The Role of Extracellular Vesicles as Shuttles of RNA and Their Clinical Significance as Biomarkers in Hepatocellular Carcinoma
Abstract
1. Extracellular Vesicles
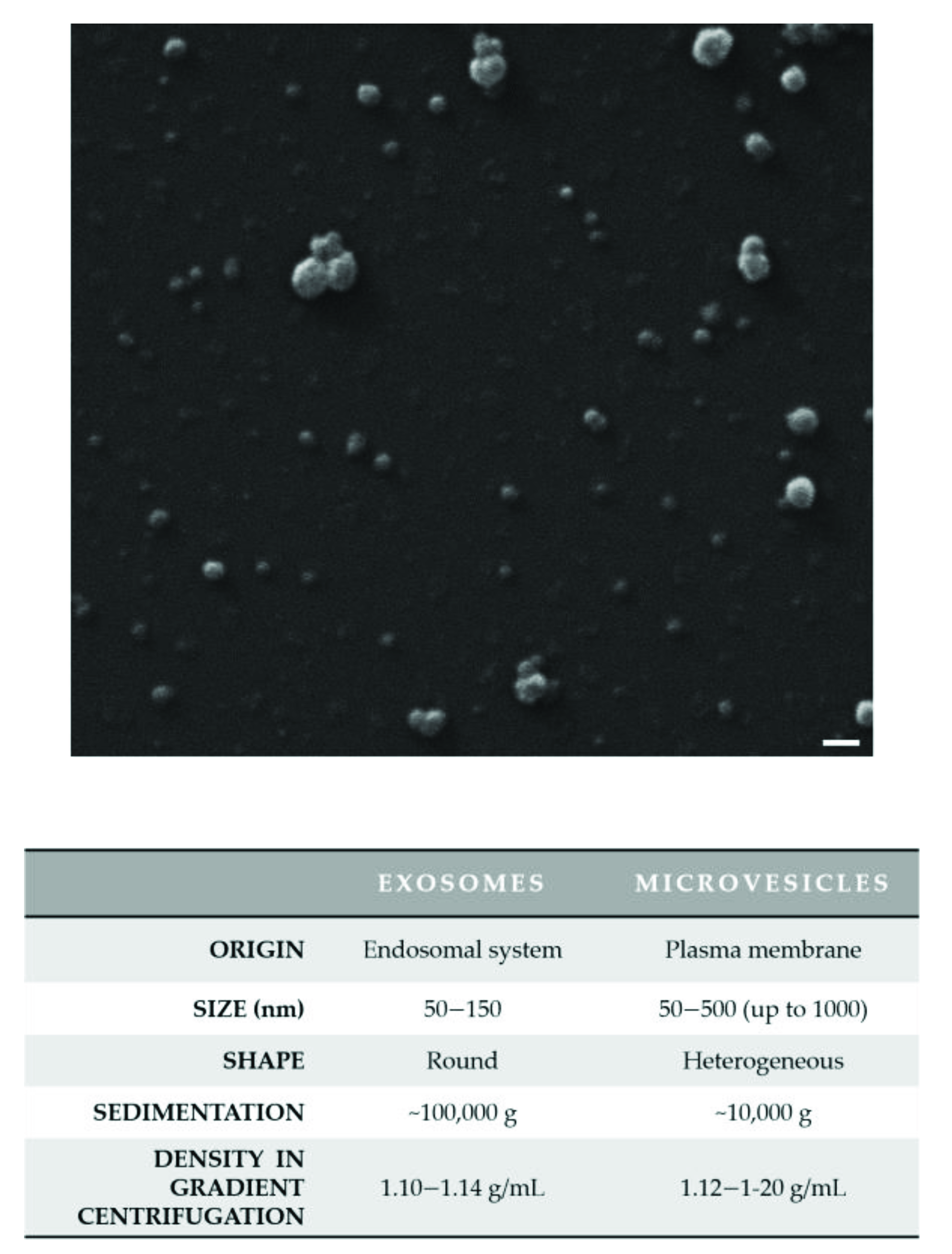
1.1. Definition and Classification
1.2. Biological and Functional Features of EVs
2. Extracellular Vesicles as Carriers of RNA Molecules
2.1. Classes of RNA Molecules Retrieved in EVs and Their Biological Function
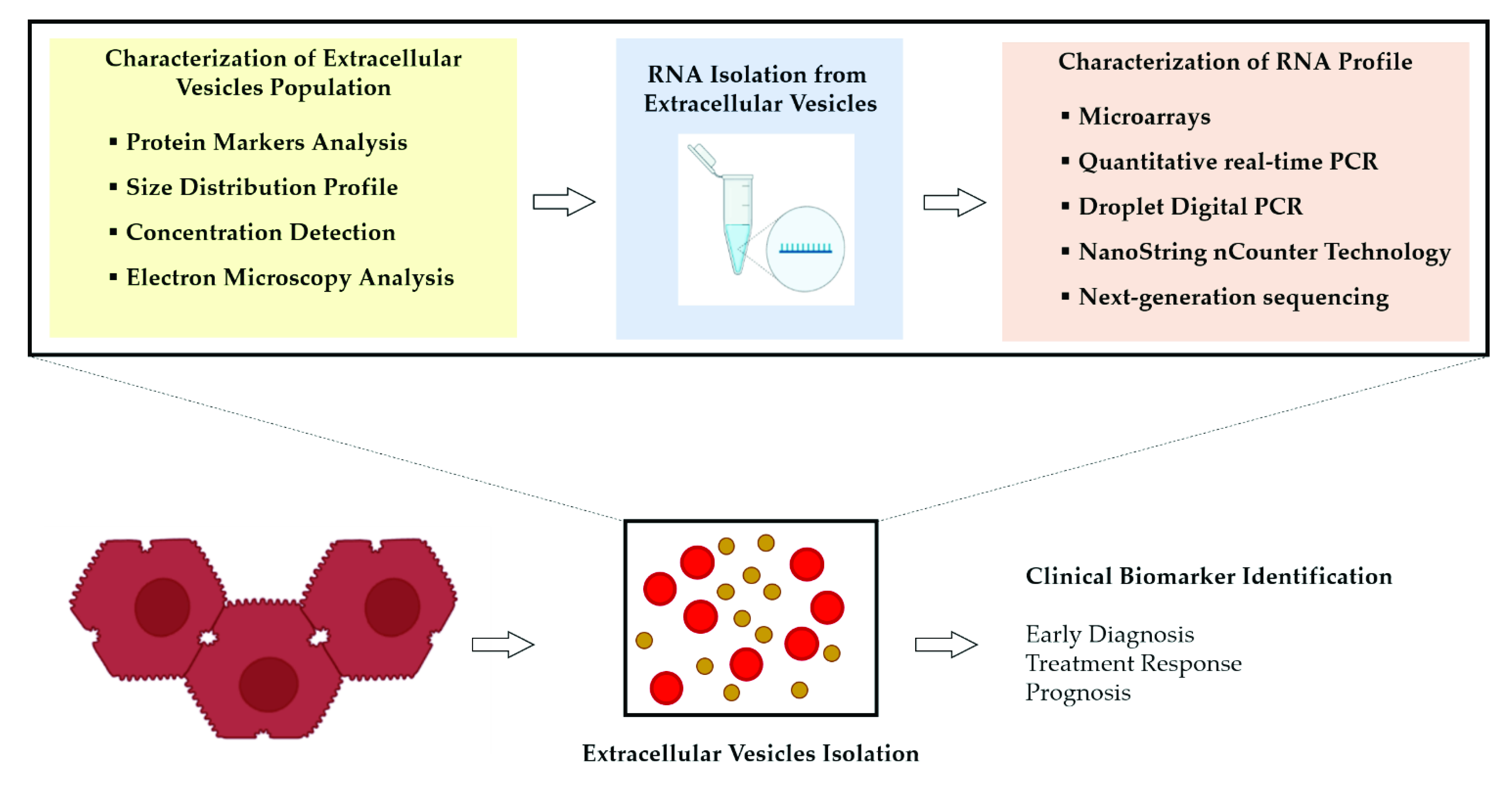
2.2. Methodological Approaches to Study RNA Molecules Carried by EVs
3. Extracellular Vesicle-Derived RNAs Correlated with Hepatocellular Carcinoma
3.1. Role of EV-Derived RNA in the HCC Microenvironment Influencing Tumor Progression, Metastasis, and Angiogenesis
3.2. Role of EV-Derived RNA as HCC Suppressors
4. Extracellular Vesicle-Derived RNAs as Potential Biomarkers in HCC
4.1. EV-Derived miRNAs as Prognostic and Diagnostic Markers for HCC
4.2. Other EV-Derived RNA Molecules as Biomarkers for HCC
| Clinical Significance | EV-Derived RNA | Expression Level | Patients | Source | EV Isolation Method | Clinical Relevance | Ref. |
|---|---|---|---|---|---|---|---|
| Detection and Diagnosis | miR-21 | Upregulated | 30 HCC 30 CHB 30 HV | Serum | Total Exosome Isolation Kit (Invitrogen, Carlsbad, CA, USA). | Discrimination between HCC and CHB or LC | [78] |
| miR-93 | Upregulated | 85 HCC 23 HV | Serum | Total Exosome Isolation Kit (Thermo Fisher Scientific, Waltham, MA, USA) | Biomarker for both diagnosis and prognosis in HCC. | [112] | |
| miR-224 | Upregulated | 89 HCC 50 HV | Serum | Total Exosome Isolation Kit (Thermo Fisher Scientific) | Biomarker of diagnosis and prognosis of HCC patient | [117] | |
| miR-718 | Downregulated | 59 HCC | Serum | Differential Centrifugation with a Final Ultracentrifugation Step | Predicting biomarker for recurrence after LT | [121] | |
| miR-18a miR-221 miR-222 miR-224 miR-101 miR-106b miR-122 miR-195 | Upregulated Downregulated | 20 HCC 20 LC 20 CHB | Serum | ExoQuick Exosome Precipitation Solution (System Biosciences, Mountain View, CA, USA) | Discrimination between HCC and CHB or LC | [115] | |
| miR-10b-5p miR-18a-5p miR-215-5p miR-940 | Upregulated | 90 HCC 60 CLD 28 HV | Serum | Differential Centrifugation with a Final Ultracentrifugation Step | miR-10b-5p as a diagnostic biomarker for early stage HCC | [118] | |
| miRNA-26a miRNA-29c miRNA-21 | Downregulated | 72 HCC 72 LC 72 HBV | Serum | ExoQuick Exosome Precipitation Solution (System Biosciences, Mountain View, CA, USA) | Diagnostic biomarkers for patients with HCC | [130] | |
| miR-122 miR-148a miR-1246 | Upregulated | 5 HCC 5 LC | Serum | 8% Polyethylene glycol (PEG) 6000 (Sigma-Aldrich, St Louis, MO, USA) | Diagnostic biomarker for patients with HCC | [131] | |
