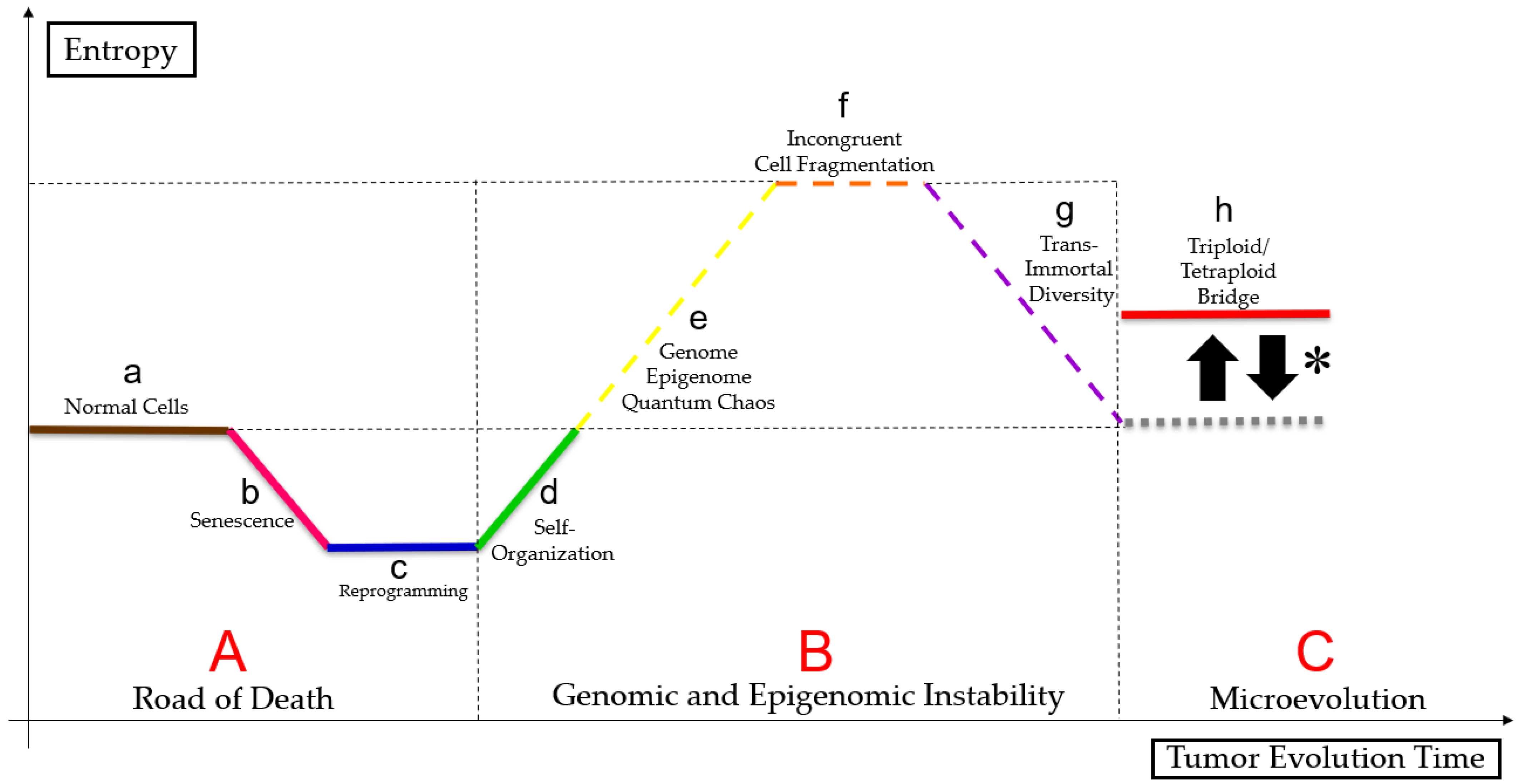
3.3. PGCC’s Role in Tumor Evolution
3.4. Genome Chaos
3.5. PGCCs Characterization in Diverse Cancer Types
3.5.1. Breast and Ovarian Cancer
3.5.2. Colorectal Cancer
3.5.3. Glioblastoma
3.5.4. Lung Cancer
3.5.5. Prostate Cancer and Melanoma
3.5.6. Only Ovarian Cancer
3.5.7. Only Breast Cancer
3.6. Autophagy, Senescence and PGCCs
4. Reaching New Paths
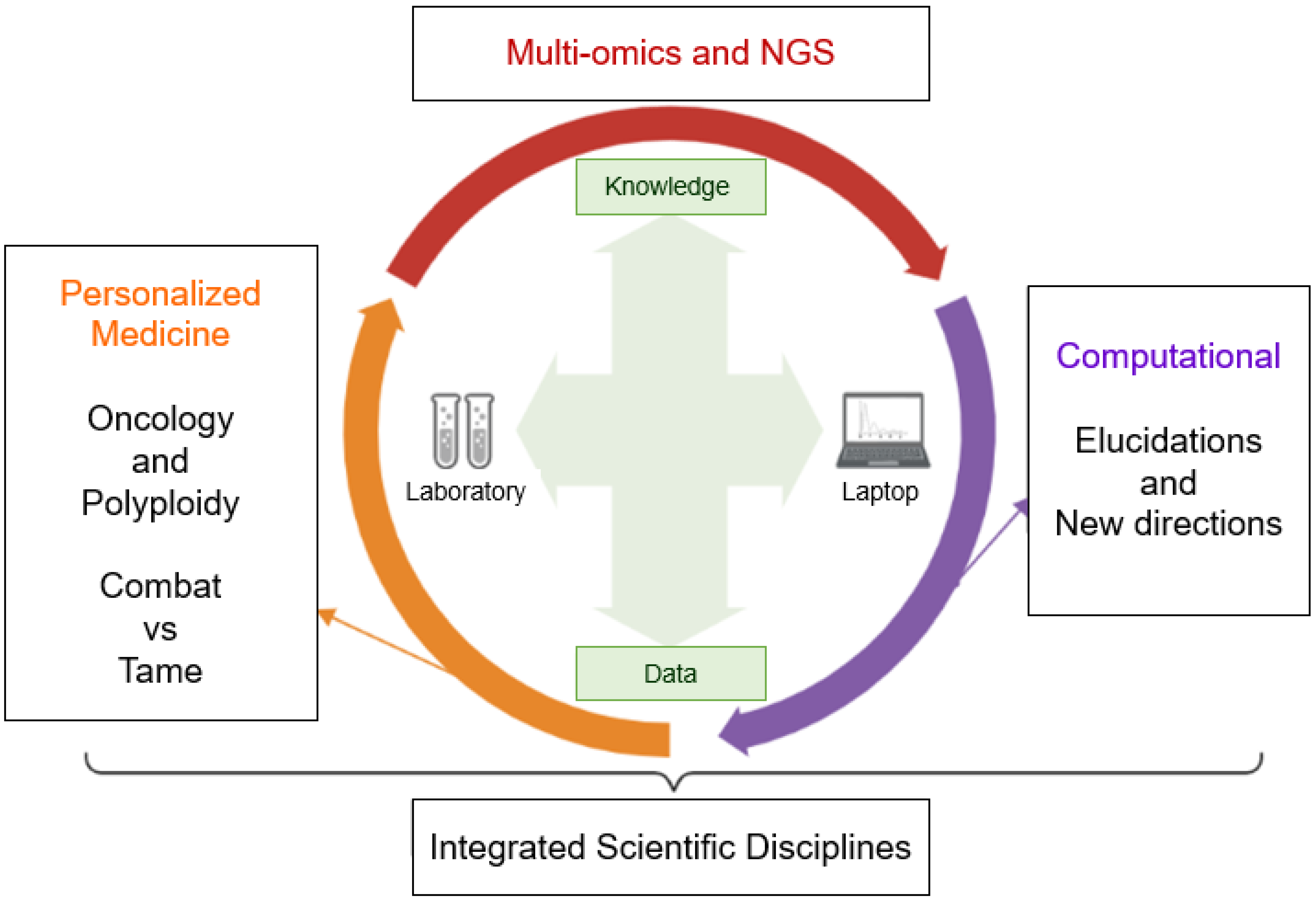
5. Computational Perspectives
5.1. Bioinformatics/Computational Studies in Oncology
5.2. Computational Studies about Polyploidization and PGCCs
5.3. Future Directions
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ASAH1 | N-acylsphingosine amidohydrolase (acid ceramidase) 1 |
| ATP | Adenosine triphosphate |
| AURKA | Aurora kinase A |
| BRAF V600E | Mutation of the BRAF gene |
| CD47 | Integrin-associated protein |
| CD9 | Motility-Related Protein |
| CDC25A | M-phase inducer phosphatase 1 |
| CDC25B | M-phase inducer phosphatase 2 |
| CDC25C | M-phase inducer phosphatase 3 |
| CENPF | Centromere Protein F |
| CK7 | Cytokeratin 7 |
| CoCl2 | Cobalt chloride |
| CSC | Cancer stem cells |
| EGFR | Epidermal growth factor receptor |
| EMT | Epithelial-Mesenchymal Transition |
| ERK | Extracellular signal-regulated kinase |
| EZH2 | Enhancer of zeste homolog 2 |
| HIF-1α | Hypoxia-inducible factor-1 |
| JNK | c-Jun N-terminal kinase |
| JNK1 | c-Jun N-terminal kinase 1 |
| LOH | Loss of heterozygosity |
| MELK | Maternal embryonic leucine zipper kinase |
| MYC | MYC Proto-Oncogene |
| NUSAP1 | Nucleolar and spindle associated protein 1 |
| p38MAPK | p38 mitogen-activated protein kinase |
| PACCs | Poly-aneuploid cancer cells |
| PD1/PD-L1 | Programmed cell death protein-1 |
| PGCCs | Polyploid giant cancer cells |
| PPARG | Peroxisome proliferator activated receptor γ |
| PTEN | Phosphatase and tensin homologue |
| S100A4 | S100 calcium binding protein A4 |
| SKP2 | S-phase kinase associated protein 2 |
| SLC7A11 | solute carrier family 7 member 11 |
| Slug | Snail family transcriptional repressor 2 |
| Snail | Snail family transcriptional repressor 1 |
| SSP | Staurosporine |
| STC1 | Stanniocalcin-1 |
| TNBC | Triple negative breast cancer |
| TOP2A | DNA Topoisomerase II α |
| TPL | Triptolide |
| Twist-1 | Twist family bHLH transcription factor 1 |
| VEGFA | Vascular endothelial growth factor A |
| VM | Vasculogenic mimicry |
| WGDs | Whole-genome duplications |
| ZEB1 | Zinc finger E-box binding homeobox 1 |
Appendix A
References
- Niu, N.; Zhang, J.; Zhang, N.; Mercado-Uribe, I.; Tao, F.; Han, Z.; Pathak, S.; Multani, A.S.; Kuang, J.; Yao, J.; et al. Linking genomic reorganization to tumor initiation via the giant cell cycle. Oncogenesis 2016, 5, e281. [Google Scholar] [CrossRef]
- Griffiths, A.J.; Wessler, S.R.; Lewontin, R.C.; Gelbart, W.M.; Suzuki, D.T.; Miller, J.H. Introdução à Genética; Grupo GEN: Guanabara Koogan, SP, Brazil, 2022; pp. 1–768. [Google Scholar]
- Fei, F.; Zhang, D.; Yang, Z.; Wang, S.; Wang, X.; Wu, Z.; Wu, Q.; Zhang, S. The number of polyploid giant cancer cells and epithelial-mesenchymal transition-related proteins are associated with invasion and metastasis in human breast cancer. J. Exp. Clin. Cancer Res. 2015, 34, 158. [Google Scholar] [CrossRef]
- Niculescu, V.F. The reproductive life cycle of cancer: Hypotheses of cell of origin, TP53 drivers and stem cell conversions in the light of the atavistic cancer cell theory. Med. Hypotheses 2019, 123, 19–23. [Google Scholar] [CrossRef]
- Glassmann, A.; Garcia, C.C.; Janzen, V.; Kraus, D.; Veit, N.; Winter, J.; Probstmeier, R. Staurosporine induces the generation of polyploid giant cancer cells in non-small-cell lung carcinoma A549 cells. Anal. Cell. Pathol. 2018, 2018, 7. [Google Scholar] [CrossRef]
- Fei, F.; Liu, K.; Li, C.; Du, J.; Wei, Z.; Li, B.; Li, Y.; Zhang, Y.; Zhang, S. Molecular mechanisms by which S100A4 regulates the migration and invasion of PGCCs with their daughter cells in human colorectal cancer. Front. Oncol. 2020, 10, 182. [Google Scholar] [CrossRef]
- Qu, Y.; Zhang, L.; Rong, Z.; He, T.; Zhang, S. Number of glioma polyploid giant cancer cells (PGCCs) associated with vasculogenic mimicry formation and tumor grade in human glioma. J. Exp. Clin. Cancer Res. 2013, 32, 75. [Google Scholar] [CrossRef]
- Chen, J.; Niu, N.; Zhang, J.; Qi, L.; Shen, W.; Donkena, K.V.; Feng, Z.; Liu, J. Polyploid giant cancer cells (PGCCs): The evil roots of cancer. Curr. Cancer Drug Targets 2019, 19, 360–367. [Google Scholar] [CrossRef]
- Liu, G.; Wang, Y.; Fei, F.; Wang, X.; Li, C.; Liu, K.; Du, J.; Cao, Y.; Zhang, S. Clinical characteristics and preliminary morphological observation of 47 cases of primary anorectal malignant melanomas. Melanoma Res. 2018, 28, 592–599. [Google Scholar] [CrossRef]
- Arun, R.P.; Sivanesan, D.; Patra, B.; Varadaraj, S.; Verma, R.S. Simulated microgravity increases polyploid giant cancer cells and nuclear localization of YAP. Sci. Rep. 2019, 9, 10684. [Google Scholar] [CrossRef]
- Liu, J. The “life code”: A theory that unifies the human life cycle and the origin of human tumors. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2020; Volume 60, pp. 380–397. [Google Scholar]
- Pienta, K.J.; Hammarlund, E.U.; Austin, R.H.; Axelrod, R.; Brown, J.S.; Amend, S.R. Cancer cells employ an evolutionarily conserved polyploidization program to resist therapy. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2020; Volume 81, pp. 145–159. [Google Scholar]
- Zhang, S.; Mercado-Uribe, I.; Liu, J. Tumor stroma and differentiated cancer cells can be originated directly from polyploid giant cancer cells induced by paclitaxel. Int. J. Cancer 2014, 134, 508–518. [Google Scholar] [CrossRef]
- Tagal, V.; Roth, M.G. Loss of Aurora kinase signaling allows lung cancer cells to adopt endoreplication and form polyploid giant cancer cells that resist antimitotic drugs. Cancer Res. 2021, 81, 400–413. [Google Scholar] [CrossRef]
