Approach to Cohort-Wide Re-Analysis of Exome Data in 1000 Individuals with Neurodevelopmental Disorders
Abstract
1. Introduction
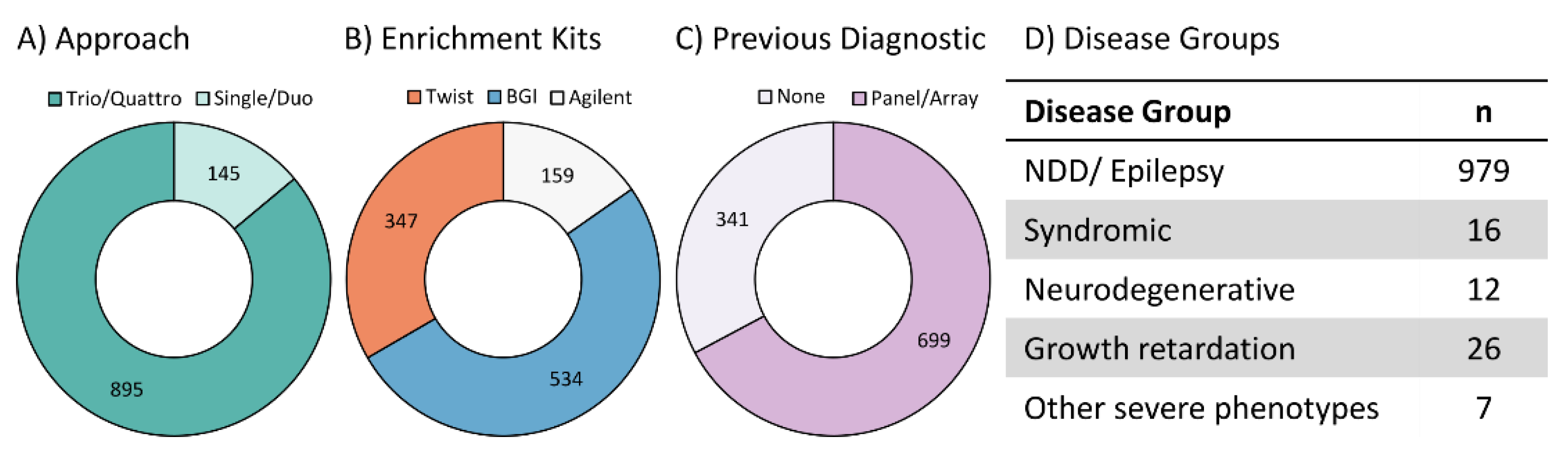
2. Materials and Methods
3. Results
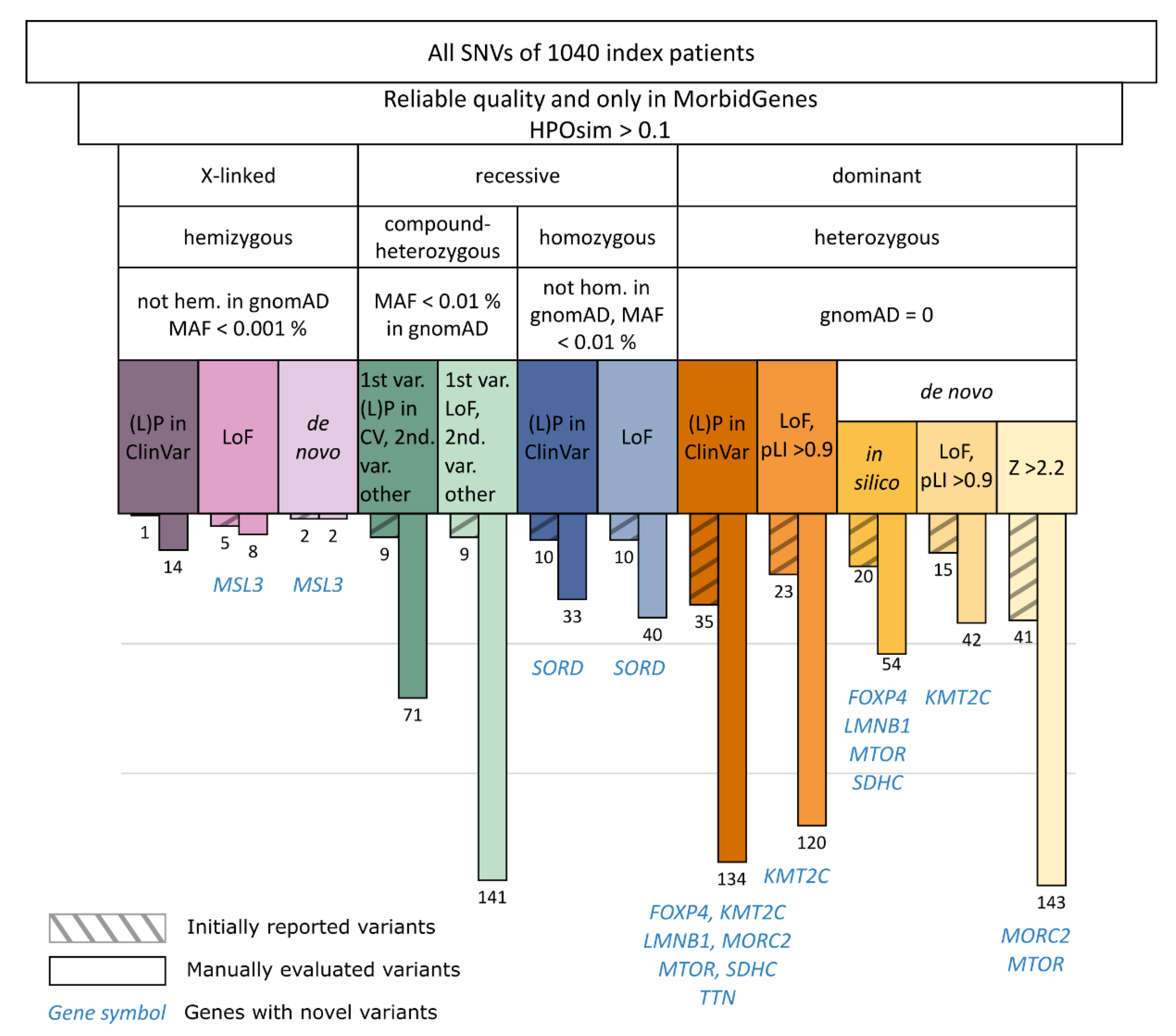
3.1. Manual Evaluation of Filtered Variants
3.2. Novel Diagnoses through Re-Analysis
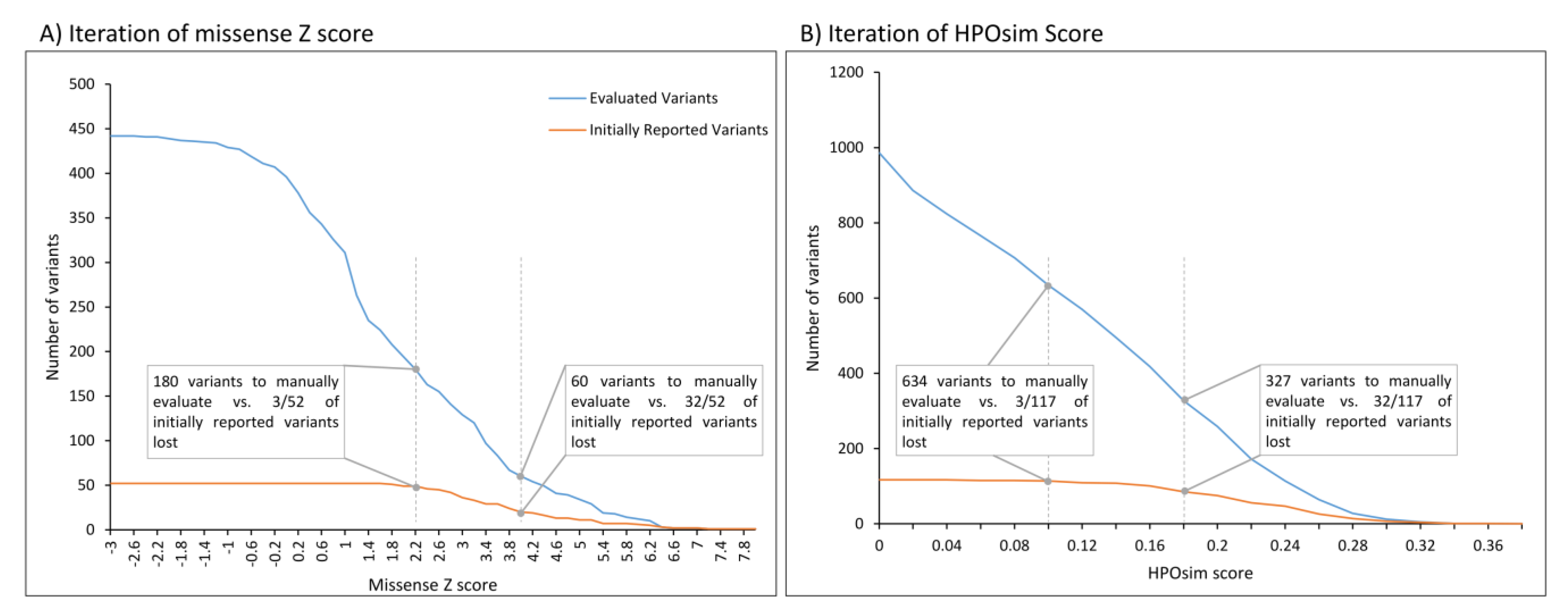
3.3. Sensitivity and Filter Adjustments
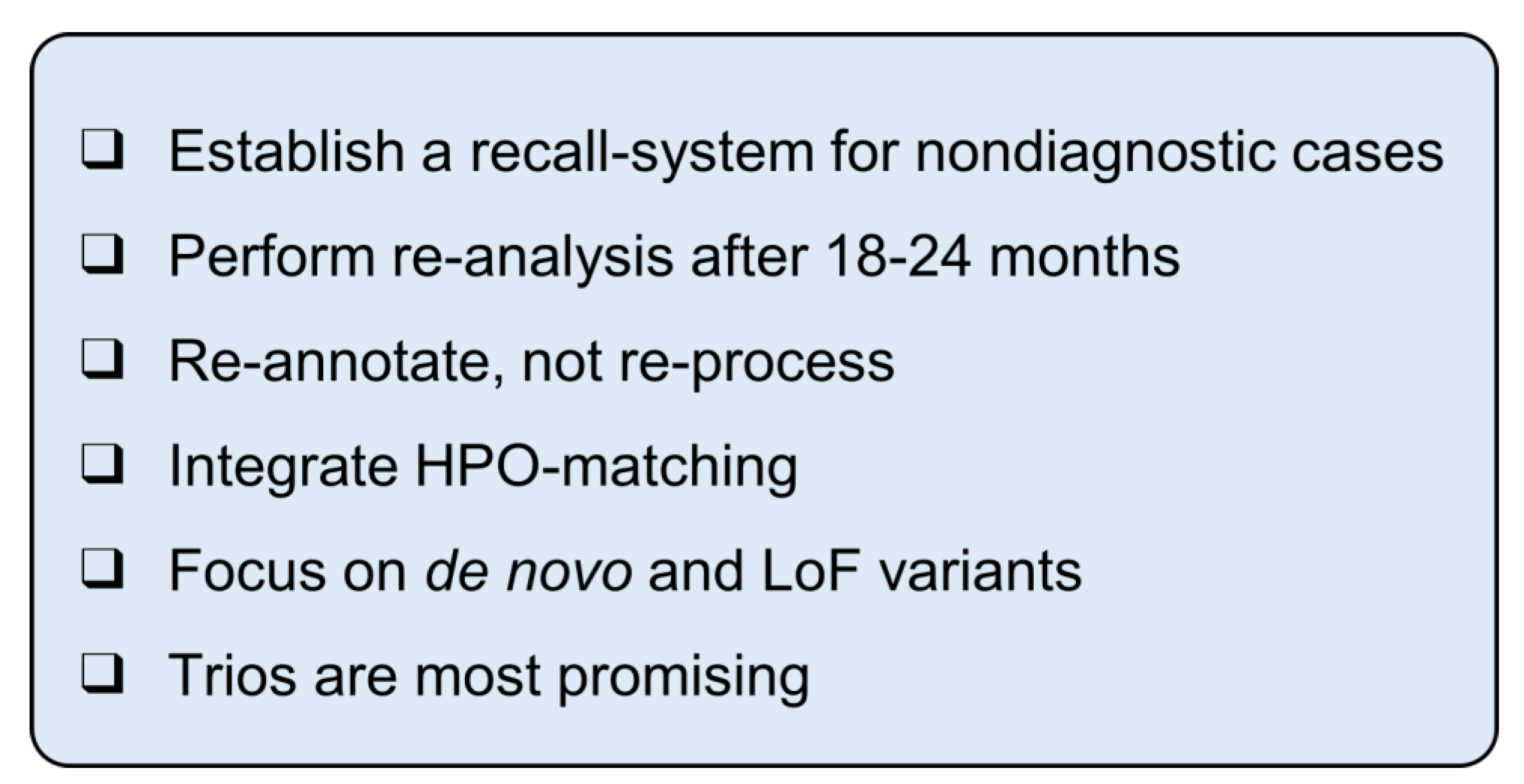
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Seo, G.H.; Lee, H.; Lee, J.; Han, H.; Cho, Y.K.; Kim, M.; Choi, Y.; Choi, J.; Choi, I.H.; Rhie, S.; et al. Diagnostic Performance of Automated, Streamlined, Daily Updated Exome Analysis in Patients with Neurodevelopmental Delay. Mol. Med. 2022, 28, 38. [Google Scholar] [CrossRef] [PubMed]
- Landrum, M.J.; Chitipiralla, S.; Brown, G.R.; Chen, C.; Gu, B.; Hart, J.; Hoffman, D.; Jang, W.; Kaur, K.; Liu, C.; et al. ClinVar: Improvements to Accessing Data. Nucleic Acids Res. 2020, 48, D835–D844. [Google Scholar] [CrossRef] [PubMed]
- Deignan, J.L.; Chung, W.K.; Kearney, H.M.; Monaghan, K.G.; Rehder, C.W.; Chao, E.C. Points to Consider in the Reevaluation and Reanalysis of Genomic Test Results: A Statement of the American College of Medical Genetics and Genomics (ACMG). Genet. Med. 2019, 21, 1267–1270. [Google Scholar] [CrossRef] [PubMed]
- Dai, P.; Honda, A.; Ewans, L.; McGaughran, J.; Burnett, L.; Law, M.; Phan, T.G. Recommendations for next Generation Sequencing Data Reanalysis of Unsolved Cases with Suspected Mendelian Disorders: A Systematic Review and Meta-Analysis. Genet. Med. 2022, 24, 1618–1629. [Google Scholar] [CrossRef] [PubMed]
