Aggregation of Genome-Wide Association Data from FinnGen and UK Biobank Replicates Multiple Risk Loci for Pregnancy Complications
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection
2.2. GWAS Meta-Analysis
2.3. Functional Annotation and Genetic Correlation Analysis
2.4. Data Availability
3. Results
3.1. Characterization of Significant Associations for Pregnancy Complications
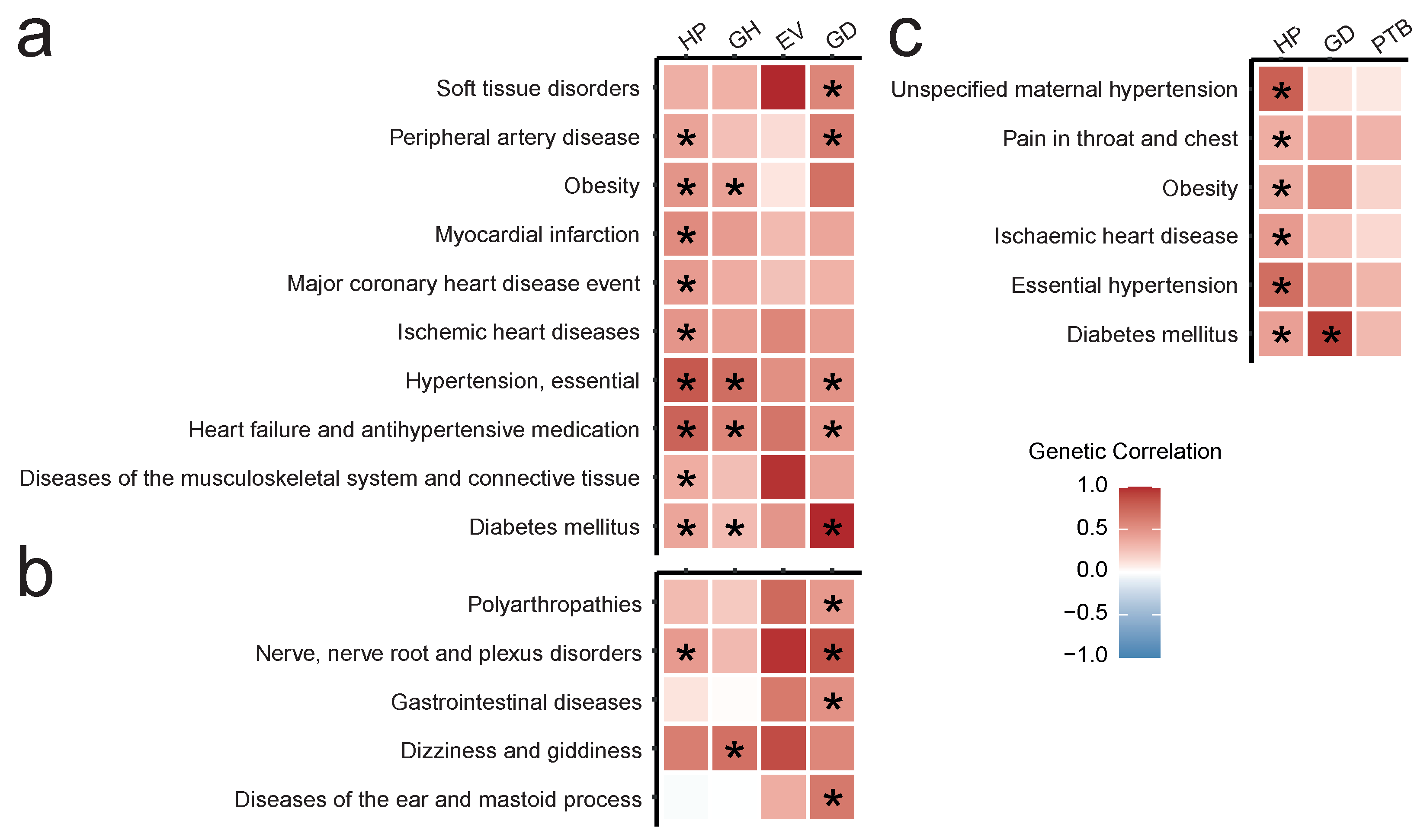
3.2. Annotation of GWAS Results for Pregnancy Complications
3.3. Replication of Known Associations in the FG and UKB Data
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| GWAS | Genome-wide association study |
| SNP | Single-nucleotide polymorphism |
| UKB | UK Biobank |
| FG | FinnGen |
| T2D | Type 2 Diabetes |
| HP | Hypertension complicating pregnancy, childbirth, and the puerperium |
| GH | Gestational hypertension |
| EV | Excessive vomiting in pregnancy |
| HG | Hyperemesis gravidarum |
| GD | Gestational diabetes |
| PTB | Preterm birth |
References
- Dalfrà, M.G.; Burlina, S.; Del Vescovo, G.G.; Lapolla, A. Genetics and Epigenetics: New Insight on Gestational Diabetes Mellitus. Front. Endocrinol. 2020, 11, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Sibai, B.; Dekker, G.; Kupferminc, M. Pre-eclampsia. Lancet 2005, 365, 785–799. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Moley, K.H.; Gronowski, A.M. Diagnostic potential for miRNAs as biomarkers for pregnancy-specific diseases. Clin. Biochem. 2013, 46, 953–960. [Google Scholar] [CrossRef] [PubMed]
- Melchiorre, K.; Thilaganathan, B.; Giorgione, V.; Ridder, A.; Memmo, A.; Khalil, A. Hypertensive Disorders of Pregnancy and Future Cardiovascular Health. Front. Cardiovasc. Med. 2020, 7, 59. [Google Scholar] [CrossRef]
- Phipps, E.A.; Thadhani, R.; Benzing, T.; Karumanchi, S.A. Pre-eclampsia: Pathogenesis, novel diagnostics and therapies. Nat. Rev. Nephrol. 2019, 15, 275–289. [Google Scholar] [CrossRef]
- Guriev, D.L.; Stepanova, E.M.; Soloukhina, O.E.; Sinkevich, L.A.; Muradova, M.A.; Ershova, Y.V. Implementing Federal Clinical Protocol “Gestational Diabetes Mellitus” in Yaroslavl Region. Doctor Ru 2016, 7, 61–66. [Google Scholar]
- Dreval’, A.V.; Shestakova, T.P.; Bunak, I.V. Gestational Diabetes Mellitus (Based on the Results of a Screening Study in the Moscow Region). Alm. Clin. Med. 2016, 44, 406–413. [Google Scholar] [CrossRef][Green Version]