| lncRNA-HEIH | Upregulated | 35 CHC 22 HCV 10 HCC | Serum | Total Exosome Isolation reagent (GS0301; Guangzhou 141 Geneseed Biotech Co., Guangzhou, China) with a final centrifugation passage | Biomarker in the HCV-related hepatocellular carcinoma | [128] | |
| LINC00853 | Upregulated | 90 HCC 28 CH 35 LC 29 HV | Serum | ExoQuick Exosome Precipitation Solution (System Biosciences, Mountain View, CA, USA) | Diagnostic biomarker discriminating both all-stage HCC and early HCC | [129] | |
| LINC00161 | Upregulated | 56 HCC 56 HV 15 HCC 15 HV | SerumUrine | Total Exosome Isolation Kit (Invitrogen, USA) | Diagnostic biomarker for patients with HCC | [126] | |
| ENSG00000258332.1 LINC00635 | Upregulated | 60 HCC 85 LC 96 CHB 60 HV | Serum | Total Exosome Isolation Kit (Thermo Fisher Scientific) | Biomarker of diagnosis and prognosis of HCC patients | [127] | |
| Jpx | Upregulated | 74 HCC 26 LC 34 CHB 72 HV | Serum | ExoQuick Exosome Precipitation Solution (System Biosciences, Mountain View, CA, USA) | Biomarkers for diagnosis of female patients with HCC | [125] | |
| 8 EV-ncRNA (circRNA and lncRNA) | Upregulated | 71 HCC (early stage, n = 45; advanced stage, n = 26) 94 HV 18 benign HCC 11 CHB 8 LC | Serum | exoRNeasy Serum/Plasma kit (Qiagen, Hilden, Germany) | Biomarkers for hepatocellular carcinoma (HCC) diagnosis | [124] | |
| Detection and Prognosis | miR-125b | Upregulated | 158 HCC 30 CHB 30 LC | Serum | ExoQuick Exosome Precipitation Solution (System Biosciences, Palo Alto, CA, USA) | Predicting biomarker for recurrence and survival | [120] |
| miR-638 | Downregulated | 126 HCC 21 HV | Serum | Total exosome isolation kit (Invitrogen, Carlsbad, CA, USA). | Poor prognosis marker for patients with HCC | [123] | |
| miR-215-5p | Upregulated | 90 HCC 60 CLD 28 HV | Serum | Differential Centrifugation with a Final Ultracentrifugation Step | miR-215-5p: prognostic biomarker for HCC | [118] | |
| miR-744 | Downregulated | 68 HCC 52 normal liver tissue samples | Serum | Differential Centrifugation with a Final Ultracentrifugation Step | Inhibition of Proliferation and chemoresistance | [132] | |
| miR-224 | Upregulated | 89 HCC 50 HV | Serum | Total Exosome Isolation Kit (Thermo Fisher Scientific) | Biomarker of diagnosis and prognosis of HCC patient | [117] | |
| miR-21 miR-10b | Upregulated | 124 HCC N.A. HV | Serum | ExoQuick-TC exosome precipitation solution (System Biosciences, CA, USA) | Prognostic molecular markers and therapeutic targets for HCC. | [114] | |
| miR-9-3p | Downregulated | N.A. HCC N.A. HV | Serum | Differential Centrifugation with a Final Ultracentrifugation Step | Potential therapeutic target for HCC. | [116] | |
