1. Introduction
In the typical form of scrapie, the risk of infection is determined by variation in amino acid sequence encoded in the
PrP gene [
1,
2,
3,
4]. There are five common haplotypes (ARR, AHQ, ARH, ARQ, and VRQ) associated with the scrapie risk of infection; in which the haplotype ARR is associated with lowest risk, and the haplotype VRQ is associated with highest risk of scrapie infection [
2,
5,
6,
7]. Other additional haplotypes have been observed in sheep, but due to their extreme rarity were not considered important for breeding programs [
2,
8]. A total of 15 common possible genotype combinations are associated with five risk groups (R1, R2, R3, R4, and R5), in which R1 genotypes are associated with low risk for scrapie infection (i.e., the most favorable genotypes) and R5 are associated with highest risk of infection [
3,
8,
9]. Due to the association of genotypes with the risk of scrapie, the use of genotyping for breeding programs is appealing for scrapie eradication programs [
5,
8,
10,
11]. However, in practice, not all animals are being genotyped with breeding rams, which are more likely to be genotyped than ewes. Thus, genotypic information is limited, as only a small fraction of the total sheep population is genotyped. Gengler et al. [
12] proposed the use of a practical method to predict the allele content of bi-allelic locus in ungenotyped animals by using the Best Linear Unbiased Predictor (BLUP), in which the number of observed alleles in the genotype (0, 1, or 2) are used as a response variable, assuming complete heritability.
Selection based on risk group of the genotypes could be practiced [
3,
5,
8,
9]. However, the genotypes corresponding to the risk groups do not act additively. For animal breeding purposes, the additive genetic effect is important since offspring inherit the alleles, rather than the genotype. There is no study that accounts for the non-additivity of risk groups corresponding to the genotypes. Therefore, adjusting for the non-additive effect is needed to account for the differences in contribution of the five different haplotypes alleles to scrapie resistance in sheep.
The objectives of this research were to: (1) define numeric values of scrapie resistance genotypes and adjust them for their non-additive genetic effect; (2) evaluate the accuracy of using BLUP for prediction of scrapie resistance and allele content of ungenotyped animals; and (3) predict selection response and assess the change of genetic merit of selected ungenotyped animals based on estimated breeding value. The hypothesis of this research was that it is possible use a linear animal model for genetic evaluation and selection of ungenotyped sheep for scrapie resistance based on a small proportion of genotyped animals.
4. Discussion
The accuracy for prediction of haplotype allele contents ranged between 0.41 and 0.71 (
Table 6). Gengler et al. [
12], who first proposed the allele content model, used it for the
myostatin gene in Belgium blue cattle, with prediction accuracies between 0.47 and 0.50. In another study in Canadian Holstein cattle, the prediction accuracy for allele content was as high as 0.93 [
15]. Legarra and Vitezica [
16] reported accuracies between 0.52 and 0.56 using the allele content model. The allele content model can be applicable for prediction in bi-allelic major
genes, such as for maedi-visna [
17] and for litter size [
18]. Applying allele content model to scrapie as first proposed by Gengler et al. [
12] is possible when only considering the number ARR haplotypes and disregarding the importance of the other haplotypes. However, the
PrP gene underlying scrapie phenotypes is multi-allelic with different contributions from the different haplotypes to the level of scrapie resistance in sheep [
3,
8,
9]. In this study, the different contributions of haplotypes in the SR genotypes were considered by defining numeric values for SR and adjusting them to non-additive genetic effects prior to their use in the linear animal model. The accuracy for prediction of SR in high and low SR populations were 0.60 and 0.54, respectively. There were no previous studies that considered the same approach to construct the SR values to compare to.