- Wenger, A.M.; Guturu, H.; Bernstein, J.A.; Bejerano, G. Systematic Reanalysis of Clinical Exome Data Yields Additional Diagnoses: Implications for Providers. Genet. Med. 2017, 19, 209–214. [Google Scholar] [CrossRef]
- Al-Nabhani, M.; Al-Rashdi, S.; Al-Murshedi, F.; Al-Kindi, A.; Al-Thihli, K.; Al-Saegh, A.; Al-Futaisi, A.; Al-Mamari, W.; Zadjali, F.; Al-Maawali, A. Reanalysis of Exome Sequencing Data of Intellectual Disability Samples: Yields and Benefits. Clin. Genet. 2018, 94, 495–501. [Google Scholar] [CrossRef]
- Costain, G.; Jobling, R.; Walker, S.; Reuter, M.S.; Snell, M.; Bowdin, S.; Cohn, R.D.; Dupuis, L.; Hewson, S.; Mercimek-Andrews, S.; et al. Periodic Reanalysis of Whole-Genome Sequencing Data Enhances the Diagnostic Advantage over Standard Clinical Genetic Testing. Eur. J. Hum. Genet. 2018, 26, 740–744. [Google Scholar] [CrossRef]
- Matalonga, L.; Hernández-Ferrer, C.; Piscia, D.; Schüle, R.; Synofzik, M.; Töpf, A.; Vissers, L.E.L.M.; de Voer, R.; Tonda, R.; Johari, M.; et al. Solving Patients with Rare Diseases through Programmatic Reanalysis of Genome-Phenome Data. Eur. J. Hum. Genet. 2021, 29, 1337–1347. [Google Scholar] [CrossRef]
- Liu, P.; Meng, L.; Normand, E.A.; Xia, F.; Song, X.; Ghazi, A.; Rosenfeld, J.; Magoulas, P.L.; Braxton, A.; Ward, P.; et al. Reanalysis of Clinical Exome Sequencing Data. N. Engl. J. Med. 2019, 380, 2478–2480. [Google Scholar] [CrossRef]
- Poplin, R.; Ruano-Rubio, V.; DePristo, M.A.; Fennell, T.J.; Carneiro, M.O.; Van der Auwera, G.A.; Kling, D.E.; Gauthier, L.D.; Levy-Moonshine, A.; Roazen, D.; et al. Scaling Accurate Genetic Variant Discovery to Tens of Thousands of Samples. BioRxiv 2017, 201178. [Google Scholar] [CrossRef]