| circPTGR1 | Upregulated | 82 HCC 47 HV | Serum | ExoQuick-TC exosome precipitation solution (System Biosciences, CA, USA) | Prognostic biomarker and therapeutic target in HCC | [95] |
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Maas, S.L.N.; Breakefield, X.O.; Weaver, A.M. Extracellular Vesicles: Unique Intercellular Delivery Vehicles. Trends Cell Biol. 2017, 27, 172–188. [Google Scholar] [CrossRef] [PubMed]
- Raposo, G.; Stoorvogel, W. Extracellular Vesicles: Exosomes, Microvesicles, and Friends. J. Cell Biol. 2013, 200, 373–383. [Google Scholar] [CrossRef] [PubMed]
- Colombo, M.; Raposo, G.; Théry, C. Biogenesis, Secretion, and Intercellular Interactions of Exosomes and Other Extracellular Vesicles. Annu. Rev. Cell Dev. Biol. 2014, 30, 255–289. [Google Scholar] [CrossRef] [PubMed]
- Lo Cicero, A.; Stahl, P.D.; Raposo, G. Extracellular Vesicles Shuffling Intercellular Messages: For Good or for Bad. Curr. Opin. Cell Biol. 2015, 35, 69–77. [Google Scholar] [CrossRef]
- Greening, D.W.; Simpson, R.J. Understanding Extracellular Vesicle Diversity–Current Status. Expert. Rev. Proteom. 2018, 15, 887–910. [Google Scholar] [CrossRef]
- Kalra, H.; Gangoda, L.; Fonseka, P.; Chitti, S.V.; Liem, M.; Keerthikumar, S.; Samuel, M.; Boukouris, S.; Al Saffar, H.; Collins, C.; et al. Extracellular Vesicles Containing Oncogenic Mutant β-Catenin Activate Wnt Signalling Pathway in the Recipient Cells. J. Extracell. Vesicles 2019, 8, 1690217. [Google Scholar] [CrossRef]
- Sagini, K.; Costanzi, E.; Emiliani, C.; Buratta, S.; Urbanelli, L. Extracellular Vesicles as Conveyors of Membrane-Derived Bioactive Lipids in Immune System. Int. J. Mol. Sci. 2018, 19, 1227. [Google Scholar] [CrossRef]
- O’Brien, K.; Breyne, K.; Ughetto, S.; Laurent, L.C.; Breakefield, X.O. RNA Delivery by Extracellular Vesicles in Mammalian Cells and Its Applications. Nat. Rev. Mol. Cell Biol. 2020, 21, 585–606. [Google Scholar] [CrossRef]
- György, B.; Szabó, T.G.; Pásztói, M.; Pál, Z.; Misják, P.; Aradi, B.; László, V.; Pállinger, É.; Pap, E.; Kittel, Á.; et al. Membrane Vesicles, Current State-of-the-Art: Emerging Role of Extracellular Vesicles. Cell. Mol. Life Sci. 2011, 68, 2667–2688. [Google Scholar] [CrossRef]
- Van Niel, G.; D’Angelo, G.; Raposo, G. Shedding Light on the Cell Biology of Extracellular Vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef]
- Noble, J.M.; Roberts, L.D.M.; Vidavsky, N.; Chiou, A.E.; Fischbach, C.; Paszek, M.J.; Estroff, L.A.; Kourkoutis, L.F. Direct Comparison of Optical and Electron Microscopy Methods for Structural Characterization of Extracellular Vesicles. J. Struct. Biol. 2020. [Google Scholar] [CrossRef]
