A Comparative Genome-Wide Transcriptome Analysis of Glucocorticoid Responder and Non-Responder Primary Human Trabecular Meshwork Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Human Donor Eyes
2.2. Human Organ-Cultured Anterior Segment (HOCAS)
2.3. Primary Human TM Cell Strain with Known GC Responsiveness
2.4. RNA Extraction and mRNA Sequencing
2.5. Mapping and Differential Expression Analysis
2.6. Pathway Enrichment Analysis
2.7. Validation of RNA Seq by RT2-Profiler PCR Array
2.8. Statistical Analysis
3. Results
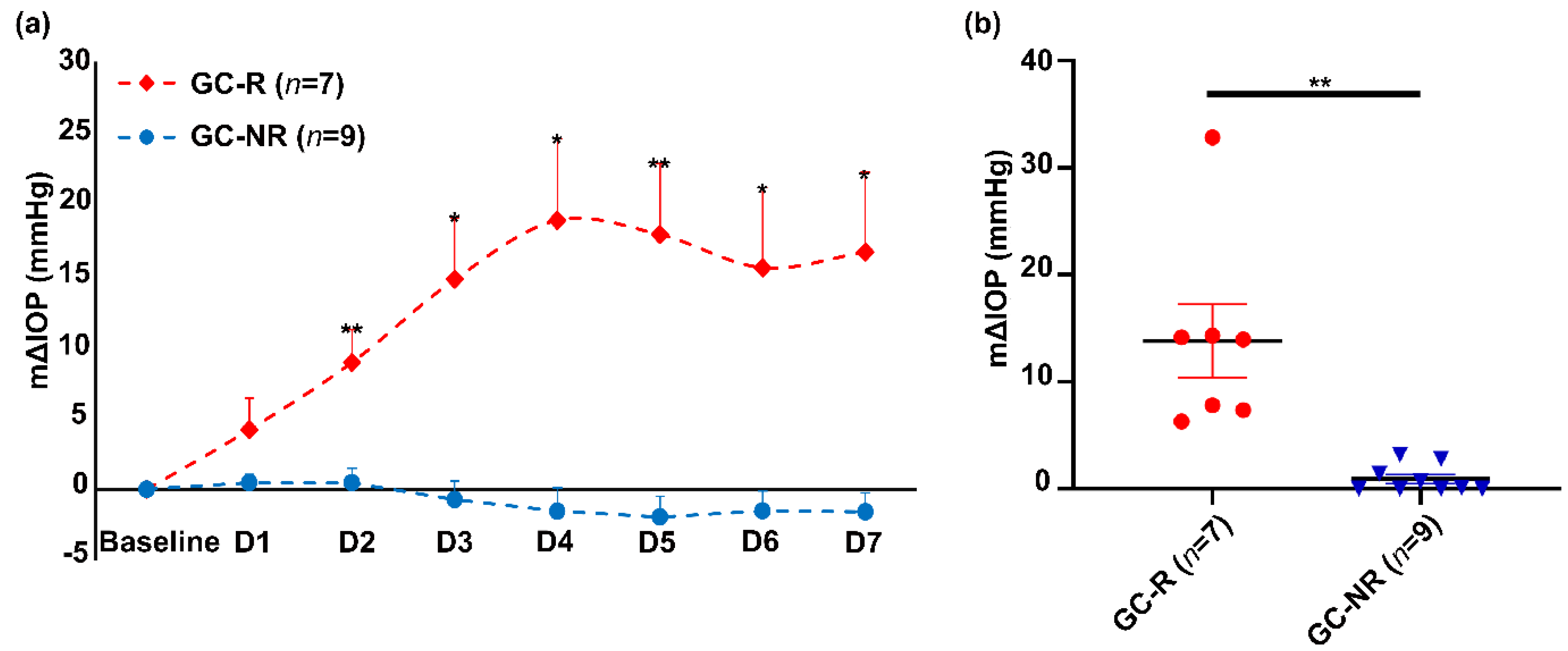
3.1. Establishment of GC-R and GC-NR HTM Cells
3.2. RNA Seq Data Quality
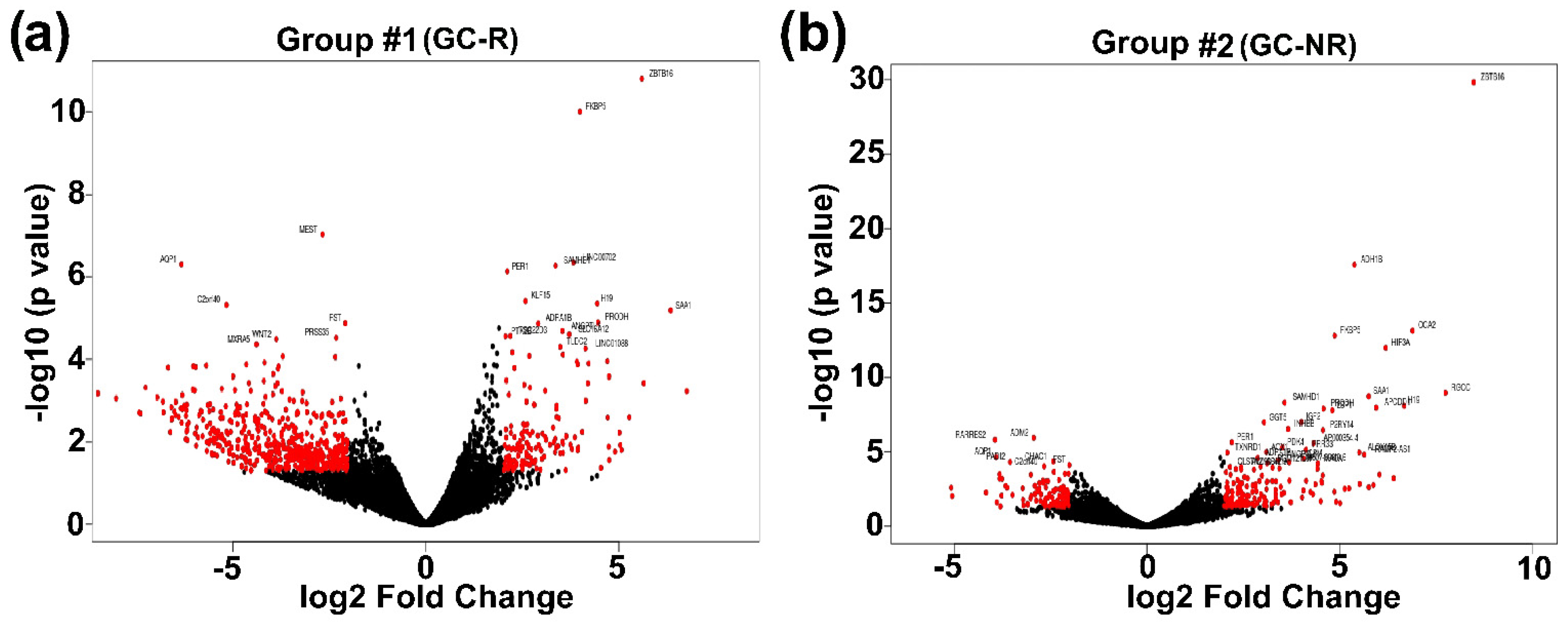
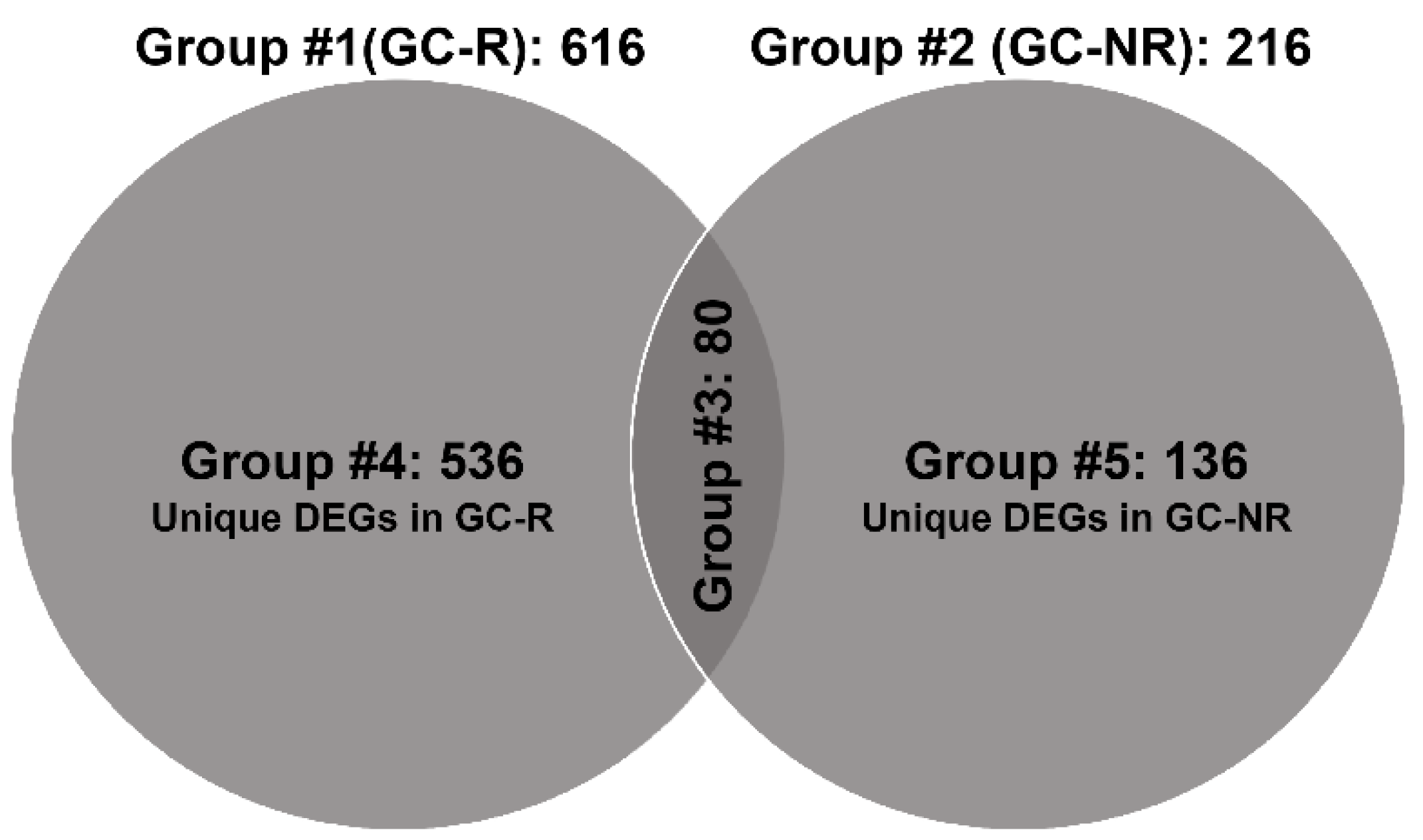
3.3. Differentially Expressed Genes of GC-R and GC-NR HTM Cells
3.4. Pathway Analysis
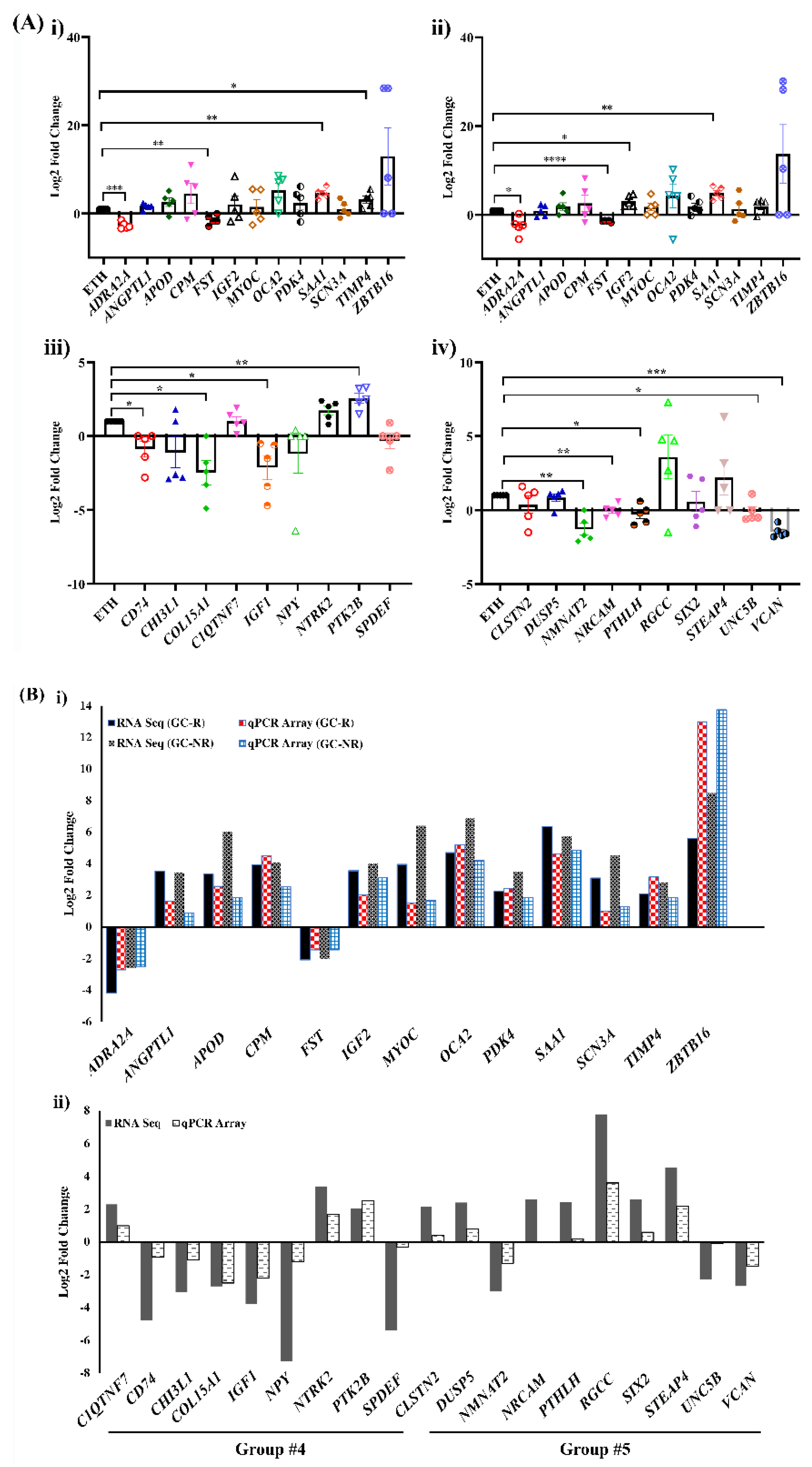
3.5. Validation of DE genes by RT2-PCR Array
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Armaly, M.F. Effect of Corticosteroids on Intraocular Pressure and Fluid Dynamics: I. The Effect of Dexamethasone * in the Normal Eye. Arch. Ophthalmol. 1963, 70, 482–491. [Google Scholar] [CrossRef] [PubMed]
- Lewis, J.M.; Priddy, T.; Judd, J.; Gordon, M.O.; Kass, M.A.; Kolker, A.E.; Becker, B. Intraocular pressure response to topical dexamethasone as a predictor for the development of primary open-angle glaucoma. Am. J. Ophthalmol. 1988, 106, 607–612. [Google Scholar] [CrossRef]
- Bartlett, J.D.; Woolley, T.W.; Adams, C.M. Identification of High Intraocular Pressure Responders to Topical Ophthalmic Corticosteroids. J. Ocul. Pharmacol. 1993, 9, 35–45. [Google Scholar] [CrossRef] [PubMed]