- Köhler, S.; Gargano, M.; Matentzoglu, N.; Carmody, L.C.; Lewis-Smith, D.; Vasilevsky, N.A.; Danis, D.; Balagura, G.; Baynam, G.; Brower, A.M.; et al. The Human Phenotype Ontology in 2021. Nucleic Acids Res. 2021, 49, D1207–D1217. [Google Scholar] [CrossRef] [PubMed]
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, P.; Gauthier, L.D.; et al. Variation across 141,456 Human Exomes and Genomes Reveals the Spectrum of Loss-of-Function Intolerance across Human Protein-Coding Genes. Nature 2020, 581, 434–443. [Google Scholar] [CrossRef] [PubMed]
- Hamosh, A.; Scott, A.F.; Amberger, J.S.; Bocchini, C.A.; McKusick, V.A. Online Mendelian Inheritance in Man (OMIM), a Knowledgebase of Human Genes and Genetic Disorders. Nucleic Acids Res. 2005, 33, D514–D517. [Google Scholar] [CrossRef] [PubMed]
- Lek, M.; Karczewski, K.J.; Minikel, E.V.; Samocha, K.E.; Banks, E.; Fennell, T.; O’Donnell-Luria, A.H.; Ware, J.S.; Hill, A.J.; Cummings, B.B.; et al. Analysis of Protein-Coding Genetic Variation in 60,706 Humans. Nature 2016, 536, 285–291. [Google Scholar] [CrossRef] [PubMed]
- Samocha, K.E.; Robinson, E.B.; Sanders, S.J.; Stevens, C.; Sabo, A.; McGrath, L.M.; Kosmicki, J.A.; Rehnström, K.; Mallick, S.; Kirby, A.; et al. A Framework for the Interpretation of de Novo Mutation in Human Disease. Nat. Genet. 2014, 46, 944–950. [Google Scholar] [CrossRef]
