Current Epigenetic Insights in Kidney Development
Abstract
1. Introduction
2. Overview of Epigenetic Mechanisms
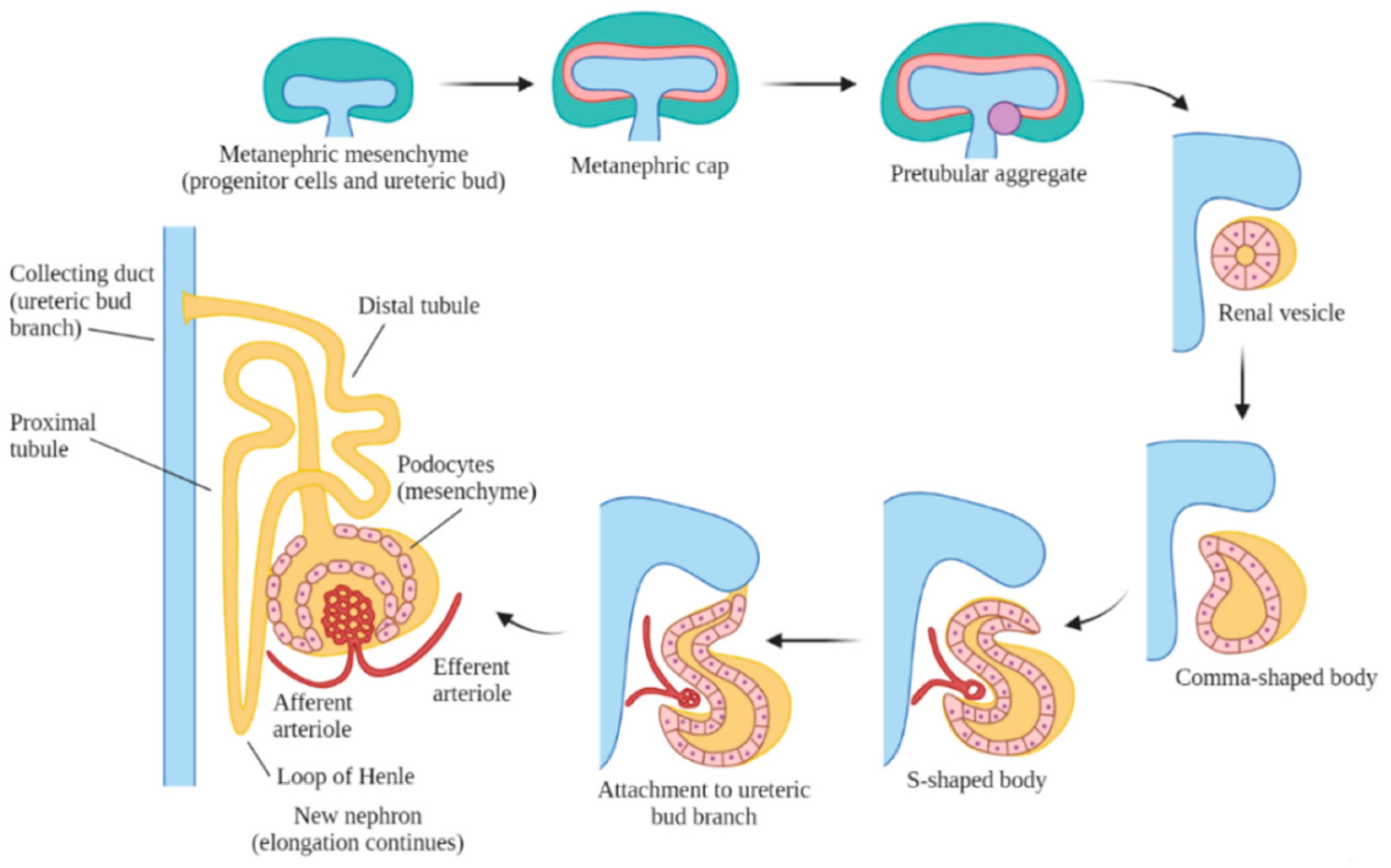
3. Overview of Human Kidney Development
4. Major Genetic Markers and Signaling Pathways in Kidney Development
5. Epigenetics in Embryonic Development
6. Major Epigenetic Markers and Epigenetic-Mediated Signaling Pathways in Kidney Development
7. The Application of Single-Cell Sequencing Techniques in Studying Kidney Development
8. Conclusions and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Acharya, V.; Olivero, J. The Kidney as an Endocrine Organ. Methodist Debakey Cardiovasc. J. 2018, 14, 305–307. [Google Scholar] [CrossRef] [PubMed]
- Al-Awqati, Q.; Oliver, J.A. Stem cells in the kidney. Kidney Int. 2002, 61, 387–395. [Google Scholar] [CrossRef] [PubMed]
- Matsell, D.G.; Hiatt, M.J. Functional Development of the Kidney in Utero. In Fetal and Neonatal Physiology; Polin, R.A., Abman, S.H., Rowitch, D.H., Benitz, W.E., Fox, W.W., Eds.; Elsevier Inc.: Philadelphia, PA, USA, 2017; pp. 965–976. [Google Scholar] [CrossRef]
- Dressler, G.R. The cellular basis of kidney development. Annu. Rev. Cell Dev. Biol. 2006, 22, 509–529. [Google Scholar] [CrossRef] [PubMed]
- Smyth, I.M.; Cullen-McEwen, L.A.; Caruana, G.; Black, M.J.; Bertram, J.F. Development of the Kidney: Morphology and Mechanisms. In Fetal and Neonatal Physiology; Polin, R.A., Abman, S.H., Rowitch, D.H., Benitz, W.E., Fox, W.W., Eds.; Elsevier Inc.: Philadelphia, PA, USA, 2017; pp. 953–964. ISBN 978-0-323-35214-7. [Google Scholar]
- Patel, S.R.; Dressler, G.R. The Genetics and Epigenetics of Kidney Development. Semin. Nephrol. 2013, 33, 314–326. [Google Scholar] [CrossRef] [PubMed]
- Saxen, L.; Sariola, H. Early organogenesis of the kidney. Pediatr. Nephrol. 1987, 1, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Dupont, C.; Armant, D.R.; Brenner, C.A. Epigenetics: Definition, mechanisms and clinical perspective. Semin. Reprod. Med. 2009, 27, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Burggren, W. Epigenetic inheritance and its role in evolutionary biology: Re-evaluation and new perspectives. Biology 2016, 5, 24. [Google Scholar] [CrossRef] [PubMed]
- Skvortsova, K.; Iovino, N.; Bogdanović, O. Functions and mechanisms of epigenetic inheritance in animals. Nat. Rev. Mol. Cell Biol. 2018, 19, 774–790. [Google Scholar] [CrossRef] [PubMed]
- Peixoto, P.; Cartron, P.F.; Serandour, A.A.; Hervouet, E. From 1957 to nowadays: A brief history of epigenetics. Int. J. Mol. Sci. 2020, 21, 7571. [Google Scholar] [CrossRef] [PubMed]
- Safi-Stibler, S.; Gabory, A. Epigenetics and the Developmental Origins of Health and Disease: Parental environment signalling to the epigenome, critical time windows and sculpting the adult phenotype. Semin. Cell Dev. Biol. 2020, 97, 172–180. [Google Scholar] [CrossRef] [PubMed]
- Susiarjo, M. Introduction to Epigenetic Mechanisms: The Probable Common Thread for Various Developmental Origins of Health and Diseases Effects. In The Epigenome and Developmental Origins of Health and Disease; Rosenfeld, C.S., Ed.; Elsevier Inc.: London, UK, 2015; pp. 49–62. ISBN 9780128013830. [Google Scholar] [CrossRef]
