A Case Report of a Feto-Placental Mosaicism Involving a Segmental Aneuploidy: A Challenge for Genome Wide Screening by Non-Invasive Prenatal Testing of Cell-Free DNA in Maternal Plasma
Abstract
1. Introduction
2. Case Report
3. Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Norton, M.E.; Jacobsson, B.; Swamy, G.K.; Laurent, L.C.; Ranzini, A.C.; Brar, H.; Tomlinson, M.W.; Pereira, L.; Spitz, J.L.; Hollemon, D.; et al. Cell-free DNA Analysis for Noninvasive Examination of Trisomy. N. Engl. J. Med. 2015, 372, 1589–1597. [Google Scholar] [CrossRef] [PubMed]
- Taylor-Phillips, S.; Freeman, K.; Geppert, J.; Agbebiyi, A.; Uthman, O.A.; Madan, J.; Clarke, A.; Quenby, S.; Clarke, A. Accuracy of non-invasive prenatal testing using cell-free DNA for detection of Down, Edwards and Patau syndromes: A systematic review and meta-analysis. BMJ Open 2016, 6, e010002. [Google Scholar] [CrossRef] [PubMed]
- Mackie, F.L.; Hemming, K.; Allen, S.; Morris, R.K.; Kilby, M.D. The accuracy of cell-free fetal DNA-based non-invasive prenatal testing in singleton pregnancies: A systematic review and bivariate meta-analysis. BJOG Int. J. Obstet. Gynaecol. 2017, 124, 32–46. [Google Scholar] [CrossRef]
- Dar, P.; Curnow, K.J.; Gross, S.J.; Hall, M.P.; Stosic, M.; Demko, Z.; Zimmermann, B.; Hill, M.; Sigurjonsson, S.; Ryan, A.; et al. Clinical experience and follow-up with large scale single-nucleotide polymorphism–based noninvasive prenatal aneuploidy testing. Am. J. Obstet. Gynecol. 2014, 211, 527.e1–527.e17. [Google Scholar] [CrossRef]
- Zhang, H.; Gao, Y.; Jiang, F.; Fu, M.; Yuan, Y.; Guo, Y.; Zhu, Z.; Lin, M.; Liu, Q.; Tian, Z.; et al. Non-invasive prenatal testing for trisomies 21, 18 and 13: Clinical experience from 146 958 pregnancies. Ultrasound Obstet. Gynecol. 2015, 45, 530–538. [Google Scholar] [CrossRef]
- Liehr, T. False-positives and false-negatives in non-invasive prenatal testing (NIPT): What can we learn from a meta-analyses on > 750,000 tests? Mol. Cytogenet. 2022, 15, 36. [Google Scholar] [CrossRef] [PubMed]
- Pescia, G.; Guex, N.; Iseli, C.; Brennan, L.; Osteras, M.; Xenarios, I.; Farinelli, L.; Conrad, B. Cell-free DNA testing of an extended range of chromosomal anomalies: Clinical experience with 6,388 consecutive cases. Anesthesia Analg. 2017, 19, 169–175. [Google Scholar] [CrossRef] [PubMed]
- Pertile, M.D.; Flowers, N.; Vavrek, D.; Andrews, D.; Kalista, T.; Craig, A.; Deciu, C.; Duenwald, S.; Meier, K.; Bhatt, S. Performance of a Paired-End Sequencing-Based Noninvasive Prenatal Screening Test in the Detection of Genome-Wide Fetal Chromosomal Anomalies. Clin. Chem. 2021, 67, 1210–1219. [Google Scholar] [CrossRef] [PubMed]