- Say, L.; Chou, D.; Gemmill, A.; Tunçalp, Ö.; Moller, A.B.; Daniels, J.; Gülmezoglu, A.M.; Temmerman, M.; Alkema, L. Global causes of maternal death: A WHO systematic analysis. Lancet Glob. Health 2014, 2, e323–e333. [Google Scholar] [CrossRef]
- Neiger, R. Long-term effects of pregnancy complications on maternal health: A review. J. Clin. Med. 2017, 6, 76. [Google Scholar] [CrossRef]
- Schaid, D.J.; Chen, W.; Larson, N.B. From genome-wide associations to candidate causal variants by statistical fine-mapping. Nat. Rev. Genet. 2018, 19, 491–504. [Google Scholar] [CrossRef]
- Steinthorsdottir, V.; McGinnis, R.; Williams, N.O.; Stefansdottir, L.; Thorleifsson, G.; Shooter, S.; Fadista, J.; Sigurdsson, J.K.; Auro, K.M.; Berezina, G.; et al. Genetic predisposition to hypertension is associated with preeclampsia in European and Central Asian women. Nat. Commun. 2020, 11, 5976. [Google Scholar] [CrossRef] [PubMed]
- Kwak, S.H.; Kim, S.H.; Cho, Y.M.; Go, M.J.; Cho, Y.S.; Choi, S.H.; Moon, M.K.; Jung, H.S.; Shin, H.D.; Kang, H.M.; et al. A genome-wide association study of gestational diabetes mellitus in Korean women. Diabetes 2012, 61, 531–541. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Feenstra, B.; Bacelis, J.; Liu, X.; Muglia, L.M.; Juodakis, J.; Miller, D.E.; Litterman, N.; Jiang, P.P.; Russell, L.; et al. Genetic Associations with Gestational Duration and Spontaneous Preterm Birth. N. Engl. J. Med. 2017, 377, 1156–1167. [Google Scholar] [CrossRef] [PubMed]
- Barbitoff, Y.A.; Tsarev, A.A.; Vashukova, E.S.; Maksiutenko, E.M.; Kovalenko, L.V.; Belotserkovtseva, L.D.; Glotov, A.S. A Data-Driven Review of the Genetic Factors of Pregnancy Complications. Int. J. Mol. Sci. 2020, 21, 3384. [Google Scholar] [CrossRef] [PubMed]
- Bycroft, C.; Freeman, C.; Petkova, D.; Band, G.; Elliott, L.T.; Sharp, K.; Motyer, A.; Vukcevic, D.; Delaneau, O.; O’Connell, J.; et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 2018, 562, 203–209. [Google Scholar] [CrossRef] [PubMed]
- Kurki, M.I.; Karjalainen, J.; Palta, P.; Sipilä, T.P.; Kristiansson, K.; Donner, K.; Reeve, M.P.; Laivuori, H.; Aavikko, M.; Kaunisto, M.A.; et al. FinnGen: Unique genetic insights from combining isolated population and national health register data. medRxiv 2022. [Google Scholar] [CrossRef]
- Hinrichs, A.S.; Karolchik, D.; Baertsch, R.; Barber, G.P.; Bejerano, G.; Clawson, H.; Diekhans, M.; Furey, T.S.; Harte, R.A.; Hsu, F.; et al. The UCSC Genome Browser Database: Update 2006. Nucleic Acids Res. 2006, 34, D590–D598. [Google Scholar] [CrossRef]