- Fei, F.; Li, C.; Cao, Y.; Liu, K.; Du, J.; Gu, Y.; Wang, X.; Li, Y.; Zhang, S. CK7 expression associates with the location, differentiation, lymph node metastasis, and the Dukes’ stage of primary colorectal cancers. J. Cancer 2019, 10, 2510. [Google Scholar] [CrossRef]
- Bharadwaj, D.; Parekh, A.; Das, S.; Jena, B.C.; Mandal, M. Polyploid giant cancer cells induce growth arrest and cytoskeletal rearrangement in breast cancer cells. New Biotechnol. 2018, 44, S141. [Google Scholar] [CrossRef]
- Mirzayans, R.; Andrais, B.; Murray, D. Roles of polyploid/multinucleated giant cancer cells in metastasis and disease relapse following anticancer treatment. Cancers 2018, 10, 118. [Google Scholar] [CrossRef]
- Mirzayans, R.; Murray, D. Do TUNEL and other apoptosis assays detect cell death in preclinical studies? Int. J. Mol. Sci. 2020, 21, 9090. [Google Scholar] [CrossRef]
- Mirzayans, R.; Murray, D. Intratumor heterogeneity and therapy resistance: Contributions of dormancy, apoptosis reversal (anastasis) and cell fusion to disease recurrence. Int. J. Mol. Sci. 2020, 21, 1308. [Google Scholar] [CrossRef]
- Bharadwaj, D.; Mandal, M. Senescence in polyploid giant cancer cells: A road that leads to chemoresistance. Cytokine Growth Factor Rev. 2020, 52, 68–75. [Google Scholar] [CrossRef]
- Zhang, D.; Yang, X.; Yang, Z.; Fei, F.; Li, S.; Qu, J.; Zhang, M.; Li, Y.; Zhang, X.; Zhang, S. Daughter cells and erythroid cells budding from PGCCs and their clinicopathological significances in colorectal cancer. J. Cancer 2017, 8, 469. [Google Scholar] [CrossRef]
- Zhang, H.; Ma, H.; Yang, X.; Fan, L.; Tian, S.; Niu, R.; Yan, M.; Zheng, M.; Zhang, S. Cell Fusion-Related Proteins and Signaling Pathways, and Their Roles in the Development and Progression of Cancer. Front. Cell Dev. Biol. 2021, 9, 809668. [Google Scholar] [CrossRef]
- Lin, K.C.; Torga, G.; Sun, Y.; Axelrod, R.; Pienta, K.J.; Sturm, J.C.; Austin, R.H. The role of heterogeneous environment and docetaxel gradient in the emergence of polyploid, mesenchymal and resistant prostate cancer cells. Clin. Exp. Metastasis 2019, 36, 97–108. [Google Scholar] [CrossRef]
- Sirois, I.; Aguilar-Mahecha, A.; Lafleur, J.; Fowler, E.; Vu, V.; Scriver, M.; Buchanan, M.; Chabot, C.; Ramanathan, A.; Balachandran, B.; et al. A unique morphological phenotype in chemoresistant triple-negative breast cancer reveals metabolic reprogramming and PLIN4 expression as a molecular vulnerability. Mol. Cancer Res. 2019, 17, 2492–2507. [Google Scholar] [CrossRef]
- White-Gilbertson, S.; Lu, P.; Jones, C.M.; Chiodini, S.; Hurley, D.; Das, A.; Delaney, J.R.; Norris, J.S.; Voelkel-Johnson, C. Tamoxifen is a candidate first-in-class inhibitor of acid ceramidase that reduces amitotic division in polyploid giant cancer cells—Unrecognized players in tumorigenesis. Cancer Med. 2020, 9, 3142–3152. [Google Scholar] [CrossRef]
- Voelkel-Johnson, C. Sphingolipids in embryonic development, cell cycle regulation, and stemness–Implications for polyploidy in tumors. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2021; Volume 81, pp. 206–219. [Google Scholar]
- Shu, Z.; Row, S.; Deng, W.-M. Endoreplication: The good, the bad, and the ugly. Trends Cell Biol. 2018, 28, 465–474. [Google Scholar] [CrossRef]
- Zhang, D.; Wang, Y.; Zhang, S. Asymmetric cell division in polyploid giant cancer cells and low eukaryotic cells. BioMed Res. Int. 2014, 2014, 432652. [Google Scholar] [CrossRef]
- Liu, J. The dualistic origin of human tumors. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2018; Volume 53, pp. 1–16. [Google Scholar]
- Anatskaya, O.V.; Vinogradov, A.E. Polyploidy as a Fundamental Phenomenon in Evolution, Development, Adaptation and Diseases. Int. J. Mol. Sci. 2022, 23, 3542. [Google Scholar] [CrossRef]
- Archetti, M. Polyploidy as an Adaptation against Loss of Heterozygosity in Cancer. Int. J. Mol. Sci. 2022, 23, 8528. [Google Scholar] [CrossRef]
- Wu, C.S.; Lu, W.H.; Hung, M.C.; Huang, Y.S.; Chao, H.W. From polyploidy to polyploidy reversal: Its role in normal and disease states. Trends Genet. 2022, 38, 991–995. [Google Scholar] [CrossRef]
- Bukkuri, A.; Pienta, K.J.; Austin, R.H.; Hammarlund, E.U.; Amend, S.R.; Brown, J.S. A life history model of the ecological and evolutionary dynamics of polyaneuploid cancer cells. Sci. Rep. 2022, 12, 13713. [Google Scholar] [CrossRef]
- Anatskaya, O.V.; Vinogradov, A.E. Whole-Genome Duplications in Evolution, Ontogeny, and Pathology: Complexity and Emergency Reserves. Mol. Biol. 2021, 55, 813–827. [Google Scholar] [CrossRef]
- Lukow, D.A.; Sheltzer, J.M. Chromosomal instability and aneuploidy as causes of cancer drug resistance. Trends Cancer 2022, 8, 43–53. [Google Scholar] [CrossRef]
- Mukherjee, S.; Ali, A.M.; Murty, V.V.; Raza, A. Mutation in SF3B1 gene promotes formation of polyploid giant cells in Leukemia cells. Med. Oncol. 2022, 39, 65. [Google Scholar] [CrossRef]
- Kasperski, A. Life Entrapped in a Network of Atavistic Attractors: How to Find a Rescue. Int. J. Mol. Sci. 2022, 23, 4017. [Google Scholar] [CrossRef]
- Niculescu, V.F. Cancer genes and cancer stem cells in tumorigenesis: Evolutionary deep homology and controversies. Genes Dis. 2022, 9, 1234–1247. [Google Scholar] [CrossRef]
- Amend, S.R.; Torga, G.; Lin, K.C.; Kostecka, L.G.; de Marzo, A.; Austin, R.H.; Pienta, K.J. Polyploid giant cancer cells: Unrecognized actuators of tumorigenesis, metastasis, and resistance. Prostate 2019, 79, 1489–1497. [Google Scholar] [CrossRef]
- White-Gilbertson, S.; Voelkel-Johnson, C. Giants and monsters: Unexpected characters in the story of cancer recurrence. Adv. Cancer Res. 2020, 148, 201–232. [Google Scholar]
- Alameddine, R.S.; Hamieh, L.; Shamseddine, A. From sprouting angiogenesis to erythrocytes generation by cancer stem cells: Evolving concepts in tumor microcirculation. BioMed Res. Int. 2014, 2014, 986768. [Google Scholar] [CrossRef]