- Murín, R.; Abdalla, M.; Murínová, N.; Hatok, J.; Dobrota, D. The metabolism of 5-methylcytosine residues in DNA. Physiol. Res. 2018, 67, 383–389. [Google Scholar] [CrossRef]
- Kako, K.; Kim, J.D.; Fukamizu, A. Emerging impacts of biological methylation on genetic information. J. Biochem. 2019, 165, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Caiafa, P.; Zampieri, M. DNA methylation and chromatin structure: The puzzling CpG islands. J. Cell. Biochem. 2005, 94, 257–265. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Zhang, Y. TET-mediated active DNA demethylation: Mechanism, function and beyond. Nat. Rev. Genet. 2017, 18, 517–534. [Google Scholar] [CrossRef] [PubMed]
- Strahl, B.D.; Allis, C.D. The language of covalent histone modifications. Nature 2000, 403, 41–45. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Lan, F.; Matson, C.; Mulligan, P.; Whetstine, J.R.; Cole, P.A.; Casero, R.A.; Shi, Y. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 2004, 119, 941–953. [Google Scholar] [CrossRef] [PubMed]
- Marmorstein, R. Structure of SET domain proteins: A new twist on histone methylation. Trends Biochem. Sci. 2003, 28, 59–62. [Google Scholar] [CrossRef]
- McLaughlin, N.; Wang, F.; Saifudeen, Z.; El-Dahr, S.S. In situ histone landscape of nephrogenesis. Epigenetics 2014, 9, 222–235. [Google Scholar] [CrossRef] [PubMed]
- Peterson, C.L.; Laniel, M.A. Histones and histone modifications. Curr. Biol. 2004, 14, 546–551. [Google Scholar] [CrossRef] [PubMed]
- Weake, V.M.; Workman, J.L. Histone Ubiquitination: Triggering Gene Activity. Mol. Cell 2008, 29, 653–663. [Google Scholar] [CrossRef] [PubMed]
- Ryu, H.-Y.; Hochstrasser, M. Histone sumoylation and chromatin dynamics. Nucleic Acids Res. 2021, 49, 6043–6052. [Google Scholar] [CrossRef] [PubMed]
- Hottiger, M.O. ADP-ribosylation of histones by ARTD1: An additional module of the histone code? FEBS Lett. 2011, 585, 1595–1599. [Google Scholar] [CrossRef] [PubMed]
- Redon, C.; Pilch, D.; Rogakou, E.; Sedelnikova, O.; Newrock, K.; Bonner, W. Histone H2A variants H2AX and H2AZ. Curr. Opin. Genet. Dev. 2002, 12, 162–169. [Google Scholar] [CrossRef]
- Berger, S.L. Histone modifications in transcriptional regulation. Curr. Opin. Genet. Dev. 2002, 12, 142–148. [Google Scholar] [CrossRef]
- Agrawal, N.; Dasaradhi, P.V.N.; Mohmmed, A.; Malhotra, P.; Bhatnagar, R.K.; Mukherjee, S.K. RNA Interference: Biology, Mechanism, and Applications. Microbiol. Mol. Biol. Rev. 2003, 67, 657–685. [Google Scholar] [CrossRef] [PubMed]
- Dykes, I.M.; Emanueli, C. Transcriptional and Post-transcriptional Gene Regulation by Long Non-coding RNA. Genom. Proteom. Bioinform. 2017, 15, 177–186. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Webster, S.; de Wreede, R. Embryology at a Glance, 2nd ed.; John Wiley & Sons: Chichester, UK, 2016. [Google Scholar]
- Davidson, A.J.; Lewis, P.; Przepiorski, A.; Sander, V. Turning mesoderm into kidney. Semin. Cell Dev. Biol. 2019, 91, 86–93. [Google Scholar] [CrossRef] [PubMed]
- Pietilä, I.; Vainio, S.J. Kidney development: An overview. Nephron-Exp. Nephrol. 2014, 126, 40–44. [Google Scholar] [CrossRef] [PubMed]