- Jain, A.; Wordinger, R.J.; Yorio, T.; Clark, A.F. Role of the alternatively spliced glucocorticoid receptor isoform GRβ in steroid responsiveness and glaucoma. J. Ocul. Pharmacol Ther. 2014, 30, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Leung, D.Y.; Kisich, K.O. Serine-arginine-rich protein p30 directs alternative splicing of glucocorticoid receptor pre-mRNA to glucocorticoid receptor β in neutrophils. J. Biol. Chem. 2003, 278, 27112–27118. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.B.; Tang, C.H.; Huang, Y.; Fang, H.; Yu, Z.Q.; Wu, L.M.; Liu, R.Y. Alternative splicing in exon 9 of glucocorticoid receptor pre-mRNA is regulated by SRp40. Mol. Biol. Rep. 2010, 37, 1427–1433. [Google Scholar] [CrossRef]
- Ishibashi, T.; Takagi, Y.; Mori, K.; Naruse, S.; Nishino, H.; Yue, B.Y.J.T.; Kinoshita, S. cDNA microarray analysis of gene expression changes induced by dexamethasone in cultured human trabecular meshwork cells. Investig. Ophthalmol. Vis. Sci. 2002, 43, 3691–3697. [Google Scholar]
- Lo, W.R.; Rowlette, L.L.; Caballero, M.; Yang, P.; Hernandez, M.R.; Borra’s, T. Tissue Differential Microarray Analysis of Dexamethasone Induction Reveals Potential Mechanisms of Steroid Glaucoma. Investig. Opthalmol. Vis. Sci. 2003, 44, 473. [Google Scholar] [CrossRef]
- Leung, Y.F.; Tam, P.O.S.; Lee, W.S.; Lam, D.S.C.; Yam, H.F.; Fan, B.J.; Tham, C.C.Y.; Chua, J.K.H.; Pang, C.P. The dual role of dexamethasone on anti-inflammation and outflow resistance demonstrated in cultured human trabecular meshwork cells. Mol. Vis. 2003, 9, 425–439. [Google Scholar]
- Rozsa, F.W.; Reed, D.M.; Scott, K.M.; Pawar, H.; Moroi, S.E.; Kijek, T.G.; Krafchak, C.M.; Othman, M.I.; Vollrath, D.; Elner, V.M.; et al. Gene expression profile of human trabecular meshwork cells in response to long-term dexamethasone exposure. Mol. Vis. 2006, 12, 125–141. [Google Scholar]