- Kowal, J.; Arras, G.; Colombo, M.; Jouve, M.; Morath, J.P.; Primdal-Bengtson, B.; Dingli, F.; Loew, D.; Tkach, M.; Théry, C. Proteomic Comparison Defines Novel Markers to Characterize Heterogeneous Populations of Extracellular Vesicle Subtypes. Proc. Natl. Acad. Sci. USA 2016. [Google Scholar] [CrossRef] [PubMed]
- Théry, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K.; et al. Minimal Information for Studies of Extracellular Vesicles 2018 (MISEV2018): A Position Statement of the International Society for Extracellular Vesicles and Update of the MISEV2014 Guidelines. J. Extracell. Vesicles 2018. [Google Scholar] [CrossRef] [PubMed]
- Witwer, K.W.; Théry, C. Extracellular Vesicles or Exosomes? On Primacy, Precision, and Popularity Influencing a Choice of Nomenclature. J. Extracell. Vesicles 2019, 8, 1648167. [Google Scholar] [CrossRef]
- Okoye, I.S.; Coomes, S.M.; Pelly, V.S.; Czieso, S.; Papayannopoulos, V.; Tolmachova, T.; Seabra, M.C.; Wilson, M.S. MicroRNA-Containing T-Regulatory-Cell-Derived Exosomes Suppress Pathogenic T Helper 1 Cells. Immunity 2014, 41, 89–103. [Google Scholar] [CrossRef]
- Ying, W.; Riopel, M.; Bandyopadhyay, G.; Dong, Y.; Birmingham, A.; Seo, J.B.; Ofrecio, J.M.; Wollam, J.; Hernandez-Carretero, A.; Fu, W.; et al. Adipose Tissue Macrophage-Derived Exosomal MiRNAs Can Modulate In vivo and In Vitro Insulin Sensitivity. Cell 2017, 171, 372–384. [Google Scholar] [CrossRef] [PubMed]
- Webber, J.P.; Spary, L.K.; Sanders, A.J.; Chowdhury, R.; Jiang, W.G.; Steadman, R.; Wymant, J.; Jones, A.T.; Kynaston, H.; Mason, M.D.; et al. Differentiation of Tumour-Promoting Stromal Myofibroblasts by Cancer Exosomes. Oncogene 2015, 34, 319–333. [Google Scholar] [CrossRef]
- Al-Nedawi, K.; Meehan, B.; Kerbel, R.S.; Allison, A.C.; Rak, A. Endothelial Expression of Autocrine VEGF upon the Uptake of Tumor-Derived Microvesicles Containing Oncogenic EGFR. Proc. Natl. Acad. Sci. USA 2009, 106, 3794–3799. [Google Scholar] [CrossRef]
- Luga, V.; Zhang, L.; Viloria-Petit, A.M.; Ogunjimi, A.A.; Inanlou, M.R.; Chiu, E.; Buchanan, M.; Hosein, A.N.; Basik, M.; Wrana, J.L. Exosomes Mediate Stromal Mobilization of Autocrine Wnt-PCP Signaling in Breast Cancer Cell Migration. Cell 2012, 151, 1542–1556. [Google Scholar] [CrossRef]
- Ma, C.; Jiang, F.; Ma, Y.; Wang, J.; Li, H.; Zhang, J. Isolation and Detection Technologies of Extracellular Vesicles and Application on Cancer Diagnostic. Dose Response 2019, 17, 1559325819891004. [Google Scholar] [CrossRef]
- Keller, S.; Ridinger, J.; Rupp, A.K.; Janssen, J.W.G.; Altevogt, P. Body Fluid Derived Exosomes as a Novel Template for Clinical Diagnostics. J. Transl. Med. 2011. [Google Scholar] [CrossRef] [PubMed]
- Urbanelli, L.; Buratta, S.; Sagini, K.; Ferrara, G.; Lanni, M.; Emiliani, C. Exosome-Based Strategies for Diagnosis and Therapy. Recent Patents CNS Drug Discov. 2015. [Google Scholar] [CrossRef] [PubMed]