- Willer, C.J.; Li, Y.; Abecasis, G.R. METAL: Fast and efficient meta-analysis of genomewide association scans. Bioinformatics 2010, 26, 2190–2191. [Google Scholar] [CrossRef]
- Turley, P.; Walters, R.K.; Maghzian, O.; Okbay, A.; Lee, J.J.; Fontana, M.A.; Nguyen-Viet, T.A.; Wedow, R.; Zacher, M.; Furlotte, N.A.; et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat. Genet. 2018, 50, 229–237. [Google Scholar] [CrossRef]
- Watanabe, K.; Taskesen, E.; van Bochoven, A.; Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 2017, 8, 1826. [Google Scholar] [CrossRef]
- Speed, D.; Cai, N.; Johnson, M.R.; Nejentsev, S.; Balding, D.J. Reevaluation of SNP heritability in complex human traits. Nat. Genet. 2017, 49, 986–992. [Google Scholar] [CrossRef] [PubMed]
- Ganna, A.; Verweij, K.J.; Nivard, M.G.; Maier, R.; Wedow, R.; Busch, A.S.; Abdellaoui, A.; Guo, S.; Fah Sathirapongsasuti, J.; Team, R.; et al. Large-scale GWAS reveals insights into the genetic architecture of same-sex sexual behavior. Science 2019, 365, aat7693. [Google Scholar] [CrossRef] [PubMed]
- Niemi, M.E.K.; Karjalainen, J.; Liao, R.G.; Neale, B.M.; Daly, M.; Ganna, A.; Pathak, G.A.; Andrews, S.J.; Kanai, M.; Veerapen, K.; et al. Mapping the human genetic architecture of COVID-19. Nature 2021, 600, 472–477. [Google Scholar] [CrossRef]
- Benner, C.; Spencer, C.C.; Havulinna, A.S.; Salomaa, V.; Ripatti, S.; Pirinen, M. FINEMAP: Efficient variable selection using summary data from genome-wide association studies. Bioinformatics 2016, 32, 1493–1501. [Google Scholar] [CrossRef] [PubMed]
- Auton, A.; Abecasis, G.R.; Altshuler, D.M.; Durbin, R.M.; Bentley, D.R.; Chakravarti, A.; Clark, A.G.; Donnelly, P.; Eichler, E.E.; Flicek, P.; et al. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar] [CrossRef]
- Tyrmi, J.S.; Kaartokallio, T.; Lokki, I.; Jääskeläinen, T.; Kortelainen, E.; Ruotsalainen, S.; Karjalainen, J.; Ripatti, S.; Laisk, T.; Kettunen, J.; et al. GWAS of preeclampsia and hypertensive disorders of pregnancy uncovers genes related to cardiometabolic, endothelial and placental function. medRxiv 2022. [Google Scholar] [CrossRef]
- Fejzo, M.S.; Sazonova, O.V.; Sathirapongsasuti, J.F.; Hallgrímsdóttir, I.B.; Vacic, V.; MacGibbon, K.W.; Schoenberg, F.P.; Mancuso, N.; Slamon, D.J.; Mullin, P.M.; et al. Placenta and appetite genes GDF15 and IGFBP7 are associated with hyperemesis gravidarum. Nat. Commun. 2018, 9, 1178. [Google Scholar] [CrossRef] [PubMed]