- Fan, B.J.; Wang, D.Y.; Tham, C.C.Y.; Lam, D.S.C.; Pang, C.P. Gene expression profiles of human trabecular meshwork cells induced by triamcinolone and dexamethasone. Invest. Ophthalmol. Vis. Sci. 2008, 49, 1886–1897. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Nehmé, A.; Lobenhofer, E.K.; Stamer, W.D.; Edelman, J.L. Glucocorticoids with different chemical structures but similar glucocorticoid receptor potency regulate subsets of common and unique genes in human trabecular meshwork cells. BMC Med. Genom. 2009, 2, 58. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, A.; Asada, Y.; Takakuwa, K.; Sugita, J.; Murakami, A.; Ebihara, N. DNA Methylation Analysis of Human Trabecular Meshwork Cells During Dexamethasone Stimulation. Investig. Opthalmol. Vis. Sci. 2015, 56, 3801. [Google Scholar] [CrossRef] [PubMed]
- Faralli, J.A.; Desikan, H.; Peotter, J.; Kanneganti, N.; Weinhaus, B.; Filla, M.S.; Peters, D.M. Genomic/proteomic analyses of dexamethasone-treated human trabecular meshwork cells reveal a role for GULP1 and ABR in phagocytosis. Mol. Vis. 2019, 25, 237–254. [Google Scholar] [PubMed]
- Hoheisel, J.D. Microarray technology: Beyond transcript profiling and genotype analysis. Nat. Rev. Genet. 2006, 7, 200–210. [Google Scholar] [CrossRef] [PubMed]
- Bermudez, J.Y.; Webber, H.C.; Brown, B.; Braun, T.A.; Clark, A.F.; Mao, W. A Comparison of Gene Expression Profiles between Glucocorticoid Responder and Non-Responder Bovine Trabecular Meshwork Cells Using RNA Sequencing. PLoS ONE 2017, 12, e0169671. [Google Scholar] [CrossRef]
- Tripathi, R.C. Ultrastructure of the exit pathway of the aqueous in lower mammals. (A preliminary report on the “angular aqueous plexus”). Exp. Eye Res. 1971, 12, 311–314. [Google Scholar] [CrossRef]
- Erickson-Lamy, K.; Schroeder, A.M.; Bassett-Chu, S.; Epstein, D.L. Absence of time-dependent facility increase (‘washout’) in the perfused enucleated human eye. Invest. Ophthalmol. Vis. Sci. 1990, 31, 2384–2388. [Google Scholar]
- Scott, P.A.; Overby, D.R.; Freddo, T.F.; Gong, H. Comparative studies between species that do and do not exhibit the washout effect. Exp. Eye Res. 2007, 84, 435–443. [Google Scholar] [CrossRef]