- Oxburgh, L. Kidney Nephron Determination. Annu. Rev. Cell Dev. Biol. 2018, 34, 427–450. [Google Scholar] [CrossRef] [PubMed]
- Seldin, D.W.; Giebsich, G.H. Seldin and Giebsich’s The Kidney: Physiology and Pathophysiology, 5th ed.; Alpern, R.J., Moe, O.W., Caplan, M., Eds.; Elsevier Inc.: London, UK, 2013. [Google Scholar]
- Sequeira Lopez, M.L.S.; Gomez, R.A. Development of the renal arterioles. J. Am. Soc. Nephrol. 2011, 22, 2156–2165. [Google Scholar] [CrossRef] [PubMed]
- Constantini, F.; Kopan, R. Patterning a complex organ: Branching morphogenesis and nephron segmentation in kidney development. Dev. Cell 2010, 18, 698–712. [Google Scholar] [CrossRef] [PubMed]
- Koeppen, B.M.; Stanton, B.A. Renal Physiology, 6th ed.; Elsevier Inc.: Philadelphia, PA, USA, 2019. [Google Scholar]
- Ito, S.; Abe, K. Tubuloglomerular Feedback. Jpn. Heart J. 1996, 37, 153–163. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, T.; Sequeira-Lopez, M.L.S. Development of the renal vasculature. Semin. Cell Dev. Biol. 2019, 91, 132–146. [Google Scholar] [CrossRef] [PubMed]
- Wellik, D.M. Hox genes and kidney development. Pediatr. Nephrol. 2011, 26, 1559–1565. [Google Scholar] [CrossRef] [PubMed]
- Cirio, M.C.; Hui, Z.; Haldin, C.E.; Cosentino, C.C.; Stuckenholz, C.; Chen, X.; Hong, S.K.; Dawid, I.B.; Hukriede, N.A. Lhx1 is required for specification of the renal progenitor cell field. PLoS ONE 2011, 6, e18858. [Google Scholar] [CrossRef] [PubMed]
- Tena, J.J.; Neto, A.; de la Calle-Mustienes, E.; Bras-Pereira, C.; Casares, F.; Gómez-Skarmeta, J.L. Odd-skipped genes encode repressors that control kidney development. Dev. Biol. 2007, 301, 518–531. [Google Scholar] [CrossRef] [PubMed]
- Narlis, M.; Grote, D.; Gaitan, Y.; Boualia, S.K.; Bouchard, M. Pax2 and Pax8 regulate branching morphogenesis and nephron differentiation in the developing kidney. J. Am. Soc. Nephrol. 2007, 18, 1121–1129. [Google Scholar] [CrossRef] [PubMed]
- Gong, K.-Q.; Yallowitz, A.R.; Sun, H.; Dressler, G.R.; Wellik, D.M. A Hox-Eya-Pax Complex Regulates Early Kidney Developmental Gene Expression. Mol. Cell. Biol. 2007, 27, 7661–7668. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Wong, E.Y.M.; Cheng, C.; Li, J.; Sharkar, M.T.K.; Xu, C.Y.; Chen, B.; Sun, J.; Jing, D.; Xu, P.X. Eya1 interacts with Six2 and Myc to regulate expansion of the nephron progenitor pool during nephrogenesis. Dev. Cell 2014, 31, 434–447. [Google Scholar] [CrossRef] [PubMed]
- Kreidberg, J.A. WT1 and kidney progenitor cells. Organogenesis 2010, 6, 61–70. [Google Scholar] [CrossRef] [PubMed]
- El-Dahr, S.S.; Saifudeen, Z. Epigenetic Regulation of Renal Development. Semin. Cell Dev. Biol. 2019, 91, 111–118. [Google Scholar] [CrossRef] [PubMed]
- Nishinakamura, R.; Sakaguchi, M. BMP signaling and its modifiers in kidney development. Pediatr. Nephrol. 2014, 29, 681–686. [Google Scholar] [CrossRef] [PubMed]