- Landrum, M.J.; Lee, J.M.; Benson, M.; Brown, G.; Chao, C.; Chitipiralla, S.; Gu, B.; Hart, J.; Hoffman, D.; Hoover, J.; et al. ClinVar: Public Archive of Interpretations of Clinically Relevant Variants. Nucleic Acids Res. 2016, 44, D862–D868. [Google Scholar] [CrossRef]
- Stenson, P.D.; Ball, E.V.; Mort, M.; Phillips, A.D.; Shiel, J.A.; Thomas, N.S.T.; Abeysinghe, S.; Krawczak, M.; Cooper, D.N. Human Gene Mutation Database (HGMD): 2003 Update. Hum. Mutat. 2003, 21, 577–581. [Google Scholar] [CrossRef]
- Firth, H.V.; Richards, S.M.; Bevan, A.P.; Clayton, S.; Corpas, M.; Rajan, D.; Van Vooren, S.; Moreau, Y.; Pettett, R.M.; Carter, N.P. DECIPHER: Database of Chromosomal Imbalance and Phenotype in Humans Using Ensembl Resources. Am. J. Hum. Genet. 2009, 84, 524–533. [Google Scholar] [CrossRef]
- Robinson, J.T.; Thorvaldsdóttir, H.; Winckler, W.; Guttman, M.; Lander, E.S.; Getz, G.; Mesirov, J.P. Integrative Genomics Viewer. Nat. Biotechnol. 2011, 29, 24–26. [Google Scholar] [CrossRef]
- Jaganathan, K.; Panagiotopoulou, S.K.; McRae, J.F.; Darbandi, S.F.; Knowles, D.; Li, Y.I.; Kosmicki, J.A.; Arbelaez, J.; Cui, W.; Schwartz, G.B.; et al. Predicting Splicing from Primary Sequence with Deep Learning. Cell 2019, 176, 535–548. [Google Scholar] [CrossRef]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and Guidelines for the Interpretation of Sequence Variants: A Joint Consensus Recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef] [PubMed]
- Ellard, S.; Baple, E.L.; Berry, I. ACGS Best Practic Guidelines for Variant Classification 2019. 2019. Available online: https://www.acgs.uk.com/media/11631/uk-practice-guidelines-for-variant-classification-v4-01-2020.pdf (accessed on 11 April 2022).