- Yang, Z.; Yao, H.; Fei, F.; Li, Y.; Qu, J.; Li, C.; Zhang, S. Generation of erythroid cells from polyploid giant cancer cells: Re-thinking about tumor blood supply. J. Cancer Res. Clin. Oncol. 2018, 144, 617–627. [Google Scholar] [CrossRef]
- Amend, S.R.; Pienta, K.J. Ecology meets cancer biology: The cancer swamp promotes the lethal cancer phenotype. Oncotarget 2015, 6, 9669. [Google Scholar] [CrossRef]
- Liu, J.; Niu, N.; Li, X.; Zhang, X.; Sood, A.K. The life cycle of polyploid giant cancer cell and dormancy in cancer: Opportunities for novel therapeutic interventions. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2021; Volume 81, pp. 132–144. [Google Scholar]
- Liu, J.; Erenpreisa, J.; Sikora, E. Polyploid giant cancer cells: An emerging new field of cancer biology. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2021; Volume 81, pp. 1–4. [Google Scholar]
- Liu, J. Giant cells: Linking McClintock’s heredity to early embryogenesis and tumor origin throughout millennia of evolution on Earth. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2021; Volume 81, pp. 176–192. [Google Scholar]
- Song, Y.; Zhao, Y.; Deng, Z.; Zhao, R.; Huang, Q. Stress-Induced Polyploid Giant Cancer Cells: Unique Way of Formation and Non-Negligible Characteristics. Front. Oncol. 2021, 11, 3390. [Google Scholar] [CrossRef]
- Erenpreisa, J.; Salmina, K.; Anatskaya, O.; Cragg, M.S. Paradoxes of cancer: Survival at the brink. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2020; Volume 81, pp. 119–131. [Google Scholar]
- Niu, N.; Mercado-Uribe, I.; Liu, J. Dedifferentiation into blastomere-like cancer stem cells via formation of polyploid giant cancer cells. Oncogene 2017, 36, 4887–4900. [Google Scholar] [CrossRef]
- Erenpreisa, J.; Cragg, M.S. Three steps to the immortality of cancer cells: Senescence, polyploidy and self-renewal. Cancer Cell Int. 2013, 13, 92. [Google Scholar] [CrossRef]
- Erenpreisa, J.; Salmina, K.; Anatskaya, O.; Vinogradov, A.; Cragg, M.S. The Enigma of cancer resistance to treatment. Organisms. J. Biol. Sci. 2022, 5, 71–75. [Google Scholar]
- Mayfield-Jones, D.; Washburn, J.D.; Arias, T.; Edger, P.P.; Pires, J.C.; Conant, G.C. Watching the grin fade: Tracing the effects of polyploidy on different evolutionary time scales. In Seminars in Cell & Developmental Biology; Academic Press: Cambridge, MA, USA, 2013; Volume 24, pp. 320–331. [Google Scholar]
- Moein, S.; Adibi, R.; Meirelles, L.d.S.; Nardi, N.B.; Gheisari, Y. Cancer regeneration: Polyploid cells are the key drivers of tumor progression. Biochim. Biophys. Acta (BBA)-Rev. Cancer 2020, 1874, 188408. [Google Scholar] [CrossRef]
- Dasari, K.; Somarelli, J.A.; Kumar, S.; Townsend, J.P. The somatic molecular evolution of cancer: Mutation, selection, and epistasis. Prog. Biophys. Mol. Biol. 2021, 165, 56–65. [Google Scholar] [CrossRef]
- Somarelli, J.A.; Gardner, H.; Cannataro, V.L.; Gunady, E.F.; Boddy, A.M.; Johnson, N.A.; Fisk, J.N.; Gaffney, S.G.; Chuang, J.H.; Li, S.; et al. Molecular biology and evolution of cancer: From discovery to action. Mol. Biol. Evol. 2020, 37, 320–326. [Google Scholar] [CrossRef]
- Dujon, A.M.; Aktipis, A.; Alix-Panabières, C.; Amend, S.R.; Boddy, A.M.; Brown, J.S.; Capp, J.P.; DeGregori, J.; Ewald, P.; Gatenby, R.; et al. Identifying key questions in the ecology and evolution of cancer. Evol. Appl. 2021, 14, 877–892. [Google Scholar] [CrossRef]
- Casás-Selves, M.; DeGregori, J. How cancer shapes evolution and how evolution shapes cancer. Evol. Educ. Outreach 2011, 4, 624–634. [Google Scholar] [CrossRef]
- Fortunato, A.; Boddy, A.; Mallo, D.; Aktipis, A.; Maley, C.C.; Pepper, J.W. Natural selection in cancer biology: From molecular snowflakes to trait hallmarks. Cold Spring Harb. Perspect. Med. 2017, 7, a029652. [Google Scholar] [CrossRef]
- Arendt, D. The evolution of cell types in animals: Emerging principles from molecular studies. Nat. Rev. Genet. 2008, 9, 868–882. [Google Scholar] [CrossRef]
- Arendt, D.; Musser, J.M.; Baker, C.V.; Bergman, A.; Cepko, C.; Erwin, D.H.; Wagner, G.P. The origin and evolution of cell types. Nat. Rev. Genet. 2016, 17, 744–757. [Google Scholar] [CrossRef]
- Tarashansky, A.J.; Musser, J.M.; Khariton, M.; Li, P.; Arendt, D.; Quake, S.R.; Wang, B. Mapping single-cell atlases throughout Metazoa unravels cell type evolution. eLife 2021, 10, e66747. [Google Scholar] [CrossRef]
- Was, H.; Borkowska, A.; Olszewska, A.; Klemba, A.; Marciniak, M.; Synowiec, A.; Kieda, C. Polyploidy formation in cancer cells: How a Trojan horse is born. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2022; Volume 81, pp. 24–36. [Google Scholar]
- Niculescu, V.F. aCLS cancers: Genomic and epigenetic changes transform the cell of origin of cancer into a tumorigenic pathogen of unicellular organization and lifestyle. Gene 2020, 726, 144174. [Google Scholar] [CrossRef]
- Niculescu, V.F. Is an ancient genome repair mechanism the Trojan Horse of cancer. Nov Appro Can Study 2021, 5, 555–557. [Google Scholar] [CrossRef]
- Baker, S.G.; Kramer, B.S. Paradoxes in carcinogenesis: New opportunities for research directions. BMC Cancer 2007, 7, 151. [Google Scholar] [CrossRef]
- Baker, S.G. Paradoxes in carcinogenesis should spur new avenues of research: An historical perspective. Disruptive Sci. Technol. 2012, 1, 100–107. [Google Scholar] [CrossRef]
- Heng, J.; Heng, H.H. Two-phased evolution: Genome chaos-mediated information creation and maintenance. Prog. Biophys. Mol. Biol. 2021, 165, 29–42. [Google Scholar] [CrossRef]
- Heng, J.; Heng, H.H. Genome chaos: Creating new genomic information essential for cancer macroevolution. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2020; Volume 81, pp. 160–175. [Google Scholar]
- Heng, J.; Heng, H.H. Genome Chaos, Information Creation, and Cancer Emergence: Searching for New Frameworks on the 50th Anniversary of the “War on Cancer”. Genes 2021, 13, 101. [Google Scholar] [CrossRef]