- Haribalaganesh, R.; Gowri Priya, C.; Sharmila, R.; Krishnadas, S.; Muthukkaruppan, V.; Willoughby, C.E.; Senthilkumari, S. Assessment of differential intraocular pressure response to dexamethasone treatment in perfusion cultured Indian cadaveric eyes. Sci. Rep. 2021, 11, 605. [Google Scholar] [CrossRef]
- Mao, W.; Tovar-Vidales, T.; Yorio, T.; Wordinger, R.J.; Clark, A.F. Perfusion-Cultured Bovine Anterior Segments as an Ex Vivo Model for Studying Glucocorticoid-Induced Ocular Hypertension and Glaucoma. Investig. Ophthalmol. Vis. Sci. 2011, 52, 8068–8075. [Google Scholar] [CrossRef] [PubMed]
- Clark, A.F.; Wilson, K.; De Kater, A.W.; Allingham, R.R.; McCartney, M.D. Dexamethasone-induced ocular hypertension in perfusion-cultured human eyes. Investig. Ophthalmol. Vis. Sci. 1995, 36, 478–489. [Google Scholar]
- Stamer, W.D.; Seftor, R.E.B.; Williams, S.K.; Samaha, H.A.M.; Snyder, R.W. Isolation and culture of human trabecular meshwork cells by extracellular matrix digestion. Curr. Eye Res. 1995, 14, 611–617. [Google Scholar] [CrossRef] [PubMed]
- Ashwinbalaji, S.; Senthilkumari, S.; Gowripriya, C.; Krishnadas, S.; Gabelt, B.A.T.; Kaufman, P.L.; Muthukkaruppan, V. SB772077B, A New Rho Kinase Inhibitor Enhances Aqueous Humour Outflow Facility in Human Eyes. Sci. Rep. 2018, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Keller, K.E.; Bhattacharya, S.K.; Borrás, T.; Brunner, T.M.; Chansangpetch, S.; Clark, A.F.; Dismuke, W.M.; Du, Y.; Elliott, M.H.; Ethier, C.R.; et al. Consensus recommendations for trabecular meshwork cell isolation, characterization and culture. Exp. Eye Res. 2018, 171, 164–173. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef]
- Dennis, G.; Sherman, B.T.; Hosack, D.A.; Yang, J.; Gao, W.; Lane, H.C.; Lempicki, R.A. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 2003, 4, 1–11. [Google Scholar] [CrossRef]