- Merchant, M.L.; Rood, I.M.; Deegens, J.K.J.; Klein, J.B. Isolation and Characterization of Urinary Extracellular Vesicles: Implications for Biomarker Discovery. Nat. Rev. Nephrol. 2017, 13, 731–749. [Google Scholar] [CrossRef] [PubMed]
- Zöller, M. Tetraspanins: Push and Pull in Suppressing and Promoting Metastasis. Nat. Rev. Cancer. 2009, 9, 40–55. [Google Scholar] [CrossRef] [PubMed]
- Quinn, J.F.; Patel, T.; Wong, D.; Das, S.; Freedman, J.E.; Laurent, L.C.; Carter, B.S.; Hochberg, F.; Keuren-Jensen, K.V.; Huentelman, M.; et al. Extracellular RNAs: Development as Biomarkers of Human Disease. J. Extracell. Vesicles 2015. [Google Scholar] [CrossRef]
- Ragusa, M.; Barbagallo, C.; Cirnigliaro, M.; Battaglia, R.; Brex, D.; Caponnetto, A.; Barbagallo, D.; Di Pietro, C.D.; Purrello, M. Asymmetric RNA Distribution among Cells and Their Secreted Exosomes: Biomedical Meaning and Considerations on Diagnostic Applications. Front. Mol. Biosci. 2017, 4, 1–14. [Google Scholar] [CrossRef]
- Ferreri, C.; Sansone, A.; Buratta, S.; Urbanelli, L.; Costanzi, E.; Emiliani, C.; Chatgilialoglu, C. The n-10 Fatty Acids Family in the Lipidome of Human Prostatic Adenocarcinoma Cell Membranes and Extracellular Vesicles. Cancers 2020, 12, 900. [Google Scholar] [CrossRef]
- Fais, S.; Logozzi, M. A New Method to Measure and Characterize Microvesicles in the Human Body Fluids. WO Patent 2009092386, 30 July 2009. [Google Scholar]
- Moon, P.G.; Lee, J.E.; Cho, Y.E.; Lee, S.J.; Chae, Y.S.; Jung, J.H.; Kim, I.S.; Park, H.Y.; Baek, M.C. Fibronectin on Circulating Extracellular Vesicles as a Liquid Biopsy to Detect Breast Cancer. Oncotarget 2016, 7, 40189–40199. [Google Scholar] [CrossRef]
- Tao, L.; Zhou, J.; Yuan, C.; Zhang, L.; Li, D.; Si, D.; Xiu, D.; Zhong, L. Metabolomics Identifies Serum and Exosomes Metabolite Markers of Pancreatic Cancer. Metabolomics 2019, 15, 86. [Google Scholar] [CrossRef]
- Valadi, H.; Ekström, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-Mediated Transfer of MRNAs and MicroRNAs Is a Novel Mechanism of Genetic Exchange between Cells. Nat. Cell Biol. 2007. [Google Scholar] [CrossRef]
- Di Liegro, C.M.; Schiera, G.; Di Liegro, I. Extracellular Vesicle-Associated RNA as a Carrier of Epigenetic Information. Genes 2017, 8, 240. [Google Scholar] [CrossRef]
- Iraci, N.; Leonardi, T.; Gessler, F.; Vega, B.; Pluchino, S. Focus on Extracellular Vesicles: Physiological Role and Signalling Properties of Extracellular Membrane Vesicles. Int. J. Mol. Sci. 2016, 17, 171. [Google Scholar] [CrossRef]
- Fatima, F.; Nawaz, M. Vesiculated Long Non-Coding RNAs: Offshore Packages Deciphering Trans-Regulation between Cells, Cancer Progression and Resistance to Therapies. Noncoding RNA 2017, 3, 10. [Google Scholar] [CrossRef]