- Liu, J.; Mercado-Uribe, I.; Niu, N.; Sun, B.; Kuang, J.; Zhang, S. Re-thinking the concept of cancer stem cells: Polyploid giant cancer cells as mother cancer stem cells. Cancer Res. 2014, 74, 1917. [Google Scholar] [CrossRef]
- Baramiya, M.G.; Baranov, E. From cancer to rejuvenation: Incomplete regeneration as the missing link (Part I: The same origin, different outcomes). Future Sci. AO 2020, 6, FSO450. [Google Scholar] [CrossRef]
- Baramiya, M.G.; Baranov, E.; Saburina, I.; Salnikov, L. From cancer to rejuvenation: Incomplete regeneration as the missing link (part II: Rejuvenation circle). Future Sci. AO 2020, 6, FSO610. [Google Scholar] [CrossRef]
- Liu, K.; Zheng, M.; Zhao, Q.; Zhang, K.; Li, Z.; Fu, F.; Zhang, H.; Du, J.; Li, Y.; Zhang, S. Different p53 genotypes regulating different phosphorylation sites and subcellular location of CDC25C associated with the formation of polyploid giant cancer cells. J. Exp. Clin. Cancer Res. 2020, 39, 83. [Google Scholar] [CrossRef]
- Wang, X.; Zheng, M.; Fei, F.; Li, C.; Du, J.; Liu, K.; Li, Y.; Zhang, S. EMT-related protein expression in polyploid giant cancer cells and their daughter cells with different passages after triptolide treatment. Med. Oncol. 2019, 36, 82. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Sanchez, L.M.; Jimenez, C.; Valverde, A.; Hernandez, V.; Penarando, J.; Martinez, A.; Lopez-Pedrera, C.; Muñoz-Castañeda, J.R.; Haba-Rodríguez, J.R.D.; Aranda, E.; et al. CoCl2, a mimic of hypoxia, induces formation of polyploid giant cells with stem characteristics in colon cancer. PLoS ONE 2014, 9, e99143. [Google Scholar] [CrossRef]
- Fei, F.; Zhang, M.; Li, B.; Zhao, L.; Wang, H.; Liu, L.; Li, Y.; Ding, P.; Gu, Y.; Zhang, X.; et al. Formation of polyploid giant cancer cells involves in the prognostic value of neoadjuvant chemoradiation in locally advanced rectal cancer. J. Oncol. 2019, 2019, 2316436. [Google Scholar] [CrossRef]
- Fei, F.; Li, C.; Wang, X.; Du, J.; Liu, K.; Li, B.; Yao, P.; Li, Y.; Zhang, S. Syncytin 1, CD9, and CD47 regulating cell fusion to form PGCCs associated with cAMP/PKA and JNK signaling pathway. Cancer Med. 2019, 8, 3047–3058. [Google Scholar] [CrossRef]
- Fei, F.; Qu, J.; Liu, K.; Li, C.; Wang, X.; Li, Y.; Zhang, S. The subcellular location of cyclin B1 and CDC25 associated with the formation of polyploid giant cancer cells and their clinicopathological significance. Lab. Investig. 2019, 99, 483–498. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Shi, Y.; Wu, M.; Liu, J.; Wu, H.; Xu, C.; Chen, L. Hypoxia-induced polypoid giant cancer cells in glioma promote the transformation of tumor-associated macrophages to a tumor-supportive phenotype. CNS Neurosci. Ther. 2022, 28, 1326–1338. [Google Scholar] [CrossRef]
- White-Gilbertson, S.; Lu, P.; Esobi, I.; Echesabal-Chen, J.; Mulholland, P.J.; Gooz, M.; Ogretmen, B.; Stamatikos, A.; Voelkel-Johnson, C. Polyploid giant cancer cells are dependent on cholesterol for progeny formation through amitotic division. Sci. Rep. 2022, 12, 8971. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Mercado-Uribe, I.; Hanash, S.; Liu, J. iTRAQ-based proteomic analysis of polyploid giant cancer cells and budding progeny cells reveals several distinct pathways for ovarian cancer development. PLoS ONE 2013, 8, e80120. [Google Scholar] [CrossRef]
- Lv, H.; Shi, Y.; Zhang, L.; Zhang, D.; Liu, G.; Yang, Z.; Li, Y.; Fei, F.; Zhang, S. Polyploid giant cancer cells with budding and the expression of cyclin E, S-phase kinase-associated protein 2, stathmin associated with the grading and metastasis in serous ovarian tumor. BMC Cancer 2014, 14, 576. [Google Scholar] [CrossRef]
- Zhang, S.; Mercado-Uribe, I.; Xing, Z.; Sun, B.; Kuang, J.; Liu, J. of cancer stem-like cells through the formation of polyploid giant cancer cells. Oncogene 2014, 33, 116–128. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Ding, P.; Lv, H.; Zhang, D.; Liu, G.; Yang, Z.; Li, Y.; Liu, J.; Zhang, S. Number of polyploid giant cancer cells and expression of EZH2 are associated with VM formation and tumor grade in human ovarian tumor. BioMed Res. Int. 2014, 2014, 903542. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Lu, R.; Zhao, Q.; Du, J.; Li, Y.; Zheng, M.; Zhang, S. Association and clinicopathologic significance of p38MAPK-ERK-JNK-CDC25C with polyploid giant cancer cell formation. Med. Oncol. 2020, 37, 6. [Google Scholar] [CrossRef]
- You, B.; Xia, T.; Gu, M.; Zhang, Z.; Zhang, Q.; Shen, J.; Fan, Y.; Yao, H.; Pan, S.; Lu, Y.; et al. AMPK–mTOR–Mediated Activation of Autophagy Promotes Formation of Dormant Polyploid Giant Cancer Cells. Cancer Res. 2022, 82, 846–858. [Google Scholar] [CrossRef]
- Bowers, R.R.; Andrade, M.F.; Jones, C.M.; White-Gilbertson, S.; Voelkel-Johnson, C.; Delaney, J.R. Autophagy modulating therapeutics inhibit ovarian cancer colony generation by polyploid giant cancer cells (PGCCs). BMC Cancer 2022, 22, 410. [Google Scholar] [CrossRef] [PubMed]
- Czarnecka-Herok, J.; Sliwinska, M.A.; Herok, M.; Targonska, A.; Strzeszewska-Potyrala, A.; Bojko, A.; Wolny, A.; Mosieniak, G.; Sikora, E. Therapy-Induced Senescent/Polyploid Cancer Cells Undergo Atypical Divisions Associated with Altered Expression of Meiosis, Spermatogenesis and EMT Genes. Int. J. Mol. Sci. 2022, 23, 8288. [Google Scholar] [CrossRef]
- Ye, J.C.; Horne, S.; Zhang, J.Z.; Jackson, L.; Heng, H.H. Therapy induced genome chaos: A novel mechanism of rapid cancer drug resistance. Front. Cell Dev. Biol. 2021, 9, 676344. [Google Scholar] [CrossRef]
- Paul, D. The systemic hallmarks of cancer. J. Cancer Metastasis Treat. 2020, 6, 29. [Google Scholar] [CrossRef]
- Zhang, J.; Qiao, Q.; Xu, H.; Zhou, R.; Liu, X. Human cell polyploidization: The good and the evil. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2021; Volume 81, pp. 54–63. [Google Scholar]
- Zhang, S.; Xu, X.; Zhu, S.; Liu, J. PGCCS generating erythrocytes to form VM structure contributes to tumor blood supply. BioMed Res. Int. 2015, 2015, 402619. [Google Scholar] [CrossRef]