- Fini, M.E.; Schwartz, S.G.; Gao, X.; Jeong, S.; Patel, N.; Itakura, T.; Price, M.O.; Price, F.W.; Varma, R.; Stamer, W.D. Steroid-induced ocular hypertension/glaucoma: Focus on pharmacogenomics and implications for precision medicine. Prog. Retin. Eye Res. 2017, 56, 58–83. [Google Scholar] [CrossRef]
- Al-Dabbagh, N.M.; Al-Dohayan, N.; Arfin, M.; Tariq, M. Apolipoprotein E polymorphisms and primary glaucoma in Saudis. Mol. Vis. 2009, 15, 912–919. [Google Scholar]
- Liao, R.; Ye, M.; Xu, X. An updated meta-analysis: Apolipoprotein E genotypes and risk of primary open-angle glaucoma. Mol. Vis. 2014, 20, 1025–1036. [Google Scholar]
- Santos-Carvalho, A.; Elvas, F.; Álvaro, A.R.; Ambrósio, A.F.; Cavadas, C. Neuropeptide Y receptors activation protects rat retinal neural cells against necrotic and apoptotic cell death induced by glutamate. Cell Death Dis. 2013, 4, e636. [Google Scholar] [CrossRef]
- Ohuchi, T.; Tanihara, H.; Yoshimura, N.; Kuriyama, S.; Ito, S.; Honda, Y. Neuropeptide-induced [Ca2+]i transients in cultured bovine trabecular cells. Investig. Ophthalmol. Vis. Sci. 1992, 33, 1676–1684. [Google Scholar]
- Liesenborghs, I.; Eijssen, L.M.T.; Kutmon, M.; Gorgels, T.G.M.F.; Evelo, C.T.; Beckers, H.J.M.; Webers, C.A.B.; Schouten, J.S.A.G. The Molecular Processes in the Trabecular Meshwork After Exposure to Corticosteroids and in Corticosteroid-Induced Ocular Hypertension. Investig. Opthalmol. Vis. Sci. 2020, 61, 24. [Google Scholar] [CrossRef]
- Wang, W.-H.; McNatt, L.G.; Pang, I.-H.; Millar, J.C.; Hellberg, P.E.; Hellberg, M.H.; Steely, H.T.; Rubin, J.S.; Fingert, J.H.; Sheffield, V.C.; et al. Increased expression of the WNT antagonist sFRP-1 in glaucoma elevates intraocular pressure. J. Clin. Investig. 2008, 118, 1056–1064. [Google Scholar] [CrossRef]