- Soster, E.; Boomer, T.; Hicks, S.; Caldwell, S.; Dyr, B.; Chibuk, J.; Almasri, E. Three years of clinical experience with a genome-wide cfDNA screening test for aneuploidies and copy-number variants. Anesthesia Analg. 2021, 23, 1349–1355. [Google Scholar] [CrossRef]
- Mossfield, T.; Soster, E.; Menezes, M.; Agenbag, G.; Dubois, M.-L.; Gekas, J.; Hardy, T.; Jurkowska, M.; Kleinfinger, P.; Loggenberg, K.; et al. Multisite assessment of the impact of cell-free DNA-based screening for rare autosomal aneuploidies on pregnancy management and outcomes. Front. Genet. 2022, 13, 975987. [Google Scholar] [CrossRef]
- Van Prooyen Schuurman, L.; Sistermans, E.A.; Van Opstal, D.; Henneman, L.; Bekker, M.N.; Bax, C.J.; Pieters, M.J.; Bouman, K.; de Munnik, S.; den Hollander, N.S.; et al. Clinical impact of additional findings detected by genome-wide non-invasive prenatal testing: Follow-up results of the TRIDENT-2 study. Am. J. Hum. Genet. 2022, 109, 1140–1152. [Google Scholar] [CrossRef] [PubMed]
- Rafalko, J.; Soster, E.; Caldwell, S.; Almasri, E.; Westover, T.; Weinblatt, V.; Cacheris, P. Genome-wide cell-free DNA screening: A focus on copy-number variants. Anesthesia Analg. 2021, 23, 1847–1853. [Google Scholar] [CrossRef] [PubMed]
- Grati, F.R.; Malvestiti, F.; Ferreira, J.C.; Bajaj, K.; Gaetani, E.; Agrati, C.; Grimi, B.; Dulcetti, F.; Ruggeri, A.M.; De Toffol, S.; et al. Fetoplacental mosaicism: Potential implications for false-positive and false-negative noninvasive prenatal screening results. Anesthesia Analg. 2014, 16, 620–624. [Google Scholar] [CrossRef] [PubMed]
- Hartwig, T.S.; Ambye, L.; Sørensen, S.; Jørgensen, F.S. Discordant non-invasive prenatal testing (NIPT)—A systematic review. Prenat. Diagn. 2017, 37, 527–539. [Google Scholar] [CrossRef]
- Cai, Y.-H.; Yao, G.-Y.; Chen, L.-J.; Gan, H.-Y.; Ye, C.-S.; Yang, X.-X. The Combining Effects of Cell-Free Circulating Tumor DNA of Breast Tumor to the Noninvasive Prenatal Testing Results: A Simulating Investigation. DNA Cell Biol. 2018, 37, 626–633. [Google Scholar] [CrossRef]
- Koumbaris, G.; Achilleos, A.; Nicolaou, M.; Loizides, C.; Tsangaras, K.; Kypri, E.; Mina, P.; Sismani, C.; Velissariou, V.; Christopoulou, G.; et al. Targeted capture enrichment followed by NGS: Development and validation of a single comprehensive NIPT for chromosomal aneuploidies, microdeletion syndromes and monogenic diseases. Mol. Cytogenet. 2019, 12, 48. [Google Scholar] [CrossRef]