- Zhang, S.; Zhang, D.; Yang, Z.; Zhang, X. Tumor budding, micropapillary pattern, and polyploidy giant cancer cells in colorectal cancer: Current status and future prospects. Stem Cells Int. 2016, 2016, 4810734. [Google Scholar] [CrossRef]
- Zhang, Z.; Feng, X.; Deng, Z.; Cheng, J.; Wang, Y.; Zhao, M.; Zhao, Y.; He, S.; Huang, Q. Irradiation-induced polyploid giant cancer cells are involved in tumor cell repopulation via neosis. Mol. Oncol. 2021, 15, 2219–2234. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.T.; Xia, T.; You, Y.W.; Zhang, Q.C.; Ni, H.S.; Liu, Y.F.; Liu, Y.R.; Xu, Y.Q.; You, B.; Zhang, Z.X. Characteristics and clinical significance of polyploid giant cancer cells in laryngeal carcinoma. Laryngoscope Investig. Otolaryngol. 2021, 6, 1228–1234. [Google Scholar] [CrossRef]
- Liu, L.L.; Long, Z.J.; Wang, L.X.; Zheng, F.M.; Fang, Z.G.; Yan, M.; Xu, D.F.; Chen, J.J.; Wang, S.W.; Lin, D.J.; et al. Inhibition of mTOR Pathway Sensitizes Acute Myeloid Leukemia Cells to Aurora Inhibitors by Suppression of Glycolytic Metabolism. Mol. Cancer Res. 2013, 11, 1326–1336. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Hanahan, D. Hallmarks of cancer: New dimensions. Cancer Discov. 2022, 12, 31–46. [Google Scholar] [CrossRef]
- Nam, A.S.; Chaligne, R.; Landau, D.A. Integrating genetic and non-genetic determinants of cancer evolution by single-cell multi-omics. Nat. Rev. Genet. 2021, 22, 3–18. [Google Scholar] [CrossRef]
- Hasanzad, M.; Sarhangi, N.; Ehsani Chimeh, S.; Ayati, N.; Afzali, M.; Khatami, F.; Nikfar, S.; Aghaei Meybodi, H.R. Precision medicine journey through omics approach. J. Diabetes Metab. Disord. 2021, 21, 881–888. [Google Scholar] [CrossRef]
- Lewis, S.M.; Asselin-Labat, M.L.; Nguyen, Q.; Berthelet, J.; Tan, X.; Wimmer, V.C.; Merino, D.; Rogers, K.L.; Naik, S.H. Spatial omics and multiplexed imaging to explore cancer biology. Nat. Methods 2021, 18, 997–1012. [Google Scholar] [CrossRef]
- Dotolo, S.; Esposito, A.R.; Roma, C.; Guido, D.; Preziosi, A.; Tropea, B.; Palluzzi, F.; Giacò, L.; Normanno, N. Bioinformatics: From NGS Data to Biological Complexity in Variant Detection and Oncological Clinical Practice. Biomedicines 2022, 10, 2074. [Google Scholar] [CrossRef] [PubMed]
- Jiang, P.; Sinha, S.; Aldape, K.; Hannenhalli, S.; Sahinalp, C.; Ruppin, E. Big data in basic and translational cancer research. Nat. Rev. Cancer 2022, 22, 625–639. [Google Scholar] [CrossRef] [PubMed]
- Bourke, P.M.; Voorrips, R.E.; Visser, R.G.; Maliepaard, C. Tools for genetic studies in experimental populations of polyploids. Front. Plant Sci. 2018, 9, 513. [Google Scholar] [CrossRef] [PubMed]
- Desany, B.; Zhang, Z. Bioinformatics and cancer target discovery. Drug Discov. Today 2004, 9, 795–802. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Rice, C.M.; Wang, X. Cancer bioinformatics: A new approach to systems clinical medicine. BMC Bioinform. 2012, 1, 71. [Google Scholar] [CrossRef]
- Dopazo, J. Bioinformatics and cancer: An essential alliance. Clin. Transl. Oncol. 2006, 8, 409–415. [Google Scholar] [CrossRef]
- Zheng, H.; Zhang, G.; Zhang, L.; Wang, Q.; Li, H.; Han, Y.; Xie, L.; Yan, Z.; Li, Y.; An, Y.; et al. Comprehensive review of web servers and bioinformatics tools for cancer prognosis analysis. Front. Oncol. 2020, 10, 68. [Google Scholar] [CrossRef]
- Gómez-López, G.; Valencia, A. Bioinformatics and cancer research: Building bridges for translational research. Clin. Transl. Oncol. 2008, 10, 85–95. [Google Scholar] [CrossRef]
- Menyhárt, O.; Győrffy, B. Multi-omics approaches in cancer research with applications in tumor subtyping, prognosis, and diagnosis. Comput. Struct. Biotechnol. J. 2021, 19, 949–960. [Google Scholar] [CrossRef]
- Brenner, C. Applications of bioinformatics in cancer. Cancers 2019, 11, 1630. [Google Scholar] [CrossRef]
- Voshall, A.; Moriyama, E.N. Next-generation transcriptome assembly and analysis: Impact of ploidy. Methods 2020, 1, 14–24. [Google Scholar] [CrossRef]
- Jiménez-Santos, M.J.; García-Martín, S.; Fustero-Torre, C.; Di Domenico, T.; Gómez-López, G.; Al-Shahrour, F. Bioinformatics roadmap for therapy selection in cancer genomics. Mol. Oncol. 2022, 16, 3881–3908. [Google Scholar] [CrossRef]
- Hanauer, D.A.; Rhodes, D.R.; Sinha-Kumar, C.; Chinnaiyan, A.M. Bioinformatics approaches in the study of cancer. Curr. Mol. Med. 2007, 7, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Lemus, E.; Martínez-García, M. Pathway-based drug-repurposing schemes in cancer: The role of translational bioinformatics. Front. Oncol. 2021, 10, 605680. [Google Scholar] [CrossRef] [PubMed]
- Nasser, F.K.; Behadili, S.F. A Review of Data Mining and Knowledge Discovery Approaches for Bioinformatics. Iraqi J. Sci. 2022, 63, 3169–3188. [Google Scholar] [CrossRef]
- Yang, H.H.; Lee, M.P. Application of bioinformatics in cancer epigenetics. Ann. N. Y. Acad. Sci. 2004, 1020, 67–76. [Google Scholar] [CrossRef]
- Alam, M.S.; Sultana, A.; Reza, M.S.; Amanullah, M.; Kabir, S.R.; Mollah, M.N.H. Integrated bioinformatics and statistical approaches to explore molecular biomarkers for breast cancer diagnosis, prognosis and therapies. PLoS ONE 2022, 17, e0268967. [Google Scholar] [CrossRef] [PubMed]
- Kaur, B.; Mukhlis, Y.; Natesh, J.; Penta, D.; Musthapa Meeran, S. Identification of hub genes associated with EMT-induced chemoresistance in breast cancer using integrated bioinformatics analysis. Gene 2022, 809, 146016. [Google Scholar] [CrossRef]
- Mukherjee, N.; Browne, A.; Ivers, L.; Santra, T.; Cremona, M.; Hennessy, B.T.; O’Donovan, N.; Crown, J.; Kolch, W.; Fey, D.; et al. A Systems Biology Approach to Investigate Kinase Signal Transduction Networks That Are Involved in Triple Negative Breast Cancer Resistance to Cisplatin. J. Pers. Med. 2022, 12, 1277. [Google Scholar] [CrossRef]
- Yadav, D.K.; Sharma, A.; Dube, P.; Shaikh, S.; Vaghasia, H.; Rawal, R.M. Identification of crucial hub genes and potential molecular mechanisms in breast cancer by integrated bioinformatics analysis and experimental validation. Comput. Biol. Med. 2022, 149, 106036. [Google Scholar] [CrossRef]
