Association of Genetically Predicted BCAA Levels with Muscle Fiber Size in Athletes Consuming Protein
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethical Approval
2.2. Study Participants
2.3. Anthropometry and Body Composition
2.4. Physical Activity and Training Parameters
2.5. Assessment of Dietary Protein Intake
2.6. Evaluation of Muscle Fiber Composition by Immunohistochemistry
2.7. Genotyping
2.8. Polygenic Analysis
2.9. Statistical Analyses
3. Results
3.1. Hardy–Weinberg Equilibrium (HWE) and Genotype Distribution
3.2. Anthropometry
3.3. Muscle Fiber Distribution and Cross-Sectional Area (CSA)
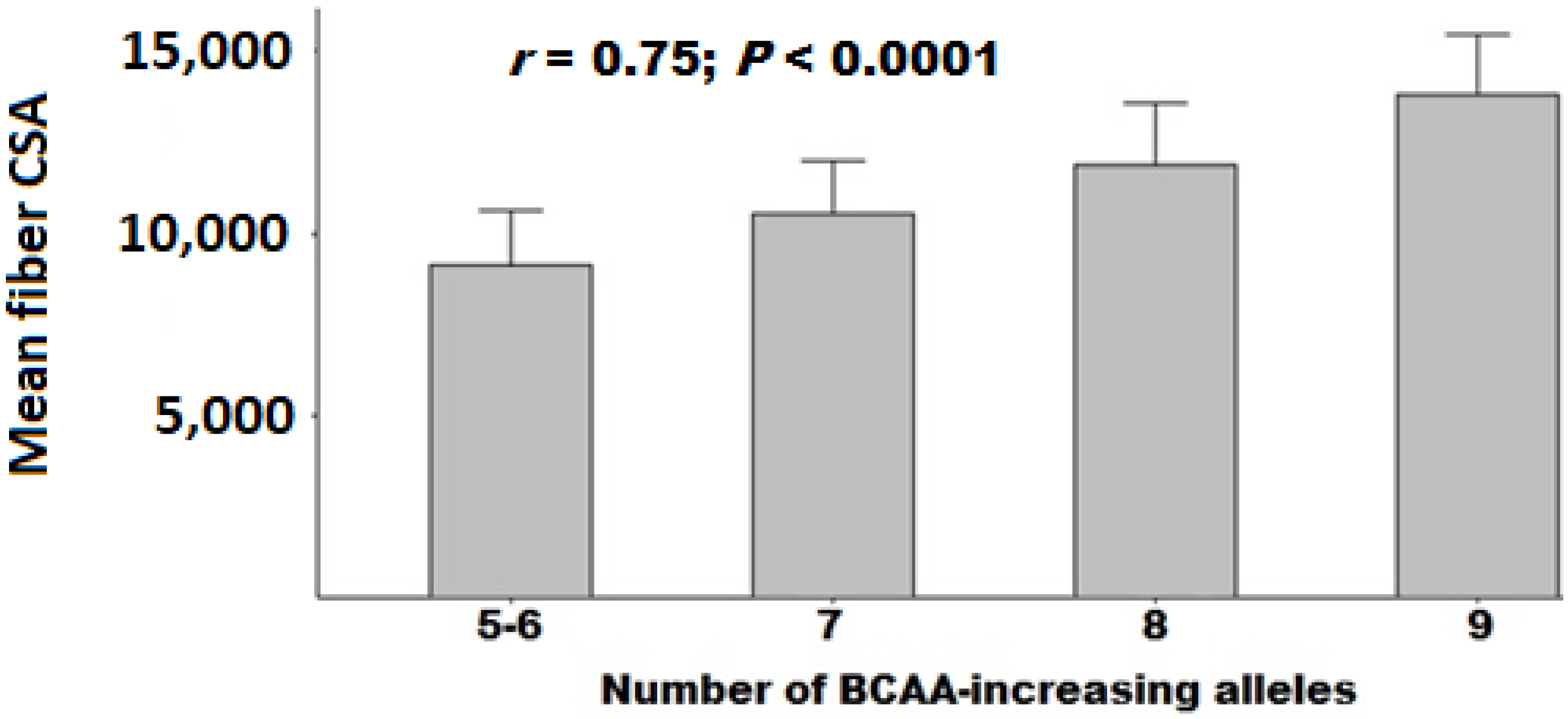
3.4. Polygenic Association with Muscle Fiber CSA
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Anthony, T.G.; Vary, T.C.; Jefferson, L.S.; Kimball, S.R. Leucine stimulates translation initiation in skeletal muscle of postabsorptive rats via a rapamycin-sensitive pathway. J. Nutr. 2000, 130, 2413–2419. [Google Scholar] [CrossRef] [PubMed]
- Bohé, J.; Low, A.; Wolfe, R.R.; Rennie, M.J. Human muscle protein synthesis is modulated by extracellular, not intramuscular amino acid availability: A dose-response study. J. Phys. 2003, 552, 315–324. [Google Scholar] [CrossRef] [PubMed]
- Tipton, K.D.; Ferrando, A.A.; Phillips, S.M.; Doyle, D., Jr.; Wolfe, R.R. Postexercise net protein synthesis in human muscle from orally administered amino acids. Am. J. Physiol. 1999, 276, 628–634. [Google Scholar] [CrossRef]
- Lynch, C.J.; Halle, B.; Fujii, H.; Vary, T.C.; Wallin, R.; Damuni, Z.; Hutson, S.M. Potential role of leucine metabolism in the leucine-signaling pathway involving mTOR. Am. J. Physiol. Endocrinol. Metab. 2003, 285, E854–E863. [Google Scholar] [CrossRef] [PubMed]
- Howatson, G.; Hoad, M.; Goodall, S.; Tallent, J.; Bell, P.G.; French, D.N. Exercise-induced muscle dam- age is reduced in resistance-trained males by branched chain amino acids: A randomized, double- blind, placebo controlled study. J. Int. Soc. Sports Nutr. 2012, 9, 20. [Google Scholar] [CrossRef]