- Malvestiti, F.; Agrati, C.; Grimi, B.; Pompilii, E.; Izzi, C.; Martinoni, L.; Gaetani, E.; Liuti, M.R.; Trotta, A.; Maggi, F.; et al. Interpreting mosaicism in chorionic villi: Results of a monocentric series of 1001 mosaics in chorionic villi with follow-up amniocentesis. Prenat. Diagn. 2015, 35, 1117–1127. [Google Scholar] [CrossRef]
- Grati, F.R.; Malvestiti, F.; Branca, L.; Agrati, C.; Maggi, F.; Simoni, G. Chromosomal mosaicism in the fetoplacental unit. Best Pract. Res. Clin. Obstet. Gynaecol. 2017, 42, 39–52. [Google Scholar] [CrossRef]
- Grati, F.R. Chromosomal Mosaicism in Human Feto-Placental Development: Implications for Prenatal Diagnosis. J. Clin. Med. 2014, 3, 809–837. [Google Scholar] [CrossRef]
- Levy, B.; Hoffmann, E.R.; McCoy, R.C.; Grati, F.R. Chromosomal mosaicism: Origins and clinical implications in preimplantation and prenatal diagnosis. Prenat. Diagn. 2021, 41, 631–641. [Google Scholar] [CrossRef]
- Eggenhuizen, G.M.; Go, A.; Koster, M.P.H.; Baart, E.B.; Galjaard, R.J. Confined placental mosaicism and the association with pregnancy outcome and fetal growth: A review of the literature. Hum. Reprod. Updat. 2021, 27, 885–903. [Google Scholar] [CrossRef]
- Brady, P.; Brison, N.; Van Den Bogaert, K.; de Ravel, T.; Peeters, H.; Van Esch, H.; Devriendt, K.; Legius, E.; Vermeesch, J.R. Clinical implementation of NIPT—Technical and biological challenges. Clin. Genet. 2015, 89, 523–530. [Google Scholar] [CrossRef] [PubMed]
- Van Opstal, D.; Eggenhuizen, G.M.; Joosten, M.; Diderich, K.; Govaerts, L.; Galjaard, R.; Go, A.; Knapen, M.; Boter, M.; Cheung, W.Y.; et al. Noninvasive prenatal testing as compared to chorionic villus sampling is more sensitive for the detection of confined placental mosaicism involving the cytotrophoblast. Prenat. Diagn. 2020, 40, 1338–1342. [Google Scholar] [CrossRef] [PubMed]
- Committee on Genetics Society for Maternal–Fetal Medicine. Committee Opinion No. 640. Obstet. Gynecol. 2015, 126, e31–e37. [Google Scholar] [CrossRef] [PubMed]
- Cherry, A.M.; Akkari, Y.M.; Barr, K.M.; Kearney, H.M.; Rose, N.C.; South, S.T.; Tepperberg, J.H.; Meck, J.M. Diagnostic cytogenetic testing following positive noninvasive prenatal screening results: A clinical laboratory practice resource of the American College of Medical Genetics and Genomics (ACMG). Anesthesia Analg. 2017, 19, 845–850. [Google Scholar] [CrossRef]
- Kleinfinger, P.; Lohmann, L.; Luscan, A.; Trost, D.; Bidat, L.; Debarge, V.; Castaigne, V.; Senat, M.-V.; Brechard, M.-P.; Guilbaud, L.; et al. Strategy for Use of Genome-Wide Non-Invasive Prenatal Testing for Rare Autosomal Aneuploidies and Unbalanced Structural Chromosomal Anomalies. J. Clin. Med. 2020, 9, 2466. [Google Scholar] [CrossRef]