- Yang, G.; Lu, T.; Weisenberger, D.J.; Liang, G. The Multi-Omic Landscape of Primary Breast Tumors and Their Metastases: Expanding the Efficacy of Actionable Therapeutic Targets. Genes 2022, 13, 1555. [Google Scholar] [CrossRef]
- Khan, M.M.; Mohsen, M.T.; Malik, M.Z.; Bagabir, S.A.; Alkhanani, M.F.; Haque, S.; Serajuddin, M.; Bharadwaj, M. Identification of Potential Key Genes in Prostate Cancer with Gene Expression, Pivotal Pathways and Regulatory Networks Analysis Using Integrated Bioinformatics Methods. Genes 2022, 13, 655. [Google Scholar] [CrossRef]
- Kumari, B.; Sakode, C.; Raghavendran, L.; Kumar Roy, P. Systems biology basis of permanent tumor regression with normal tissue protection: Experimentally validated signaling pathway framework. ASCO Annu. Meet. I 2022, 40, e15066. [Google Scholar] [CrossRef]
- Lin, Y.; Dong, Y.; Liu, W.; Fan, X.; Sun, Y. Pan-Cancer Analyses Confirmed the Ferroptosis-Related Gene SLC7A11 as a Prognostic Biomarker for Cancer. Int. J. Gen. Med. 2022, 15, 2501. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.B.; Do, D.N.; Nguyen-Thi, M.L.; Hoang-The, H.; Tran, T.T.; Nguyen-Thanh, T. Identification of potential crucial genes and key pathways shared in Inflammatory Bowel Disease and cervical cancer by machine learning and integrated bioinformatics. Comput. Biol. Med. 2022, 149, 105996. [Google Scholar] [CrossRef]
- Reza, M.S.; Harun-Or-Roshid, M.; Islam, M.A.; Hossen, M.A.; Hossain, M.T.; Feng, S.; Xi, W.; Mollah, M.N.H.; Wei, Y. Bioinformatics Screening of Potential Biomarkers from mRNA Expression Profiles to Discover Drug Targets and Agents for Cervical Cancer. Int. J. Mol. Sci. 2022, 23, 3968. [Google Scholar] [CrossRef]
- Roudi, R.; Beikzadeh, B.; Roviello, G.; D’angelo, A.; Hadizadeh, M. Identification of hub genes, modules and biological pathways associated with lung adenocarcinoma: A system biology approach. Gene Rep. 2022, 27, 101638. [Google Scholar] [CrossRef]
- Zhuang, Z.; Zhong, X.; Chen, Q.; Chen, H.; Liu, Z. Bioinformatics and System Biology Approach to Reveal the Interaction Network and the Therapeutic Implications for Non-Small Cell Lung Cancer Patients With COVID-19. Front. Pharmacol. 2022, 13, 857730. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, K.K. Robustness and the generalist niche of polyploid species: Genome shock or gradual evolution? Curr. Opin. Plant Biol. 2022, 69, 102292. [Google Scholar] [CrossRef]
- Baudoin, N.C.; Bloomfield, M. Karyotype aberrations in action: The evolution of cancer genomes and the tumor microenvironment. Genes 2021, 12, 558. [Google Scholar] [CrossRef]
- Kaigorodova, E.V.; Kozik, A.V.; Zavaruev, I.S.; Grishchenko, M.Y. Hybrid/Atypical Forms of Circulating Tumor Cells: Current State of the Art. Biochemistry 2022, 87, 380–390. [Google Scholar] [CrossRef]
- Soraggi, S.; Rhodes, J.; Altinkaya, I.; Tarrant, O.; Balloux, F.; Fisher, M.C.; Fumagalli, M. HMMploidy: Inference of ploidy levels from short-read sequencing data. Peer Community J. 2022, 2, e60. [Google Scholar] [CrossRef]
- Van Belzen, I.A.E.M.; Schönhuth, A.; Kemmeren, P.; Hehir-Kwa, J.Y. Structural variant detection in cancer genomes: Computational challenges and perspectives for precision oncology. NPJ Precis. Oncol. 2021, 5, 15. [Google Scholar] [CrossRef] [PubMed]
- Spence, T.; Dubuc, A.M. Copy Number Analysis in Cancer Diagnostic Testing. Clin. Lab. Med. 2022, 42, 451–468. [Google Scholar] [CrossRef]
- Zhang, T.; Zhou, J.; Gao, W.; Jia, Y.; Wei, Y.; Wang, G. Complex genome assembly based on long-read sequencing. Brief. Bioinform. 2022, 23, bbac305. [Google Scholar] [CrossRef] [PubMed]
- Anatskaya, O.V.; Vinogradov, A.E. Polyploidy and Myc Proto-Oncogenes Promote Stress Adaptation via Epigenetic Plasticity and Gene Regulatory Network Rewiring. Int. J. Mol. Sci. 2022, 23, 9691. [Google Scholar] [CrossRef]
- Yang, H.J.; Xue, J.M.; Li, J.; Wan, L.H.; Zhu, Y.X. Identification of key genes and pathways of diagnosis and prognosis in cervical cancer by bioinformatics analysis. Mol. Genet. Genom. Med. 2020, 8, e1200. [Google Scholar] [CrossRef]
- Yan, G.; Liu, Z. Identification of differentially expressed genes in hepatocellular carcinoma by integrated bioinformatic analysis. bioRxiv 2019, 570846. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, J.; Yan, K.; Lin, J.; Zheng, Z.; Bi, J. Identification of core genes associated with prostate cancer progression and outcome via bioinformatics analysis in multiple databases. PeerJ 2020, 8, e8786. [Google Scholar] [CrossRef]
- Anatskaya, O.V.; Vinogradov, A.E.; Vainshelbaum, N.M.; Giuliani, A.; Erenpreisa, J. Phylostratic shift of whole-genome duplications in normal mammalian tissues towards unicellularity is driven by developmental bivalent genes and reveals a link to cancer. Int. J. Mol. Sci. 2020, 21, 8759. [Google Scholar] [CrossRef]
- Kosugi, S.; Momozawa, Y.; Liu, X.; Terao, C.; Kubo, M.; Kamatani, Y. Comprehensive evaluation of structural variation detection algorithms for whole genome sequencing. Genome Biol. 2019, 20, 117. [Google Scholar] [CrossRef]
- Rahman, M.M.; Islam, M.R.; Rahman, F.; Rahaman, M.S.; Khan, M.S.; Abrar, S.; Ray, T.K.; Uddin, M.B.; Kali, M.S.K.; Dua, K.; et al. Emerging promise of computational techniques in anti-cancer research: At a glance. Bioengineering 2022, 9, 335. [Google Scholar] [CrossRef] [PubMed]
- Lobato-Delgado, B.; Priego-Torres, B.; Sanchez-Morillo, D. Combining Molecular, Imaging, and Clinical Data Analysis for Predicting Cancer Prognosis. Cancers 2022, 14, 3215. [Google Scholar] [CrossRef]
- Collin, C.B.; Gebhardt, T.; Golebiewski, M.; Karaderi, T.; Hillemanns, M.; Khan, F.M.; Salehzadeh-Yazdi, A.; Kirschner, M.; Krobitsch, S.; EU-STANDS4PM consortium; et al. Computational models for clinical applications in personalized medicine—Guidelines and recommendations for data integration and model validation. J. Pers. Med. 2022, 12, 166. [Google Scholar] [CrossRef] [PubMed]