- Greer, B.K.; Woodard, J.L.; White, J.P.; Arguello, E.M.; Haymes, E.M. Branched-chain amino acid supple- mentation and indicators of muscle damage after endurance exercise. Int. J. Sport Nutr. Exerc. Metab. 2007, 17, 595–607. [Google Scholar] [CrossRef]
- Koba, T.; Hamada, K.; Sakurai, M.; Matsumoto, K.; Hayase, H.; Imaizumi, K.; Tsujimoto, H.; Mitsuzono, R. Branched-chain amino acids supplementation attenuates the accumulation of blood lactate dehydrogenase during distance running. J. Sports Med. Phys. Fit. 2007, 47, 316–322. [Google Scholar]
- Knechtle, B.; Knechtle, P.; Mrazek, C.; Senn, O.; Rosemann, T.; Imoberdorf, R.; Ballmer, P. No effect of short- term amino acid supplementation on variables related to skeletal muscle damage in 100 km ultra-runners—A randomized controlled trial. J. Int. Soc. Sports. Nutr. 2011, 8, 6. [Google Scholar] [CrossRef]
- Devries, M.C.; Phillips, S.M. Supplemental protein in support of muscle mass and health: Advantage whey. J. Food Sci. 2015, 80, A8–A15. [Google Scholar] [CrossRef]
- Lam, F.C.; Khan, T.M.; Faidah, H.; Haseeb, A.; Khan, A.H. Effectiveness of whey protein supplements on the serum levels of amino acid, creatinine kinase and myoglobin of athletes: A systematic review and meta-analysis. Syst. Rev. 2019, 8, 130. [Google Scholar] [CrossRef]
- Lustgarten, M.S.; Price, L.L.; Chale, A.; Phillips, E.M.; Fielding, R.A. Branched chain amino acids are associated with muscle mass in functionally limited older adults. J. Gerontol. Ser. A Biol. Sci. Med. Sci. 2014, 69, 717–724. [Google Scholar] [CrossRef] [PubMed]
- Tsuji, S.; Koyama, S.; Taniguchi, R.; Fujiwara, T.; Fujiwara, H.; Sato, Y. Nutritional status of outpatients with chronic stable heart failure based on serum amino acid concentration. J. Cardiol. 2018, 72, 458–465. [Google Scholar] [CrossRef] [PubMed]