- Jain, V.G.; Monangi, N.; Zhang, G.; Muglia, L.J. Genetics, epigenetics, and transcriptomics of preterm birth. Am. J. Reprod. Immunol. 2022, 88, e13600. [Google Scholar] [CrossRef] [PubMed]
- Gupta, J.K.; Care, A.; Goodfellow, L.; Alfirevic, Z.; Müller-Myhsok, B.; Alfirevic, A. Genome and transcriptome profiling of spontaneous preterm birth phenotypes. Sci. Rep. 2022, 12, 1003. [Google Scholar] [CrossRef]
- Hong, X.; Hao, K.; Ji, H.; Peng, S.; Sherwood, B.; Di Narzo, A.; Tsai, H.J.; Liu, X.; Burd, I.; Wang, G.; et al. Genome-wide approach identifies a novel gene-maternal pre-pregnancy BMI interaction on preterm birth. Nat. Commun. 2017, 8, 15608. [Google Scholar] [CrossRef]
- Rappoport, N.; Toung, J.; Hadley, D.; Wong, R.J.; Fujioka, K.; Reuter, J.; Abbott, C.W.; Oh, S.; Hu, D.; Eng, C.; et al. A genome-wide association study identifies only two ancestry specific variants associated with spontaneous preterm birth. Sci. Rep. 2018, 8, 226. [Google Scholar] [CrossRef] [PubMed]
- Laisk, T.; Soares, A.L.G.; Ferreira, T.; Painter, J.N.; Censin, J.C.; Laber, S.; Bacelis, J.; Chen, C.Y.; Lepamets, M.; Lin, K.; et al. The genetic architecture of sporadic and multiple consecutive miscarriage. Nat. Commun. 2020, 11, 5980. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Helenius, D.; Skotte, L.; Beaumont, R.N.; Wielscher, M.; Geller, F.; Juodakis, J.; Mahajan, A.; Bradfield, J.P.; Lin, F.T.J.; et al. Variants in the fetal genome near pro-inflammatory cytokine genes on 2q13 associate with gestational duration. Nat. Commun. 2019, 10, 3927. [Google Scholar] [CrossRef] [PubMed]
- Schierding, W.; Antony, J.; Karhunen, V.; Vääräsmäki, M.; Franks, S.; Elliott, P.; Kajantie, E.; Sebert, S.; Blakemore, A.; Horsfield, J.A.; et al. GWAS on prolonged gestation (post-term birth): Analysis of successive Finnish birth cohorts. J. Med. Genet. 2018, 55, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Pervjakova, N.; Moen, G.h.; Borges, M.c.; Ferreira, T.; Cook, J.P.; Allard, C.; Beaumont, R.N.; Canouil, M.; Hatem, G.; Heiskala, A.; et al. Multi-ancestry genome-wide association study of gestational diabetes mellitus highlights genetic links with type 2 diabetes. Hum. Mol. Genet. 2022, 31, 3377–3391. [Google Scholar] [CrossRef]
- Buniello, A.; MacArthur, J.A.L.; Cerezo, M.; Harris, L.W.; Hayhurst, J.; Malangone, C.; McMahon, A.; Morales, J.; Mountjoy, E.; Sollis, E.; et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 2019, 47, D1005–D1012. [Google Scholar] [CrossRef]