Selection response on SR and allele content was predicted for ungenotyped sheep (
Table 7). The selection response depends on selection intensity, accuracy, and the additive genetic standard deviation [
14]. The differences between the trait prediction accuracies (
Table 6) and the additive genetic standard deviations (
Table 4) explain the differences in predicted responses. Selecting ungenotyped animals with EBV ≥ mean resulted in increase in genetic merit for SR and allele content (
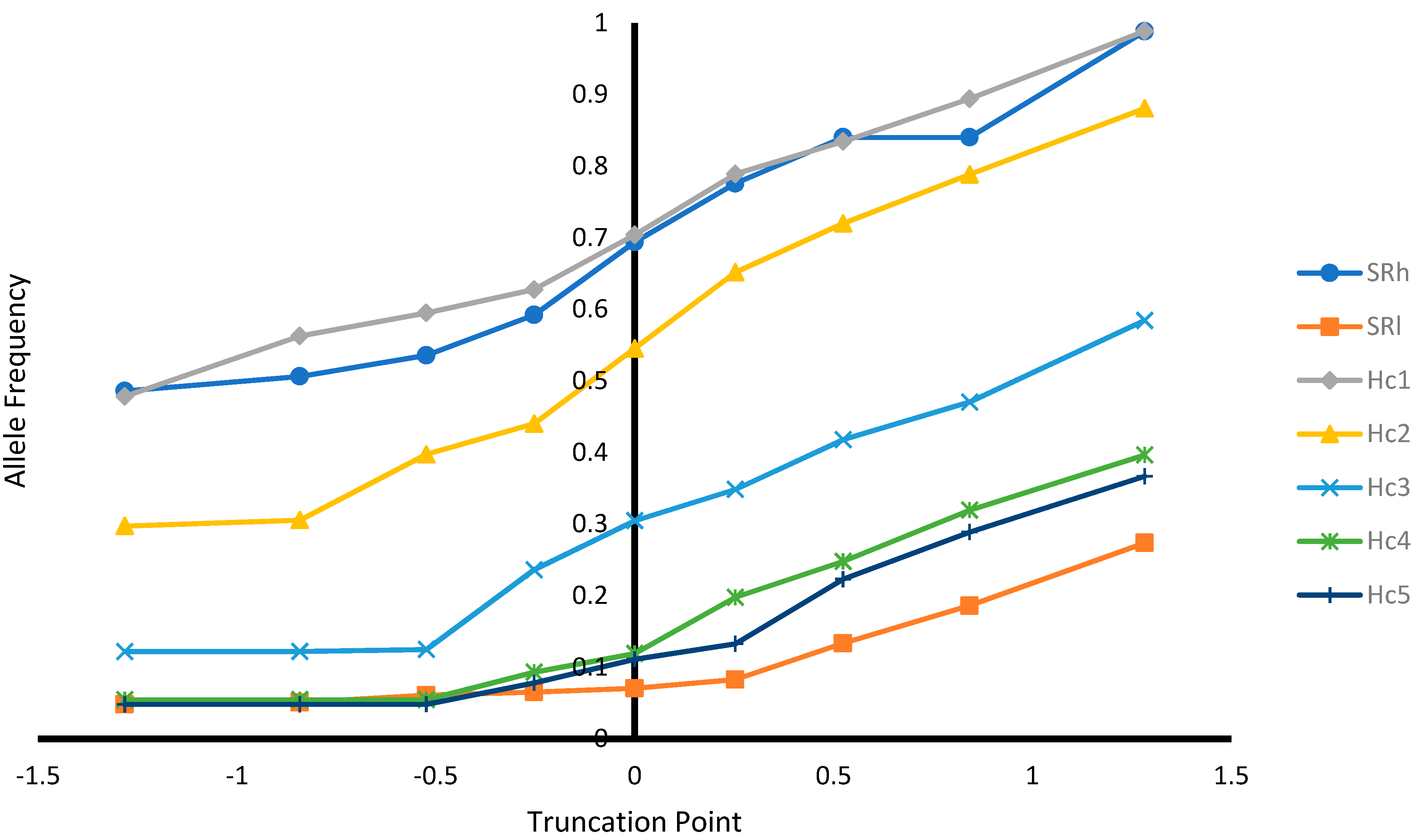
Table 8) and increased favorable haplotype allele frequency (
Figure 3). Breeding programs for genetic improvement for SR involve selection based on scrapie genotypes. Such breeding programs were successful in increasing ARR frequency and SR in Czech Republic [
8], Netherland [
19,
20], Belguim [
21], and Hungary [
5]. However, in all previous studies, the genetic change was limited to the animals being genotyped. In this research, the genetic improvement for SR was possible for the ungenotyped animals when using predictions from a linear animal model.