- Shao, D.; Dai, Y.; Li, N.; Cao, X.; Zhao, W.; Cheng, L.; Rong, Z.; Huang, L.; Wang, Y.; Zhao, J. Artificial intelligence in clinical research of cancers. Brief. Bioinform. 2022, 23, bbab523. [Google Scholar] [CrossRef] [PubMed]
- Adibi, R.; Moein, S.; Gheisari, Y. Cisplatin resistant ovarian cancer cells reveal a polyploid phenotype with remarkable activation of nuclear processes. Res. Sq. 2021, 1, rs-440506. [Google Scholar]
- Quinton, R.J.; DiDomizio, A.; Vittoria, M.A.; Kotýnková, K.; Ticas, C.J.; Patel, S.; Koga, Y.; Vakhshoorzadeh, J.; Hermance, N.; Kuroda, T.; et al. Whole-genome doubling confers unique genetic vulnerabilities on tumour cells. Nature 2021, 590, 492–497. [Google Scholar] [CrossRef] [PubMed]
- Rohnalter, V.; Roth, K.; Finkernagel, F.; Adhikary, T.; Obert, J.; Dorzweiler, K.; Bensberg, M.; Müller-Brüsselbach, S.; Müller, R. A multi-stage process including transient polyploidization and EMT precedes the emergence of chemoresistent ovarian carcinoma cells with a dedifferentiated and pro-inflammatory secretory phenotype. Oncotarget 2015, 6, 40005. [Google Scholar] [CrossRef]
- Alwash, B.H.; Al-Rubaye, J.R.A.; Alaaraj, M.M.; Ebrahim, A.Y. Control of Cytoskeletal Dynamics in Cancer through a Combination of Cytoskeletal Components. In Cancer Bioinformatics; Intechopen: London, UK, 2022; Chapter 3; pp. 167–211. [Google Scholar]
- Anatskaya, O.V.; Erenpreisa, J.A.; Nikolsky, N.N.; Vinogradov, A.E. Pairwise comparison of mammalian transcriptomes associated with the effect of polyploidy on the expression activity of developmental gene modules. Cell Tissue Biol. 2016, 10, 122–132. [Google Scholar] [CrossRef]
- Kimmel, G.J.; Dane, M.; Heiser, L.M.; Altrock, P.M.; Andor, N. Integrating mathematical modeling with high-throughput imaging explains how polyploid populations behave in nutrient-sparse environments. Cancer Res. 2020, 80, 5109–5120. [Google Scholar] [CrossRef]
- Potapova, T.A.; Seidel, C.W.; Box, A.C.; Rancati, G.; Li, R. Transcriptome analysis of tetraploid cells identifies cyclin D2 as a facilitator of adaptation to genome doubling in the presence of p53. Mol. Biol. Cell 2016, 27, 3065–3084. [Google Scholar] [CrossRef]
- Buhler, A.; Kruger, R.; Monavari, M.; Fuentes-Chandia, M.; Palmisano, R.; Schodel, J.; Boccaccini, A.R.; Boberhoff, A.K.; Kappelmann-Fenzl, M.; Letort, G.; et al. When Mechanical Stress Matters: Generation of Polyploid Giant Cancer Cells in Tumor-like Microcapsules. BioRxiv 2022. [Google Scholar] [CrossRef]
- Rantala, J.K.; Edgren, H.; Lehtinen, L.; Wolf, M.; Kleivi, K.; Vollan, H.K.; Aaltola, A.R.; Laasola, P.; Kilpinen, S.; Saviranta, P.; et al. Integrative functional genomics analysis of sustained polyploidy phenotypes in breast cancer cells identifies an oncogenic profile for GINS2. Neoplasia 2010, 12, 877-IN14. [Google Scholar] [CrossRef] [PubMed]
- Fu, F.; Chen, L.; Yang, X.; Fan, L.; Zhang, M.; Chen, S.; Zheng, M.; Gao, M.; Zhang, S. SPLK4 is a key molecule in the formation of PGCCs and promotes invasion and migration of progeny cells derived from PGCCs. J. Cancer 2022, 13, 2954. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Zhang, K.; Li, Z.; Zhang, H.; Fu, F.; Fu, J.; Zheng, M.; Zhang, S. High migration and invasion ability of PGCCs and their daughter cells associated with the nuclear localization of S100A10 modified by SUMOylation. Front. Cell Dev. Biol. 2021, 9, 696871. [Google Scholar] [CrossRef] [PubMed]
- Gomaa, A.; Peng, D.; Chen, Z.; Soutto, M.; Abouelezz, K.; Corvalan, A.; El-Rifai, W. Epigenetic regulation of AURKA by miR-4715-3p in upper gastrointestinal cancers. Sci. Rep. 2019, 9, 1–11. [Google Scholar] [CrossRef]
- Salmina, K.; Gerashchenko, B.I.; Hausmann, M.; Vainshelbaum, N.M.; Zayakin, P.; Erenpreiss, J.; Freivalds, T.; Cragg, M.S.; Erenpreisa, J. When three isn’ta crowd: A digyny concept for treatment-resistant, near-triploid human cancers. Genes 2019, 10, 2019. [Google Scholar] [CrossRef] [PubMed]
- Hambali, M.A.; Oladele, T.O.; Adewole, K.S. Microarray cancer feature selection: Review, challenges and research directions. Int. J. Cogn. Comput. Eng. 2020, 1, 78–97. [Google Scholar] [CrossRef]
- Sebastian, A.M.; Peter, D. Artificial Intelligence in Cancer Research: Trends, Challenges and Future Directions. Life 2022, 12, 1991. [Google Scholar] [CrossRef]
- Csikasz-Nagy, A. Computational systems biology of the cell cycle. Brief. Bioinform. 2009, 10, 424–434. [Google Scholar] [CrossRef]
- Malhotra, A.; Wang, Y.; Waters, J.; Chen, K.; Meric-Bernstam, F.; Hall, I.M.; Navin, N.E. Ploidy-Seq: Inferring mutational chronology by sequencing polyploid tumor subpopulations. Genome Med. 2015, 7, 6. [Google Scholar] [CrossRef]
- Peng, H.; Tan, H.; Zhao, W.; Jin, G.; Sharma, S.; Xing, F.; Watabe, K.; Zhou, X. Computational systems biology in cancer brain metastasis. Front. Biosci. 2016, 8, 169. [Google Scholar]
- Arjmand, B.; Hamidpour, S.K.; Tayanloo-Beik, A.; Goodarzi, P.; Aghayan, H.R.; Adibi, H.; Larijani, B. Machine learning: A new prospect in multi-omics data analysis of cancer. Front. Genet. 2022, 13, 76. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.B.; Okuno, Y. Systems biology and systems chemistry: New directions for drug discovery. Chem. Biol. 2012, 19, 23–28. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, D.; Thakur, A.D.; Thakur, A. A feedback-based manoeuvre planner for nonprehensile magnetic micromanipulation of large microscopic biological objects. Robot. Auton. Syst. 2022, 148, 103941. [Google Scholar] [CrossRef]
- Olsen, T.R.; Talla, P.; Furnari, J.; Bruce, J.N.; Canoll, P.; Zha, S.; Sims, P.A. Scalable co-sequencing of RNA and DNA from individual nuclei. bioRxiv 2023, 2023, 527940. [Google Scholar]
- Zhang, C.Z.; Pellman, D. Cancer genomic rearrangements and copy number alterations from errors in cell division. Annu. Rev. Cancer Biol. 2022, 6, 245–268. [Google Scholar] [CrossRef]