- Borth, H.; Teubert, A.; Glaubitz, R.; Knippenberg, S.; Kutur, N.; Winkler, T.; Eiben, B. Analysis of cell-free DNA in a consecutive series of 13,607 routine cases for the detection of fetal chromosomal aneuploidies in a single center in Germany. Arch. Gynecol. Obstet. 2020, 303, 1407–1414. [Google Scholar] [CrossRef]
- La Verde, M.; De Falco, L.; Torella, A.; Savarese, G.; Savarese, P.; Ruggiero, R.; Conte, A.; Fico, V.; Torella, M.; Fico, A. Performance of cell-free DNA sequencing-based non-invasive prenatal testing: Experience on 36,456 singleton and multiple pregnancies. BMC Med. Genom. 2021, 14, 93. [Google Scholar] [CrossRef] [PubMed]
- Brison, N.; Neofytou, M.; Dehaspe, L.; Bayindir, B.; Van Den Bogaert, K.; Dardour, L.; Peeters, H.; Van Esch, H.; Van Buggenhout, G.; Vogels, A.; et al. Predicting fetoplacental chromosomal mosaicism during non-invasive prenatal testing. Prenat. Diagn. 2018, 38, 258–266. [Google Scholar] [CrossRef]
- Porter, S.; Wilson, E.; Tyler, X.; Warren, R.; Ffrench-Constant, C.; Pearson, J. A case of discordant related abnormal karyotypes from chorionic villi and amniocytes. Prenat. Diagn. 1999, 19, 887–890. [Google Scholar] [CrossRef]
- Brisset, S.; Aboura, A.; Audibert, F.; Costa, J.-M.; L’Herminé, A.C.; Gautier, V.; Frydman, R.; Tachdjian, G. Discordant prenatal diagnosis of trisomy 21 due to mosaic structural rearrangements of chromosome 21. Prenat. Diagn. 2003, 23, 461–469. [Google Scholar] [CrossRef]
- Soler, A.; Sánchez, A.; Carrió, A.; Badenas, C.; Milà, M.; Borrell, A. Fetoplacental discrepancy involving structural abnormalities of chromosome 8 detected by prenatal diagnosis. Prenat. Diagn. 2003, 23, 319–322. [Google Scholar] [CrossRef]
- Ledbetter, D.H.; Zachary, J.M.; Simpson, J.L.; Golbus, M.S.; Pergament, E.; Jackson, L.; Mahoney, M.J.; Desnick, R.J.; Schulman, J.; Copeland, K.L.; et al. Cytogenetic results from the U.S. collaborative study on CVS. Prenat. Diagn. 1992, 12, 317–345. [Google Scholar] [CrossRef]
- Smidt-Jensen, S.; Lind, A.-M.; Permin, M.; Zachary, J.M.; Lundsteen, C.; Philip, J. Cytogenetic analysis of 2928 CVS samples and 1075 amniocenteses from randomized studies. Prenat. Diagn. 1993, 13, 723–740. [Google Scholar] [CrossRef]
- Grati, F.R. Implications of fetoplacental mosaicism on cell-free DNA testing: A review of a common biological phenomenon. Ultrasound Obstet. Gynecol. 2016, 48, 415–423. [Google Scholar] [CrossRef]