- Miller, D.T.; Lee, K.; Chung, W.K.; Gordon, A.S.; Herman, G.E.; Klein, T.E.; Stewart, D.R.; Amendola, L.M.; Adelman, K.; Bale, S.J.; et al. ACMG SF v3.0 List for Reporting of Secondary Findings in Clinical Exome and Genome Sequencing: A Policy Statement of the American College of Medical Genetics and Genomics (ACMG). Genet. Med. 2021, 23, 1381–1390. [Google Scholar] [CrossRef] [PubMed]
- Garrison, E.; Marth, G. Haplotype-Based Variant Detection from Short-Read Sequencing. arXiv 2012. [Google Scholar] [CrossRef]
- Tan, N.B.; Stapleton, R.; Stark, Z.; Delatycki, M.B.; Yeung, A.; Hunter, M.F.; Amor, D.J.; Brown, N.J.; Stutterd, C.A.; McGillivray, G.; et al. Evaluating Systematic Reanalysis of Clinical Genomic Data in Rare Disease from Single Center Experience and Literature Review. Mol. Genet. Genom. Med. 2020, 8, e1508. [Google Scholar] [CrossRef] [PubMed]
- Wright, C.F.; Fitzgerald, T.W.; Jones, W.D.; Clayton, S.; McRae, J.F.; van Kogelenberg, M.; King, D.A.; Ambridge, K.; Barrett, D.M.; Bayzetinova, T.; et al. Genetic Diagnosis of Developmental Disorders in the DDD Study: A Scalable Analysis of Genome-Wide Research Data. Lancet 2015, 385, 1305–1314. [Google Scholar] [CrossRef]
- Jalkh, N.; Corbani, S.; Haidar, Z.; Hamdan, N.; Farah, E.; Ghoch, J.A.; Ghosn, R.; Salem, N.; Fawaz, A.; Khayat, C.D.; et al. The Added Value of WES Reanalysis in the Field of Genetic Diagnosis: Lessons Learned from 200 Exomes in the Lebanese Population. BMC Med. Genom. 2019, 12, 11. [Google Scholar] [CrossRef] [PubMed]
- Wright, C.F.; McRae, J.F.; Clayton, S.; Gallone, G.; Aitken, S.; FitzGerald, T.W.; Jones, P.; Prigmore, E.; Rajan, D.; Lord, J.; et al. Making New Genetic Diagnoses with Old Data: Iterative Reanalysis and Reporting from Genome-Wide Data in 1,133 Families with Developmental Disorders. Genet. Med. 2018, 20, 1216–1223. [Google Scholar] [CrossRef]
- Chen, Y.; Chen, D.; Zhao, S.; Liu, G.; Li, H.; Wu, Z.-Y. Penetrance Estimation of PRRT2 Variants in Paroxysmal Kinesigenic Dyskinesia and Infantile Convulsions. Front. Med. 2021, 15, 877–886. [Google Scholar] [CrossRef]
- Richmond, C.M.; Savarirayan, R. Schmid Metaphyseal Chondrodysplasia. In GeneReviews®; Adam, M.P., Everman, D.B., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 2019. [Google Scholar]
- Jamra, R. Genetics of Autosomal Recessive Intellectual Disability. Med. Genet. 2018, 30, 323–327. [Google Scholar] [CrossRef]
- Köhler, S.; Schulz, M.H.; Krawitz, P.; Bauer, S.; Dölken, S.; Ott, C.E.; Mundlos, C.; Horn, D.; Mundlos, S.; Robinson, P.N. Clinical Diagnostics in Human Genetics with Semantic Similarity Searches in Ontologies. Am. J. Hum. Genet. 2009, 85, 457–464. [Google Scholar] [CrossRef]
- Kochinke, K.; Zweier, C.; Nijhof, B.; Fenckova, M.; Cizek, P.; Honti, F.; Keerthikumar, S.; Oortveld, M.A.W.; Kleefstra, T.; Kramer, J.M.; et al. Systematic Phenomics Analysis Deconvolutes Genes Mutated in Intellectual Disability into Biologically Coherent Modules. Am. J. Hum. Genet. 2016, 98, 149–164. [Google Scholar] [CrossRef] [PubMed]
- Martin, A.R.; Williams, E.; Foulger, R.E.; Leigh, S.; Daugherty, L.C.; Niblock, O.; Leong, I.U.S.; Smith, K.R.; Gerasimenko, O.; Haraldsdottir, E.; et al. PanelApp Crowdsources Expert Knowledge to Establish Consensus Diagnostic Gene Panels. Nat. Genet. 2019, 51, 1560–1565. [Google Scholar] [CrossRef] [PubMed]
- Snijders Blok, L.; Vino, A.; den Hoed, J.; Underhill, H.R.; Monteil, D.; Li, H.; Reynoso Santos, F.J.; Chung, W.K.; Amaral, M.D.; Schnur, R.E.; et al. Heterozygous Variants That Disturb the Transcriptional Repressor Activity of FOXP4 Cause a Developmental Disorder with Speech/Language Delays and Multiple Congenital Abnormalities. Genet. Med. 2021, 23, 534–542. [Google Scholar] [CrossRef]