- Li, Y.; Zheng, Q.; Bao, C.; Li, S.; Guo, W.; Zhao, J.; Chen, D.; Gu, J.; He, X.; Huang, S. Circular RNA Is Enriched and Stable in Exosomes: A Promising Biomarker for Cancer Diagnosis. Cell Res. 2015, 25, 981–984. [Google Scholar] [CrossRef]
- Yu, F.-J.; Zheng, J.-J.; Dong, P.-H.; Fan, X.-M. Long Non-Coding RNAs and Hepatocellular Carcinoma. Mol. Clin. Oncol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Gezer, U.; Özgür, E.; Cetinkaya, M.; Isin, M.; Dalay, N. Long Non-Coding RNAs with Low Expression Levels in Cells Are Enriched in Secreted Exosomes. Cell Biol. Int. 2014, 38, 1076–1079. [Google Scholar] [CrossRef]
- Conti, I.; Varano, G.; Simioni, C.; Laface, I.; Milani, D.; Rimondi, E.; Neri, L.M. MiRNAs as Influencers of Cell-Cell Communication in Tumor Microenvironment. Cells 2020, 9, 220. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.; Chen, C.Z. Micromanagers of Gene Expression: The Potentially Widespread Influence of Metazoan MicroRNAs. Nat. Rev. Genet 2004, 5, 396–400. [Google Scholar] [CrossRef] [PubMed]
- Thomson, J.M.; Newman, M.; Parker, J.S.; Morin-Kensicki, E.M.; Wright, T.; Hammond, S.M. Extensive Post-Transcriptional Regulation of MicroRNAs and Its Implications for Cancer. Genes Dev. 2006. [Google Scholar] [CrossRef]
- Fabbri, M.; Garzon, R.; Andreeff, M.; Kantarjian, H.M.; Garcia-Manero, G.; Calin, G.A. MicroRNAs and Noncoding RNAs in Hematological Malignancies: Molecular, Clinical and Therapeutic Implications. Leukemia 2008, 22, 1095–1105. [Google Scholar] [CrossRef]
- Calin, G.A.; Croce, C.M. Genomics of Chronic Lymphocytic Leukemia MicroRNAs as New Players with Clinical Significance. Semin. Oncol. 2006, 33, 167–173. [Google Scholar] [CrossRef] [PubMed]
- Ebert, M.S.; Sharp, P.A. MicroRNA Sponges: Progress and Possibilities. RNA 2010, 16, 2043–2050. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; Wang, B.; Feng, X.; Xu, Y.; Lu, K.; Sun, M. CircRNAs and Exosomes: A Mysterious Frontier for Human Cancer. Mol. Ther. Nucleic Acids 2020, 19, 384–392. [Google Scholar] [CrossRef] [PubMed]
- Bao, C.; Lyu, D.; Huang, S. Circular RNA Expands Its Territory. Mol. Cell. Oncol. 2015, 3, E1084443. [Google Scholar] [CrossRef]
- Dou, Y.; Cha, D.J.; Franklin, J.L.; Higginbotham, J.N.; Jeppesen, D.K.; Weaver, A.M.; Prasad, N.; Levy, S.; Coffey, R.J.; Patton, J.G.; et al. Circular RNAs Are down-Regulated in KRAS Mutant Colon Cancer Cells and Can Be Transferred to Exosomes. Sci. Rep. 2016, 6, 37982. [Google Scholar] [CrossRef]