- Costantini, F.; Shakya, R. GDNF/Ret signaling and the development of the kidney. BioEssays 2006, 28, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Marlier, A.; Schmidt-Ott, K.M.; Gallagher, A.R.; Barasch, J.; Karihaloo, A. Vegf as an epithelial cell morphogen modulates branching morphogenesis of embryonic kidney by directly acting on the ureteric bud. Mech. Dev. 2009, 126, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Trueb, B.; Amann, R.; Gerber, S.D. Role of FGFRL1 and other FGF signaling proteins in early kidney development. Cell. Mol. Life Sci. 2013, 70, 2505–2518. [Google Scholar] [CrossRef] [PubMed]
- Tufro, A.; Teichman, J.; Banu, N.; Villegas, G. Crosstalk between VEGF-A/VEGFR2 and GDNF/RET signaling pathways. Biochem. Biophys. Res. Commun. 2007, 358, 410–416. [Google Scholar] [CrossRef] [PubMed]
- Dudley, A.T.; Godin, R.E.; Robertson, E.J. Interaction between FGF and BMP signaling pathways regulates development of metanephric mesenchyme. Genes Dev. 1999, 13, 1601–1613. [Google Scholar] [CrossRef]
- Khoshdel Rad, N.; Aghdami, N.; Moghadasali, R. Cellular and Molecular Mechanisms of Kidney Development: From the Embryo to the Kidney Organoid. Front. Cell Dev. Biol. 2020, 8, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Church, R.H.; Ali, I.; Tate, M.; Lavin, D.; Krishnakumar, A.; Kok, H.M.; Hombrebueno, J.R.; Dunne, P.D.; Bingham, V.; Goldschmeding, R.; et al. Gremlin1 plays a key role in kidney development and renal fibrosis. Am. J. Physiol. Ren. Physiol. 2017, 312, F1141–F1157. [Google Scholar] [CrossRef] [PubMed]
- Huh, S.H.; Ha, L.; Jang, H.S. Nephron progenitor maintenance is controlled through fibroblast growth factors and sprouty1 interaction. J. Am. Soc. Nephrol. 2020, 31, 2559–2572. [Google Scholar] [CrossRef] [PubMed]
- Piper, M.; Georgas, K.; Yamada, T.; Little, M. Expression of the vertebrate Slit Gene family and their putative receptors, the Robo genes, in the developing murine kidney. Mech. Dev. 2000, 94, 213–217. [Google Scholar] [CrossRef]
- Chen, J.K.; Nagai, K.; Chen, J.; Plieth, D.; Hino, M.; Xu, J.; Sha, F.; Ikizler, T.A.; Quarles, C.C.; Threadgill, D.W.; et al. Phosphatidylinositol 3-kinase signaling determines kidney size. J. Clin. Investig. 2015, 125, 2429–2444. [Google Scholar] [CrossRef] [PubMed]
- Di Giovanni, V.; Walker, K.A.; Bushnell, D.; Schaefer, C.; Sims-Lucas, S.; Puri, P.; Bates, C.M. Fibroblast growth factor receptor-Frs2α signaling is critical for nephron progenitors. Dev. Biol. 2015, 400, 82–93. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Xu, J.; Xu, P.X. Eya-six are necessary for survival of nephrogenic cord progenitors and inducing nephric duct development before ureteric bud formation. Dev. Dyn. 2015, 244, 866–873. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhou, C.J.; Liu, Y. Wnt Signaling in Kidney Development and Disease. Prog. Mol. Biol. Transl. Sci. 2018, 153, 181–207. [Google Scholar] [CrossRef] [PubMed]
- Song, R.; Lopez, M.L.S.S.; Yosypiv, I.V. Foxd1 is an upstream regulator of the renin-angiotensin system during metanephric kidney development. Pediatr. Res. 2017, 82, 855–862. [Google Scholar] [CrossRef] [PubMed]
- Cui, S.; Schwartz, L.; Quaggin, S.E. Pod1 is required in stromal cells for glomerulogenesis. Dev. Dyn. 2003, 226, 512–522. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, M.; Fogarty, E.; Janga, M.; Surendran, K. Notch signaling in kidney development, maintenance, and disease. Biomolecules 2019, 9, 692. [Google Scholar] [CrossRef] [PubMed]