- Kujala, U.M.; Mäkinen, V.-P.; Heinonen, I.; Soininen, P.; Kangas, A.J.; Leskinen, T.H.; Rahkila, P.; Würtz, P.; Kovanen, V.; Cheng, S.; et al. Long-term Leisure-time Physical Activity and Serum Metabolome. Circulation 2013, 127, 340–348. [Google Scholar] [CrossRef] [PubMed]
- Leskinen, T.; Rinnankoski-Tuikka, R.; Rintala, M.; Seppänen-Laakso, T.; Pöllänen, E.L.; Alen, M.; Sipilä, S.; Kaprio, J.; Kovanen, V.; Rahkila, P.; et al. Differences in Muscle and Adipose Tissue Gene Expression and Cardio-Metabolic Risk Factors in the Members of Physical Activity Discordant Twin Pairs. PLoS ONE 2010, 5, e12609. [Google Scholar] [CrossRef] [PubMed]
- Kujala, U.M.; Peltonen, M.; Laine, M.K.; Kaprio, J.; Heinonen, O.J.; Sundvall, J.; Eriksson, J.G.; Jula, A.; Sarna, S.; Kainulainen, H. Branched-Chain Amino Acid Levels Are Related with Surrogates of Disturbed Lipid Metabolism among Older Men. Front. Med. 2016, 3, 57. [Google Scholar] [CrossRef] [PubMed]
- Newgard, C.B.; An, J.; Bain, J.R.; Muehlbauer, M.J.; Stevens, R.D.; Lien, L.F.; Haqq, A.M.; Shah, S.H.; Arlotto, M.; Slentz, C.A.; et al. A branched-chain amino acid- related metabolic signature that differentiates obese and lean humans and contributes to insulin resistance. Cell Metab. 2009, 9, 311–326. [Google Scholar] [CrossRef]
- Palmer, N.D.; Stevens, R.D.; Antinozzi, P.A.; Anderson, A.; Bergman, R.N.; Wagenknecht, L.E.; Newgard, C.B.; Bowden, D.W. Metabolomic profile associated with insulin resistance and conversion to diabetes in the Insulin Resistance Atherosclerosis Study. J. Clin. Endocr. Metab. 2015, 100, E463–E468. [Google Scholar] [CrossRef]
- Lotta, L.A.; Scott, R.A.; Sharp, S.J.; Burgess, S.; Luan, J.A.; Tillin, T.; Schmidt, A.F.; Imamura, F.; Stewart, I.D.; Perry, J.R.; et al. Genetic predisposition to an impaired metabolism of the branched-chain amino acids and risk of type 2 diabetes: A Mendelian randomisation analysis. PLoS Med. 2016, 13, e1002179. [Google Scholar] [CrossRef]
- UK Biobank Database. Open Targets Genetics. Available online: https://genetics.opentargets.org (accessed on 11 December 2021).