- Ardissino, M.; Slob, E.A.; Millar, O.; Reddy, R.K.; Lazzari, L.; Patel, K.H.K.; Ryan, D.; Johnson, M.R.; Gill, D.; Ng, F.S. Maternal Hypertension Increases Risk of Preeclampsia and Low Fetal Birthweight: Genetic Evidence From a Mendelian Randomization Study. Hypertension 2022, 79, 588–598. [Google Scholar] [CrossRef]
- Xin, Q.; Han, Y.; Jiang, W.; Wang, J.; Luan, Y.; Ji, Q.; Sun, W. Genetic susceptibility analysis of FGF5 polymorphism to preeclampsia in Chinese Han population. Mol. Genet. Genom. 2022, 297, 791–800. [Google Scholar] [CrossRef]
- Yu, S.; Choi, W.I.; Choi, Y.J.; Kim, H.Y.; Hildebrandt, F.; Gee, H.Y. PLCE1 regulates the migration, proliferation, and differentiation of podocytes. Exp. Mol. Med. 2020, 52, 594–603. [Google Scholar] [CrossRef]
- Shao, H.; Andres, D.A. A Novel RalGEF-like Protein, RGL3, as a Candidate Effector for Rit and Ras. J. Biol. Chem. 2000, 275, 26914–26924. [Google Scholar] [CrossRef]
- Kichaev, G.; Bhatia, G.; Loh, P.R.; Gazal, S.; Burch, K.; Freund, M.K.; Schoech, A.; Pasaniuc, B.; Price, A.L. Leveraging Polygenic Functional Enrichment to Improve GWAS Power. Am. J. Hum. Genet. 2019, 104, 65–75. [Google Scholar] [CrossRef] [PubMed]
- Huopio, H.; Cederberg, H.; Vangipurapu, J.; Hakkarainen, H.; Paakkonen, M.; Kuulasmaa, T.; Heinonen, S.; Laakso, M. Association of risk variants for type 2 diabetes and hyperglycemia with gestational diabetes. Eur. J. Endocrinol. 2013, 169, 291–297. [Google Scholar] [CrossRef] [PubMed]
- Popova, P.V.; Klyushina, A.A.; Vasilyeva, L.B.; Tkachuk, A.S.; Vasukova, E.A.; Anopova, A.D.; Pustozerov, E.A.; Gorelova, I.V.; Kravchuk, E.N.; Li, O.; et al. Association of Common Genetic Risk Variants With Gestational Diabetes Mellitus and Their Role in GDM Prediction. Front. Endocrinol. 2021, 12, 628582. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Yang, Y.; Liu, X.; Li, Z.; Liu, H.; Tan, Q. Glucose metabolism-related gene polymorphisms as the risk predictors of type 2 diabetes. Diabetol. Metab. Syndr. 2020, 12, 97. [Google Scholar] [CrossRef]
- McElwain, C.J.; McCarthy, F.P.; McCarthy, C.M. Gestational diabetes mellitus and maternal immune dysregulation: What we know so far. Int. J. Mol. Sci. 2021, 22, 4261. [Google Scholar] [CrossRef]
- Shikov, A.E.; Skitchenko, R.K.; Predeus, A.V.; Barbitoff, Y.A. Phenome-wide functional dissection of pleiotropic effects highlights key molecular pathways for human complex traits. Sci. Rep. 2020, 10, 1037. [Google Scholar] [CrossRef]