- Funnell, T.; O’Flanagan, C.H.; Williams, M.J.; McPherson, A.; McKinney, S.; Kabeer, F.; Lee, H.; Salehi, S.; Vázquez-García, I.; Shi, H.; et al. Single-cell genomic variation induced by mutational processes in cancer. Nature 2022, 612, 106–115. [Google Scholar] [CrossRef]
- Erenpreisa, J.; Giuliani, A.; Yoshikawa, K.; Falk, M.; Hildenbrand, G.; Salmina, K.; Freivalds, T.; Vainshelbaum, N.; Weidner, J.; Sievers, A.; et al. Spatial-Temporal Genome Regulation in Stress-Response and Cell-Fate Change. Int. J. Mol. Sci. 2023, 24, 2658. [Google Scholar] [CrossRef]
- Mirzayans, R.; Murray, D. What Are the Reasons for Continuing Failures in Cancer Therapy? Are Misleading/Inappropriate Preclinical Assays to Be Blamed? Might Some Modern Therapies Cause More Harm than Benefit? Int. J. Mol. Sci. 2022, 23, 13217. [Google Scholar] [CrossRef]
- Kim, Y.K.; Cho, B.; Cook, D.P.; Trcka, D.; Wrana, J.L.; Ramalho-Santos, M. Absolute scaling of single-cell transcriptomes identifies pervasive hypertranscription in adult stem and progenitor cells. Cell Rep. 2023, 42, 111978. [Google Scholar] [CrossRef]
| Cancer Type | Study Description | Author |
|---|---|---|
| Breast Cancer | Computational studies about differentially expressed genes, omics, and systems biology data provided the identification of new gene signatures-NUSAP1, MELK, CENPF, TOP2A, and PPARG-genes related to chemoresistance, potential biomarkers, and new therapeutic targets associated with tumor polyploidy. | Alam et al. [119], Kaur et al. [120], Mukherjee et al. [121], Yadav et al. [122], and Yang et al. [123] |
| Prostate Cancer | Systems biology and bioinformatics identification of key genes as biomarkers of diagnostics, prognosis, and treatment (EGFR, MYC, VEGFA, PTEN). | Khan et al. [124] |
| Melanoma | Mathematical modeling after therapeutic regression was able to identify a triphasic signaling pathway in tumor regression. | Kumari et al. [125] |
| Pan-cancer | Identified the relationship between SLC7A11 expression and tumor microenvironment, using bioinformatics in 20 tumors. | Lin et al. [126] |
| Colorectal and Uterine Cancer | Found potential genes and their signaling pathways using bioinformatics and systems biology. | Nguyen et al. [127] andReza et al. [128] |
| Lung Cancer | An integrated systems biology and bioinformatics approach provided the detection of genetic correlations between COVID-19 and small cell lung cancer and the interaction of biological pathways associated with tumor polyploidization. | Roudi et al. [129] and Zhuang et al. [130] |
| Cancer Type | Study Description and Key Findings | Author |
|---|---|---|
| Ovary Cancer | Computational research based on NGS, total RNA, and microarray sequencing, using primary tumors, cell lines, and tumor chemoresistance highlighted the association with tumor polyploidy. | Adibi, Moein and Gheisari [148], Quinton et al. [149], Rohnalter et al. [150] |
| Lung Cancer | Investigated the role of natural and synthetic mutations in tumor migration and invasion. | Alwash et al. [151] |
| Several types of cancer | Molecular mechanisms associated with polyploidy, cell plasticity, unicellularity, energy metabolism, tumor DNA damage in tumors, phylogenetic approaches, and molecular modeling were used study the effects of PGCCs on gene expression, tumor microenvironment, and p53. | Anatskaya and Vinogradov [138], Anatskaya et al. [152], Anatskaya et al. [142], Kimmel et al. [153], and Potapova et al. [154] |
| Breast Cancer | Studied formation of PGCCs by mechanical stress.In silico studies to detect gene signatures related to PGCC formation. | Buehler et al. [155] and Rantala et al. [156] |
| Colorectal Cancer | S100A10 expression changes caused by differential SUMOylation during the migration of PGCCs. | Fu et al. [157] and Zhao et al. [158] |
| Gastric Cancer | MiRNA sequencing to study the role of epigenetic in regulation of Aurora kinase A (AURKA) expression. | Gomaa et al. [159] |
| Cervical cancer, Breast Cancer and Burkitt Lymphoma | In silico studies using Mitelman’s database to uncover PGCC’s role in DNA repair, genetic variation, and tumor survival. | Salmina et al. [160] |
| Prostate Cancer and Melanoma | Transcriptome analysis of PGCCs after ASAH1 treatment leading to cholesterol metabolism alterations. | White-Gilbertson et al. [80] |
| Nasopharyngeal Cancer | Induction of PGCC by autophagy. | You et al. [86] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Casotti, M.C.; Meira, D.D.; Zetum, A.S.S.; Araújo, B.C.d.; Silva, D.R.C.d.; Santos, E.d.V.W.d.; Garcia, F.M.; Paula, F.d.; Santana, G.M.; Louro, L.S.; et al. Computational Biology Helps Understand How Polyploid Giant Cancer Cells Drive Tumor Success. Genes 2023, 14, 801. https://doi.org/10.3390/genes14040801
Casotti MC, Meira DD, Zetum ASS, Araújo BCd, Silva DRCd, Santos EdVWd, Garcia FM, Paula Fd, Santana GM, Louro LS, et al. Computational Biology Helps Understand How Polyploid Giant Cancer Cells Drive Tumor Success. Genes. 2023; 14(4):801. https://doi.org/10.3390/genes14040801
Chicago/Turabian StyleCasotti, Matheus Correia, Débora Dummer Meira, Aléxia Stefani Siqueira Zetum, Bruno Cancian de Araújo, Danielle Ribeiro Campos da Silva, Eldamária de Vargas Wolfgramm dos Santos, Fernanda Mariano Garcia, Flávia de Paula, Gabriel Mendonça Santana, Luana Santos Louro, and et al. 2023. "Computational Biology Helps Understand How Polyploid Giant Cancer Cells Drive Tumor Success" Genes 14, no. 4: 801. https://doi.org/10.3390/genes14040801
APA StyleCasotti, M. C., Meira, D. D., Zetum, A. S. S., Araújo, B. C. d., Silva, D. R. C. d., Santos, E. d. V. W. d., Garcia, F. M., Paula, F. d., Santana, G. M., Louro, L. S., Alves, L. N. R., Braga, R. F. R., Trabach, R. S. d. R., Bernardes, S. S., Louro, T. E. S., Chiela, E. C. F., Lenz, G., Carvalho, E. F. d., & Louro, I. D. (2023). Computational Biology Helps Understand How Polyploid Giant Cancer Cells Drive Tumor Success. Genes, 14(4), 801. https://doi.org/10.3390/genes14040801