- Weiner, J.S.; Lourie, J.A. Human Biology: A Guide to Field Methods; Blackwell Scientific: Hoboken, NJ, USA, 1969; p. 621. [Google Scholar]
- Shanely, R.A.; Zwetsloot, K.A.; Triplett, N.T.; Meaney, M.P.; Farris, G.E.; Nieman, D.C. Human skeletal muscle biopsy procedures using the modified Bergström technique. J. Vis. Exp. 2014, 10, 51812. [Google Scholar]
- Xia, Z.; Cholewa, J.; Zhao, Y.; Yang, Y.Q.; Shang, H.Y.; Guimarães-Ferreira, L.; Naimo, M.A.; Su, Q.S.; Zanchi, N.E. Hypertrophy-promoting effects of leucine supplementation and moderate intensity aerobic exercise in pre-senescent mice. Nutrients 2016, 8, 246. [Google Scholar] [CrossRef]
- Ahmetov, I.I.; Mozhayskaya, I.A.; Lyubaeva, E.V.; Vinogradova, O.L.; Rogozkin, V.A. PPARG Gene polymorphism and locomotor activity in humans. Bull. Exp. Biol. Med. 2008, 146, 630–632. [Google Scholar] [CrossRef] [PubMed]
- Seaborne, R.A.; Hughes, D.C.; Turner, D.C.; Owens, D.J.; Baehr, L.M.; Gorski, P.; Semenova, E.A.; Borisov, O.V.; Larin, A.K.; Popov, D.V.; et al. UBR5 is a novel E3 ubiquitin ligase involved in skeletal muscle hypertrophy and recovery from atrophy. J. Physiol. 2019, 597, 3727–3749. [Google Scholar] [CrossRef] [PubMed]
- Guilherme, J.P.L.F.; Semenova, E.A.; Borisov, O.V.; Larin, A.K.; Moreland, E.; Generozov, E.V.; Ahmetov, I.I. Genomic predictors of testosterone levels are associated with muscle fiber size and strength. Eur. J. Appl. Physiol. 2022, 122, 415–423. [Google Scholar] [CrossRef] [PubMed]
- Grishina, E.E.; Zmijewski, P.; Semenova, E.A.; Cieszczyk, P.; Huminska-Lisowska, K.; Michalowska-Sawczyn, M.; Maculewicz, E.; Crewther, B.; Orysiak, J.; Kostryukova, E.S.; et al. Three DNA polymorphisms previously identified as markers for handgrip strength are associated with strength in weightlifters and muscle fiber hypertrophy. J. Strength Cond. Res. 2019, 33, 2602–2607. [Google Scholar] [CrossRef] [PubMed]
- Bell, R.A.; Al-Khalaf, M.; Megeney, L.A. The beneficial role of proteolysis in skeletal muscle growth and stress adaptation. Skelet. Muscle 2016, 6, 1–13. [Google Scholar]
- Areces, F.; González-Millán, C.; Salinero, J.J.; Abian-Vicen, J.; Lara, B.; Gallo-Salazar, C.; Ruiz-Vicente, D.; Del Coso, J. Changes in serum free amino acids and muscle fatigue experienced during a half-ironman triathlon. PLoS ONE 2015, 10, e0138376. [Google Scholar]
- Roelands, B.; de Koning, J.; Foster, C.; Hettinga, F.; Meeusen, R. Neurophysiological determinants of theoretical concepts and mechanisms involved in pacing. Sports Med. 2013, 43, 301–311. [Google Scholar] [CrossRef]
- Greer, B.K.; White, J.P.; Arguello, E.M.; Haymes, E.M. Branched-chain amino acid supplementation lowers perceived exertion but does not affect performance in untrained males. J. Strength Cond. Res. 2011, 25, 539–544. [Google Scholar] [CrossRef]