- Plisov, S.; Tsang, M.; Shi, G.; Boyle, S.; Yoshino, K.; Dunwoodie, S.L.; Dawid, I.B.; Shioda, T.; Perantoni, A.O.; De Caestecker, M.P. Cited1 is a bifunctional transcriptional cofactor that regulates early nephronic patterning. J. Am. Soc. Nephrol. 2005, 16, 1632–1644. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kobayashi, A.; Valerius, M.T.; Mugford, J.W.; Carroll, T.J.; Self, M.; Oliver, G.; McMahon, A.P. Six2 Defines and Regulates a Multipotent Self-Renewing Nephron Progenitor Population throughout Mammalian Kidney Development. Cell Stem Cell 2008, 3, 169–181. [Google Scholar] [CrossRef] [PubMed]
- Park, J.S.; Valerius, M.T.; McMahon, A.P. Wnt/β-catenin signaling regulates nephron induction during mouse kidney development. Development 2007, 134, 2533–2539. [Google Scholar] [CrossRef] [PubMed]
- Roy, A.; Al-Bataineh, M.M.; Pastor-Soler, N.M. Collecting duct intercalated cell function and regulation. Clin. J. Am. Soc. Nephrol. 2015, 10, 305–324. [Google Scholar] [CrossRef] [PubMed]
- Schell, C.; Wanner, N.; Huber, T.B. Glomerular development—Shaping the multi-cellular filtration unit. Semin. Cell Dev. Biol. 2014, 36, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Eckersley-Maslin, M.A.; Alda-Catalinas, C.; Reik, W. Dynamics of the epigenetic landscape during the maternal-to-zygotic transition. Nat. Rev. Mol. Cell Biol. 2018, 19, 436–450. [Google Scholar] [CrossRef] [PubMed]
- Nugent, B.M.; Bale, T.L. The omniscient placenta: Metabolic and epigenetic regulation of fetal programming. Front. Neuroendocrinol. 2015, 39, 28–37. [Google Scholar] [CrossRef] [PubMed]
- Thorrez, L.; Laudadio, I.; Van Deun, K.; Quintens, R.; Hendrickx, N.; Granvik, M.; Lemaire, K.; Schraenen, A.; Van Lommel, L.; Lehnert, S.; et al. Tissue-specific disallowance of housekeeping genes: The other face of cell differentiation. Genome Res. 2011, 21, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Hartwig, F.P.; De Mola, C.L.; Davies, N.M.; Victora, C.G.; Relton, C.L. Breastfeeding effects on DNA methylation in the offspring: A systematic literature review. PLoS ONE 2017, 12, 1–18. [Google Scholar] [CrossRef]
- Probst, A.V.; Dunleavy, E.; Almouzni, G. Epigenetic inheritance during the cell cycle. Nat. Rev. Mol. Cell Biol. 2009, 10, 192–206. [Google Scholar] [CrossRef] [PubMed]
- Hilliard, S.A.; El-Dahr, S.S. Epigenetics mechanisms in renal development. Pediatr. Nephrol. 2016, 31, 1055–1060. [Google Scholar] [CrossRef]
- Dressler, G.R.; Patel, S.R. Epigenetics in Kidney Development and Renal Disease. Transl. Res. 2015, 165, 166–176. [Google Scholar] [CrossRef] [PubMed]
- Wanner, N.; Vornweg, J.; Combes, A.; Wilson, S.; Plappert, J.; Rafflenbeul, G.; Puelles, V.G.; Rahman, R.U.; Liwinski, T.; Lindner, S.; et al. DNA methyltransferase 1 controls nephron progenitor cell renewal and differentiation. J. Am. Soc. Nephrol. 2019, 30, 63–78. [Google Scholar] [CrossRef] [PubMed]
- Gujar, H.; Weisenberger, D.J.; Liang, G. The roles of human DNA methyltransferases and their isoforms in shaping the epigenome. Genes 2019, 10, 172. [Google Scholar] [CrossRef] [PubMed]