- Grati, F.R.; Bajaj, K.; Malvestiti, F.; Agrati, C.; Grimi, B.; Malvestiti, B.; Pompilii, E.; Maggi, F.; Gross, S.; Simoni, G.; et al. The type of feto-placental aneuploidy detected by cfDNA testing may influence the choice of confirmatory diagnostic procedure. Prenat. Diagn. 2015, 35, 994–998. [Google Scholar] [CrossRef] [PubMed]
- Kalousek, D.K.; Dill, F.J. Chromosomal Mosaicism Confined to the Placenta in Human Conceptions. Science 1983, 221, 665–667. [Google Scholar] [CrossRef] [PubMed]
- Eichler, E.E. Recent duplication, domain accretion and the dynamic mutation of the human genome. Trends Genet. 2001, 17, 661–669. [Google Scholar] [CrossRef] [PubMed]
- International Human Genome Sequencing Consortium; Whitehead Institute for Biomedical Research, Center for Genome Research; Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Devon, K.; Dewar, K.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar] [CrossRef] [PubMed]
- Venter, J.C.; Adams, M.D.; Myers, E.W.; Li, P.W.; Mural, R.J.; Sutton, G.G.; Smith, H.O.; Yandell, M.; Evans, C.A.; Holt, R.A.; et al. The Sequence of the Human Genome. Science 2001, 291, 1304–1351. [Google Scholar] [CrossRef] [PubMed]
- Chavli, E.; van den Born, M.; Eleveld, C.; Boter, M.; van Marion, R.; Hoefsloot, L.; Laven, J.; Baart, E.; Van Opstal, D. Chromosomal mosaicism in human blastocysts: A cytogenetic comparison of trophectoderm and inner cell mass after next-generation sequencing. Reprod. Biomed. Online 2022, 45, 867–877. [Google Scholar] [CrossRef] [PubMed]
- McCoy, R.C. Mosaicism in Preimplantation Human Embryos: When Chromosomal Abnormalities Are the Norm. Trends Genet. 2017, 33, 448–463. [Google Scholar] [CrossRef]
- Zollino, M.; Lecce, R.; Fischetto, R.; Murdolo, M.; Faravelli, F.; Selicorni, A.; Buttè, C.; Memo, L.; Capovilla, G.; Neri, G. Mapping the Wolf-Hirschhorn Syndrome Phenotype Outside the Currently Accepted WHS Critical Region and Defining a New Critical Region, WHSCR-2. Am. J. Hum. Genet. 2003, 72, 590–597. [Google Scholar] [CrossRef]
- Battaglia, A.; Filippi, T.; Carey, J.C. Update on the clinical features and natural history of Wolf-Hirschhorn (4p-) syndrome: Experience with 87 patients and recommendations for routine health supervision. Am. J. Med. Genet. Part C: Semin. Med. Genet. 2008, 148C, 246–251. [Google Scholar] [CrossRef]
- Zollino, M.; Di Stefano, C.; Zampino, G.; Mastroiacovo, P.; Wright, T.J.; Sorge, G.; Selicorni, A.; Tenconi, R.; Battaglia, A.; Di Rocco, M.; et al. Genotype-phenotype correlations and clinical diagnostic criteria in Wolf-Hirschhorn syndrome. Am. J. Med. Genet. 2000, 94, 254–261. [Google Scholar] [CrossRef]
- Tachdjian, G.; Fondacci, G.; TAPlA, S.; Huten, Y.; Blot, P.; Nessmann, C. The Wolf-Hirschhorn syndrome in fetuses. Clin. Genet. 2008, 42, 281–287. [Google Scholar] [CrossRef] [PubMed]