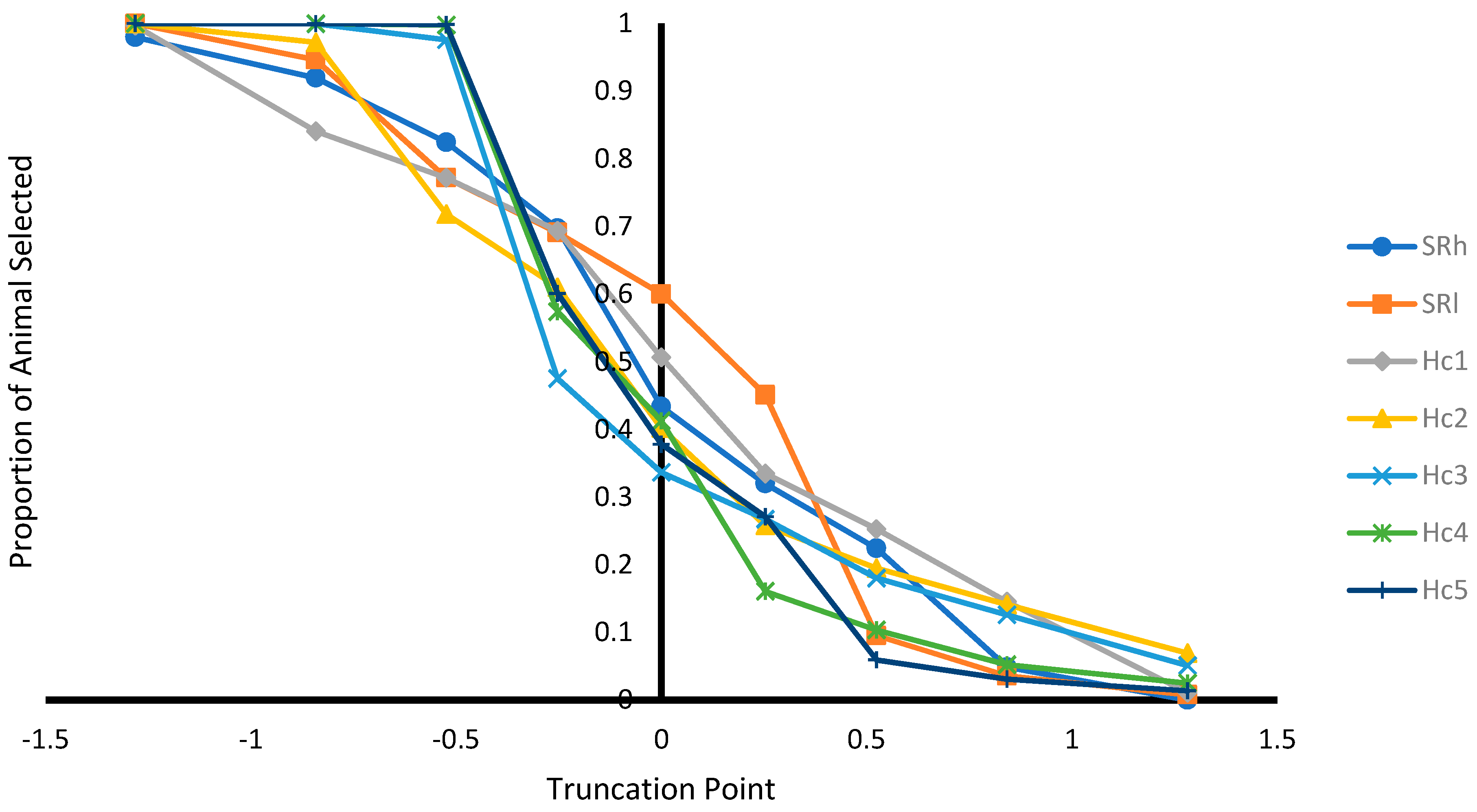
Selecting ungenotyped animals with EBV ≥ mean resulted in reduction of number of animals compare to unselected population (
Figure 1), while capturing large proportion of homozygous genotypes from the unselected population (
Figure 2). The increase of selection truncation point resulted in increased frequency for homozygous genotypes (
Figure 4), while decreasing the proportion of animals selected from the population (
Figure 1). In the breeding program in Netherlands, genotyping for selection of ARR/ARR rams was compulsory between the year 2004 and 2007 [
20]. Genotyping to identify homozygous (ARR/ARR) rams is important for breeding purposes, as they are 100% guaranteed to transmit the ARR allele to their progeny. The use of linear animal model provides another tool to reduce the number of genotyped animals by pre-selecting based on EBV prior to genotyping. The pre-selected animals with EBV ≥ mean include a large proportion of ARR/ARR genotypes from the unselected population (
Figure 2). Without pre-selection, 100% of the animals in the population must be genotyped in order to identify all the ARR/ARR animals in the population. When animals with EBV ≥ mean were pre-selected, smaller proportion (34–60%) would need to be genotyped in order to identify a large proportion (76–99%) of ARR/ARR animals (
Figure 1 and
Figure 2). Thus, genotyping cost for identifying most of the ARR/ARR animals in the population could be reduced. As pre-selection truncation point increases, the proportion of animal selected decreases (
Figure 1), but the frequency of ARR/ARR animals among the selected animals increases (
Figure 4). Thus, pre-selecting at higher truncation point would identify large proportion of ARR/ARR among the selected animals, thus saving genotyping cost to confirm the homozygous ARR/ARR status of the animals.
Breeding programs can contribute to the reduction of prevalence of the typical form of scrapie in sheep. Arnold and Rajamyagam [
22] estimated an annual reduction of 28% in scrapie prevalence cases between 2005 and 2019 in Great Britain. Hagenaars et al. [