- Yasuda, T.; Abe, T.; Sato, Y.; Midorikawa, T.; Kearns, C.F.; Inoue, K.; Ryushi, T.; Ishii, N. Muscle fiber cross-sectional area is increased after two weeks of twice daily KAATSU-resistance training. Int. J. KAATSU Train. Res. 2005, 1, 65–70. [Google Scholar] [CrossRef]
- Oyarzabal, A.; Martínez-Pardo, M.; Merinero, B.; Navarrete, R.; Desviat, L.R.; Ugarte, M.; Rodríguez-Pombo, P. A novel regulatory defect in the branched-chain α-keto acid dehydrogenase complex due to a mutation in the PPM1K gene causes a mild variant phenotype of maple syrup urine disease. Hum. Mutat. 2013, 34, 355–362. [Google Scholar] [CrossRef]
- Larsson, S.C.; Markus, H.S. Branched-chain amino acids and Alzheimer’s disease: A Mendelian randomization analysis. Sci. Rep. 2017, 7, 13604. [Google Scholar] [CrossRef] [PubMed]
- Ou, H.J.; Huang, G.; Liu, W.; Ma, X.L.; Wei, Y.; Zhou, T.; Pan, Z.M. Relationship of the APOA5/A4/C3/A1 gene cluster and APOB gene polymorphisms with dyslipidemia. Genet. Mol. Res. 2015, 14, 9277–9290. [Google Scholar] [CrossRef] [PubMed]
- Costanza, M.C.; Beer-Borst, S.; James, R.W.; Gaspoz, J.M.; Morabia, A. Consistency between cross-sectional and longitudinal SNP: Blood lipid associations. Eur. J. Epidemiol. 2012, 27, 131–138. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Singmann, P.; Baumert, J.; Herder, C.; Meisinger, C.; Holzapfel, C.; Klopp, N.; Wichmann, H.E.; Klingenspor, M.; Rathmann, W.; Illig, T.; et al. Gene-gene interaction between APOA5 and USF1: Two candidate genes for the metabolic syndrome. Obes. Facts. 2009, 2, 235–242. [Google Scholar] [CrossRef]
- Hua, S.; Ma, C.; Zhang, J.; Li, J.; Wu, W.; Xu, N.; Luo, G.; Zhao, J. Influence of APOA5 Locus on the Treatment Efficacy of Three Statins: Evidence from a Randomized Pilot Study in Chinese Subjects. Front. Pharmacol. 2018, 9, 352. [Google Scholar] [CrossRef]
- Mourier, A.; Bigard, A.X.; de Kerviler, E.; Roger, B.; Legrand, H.; Guezennec, C.Y. Combined effects of caloric restriction and branched-chain amino acid supple- mentation on body composition and exercise performance in elite wrestlers. Int. J. Sports Med. 1997, 18, 47–55. [Google Scholar] [CrossRef]
| SNP | Gene | Metabolite | Increasing Allele | p Value | Reference | Other Traits (UK Biobank) |
|---|---|---|---|---|---|---|
| rs1440580 | PPM1K | Valine | A | 1.3 × 10−31 | [19] | Whole body fat-free mass (p = 0.011) |
| Leucine | 1.9 × 10−24 | |||||
| Isoleucine | 1.7 × 10−14 | |||||
| rs12325419 | DDX19B | Leucine | G | 4.5 × 10−8 | [19] | BMI (p = 2.1 × 10−7) |
| rs2072560 | APOA5 | Valine | C | 3.3 × 10−9 | [19] | IGF1 levels (p = 0.025) |
| rs58101275 | TRMT61A | Isoleucine | G | 2.78 × 10−8 | [18] | Whole body fat-free mass (p = 3.7 × 10−7) |
| rs1420601 | CBLN1 | Isoleucine | C | 3.71 × 10−8 | [18] | Whole body fat-free mass (p = 0.0033) |
| Characteristics | Non-WPS | WPS | p Value |
|---|---|---|---|
| n = 53 | n = 22 | ||
| Height (m) | 1.81 (6.24) | 1.79 (5.72) | 0.370 |