- Villarreal, G.; Chatterjee, A.; Oh, S.S.; Oh, D.-J.; Kang, M.H.; Rhee, D.J. Canonical Wnt Signaling Regulates Extracellular Matrix Expression in the Trabecular Meshwork. Investig. Opthalmol. Vis. Sci. 2014, 55, 7433. [Google Scholar] [CrossRef]
- Morgan, J.T.; Raghunathan, V.K.; Chang, Y.-R.; Murphy, C.J.; Russell, P. Wnt inhibition induces persistent increases in intrinsic stiffness of human trabecular meshwork cells. Exp. Eye Res. 2015, 132, 174–178. [Google Scholar] [CrossRef]
- Ahadome, S.D.; Zhang, C.; Tannous, E.; Shen, J.; Zheng, J.J. Small-molecule inhibition of Wnt signaling abrogates dexamethasone-induced phenotype of primary human trabecular meshwork cells. Exp. Cell. Res. 2017, 357, 116–123. [Google Scholar] [CrossRef]
- Bolkenius, U.; Hahn, D.; Gressner, A.M.; Breitkopf, K.; Dooley, S.; Wickert, L. Glucocorticoids decrease the bioavailability of TGF-β which leads to a reduced TGF-β signaling in hepatic stellate cells. Biochem. Biophys. Res. Commun. 2004, 325, 1264–1270. [Google Scholar] [CrossRef]
- Schwartze, J.T.; Becker, S.; Sakkas, E.; Wujak, Ł.A.; Niess, G.; Usemann, J.; Reichenberger, F.; Herold, S.; Vadász, I.; Mayer, K.; et al. Glucocorticoids Recruit Tgfbr3 and Smad1 to Shift Transforming Growth Factor-β Signaling from the Tgfbr1/Smad2/3 Axis to the Acvrl1/Smad1 Axis in Lung Fibroblasts. J. Biol. Chem. 2014, 289, 3262–3275. [Google Scholar] [CrossRef] [PubMed]
- Song, C.-Z.; Tian, X.; Gelehrter, T.D. Glucocorticoid receptor inhibits transforming growth factor-β signaling by directly targeting the transcriptional activation function of Smad3. Proc. Natl. Acad. Sci. USA 1999, 96, 11776–11781. [Google Scholar] [CrossRef] [PubMed]
- Kasetti, R.B.; Maddineni, P.; Patel, P.D.; Searby, C.; Sheffield, V.C.; Zode, G.S. Transforming growth factor β2 (TGFβ2) signaling plays a key role in glucocorticoid-induced ocular hypertension. J. Biol. Chem. 2018, 293, 9854–9868. [Google Scholar] [CrossRef] [PubMed]
| List of Genes from Group #3 | ||||||||