20] reported a trend for reduction of scrapie prevalence in active scrapie surveillance in Netherland between the years 2002 and 2008. They reported 0% typical scrapie cases in genotypes with risk levels R1 and R2. The current study showed that selecting ungenotyped sheep based on EBV could increase SR (
Table 7 and
Table 8) and ARR allele frequency and its homozygous genotype frequency (
Figure 3 and
Figure 4), what could reduce typical scrapie prevalence in sheep. However, the atypical form of scrapie (Nor98) could occur in sheep that are resistant to typical scrapie [
23]. However, Nor98 scrapie type is believed to be a spontaneous disease and it is unlike to be naturally contagious among sheep and, thus, its prevalence is low [
24,
25]. Therefore, a well-established breeding program for the typical scrapie can contribute to the reduction of scrapie prevalence.
This research proposed the use of a linear animal model as a practical method for genetic evaluation and selection for SR of ungenotyped sheep. Different scrapie eradication strategies used in breeding programs were described in previous studies. For instance, Arnold et al. [
26] proposed genotyping purebred rams in the nucleus flocks used for cross-breeding and selecting the homozygous ARR/ARR and the carriers (ARR/ARQ, ARR/ARH, and ARR/AHQ) rams at the pure breeding level. Molina et al. [
3] compared different strategies in Spanish Merinos. They concluded that the optimum strategy was to genotype rams and eliminate ARQ/ARQ and VRQ carriers. According to Gáspárdy et al. [
5], in the Hungarian national breeding program, rams are genotyped and only rams at risk groups (R1, R2, and R3) are allowed to breed. In all previous studies, the genetic selection for SR was limited to the animals genotyped. This study has shown that a linear animal model can be used to provide additional information for ungenotyped animals, which will be particularly useful wherever genotyping for scrapie is not intensively practiced. Therefore, the use of an animal model could make better use of the available information to enhance breeding programs for the genetic improvement for SR in sheep.