- Koemans, T.S.; Kleefstra, T.; Chubak, M.C.; Stone, M.H.; Reijnders, M.R.F.; de Munnik, S.; Willemsen, M.H.; Fenckova, M.; Stumpel, C.T.R.M.; Bok, L.A.; et al. Functional Convergence of Histone Methyltransferases EHMT1 and KMT2C Involved in Intellectual Disability and Autism Spectrum Disorder. PLOS Genet. 2017, 13, e1006864. [Google Scholar] [CrossRef] [PubMed]
- Kleefstra, T.; de Leeuw, N. Kleefstra Syndrome. In GeneReviews®; Adam, M.P., Everman, D.B., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Li, H. Minimap2: Pairwise Alignment for Nucleotide Sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef] [PubMed]
- Cristofoli, F.; Moss, T.; Moore, H.W.; Devriendt, K.; Flanagan-Steet, H.; May, M.; Jones, J.; Roelens, F.; Fons, C.; Fernandez, A.; et al. De Novo Variants in LMNB1 Cause Pronounced Syndromic Microcephaly and Disruption of Nuclear Envelope Integrity. Am. J. Hum. Genet. 2020, 107, 753–762. [Google Scholar] [CrossRef] [PubMed]
- Parry, D.A.; Martin, C.-A.; Greene, P.; Marsh, J.A.; Ambrose, J.C.; Arumugam, P.; Baple, E.L.; Bleda, M.; Boardman-Pretty, F.; Boissiere, J.M.; et al. Heterozygous Lamin B1 and Lamin B2 Variants Cause Primary Microcephaly and Define a Novel Laminopathy. Genet. Med. 2021, 23, 408–414. [Google Scholar] [CrossRef] [PubMed]
- Guillen Sacoto, M.J.; Tchasovnikarova, I.A.; Torti, E.; Forster, C.; Andrew, E.H.; Anselm, I.; Baranano, K.W.; Briere, L.C.; Cohen, J.S.; Craigen, W.J.; et al. De Novo Variants in the ATPase Module of MORC2 Cause a Neurodevelopmental Disorder with Growth Retardation and Variable Craniofacial Dysmorphism. Am. J. Hum. Genet. 2020, 107, 352–363. [Google Scholar] [CrossRef]
- Brunet, T.; McWalter, K.; Mayerhanser, K.; Anbouba, G.M.; Armstrong-Javors, A.; Bader, I.; Baugh, E.; Begtrup, A.; Bupp, C.P.; Callewaert, B.L.; et al. Defining the Genotypic and Phenotypic Spectrum of X-Linked MSL3-Related Disorder. Genet. Med. 2021, 23, 384–395. [Google Scholar] [CrossRef]
- Basilicata, M.F.; Bruel, A.-L.; Semplicio, G.; Valsecchi, C.I.K.; Aktaş, T.; Duffourd, Y.; Rumpf, T.; Morton, J.; Bache, I.; Szymanski, W.G.; et al. De Novo Mutations in MSL3 Cause an X-Linked Syndrome Marked by Impaired Histone H4 Lysine 16 Acetylation. Nat. Genet. 2018, 50, 1442–1451. [Google Scholar] [CrossRef]
- Smith, L.; Saunders, C.; Dinwiddie, D.; Atherton, A.; Miller, N.; Soden, S.; Farrow, E.; Abdelmoity, A.; Kingsmore, S. Exome Sequencing Reveals De Novo Germline Mutation of the Mammalian Target of Rapamycin (MTOR) in a Patient with Megalencephaly and Intractable Seizures. J. Genomes Exomes 2013, 2013, 63–72. [Google Scholar] [CrossRef]
- Mroske, C.; Rasmussen, K.; Shinde, D.N.; Huether, R.; Powis, Z.; Lu, H.-M.; Baxter, R.M.; McPherson, E.; Tang, S. Germline Activating MTOR Mutation Arising through Gonadal Mosaicism in Two Brothers with Megalencephaly and Neurodevelopmental Abnormalities. BMC Med. Genet. 2015, 16, 102. [Google Scholar] [CrossRef] [PubMed]
- Cortese, A.; Zhu, Y.; Rebelo, A.P.; Negri, S.; Courel, S.; Abreu, L.; Bacon, C.J.; Bai, Y.; Bis-Brewer, D.M.; Bugiardini, E.; et al. Biallelic Mutations in SORD Cause a Common and Potentially Treatable Hereditary Neuropathy with Implications for Diabetes. Nat. Genet. 2020, 52, 473–481. [Google Scholar] [CrossRef] [PubMed]