| Gene | Group #1 | Group #2 | ||||||
| logFC | logCPM | p Value | logFC | logCPM | p Value | |||
| ZBTB16 | 5.60 | 4.66 | 0.000 | 8.48 | 4.59 | 0.000 | ||
| OCA2 | 4.71 | 1.61 | 0.000 | 6.89 | 2.24 | 0.000 | ||
| H19 | 4.44 | 7.60 | 0.000 | 6.67 | 8.47 | 0.000 | ||
| MYOC | 3.95 | 6.57 | 0.014 | 6.40 | 9.28 | 0.001 | ||
| HIF3A | 4.20 | 4.11 | 0.000 | 6.19 | 2.63 | 0.000 | ||
| APOD | 3.35 | 7.28 | 0.018 | 6.03 | 8.49 | 0.000 | ||
| SAA1 | 6.34 | 2.81 | 0.000 | 5.75 | 1.16 | 0.000 | ||
| ADH1B | 3.39 | 7.72 | 0.001 | 5.38 | 6.06 | 0.000 | ||
| FKBP5 | 3.99 | 7.23 | 0.000 | 4.87 | 7.62 | 0.000 | ||
| LSP1 | 3.91 | 5.22 | 0.000 | 4.81 | 4.97 | 0.000 | ||
| List of Genes from Group #4 | ||||||||
| Up-regulated genes | Down-regulated genes | |||||||
| Gene | logFC | logCPM | p Value | Gene | logFC | logCPM | p Value | |
| FRG2C | 5.27 | −2.07 | 0.002 | UPK3A | −8.49 | 0.36 | 0.001 | |
| NPSR1-AS1 | 5.07 | −2.19 | 0.016 | RLN1 | −8.02 | 1.24 | 0.001 | |
| BHLHE22 | 5.04 | 0.77 | 0.012 | FAM110D | −7.42 | −0.59 | 0.002 | |
| SAA4 | 4.75 | −2.34 | 0.028 | PRSS22 | −7.38 | −0.62 | 0.002 | |
| TUSC5 | 3.79 | −1.37 | 0.015 | NPY | −7.26 | 3.41 | 0.000 | |
| LEP | 3.75 | −0.16 | 0.012 | GIMAP1 | −6.96 | −0.98 | 0.001 | |
| RNA5SP111 | 3.63 | −2.00 | 0.049 | ST14 | −6.81 | 1.52 | 0.001 | |
| KCNE1 | 3.55 | 0.74 | 0.010 | LRRC26 | −6.79 | 1.53 | 0.002 | |
| RPL7P57 | 3.49 | −2.06 | 0.019 | KRT15 | −6.67 | 3.39 | 0.000 | |
| NTRK2 | 3.39 | 5.15 | 0.002 | FXYD3 | −6.59 | 2.02 | 0.001 | |
| List of Genes from Group #5 | ||||||||
| Up-regulated genes | Down-regulated genes | |||||||
| Gene | logFC | logCPM | p Value | Gene | logFC | logCPM | p Value | |
| RGCC | 7.75 | 1.95 | 0.000 | GRM5 | −5.05 | −2.43 | 0.010 | |
| APCDD1 | 5.95 | 2.15 | 0.000 | SLC24A2 | −3.89 | −1.78 | 0.025 | |
| IGF2-AS | 5.88 | −1.83 | 0.002 | GRIA2 | −3.82 | 0.29 | 0.001 | |