- Moore, A.G.; Brown, D.A.; Fairlie, W.D.; Bauskin, A.R.; Brown, P.K.; Munier, M.L.; Russell, P.K.; Salamonsen, L.A.; Wallace, E.M.; Breit, S.N. The transforming growth factor-β superfamily cytokine macrophage inhibitory cytokine-1 is present in high concentrations in the serum of pregnant women. J. Clin. Endocrinol. Metab. 2000, 85, 4781–4788. [Google Scholar] [CrossRef]
- Tsai, V.W.W.; Macia, L.; Johnen, H.; Kuffner, T.; Manadhar, R.; Jørgensen, S.B.; Lee-Ng, K.K.M.; Zhang, H.P.; Wu, L.; Marquis, C.P.; et al. TGF-b Superfamily Cytokine MIC-1/GDF15 Is a Physiological Appetite and Body Weight Regulator. PLoS ONE 2013, 8, e55174. [Google Scholar] [CrossRef]
- Källén, B. Hyperemesis during pregnancy and delivery outcome: A registry study. Eur. J. Obstet. Gynecol. Reprod. Biol. 1987, 26, 291–302. [Google Scholar] [CrossRef]
- Zhou, G.; Holzman, C.; Heng, Y.J.; Kibschull, M.; Lye, S.J.; Vazquez, A. EBF1 Gene mRNA Levels in Maternal Blood and Spontaneous Preterm Birth. Reprod. Sci. 2020, 27, 316–324. [Google Scholar] [CrossRef]
- Györy, I.; Boller, S.; Nechanitzky, R.; Mandel, E.; Pott, S.; Liu, E.; Grosschedl, R. Transcription factor EBf1 regulates differentiation stage-specific signaling, proliferation, and survival of B cells. Genes Dev. 2012, 26, 668–682. [Google Scholar] [CrossRef] [PubMed]
- Ehret, G.B.; Munroe, P.B.; Rice, K.M.; Bochud, M.; Johnson, A.D.; Chasman, D.I.; Smith, A.V.; Tobin, M.D.; Verwoert, G.C.; Hwang, S.J.; et al. Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature 2011, 478, 103–109. [Google Scholar] [CrossRef] [PubMed]

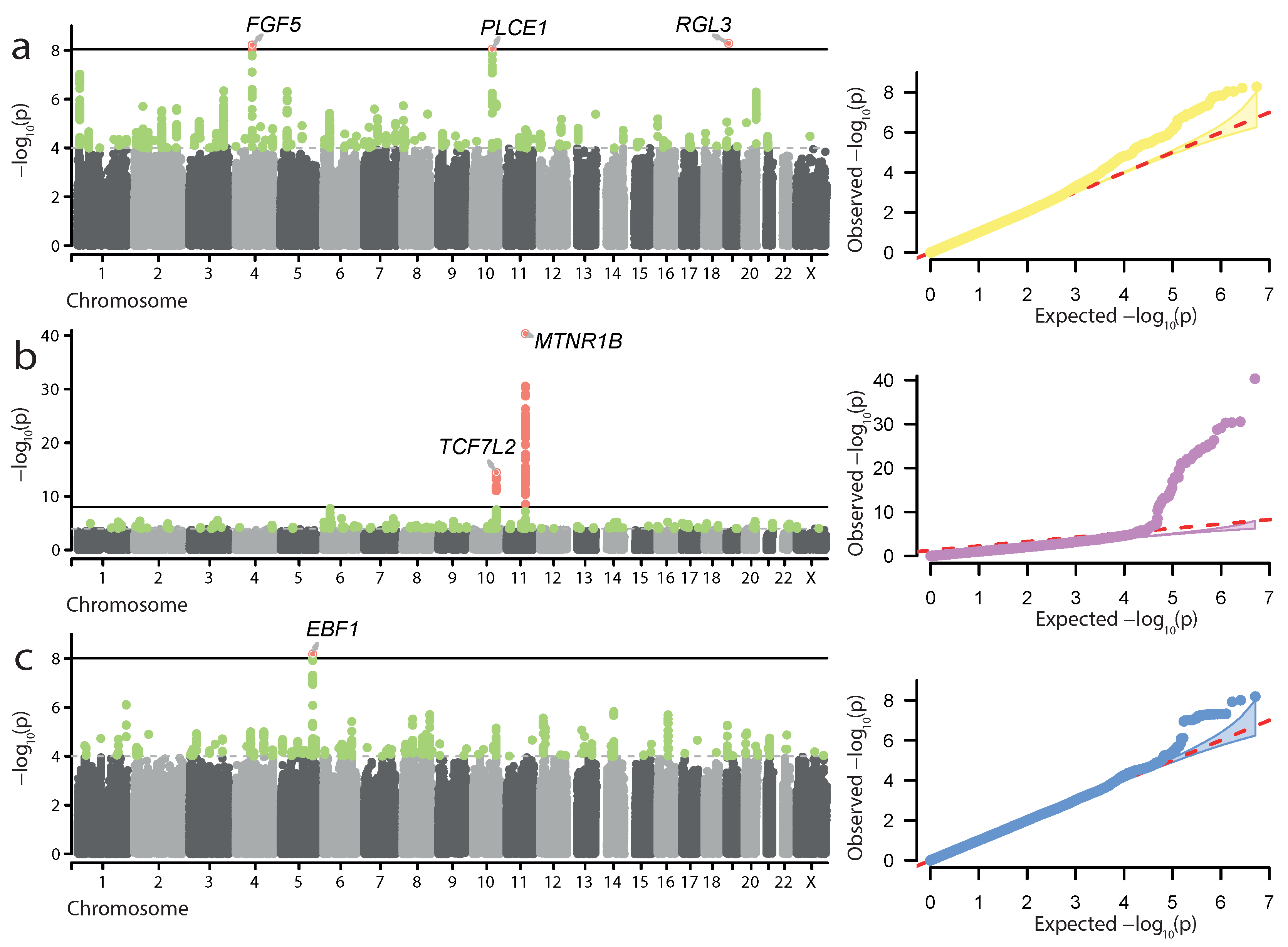
| Trait | Lead Variant | rsID | Gene | Effect | p-Value | Meta p-Value |
|---|---|---|---|---|---|---|
| HP | 1:11865804:A>G | rs13306561 | MTHFR | intron variant | ||
| 4:81188513:C>A | rs35954793 | FGF5 | intron variant | |||
| 10:95892788:T>A | rs10882398 † | PLCE1 | intron variant | |||
| 19:11526765:T>G | rs167479 † | RGL3 | missense variant | |||
| 20:59160402:A>C | rs259983 | ZNF831 | intron variant | |||
| GH | 20:47408414:A>G | rs2208589 † | PREX1 | intron variant | ||
| EV | 19:18493064:<del> | rs58835482 | GDF15 | UTR variant | n.a. ** | |
| GD | 2:27741237:T>C | rs780094 | GCKR | intron variant | ||
| 6:32668411:G>A | rs9275373 | HLA * | n.a. | |||
| 10:114782581:G>GCT | rs10659211 | TCF7L2 | intron variant | |||
| 11:92708710:C>G | rs10830963 | MTNR1B | intron variant |