- Cho, Y.W.; Hong, T.; Hong, S.H.; Guo, H.; Yu, H.; Kim, D.; Guszczynski, T.; Dressler, G.R.; Copeland, T.D.; Kalkum, M.; et al. PTIP associates with MLL3- and MLL4-containing histone H3 lysine 4 methyltransferase complex. J. Biol. Chem. 2007, 282, 20395–20406. [Google Scholar] [CrossRef]
- McLaughlin, N.; Yao, X.; Li, Y.; Saifudeen, Z.; El-Dahr, S.S. Histone signature of metanephric mesenchyme cell lines. Epigenetics 2013, 8, 970–978. [Google Scholar] [CrossRef] [PubMed]
- Kouznetsova, V.L.; Tchekanov, A.; Li, X.; Yan, X.; Tsigelny, I.F. Polycomb repressive 2 complex—Molecular mechanisms of function. Protein Sci. 2019, 28, 1387–1399. [Google Scholar] [CrossRef] [PubMed]
- Shinkai, Y.; Tachibana, M. H3K9 methyltransferase G9a and the related molecule GLP. Genes Dev. 2011, 25, 781–788. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, A.T.; Zhang, Y. The diverse functions of Dot1 and H3K79 methylation. Genes Dev. 2011, 25, 1345–1358. [Google Scholar] [CrossRef] [PubMed]
- Peters, A.H.F.M.; O’Carroll, D.; Scherthan, H.; Mechtler, K.; Sauer, S.; Schöfer, C.; Weipoltshammer, K.; Pagani, M.; Lachner, M.; Kohlmaier, A.; et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 2001, 107, 323–337. [Google Scholar] [CrossRef]
- Patel, S.R.; Kim, D.; Levitan, I.; Dressler, G.R. The BRCT-Domain Containing Protein PTIP Links PAX2 to a Histone H3, Lysine 4 Methyltransferase Complex. Dev. Cell 2007, 13, 580–592. [Google Scholar] [CrossRef]
- Zhang, X.; Liu, L.; Yuan, X.; Wei, Y.; Wei, X. JMJD3 in the regulation of human diseases. Protein Cell 2019, 10, 864–882. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.H.; Cho, Y.W.; Yu, L.R.; Yu, H.; Veenstra, T.D.; Ge, K. Identification of JmjC domain-containing UTX and JMJD3 as histone H3 lysine 27 demethylases. Proc. Natl. Acad. Sci. USA 2007, 104, 18439–18444. [Google Scholar] [CrossRef]
- Chen, S.; Bellew, C.; Yao, X.; Stefkova, J.; Dipp, S.; Saifudeen, Z.; Bachvarov, D.; El-Dahr, S.S. Histone deacetylase (HDAC) activity is critical for embryonic kidney gene expression, growth, and differentiation. J. Biol. Chem. 2011, 286, 32775–32789. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.; Liu, Z.; Vonk, A.; Zeng, Z.; Li, Z. Epigenetic regulation of kidney progenitor cells. Stem Cells Transl. Med. 2020, 9, 655–660. [Google Scholar] [CrossRef] [PubMed]
- Wen, L.; Tang, F. Boosting the power of single-cell analysis. Nat. Biotechnol. 2018, 36, 408–409. [Google Scholar] [CrossRef] [PubMed]
- Picelli, S. Single-cell RNA-sequencing: The future of genome biology is now. RNA Biol. 2017, 14, 637–650. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.W.; Chou, C.L.; Knepper, M.A. Deep sequencing in microdissected renal tubules identifies nephron segment-specific transcriptomes. J. Am. Soc. Nephrol. 2015, 26, 2669–2677. [Google Scholar] [CrossRef] [PubMed]
- Hildebrandt, F. Genetic kidney diseases. Lancet 2010, 375, 1287–1295. [Google Scholar] [CrossRef]