- Mayo, S.; Gómez-Manjón, I.; Atencia, G.; Moreno-Izquierdo, A.; Escribano, D.; Fernández-Martínez, F.J. Noninvasive prenatal testing: How far can we reach detecting fetal copy number variations. Eur. J. Obstet. Gynecol. Reprod. Biol. 2022, 272, 150–155. [Google Scholar] [CrossRef] [PubMed]


| Variable | Value |
|---|---|
| fetal_fraction | 0.108383051 |
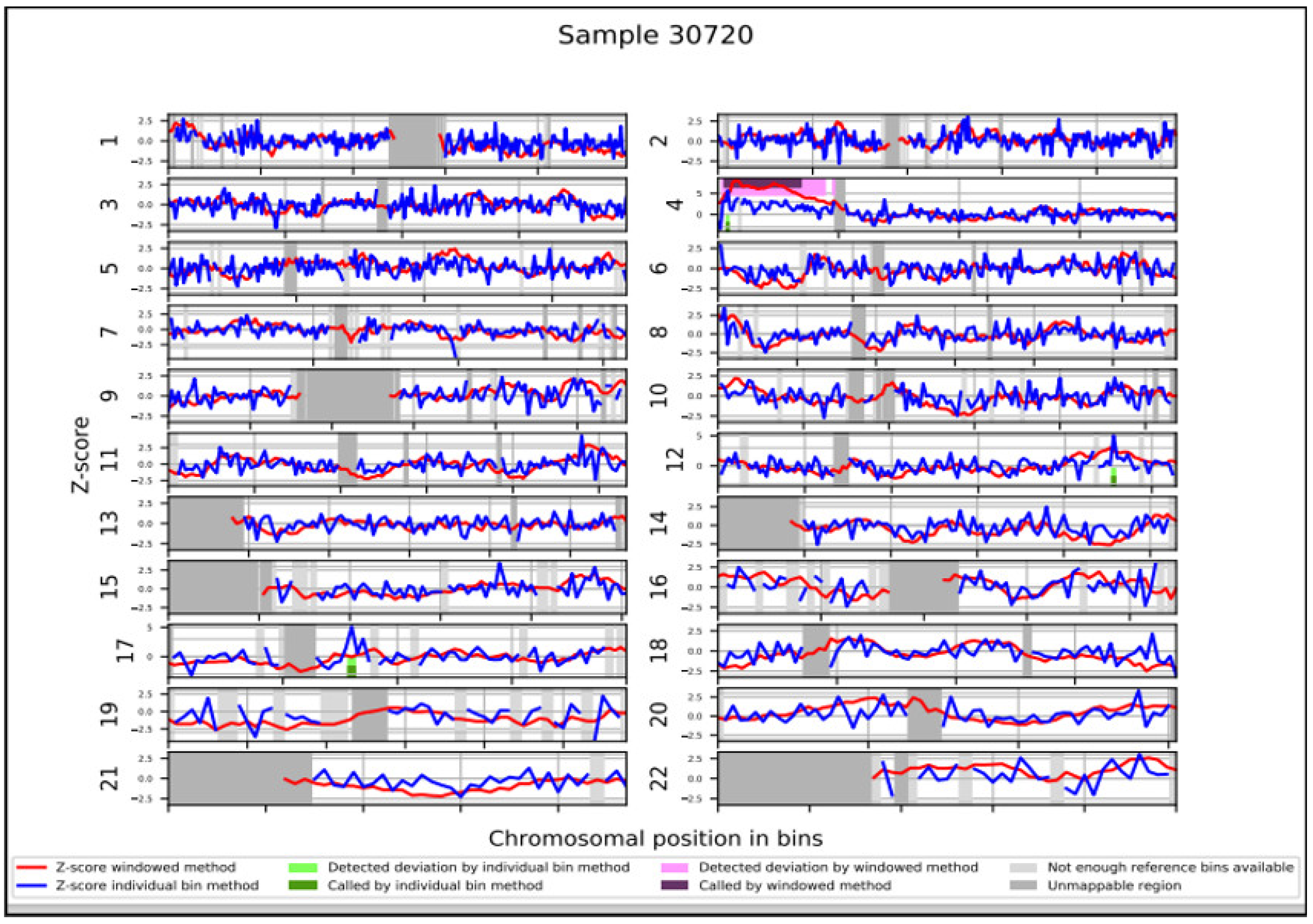
| region_classification | DETECTED: dup(4)(p16.3p12) |
| chromosome | chr4 |
| start_base | 2000001 |
| end_cytoband | p12 |
| region_size_mb | 44.1 |
| region_llr_trisomy | 148.3858609 |
| region_llr_monosomy | NA |
| region_t_stat_long_reads | 16.18982321 |
| region_mosaic_ratio | 0.792664396 |
| region_mosaic_llr_trisomy | 148.8731223 |
| region_mosaic_llr_monosomy | NA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Falco, L.; Vitiello, G.; Savarese, G.; Suero, T.; Ruggiero, R.; Savarese, P.; Ianniello, M.; Petrillo, N.; Bruno, M.; Legnante, A.; et al. A Case Report of a Feto-Placental Mosaicism Involving a Segmental Aneuploidy: A Challenge for Genome Wide Screening by Non-Invasive Prenatal Testing of Cell-Free DNA in Maternal Plasma. Genes 2023, 14, 668. https://doi.org/10.3390/genes14030668
De Falco L, Vitiello G, Savarese G, Suero T, Ruggiero R, Savarese P, Ianniello M, Petrillo N, Bruno M, Legnante A, et al. A Case Report of a Feto-Placental Mosaicism Involving a Segmental Aneuploidy: A Challenge for Genome Wide Screening by Non-Invasive Prenatal Testing of Cell-Free DNA in Maternal Plasma. Genes. 2023; 14(3):668. https://doi.org/10.3390/genes14030668
Chicago/Turabian StyleDe Falco, Luigia, Giuseppina Vitiello, Giovanni Savarese, Teresa Suero, Raffaella Ruggiero, Pasquale Savarese, Monica Ianniello, Nadia Petrillo, Mariasole Bruno, Antonietta Legnante, and et al. 2023. "A Case Report of a Feto-Placental Mosaicism Involving a Segmental Aneuploidy: A Challenge for Genome Wide Screening by Non-Invasive Prenatal Testing of Cell-Free DNA in Maternal Plasma" Genes 14, no. 3: 668. https://doi.org/10.3390/genes14030668
APA StyleDe Falco, L., Vitiello, G., Savarese, G., Suero, T., Ruggiero, R., Savarese, P., Ianniello, M., Petrillo, N., Bruno, M., Legnante, A., Passaretti, F. F., Ardisia, C., Di Spiezio Sardo, A., & Fico, A. (2023). A Case Report of a Feto-Placental Mosaicism Involving a Segmental Aneuploidy: A Challenge for Genome Wide Screening by Non-Invasive Prenatal Testing of Cell-Free DNA in Maternal Plasma. Genes, 14(3), 668. https://doi.org/10.3390/genes14030668