| PNMT | 5.76 | −1.91 | 0.002 | RBFOX1 | −3.80 | −1.82 | 0.048 | |
| RAMP2-AS1 | 5.64 | 0.83 | 0.000 | KRT17P1 | −3.77 | −1.24 | 0.009 | |
| ALOX15B | 5.51 | −0.80 | 0.000 | GRID2 | −3.67 | −0.01 | 0.002 | |
| PTGDR2 | 5.24 | −2.22 | 0.003 | VCAN-AS1 | −3.63 | −1.95 | 0.003 | |
| LINC00525 | 5.13 | −2.29 | 0.003 | PADI2 | −3.55 | 0.96 | 0.000 | |
| STEAP4 | 4.57 | 3.40 | 0.000 | KLHDC7B | −3.22 | −0.85 | 0.003 | |
| SFTPC | 4.51 | −2.59 | 0.021 | IGFL2 | −3.20 | −2.15 | 0.040 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kathirvel, K.; Haribalaganesh, R.; Krishnadas, R.; Muthukkaruppan, V.; Willoughby, C.E.; Bharanidharan, D.; Senthilkumari, S. A Comparative Genome-Wide Transcriptome Analysis of Glucocorticoid Responder and Non-Responder Primary Human Trabecular Meshwork Cells. Genes 2022, 13, 882. https://doi.org/10.3390/genes13050882
Kathirvel K, Haribalaganesh R, Krishnadas R, Muthukkaruppan V, Willoughby CE, Bharanidharan D, Senthilkumari S. A Comparative Genome-Wide Transcriptome Analysis of Glucocorticoid Responder and Non-Responder Primary Human Trabecular Meshwork Cells. Genes. 2022; 13(5):882. https://doi.org/10.3390/genes13050882
Chicago/Turabian StyleKathirvel, Kandasamy, Ravinarayanan Haribalaganesh, Ramasamy Krishnadas, Veerappan Muthukkaruppan, Colin E. Willoughby, Devarajan Bharanidharan, and Srinivasan Senthilkumari. 2022. "A Comparative Genome-Wide Transcriptome Analysis of Glucocorticoid Responder and Non-Responder Primary Human Trabecular Meshwork Cells" Genes 13, no. 5: 882. https://doi.org/10.3390/genes13050882
APA StyleKathirvel, K., Haribalaganesh, R., Krishnadas, R., Muthukkaruppan, V., Willoughby, C. E., Bharanidharan, D., & Senthilkumari, S. (2022). A Comparative Genome-Wide Transcriptome Analysis of Glucocorticoid Responder and Non-Responder Primary Human Trabecular Meshwork Cells. Genes, 13(5), 882. https://doi.org/10.3390/genes13050882