- Wu, H.; Chen, L.; Zhou, Q.; Zhang, X.; Berger, S.; Bi, J.; Lewis, D.E.; Xia, Y.; Zhang, W. Aqp2-expressing cells give rise to renal intercalated cells. J. Am. Soc. Nephrol. 2013, 24, 243–252. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Shrestha, R.; Qiu, C.; Kondo, A.; Huang, S.; Werth, M.; Li, M.; Barasch, J.; Susztak, K. Single-cell transcriptomics of the mouse kidney reveals potential cellular targets of kidney disease. Science 2018, 360, 758–763. [Google Scholar] [CrossRef] [PubMed]
- Naganuma, H.; Miike, K.; Ohmori, T.; Tanigawa, S.; Ichikawa, T.; Yamane, M.; Eto, M.; Niwa, H.; Kobayashi, A.; Nishinakamura, R. Molecular detection of maturation stages in the developing kidney. Dev. Biol. 2021, 470, 62–73. [Google Scholar] [CrossRef] [PubMed]
- Menon, R.; Otto, E.A.; Kokoruda, A.; Zhou, J.; Zhang, Z.; Yoon, E.; Chen, Y.C.; Troyanskaya, O.; Spence, J.R.; Kretzler, M.; et al. Single-cell analysis of progenitor cell dynamics and lineage specification in the human fetal kidney. Development 2018, 145. [Google Scholar] [CrossRef]
- Hochane, M.; van den Berg, P.R.; Fan, X.; Bérenger-Currias, N.; Adegeest, E.; Bialecka, M.; Nieveen, M.; Menschaart, M.; Chuva de Sousa Lopes, S.M.; Semrau, S. Single-cell transcriptomics reveals gene expression dynamics of human fetal kidney development. PLoS Biology 2019, e3000152. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Chen, Y.; Yong, J.; Cui, Y.; Wang, R.; Wen, L.; Qiao, J.; Tang, F. Dissecting the Global Dynamic Molecular Profiles of Human Fetal Kidney Development by Single-Cell RNA Sequencing. Cell Rep. 2018, 24, 3554–3567.e3. [Google Scholar] [CrossRef]
- Lindström, N.O.; Guo, J.; Kim, A.D.; Tran, T.; Guo, Q.; De Sena Brandine, G.; Ransick, A.; Parvez, R.K.; Thornton, M.E.; Basking, L.; et al. Conserved and divergent features of mesenchymal progenitor cell types within the cortical nephrogenic niche of the human and mouse kidney. J. Am. Soc. Nephrol. 2018, 29, 806–824. [Google Scholar] [CrossRef] [PubMed]
- Farlik, M.; Sheffield, N.C.; Nuzzo, A.; Datlinger, P.; Schönegger, A.; Klughammer, J.; Bock, C. Single-Cell DNA Methylome Sequencing and Bioinformatic Inference of Epigenomic Cell-State Dynamics. Cell Rep. 2015, 10, 1386–1397. [Google Scholar] [CrossRef]
- Smith, Z.D.; Gu, H.; Bock, C.; Gnirke, A.; Meissner, A. High-throughput bisulfite sequencing in mammalian genomes. Methods 2009, 48, 226–232. [Google Scholar] [CrossRef] [PubMed]
- Grosselin, K.; Durand, A.; Marsolier, J.; Poitou, A.; Marangoni, E.; Nemati, F.; Dahmani, A.; Lameiras, S.; Reyal, F.; Frenoy, O.; et al. High-throughput single-cell ChIP-seq identifies heterogeneity of chromatin states in breast cancer. Nat. Genet. 2019, 51, 1060–1066. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Ji, Z.; Fang, W.; Ji, H. Global prediction of chromatin accessibility using small-cell-number and single-cell RNA-seq. Nucleic Acids Res. 2019, 47, e121. [Google Scholar] [CrossRef] [PubMed]
- Cao, J.; Cusanovich, D.A.; Ramani, V.; Aghamirzaie, D.; Pliner, H.A.; Hill, A.J.; Daza, R.M.; McFaline-Figueroa, J.L.; Packer, J.S.; Christiansen, L.; et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 2018, 361, 1380–1385. [Google Scholar] [CrossRef] [PubMed]
- de Wit, E.; de Laat, W. A decade of 3C technologies: Insights into nuclear organization. Genes Dev. 2012, 26, 11–24. [Google Scholar] [CrossRef] [PubMed]