- Santangelo, L.; Giurato, G.; Cicchini, C.; Montaldo, C.; Mancone, C.; Tarallo, R.; Battistelli, C.; Alonzi, T.; Weisz, A.; Tripodi, M. The RNA-Binding Protein SYNCRIP Is a Component of the Hepatocyte Exosomal Machinery Controlling MicroRNA Sorting. Cell Rep. 2016. [Google Scholar] [CrossRef]
- Guduric-Fuchs, J.; O’Connor, A.; Camp, B.; O’Neill, C.L.; Medina, R.J.; Simpson, D.A. Selective Extracellular Vesicle-Mediated Export of an Overlapping Set of MicroRNAs from Multiple Cell Types. BMC Genom. 2012. [Google Scholar] [CrossRef]
- Corbeil, D.; Santos, M.F.; Karbanová, J.; Kurth, T.; Rappa, G.; Lorico, A. Uptake and Fate of Extracellular Membrane Vesicles: Nucleoplasmic Reticulum-Associated Late Endosomes as a New Gate to Intercellular Communication. Cells 2020, 9, 1931. [Google Scholar] [CrossRef]
- Esteller, M. Non-Coding RNAs in Human Disease. Nat. Rev. Genet 2011, 12, 861–874. [Google Scholar] [CrossRef]
- Flynt, A.S.; Lai, E.C. Biological Principles of MicroRNA-Mediated Regulation: Shared Themes amid Diversity. Nat. Rev. Genet 2008, 9, 831–842. [Google Scholar] [CrossRef]
- Chang, W.; Zhang, L.; Xian, Y.; Yu, Z. MicroRNA-33a Promotes Cell Proliferation and Inhibits Apoptosis by Targeting PPARα in Human Hepatocellular Carcinoma. Exp. Ther. Med. 2017. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, U.; Lai, X.; Winter, F.; Wolkenhauer, O.; Vera, J.; Gupta, S.K. Cooperative Gene Regulation by MicroRNA Pairs and Their Identification Using a Computational Workflow. Nucleic Acids Res. 2014. [Google Scholar] [CrossRef]
- Patel, T. Extracellular Vesicle Noncoding RNA: New Players in the Diagnosis and Pathogenesis of Cholangiocarcinoma. Hepatology 2014, 60, 782–784. [Google Scholar] [CrossRef]
- Coumans, F.A.W.; Brisson, A.R.; Buzas, E.I.; Dignat-George, F.; Drees, E.E.E.; El-Andaloussi, S.; Emanueli, C.; Gasecka, A.; Hendrix, A.; Hill, A.F.; et al. Methodological Guidelines to Study Extracellular Vesicles. Circ. Res. 2017, 120, 1632–1648. [Google Scholar] [CrossRef]
- Sun, N.; Lee, Y.T.; Zhang, R.Y.; Kao, R.; Teng, P.C.; Yang, Y.; Yang, P.; Wang, J.J.; Smalley, M.; Chen, P.J.; et al. Purification of HCC-Specific Extracellular Vesicles on Nanosubstrates for Early HCC Detection by Digital Scoring. Nat. Commun. 2020, 11, 4489. [Google Scholar] [CrossRef] [PubMed]
- Konoshenko, M.Y.; Lekchnov, E.A.; Bryzgunova, O.E.; Kiseleva, E.; Pyshnaya, I.A.; Laktionov, P.P. Isolation of Extracellular Vesicles from Biological Fluids via the Aggregation-Precipitation Approach for Downstream MiRNAs Detection. Diagnostics 2021, 11, 384. [Google Scholar] [CrossRef] [PubMed]
- Segundo-Val, I.S.; Sanz-Lozano, C.S. Introduction to the Gene Expression Analysis. Methods Mol. Biol. 2016, 1434, 29–43. [Google Scholar] [CrossRef]
- Narrandes, S.; Xu, W. Gene Expression Detection Assay for Cancer Clinical Use. J. Cancer 2018, 9, 2249–2265. [Google Scholar] [CrossRef]