- Else, T.; Greenberg, S.; Fishbein, L. Hereditary Paraganglioma-Pheochromocytoma Syndromes. In GeneReviews®; Adam, M.P., Everman, D.B., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Roberts, A.M.; Ware, J.S.; Herman, D.S.; Schafer, S.; Baksi, J.; Bick, A.G.; Buchan, R.J.; Walsh, R.; John, S.; Wilkinson, S.; et al. Integrated Allelic, Transcriptional, and Phenomic Dissection of the Cardiac Effects of Titin Truncations in Health and Disease. Sci. Transl. Med. 2015, 7, 270ra6. [Google Scholar] [CrossRef] [PubMed]
- Herman, D.S.; Lam, L.; Taylor, M.R.G.; Wang, L.; Teekakirikul, P.; Christodoulou, D.; Conner, L.; DePalma, S.R.; McDonough, B.; Sparks, E.; et al. Truncations of Titin Causing Dilated Cardiomyopathy. N. Engl. J. Med. 2012, 366, 619–628. [Google Scholar] [CrossRef]
- Tange, O. GNU Parallel 2018, 2018; ISBN 978-1-387-50988-1.
- Di Tommaso, P.; Chatzou, M.; Floden, E.W.; Barja, P.P.; Palumbo, E.; Notredame, C. Nextflow Enables Reproducible Computational Workflows. Nat. Biotechnol. 2017, 35, 316–319. [Google Scholar] [CrossRef]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce Framework for Analyzing next-Generation DNA Sequencing Data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef]
- Li, H. Aligning Sequence Reads, Clone Sequences and Assembly Contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997. [Google Scholar]
- Freed, D.; Aldana, R.; Weber, J.A.; Edwards, J.S. The Sentieon Genomics Tools—A Fast and Accurate Solution to Variant Calling from next-Generation Sequence Data. BioRxiv 2017, 115717. [Google Scholar] [CrossRef]
- Bonfield, J.K.; McCarthy, S.A.; Durbin, R. Crumble: Reference Free Lossy Compression of Sequence Quality Values. Bioinformatics 2019, 35, 337–339. [Google Scholar] [CrossRef]
- Kendig, K.I.; Baheti, S.; Bockol, M.A.; Drucker, T.M.; Hart, S.N.; Heldenbrand, J.R.; Hernaez, M.; Hudson, M.E.; Kalmbach, M.T.; Klee, E.W.; et al. Sentieon DNASeq Variant Calling Workflow Demonstrates Strong Computational Performance and Accuracy. Front. Genet. 2019, 10. [Google Scholar] [CrossRef]
- Resnik, P. Using Information Content to Evaluate Semantic Similarity in a Taxonomy. In Proceedings of the 14th International Joint Conference on Artificial Intelligence—Volume 1; IJCAI’95; Morgan Kaufmann Publishers Inc.: San Francisco, CA, USA, 1995; pp. 448–453. [Google Scholar]
- Lin, D. An Information-Theoretic Definition of Similarity. In Proceedings of the Fifteenth International Conference on Machine Learning; ICML’98; Morgan Kaufmann Publishers Inc.: San Francisco, CA, USA, 1998; pp. 296–304. [Google Scholar]
- Deng, Y.; Gao, L.; Wang, B.; Guo, X. HPOSim: An R Package for Phenotypic Similarity Measure and Enrichment Analysis Based on the Human Phenotype Ontology. PLOS ONE 2015, 10, e0115692. [Google Scholar] [CrossRef] [PubMed]

| Gene | Approach (Date of Initial Evaluation) | Symptoms | Variant (Transcript, c-Code, p-Code) | Zygosity | Inheritance | ACMG Criteria (Final Classification) | OMIM Phenotype | Why Not Found Initially |
|---|---|---|---|---|---|---|---|---|
| 1. FOXP4 | Trio (08/2017) | Hearing impairment, ventricular septal defect, flattened epiphysis, disproportionate short stature, craniofacial asymmetry | NM_001012426.2:c.1540 G > A, p.Ala514Thr | Heterozygous, de novo | Autosomal dominant | PS2, PS3, PS4_MOD, PM2_SUP, PP3 (pathogenic) | - | New gene–disease association after 53 months (4 years and 5 months) |
| 2. KMT2C | Trio (09/2017) | Hypothyroidism, mild intellectual disability, mild abnormality of facial shape, mild short stature | NM_170606.3:c.1829_ 1830delCA, p.Thr610Serfs * 4 | Heterozygous, de novo | Autosomal dominant | PVS1, PS2_MOD, PS4_SUP, PM2_SUP (pathogenic) | Kleefstra syndrome 2 (#617768) | Updated caller, see also Figure S1 |