- Belton, J.M.; McCord, R.P.; Gibcus, J.H.; Naumova, N.; Zhan, Y.; Dekker, J. Hi-C: A comprehensive technique to capture the conformation of genomes. Methods 2012, 58, 268–276. [Google Scholar] [CrossRef] [PubMed]

| Gene | Expression | Role(s) | Epigenetic Regulators and Markers |
|---|---|---|---|
| Mesonephric and early metanephric development | |||
| Osr1 | LPM, IM | Regulate development of posterior nephric structures | H2A.Z, HDAC, Polycomb/Trithorax |
| Lhx1 | LPM, ND | Regulate development of the metanephric duct and continued renal development | H3K9me2 and H3K27me3, HDAC |
| Pax2 | IM, ND | Regulate branching of the ureteric bud and continued renal development | H3K4 methyltransferase complex, H3K9me2 and H3K27me3, HDAC, Polycomb/Trithorax (Ash21) |
| Pax8 | IM | Regulate branching of the ureteric bud and continued renal development | H3K9me2 and H3K27me3, HDAC |
| Metanephric development | |||
| Wt1 | IM, MM | Regulates continued differentiation of metanephric progenitor cells | HDAC |
| Foxd1 | MM, SC | Regulates nephron endowment and continued branching of the ureteric bud | HDAC |
| Hox11 | MM | Regulates development of the metanephros | Polycomb/Trithorax |
| Eya1 | MM | Regulates initiation of mesoderm differentiation and formation of the initial ureteric bud | HDAC |
| Six1 | MM | Regulates formation of the initial ureteric bud and subsequent branching of the ureteric bud | |
| Six2 | MM, CM | Regulates formation of metanephric caps and subsequent nephron formation | H3K9me2 and H3K27me3, Polycomb/Trithorax (Ezh2), G9a |
| Sall1 | MM | Regulates branching of the ureteric bud and formation of new nephrons | Polycomb/Trithorax |
| Wnt9b | UB | Regulates differentiation of metanephric caps and subsequent formation of new nephrons | HDAC |
| Gdnf | MM | Regulates the formation and branching of the ureteric bud | HDAC, Ret |
| Nephron patterning | |||
| Wnt4 | CM | Regulates metanephric cap behavior and subsequent nephron formation | HDAC |
| Fgf8 | MM, CM | Regulates continued nephron formation and proper renal development | |
| Bmp7 | UB, MM | Regulates continued branching of the ureteric bud and nephron endowment | |
| Notch2 | RV, SB | Regulates proper development of proximal tubules of nephrons | HDAC |
| Tcf21 (Pod1) | SC, PC | Regulates differentiation of podocytes | |
| Pdgfr | PC | Regulates development of the glomerulus | |
| VEGF | GP | Regulates development and survival of the glomerulus | |
| Jag1 | GP, ND | Regulates notch signaling pathways | H3K9me2 and H3K27me3, H3K4me3 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chan, K.; Li, X. Current Epigenetic Insights in Kidney Development. Genes 2021, 12, 1281. https://doi.org/10.3390/genes12081281
Chan K, Li X. Current Epigenetic Insights in Kidney Development. Genes. 2021; 12(8):1281. https://doi.org/10.3390/genes12081281
Chicago/Turabian StyleChan, Katrina, and Xiaogang Li. 2021. "Current Epigenetic Insights in Kidney Development" Genes 12, no. 8: 1281. https://doi.org/10.3390/genes12081281
APA StyleChan, K., & Li, X. (2021). Current Epigenetic Insights in Kidney Development. Genes, 12(8), 1281. https://doi.org/10.3390/genes12081281