| Weight (kg) | 79.5 (8.7) | 73.8 (7.3) | 0.009 * |
| BMI (kg/m2) | 24.4 (2.2) | 23.0 (1.7) | 0.01 * |
| Age (y) | 32.1 (8.7) | 31.5 (8.3) | 0.779 |
| FTMF CSA (μm2) | 5367 (1321) | 5634.10 (1691) | 0.467 |
| STMF CSA (μm2) | 5237 (1302) | 5157.35 (1099) | 0.801 |
| All Fiber CSA (μm2) | 10,604 (2377) | 10,791.41 (2277) | 0.755 |
| FTMF (%) | 53.1 (17.0) | 45.5 (13.8) | 0.066 |
| STMF (%) | 50.6 (17.7) | 56.9 (13.9) | 0.138 |
| Physical activity (sessions) | 6.0 (4.5) | 7.7 (3.8) | 0.078 |
| Frequency of meat intake | 5.0 (1.9) | 5.8 (1.5) | 0.121 |
| Frequency of dairy intake | 4.2 (2.2) | 4.1 (2.3) | 0.879 |
| (n = 25) | (n = 14) | ||
| Fat mass (kg) | 16.0 (5.3) | 11.4 (2.8) | 0.005 * |
| Fat percentage (%) | 20.2 (5.4) | 15.6 (3.0) | 0.006 * |
| Muscle mass (kg) | 42.9 (3.9) | 46.0 (2.0) | 0.01 * |
| SNP | Genotype | Non-WPS | WPS |
|---|---|---|---|
| PPM1K | TT | 14 (26.4) | 5 (22.7) |
| rs1440580 | TA | 28 (52.8) | 10 (45.5) |
| AA | 11 (20.8) | 7 (31.8) | |
| MAF | 0.472 | 0.545 | |
| APOA5 | CC | 47 (88.7) | 20 (90.9) |
| rs2072560 | CT | 6 (11.3) | 2 (9.1) |
| TT | 0 (0.0) | 0 (0.0) | |
| MAF | 0.057 | 0.045 | |
| CBLN1 | TT | 13 (24.5) | 7 (31.8) |
| rs1420601 | TC | 28 (52.8) | 12 (54.5) |
| CC | 12 (22.6) | 3 (13.6) | |
| MAF | 0.491 | 0.409 | |
| DDX19B | GG | 39 (73.6) | 16 (72.7) |
| rs12325419 | GA | 13 (24.5) | 6 (27.3) |
| AA | 1 (1.9) | 0 (0.0) | |
| MAF | 0.142 | 0.136 | |
| TRMT61A | GG | 41 (77.4) | 13 (59.1) |
| rs58101275 | GA | 10 (18.9) | 8 (36.4) |
| AA | 2 (3.8) | 1 (4.5) | |
| MAF | 0.132 | 0.227 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hall, E.C.R.; Semenova, E.A.; Bondareva, E.A.; Andryushchenko, L.B.; Larin, A.K.; Cięszczyk, P.; Generozov, E.V.; Ahmetov, I.I. Association of Genetically Predicted BCAA Levels with Muscle Fiber Size in Athletes Consuming Protein. Genes 2022, 13, 397. https://doi.org/10.3390/genes13030397
Hall ECR, Semenova EA, Bondareva EA, Andryushchenko LB, Larin AK, Cięszczyk P, Generozov EV, Ahmetov II. Association of Genetically Predicted BCAA Levels with Muscle Fiber Size in Athletes Consuming Protein. Genes. 2022; 13(3):397. https://doi.org/10.3390/genes13030397
Chicago/Turabian StyleHall, Elliott C. R., Ekaterina A. Semenova, Elvira A. Bondareva, Liliya B. Andryushchenko, Andrey K. Larin, Pawel Cięszczyk, Edward V. Generozov, and Ildus I. Ahmetov. 2022. "Association of Genetically Predicted BCAA Levels with Muscle Fiber Size in Athletes Consuming Protein" Genes 13, no. 3: 397. https://doi.org/10.3390/genes13030397
APA StyleHall, E. C. R., Semenova, E. A., Bondareva, E. A., Andryushchenko, L. B., Larin, A. K., Cięszczyk, P., Generozov, E. V., & Ahmetov, I. I. (2022). Association of Genetically Predicted BCAA Levels with Muscle Fiber Size in Athletes Consuming Protein. Genes, 13(3), 397. https://doi.org/10.3390/genes13030397