| 3. LMNB1 | Trio (12/2019) | Microcephaly, agenesis of the corpus callosum, cerebellar hypoplasia, growth retardation (prenatal) | NM_005573.4:c.97A > G, p.Lys33Glu | Heterozygous, de novo | Autosomal dominant | PS2_VSTR, PS3, PS4_MOD, PM2_SUP, PP3 (pathogenic) | Microcephaly 26, primary, autosomal dominant (#619179) | New gene–disease association after 9 months |
| 4. MORC2 | Trio (05/2019) | Developmental delay, microcephaly | NM_001303256.3:c.79G > A, p.Glu27Lys | Heterozygous, de novo | Autosomal dominant | PS2_VSTR, PS3, PS4_MOD, PM2_SUP, PP2 (pathogenic) | Developmental delay, impaired growth, dysmorphic facies, and axonal neuropathy (#619090) | New gene–disease association after 15 months |
| 5. MSL3 | Trio (01/2018) | Global developmental delay, seizures, chylothorax, mid-aortic syndrome | NM_078629.4:c.973_ 974delAG, p.Gln326Alafs * 5 | Hemizygous, de novo | X-chromosomal dominant | PVS1, PS2, PS4_SUP, PM2_SUP (pathogenic) | Basilicata-Akhtar syndrome (#301032) | New gene–disease association after 9 months |
| 6. MTOR | Trio (9/2017) | Global developmental delay, macrocephaly | NM_004958.4:c.5911G > A, p.Ala1971Thr | Heterozygous, de novo | Autosomal dominant | PS2, PS4_SUP, PM2_SUP, PM5_SUP, PP2, PP3 (likely pathogenic) | Smith-Kingsmore syndrome (#616638) | Updated caller, see also Figure S2 |
| 7. SORD | Trio (02/2020) | Pain in both legs from the age of 17, ataxia, atrophy of the leg muscles | NM_003104.5:c.757del, p.Ala253Glnfs * 27 | Homozygous, maternal and paternal | Autosomal recessive | PVS1, PS3, PM3_VSTR (pathogenic) | Sorbitol dehydrogenase deficiency with peripheral neuropathy (#618912) | New gene–disease association after 3 months |
| 8. SHDC * | Trio (12/2017) | Mild global developmental delay, seizures, heterotopia, oral cleft, tall stature, obesity | NM_003001.5:c.377A > G, p.Tyr126Cys | Heterozygous, paternal | Autosomal dominant | PS4_MOD, PM1, PM2_SUP, PP3 (likely pathogenic) | Paragangliomas 3 (#605373) | Updated caller |
| 9. TTN * | Trio (09/2020) | Panhypopituitarism, developmental delay, patent ductus arteriosus, scoliosis, short stature, median cleft lip and palate | NM_001267550.2:c.80762_80765delAACA, p.Lys26921Argfs * 5 | Heterozygous, maternal | Autosomal dominant | PVS1, PM2_SUP (likely pathogenic) | Cardiomyopathy, dilated, 1G (#604145) | Updated AMCG secondary findings list |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Halfmeyer, I.; Bartolomaeus, T.; Popp, B.; Radtke, M.; Helms, T.; Hentschel, J.; Popp, D.; Jamra, R.A. Approach to Cohort-Wide Re-Analysis of Exome Data in 1000 Individuals with Neurodevelopmental Disorders. Genes 2023, 14, 30. https://doi.org/10.3390/genes14010030
Halfmeyer I, Bartolomaeus T, Popp B, Radtke M, Helms T, Hentschel J, Popp D, Jamra RA. Approach to Cohort-Wide Re-Analysis of Exome Data in 1000 Individuals with Neurodevelopmental Disorders. Genes. 2023; 14(1):30. https://doi.org/10.3390/genes14010030
Chicago/Turabian StyleHalfmeyer, Insa, Tobias Bartolomaeus, Bernt Popp, Maximilian Radtke, Tobias Helms, Julia Hentschel, Denny Popp, and Rami Abou Jamra. 2023. "Approach to Cohort-Wide Re-Analysis of Exome Data in 1000 Individuals with Neurodevelopmental Disorders" Genes 14, no. 1: 30. https://doi.org/10.3390/genes14010030
APA StyleHalfmeyer, I., Bartolomaeus, T., Popp, B., Radtke, M., Helms, T., Hentschel, J., Popp, D., & Jamra, R. A. (2023). Approach to Cohort-Wide Re-Analysis of Exome Data in 1000 Individuals with Neurodevelopmental Disorders. Genes, 14(1), 30. https://doi.org/10.3390/genes14010030