| Trait | Led Variant | rsID | Gene | Effect | FG p-Value | UKB p-Value | Direction | Meta p-Value |
|---|---|---|---|---|---|---|---|---|
| HP | 4:81188513:C>A | rs35954793 | FGF5 | intron variant | – – | |||
| 10:95892788:T>A | rs10882398 † | PLCE1 | intron variant | – – | ||||
| 19:11526765:T>G | rs167479 † | RGL3 | missense variant | ++ | ||||
| GD | 10:114774433:A>C | rs36090025 | TCF7L2 | intron variant | ++ | |||
| 11:92708710:C>G | rs10830963 | MTNR1B | intron variant | ++ | ||||
| PTB | 5:157907974:C>T | rs2963457 | EBF1 | intergenic variant | ++ |
| Variant * | rsID | Gene | Trait ** | Reference | UKB p-Value | FG p-Value | Meta p-Value |
|---|---|---|---|---|---|---|---|
| 4:57484899:<del> | rs143409503 | IGFBP7 | HG | [27,36] | |||
| 11:101390067:G>A | rs2508362 | TRPC6 | [27] | ||||
| 13:28564361:C>T | rs4769612 | FLT1 | PE | [36] | |||
| 3:169462088:C>A | rs1918975 † | MECOM | [11,36] | ||||
| 12:111395984:G>A | rs10774624 | SH2B3 | |||||
| 16:53767042:T>C | rs1421085 | FTO | |||||
| 6:20661019:G>C | rs7754840 | CDKAL1 | GD | [12,36] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Changalidis, A.I.; Maksiutenko, E.M.; Barbitoff, Y.A.; Tkachenko, A.A.; Vashukova, E.S.; Pachuliia, O.V.; Nasykhova, Y.A.; Glotov, A.S. Aggregation of Genome-Wide Association Data from FinnGen and UK Biobank Replicates Multiple Risk Loci for Pregnancy Complications. Genes 2022, 13, 2255. https://doi.org/10.3390/genes13122255
Changalidis AI, Maksiutenko EM, Barbitoff YA, Tkachenko AA, Vashukova ES, Pachuliia OV, Nasykhova YA, Glotov AS. Aggregation of Genome-Wide Association Data from FinnGen and UK Biobank Replicates Multiple Risk Loci for Pregnancy Complications. Genes. 2022; 13(12):2255. https://doi.org/10.3390/genes13122255
Chicago/Turabian StyleChangalidis, Anton I., Evgeniia M. Maksiutenko, Yury A. Barbitoff, Alexander A. Tkachenko, Elena S. Vashukova, Olga V. Pachuliia, Yulia A. Nasykhova, and Andrey S. Glotov. 2022. "Aggregation of Genome-Wide Association Data from FinnGen and UK Biobank Replicates Multiple Risk Loci for Pregnancy Complications" Genes 13, no. 12: 2255. https://doi.org/10.3390/genes13122255
APA StyleChangalidis, A. I., Maksiutenko, E. M., Barbitoff, Y. A., Tkachenko, A. A., Vashukova, E. S., Pachuliia, O. V., Nasykhova, Y. A., & Glotov, A. S. (2022). Aggregation of Genome-Wide Association Data from FinnGen and UK Biobank Replicates Multiple Risk Loci for Pregnancy Complications. Genes, 13(12), 2255. https://doi.org/10.3390/genes13122255