- Mohankumar, S.; Patel, T. Extracellular Vesicle Long Noncoding RNA as Potential Biomarkers of Liver Cancer. Brief Funct. Genom. 2016, 15, 249–256. [Google Scholar] [CrossRef]
- Zhao, B.; Ding, S.; Li, W.; Jin, Y. Hybridization Kinetics Analysis of an Oligonucleotide Microarray for MicroRNA Detection. Acta Biochim. Biophys. Sin. 2011, 43, 551–555. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Moya, J.M.; Vilella, F.; Simón, C. MicroRNA: Key Gene Expression Regulators. Fertil. Steril. 2014, 101, 1516–1523. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Jing, F.; Li, G.; Wu, Z.; Cheng, Z.; Zhang, J.; Zhang, H.; Jia, C.; Jin, Q.; Mao, H.; et al. Absolute Quantification of Lung Cancer Related MicroRNA by Droplet Digital PCR. Biosen. Bioelectron. 2015, 74, 836–842. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yan, I.K.; Kim, C.; Kim, J.; Patel, T. Analysis of Extracellular RNA by Digital PCR. Front. Oncol. 2014. [Google Scholar] [CrossRef]
- Shukla, N.; Yan, I.K.; Patel, T. Multiplexed Detection and Quantitation of Extracellular Vesicle RNA Expression Using NanoString. Methods Mol. Biol. 2018, 1740, 177–185. [Google Scholar] [CrossRef]
- Geiss, G.K.; Bumgarner, R.E.; Birditt, B.; Dahl, T.; Dowidar, N.; Dunaway, D.L.; Fell, H.P.; Ferree, S.; George, R.D.; Grogan, T.; et al. Direct Multiplexed Measurement of Gene Expression with Color-Coded Probe Pairs. Nat. Biotechnol. 2008, 26, 317–325. [Google Scholar] [CrossRef] [PubMed]
- Bracht, J.W.; Gimenez-Capitan, A.; Huang, C.Y.; Potie, N.; Pedraz-Valdunciel, C.; Warren, S.; Rosell, R.; Molina-Vila, M.A. Analysis of Extracellular Vesicle MRNA Derived from Plasma Using the NCounter Platform. Sci. Rep. 2021, 11, 3712. [Google Scholar] [CrossRef]
- Lopez, J.P.; Cruceanu, C.; Fiori, L.M.; Laboissiere, S.; Guillet, I.; Fontaine, J.; Ragoussis, J.; Benes, V.; Turecki, G.; Ernst, C. Biomarker Discovery: Quantification of MicroRNAs and Other Small Non-Coding RNAs Using Next Generation Sequencing. BMC Med. Genom. 2015, 8, 35. [Google Scholar] [CrossRef]
- Craig, A.J.; von Felden, J.; Garcia-Lezana, T.; Sarcognato, S.; Villanueva, A. Tumour Evolution in Hepatocellular Carcinoma. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 139–152. [Google Scholar] [CrossRef]
- Cervello, M.; Emma, M.R.; Augello, G.; Cusimano, A.; Giannitrapani, L.; Soresi, M.; Akula, S.M.; Abrams, S.L.; Steelman, L.S.; Gulino, A.; et al. New Landscapes and Horizons in Hepatocellular Carcinoma Therapy. Aging 2020, 12, 3053–3094. [Google Scholar] [CrossRef]
- D’Agnano, I.; Berardi, A.C. Extracellular Vesicles, A Possible Theranostic Platform Strategy for Hepatocellular Carcinoma-An Overview. Cancers 2020, 12, 261. [Google Scholar] [CrossRef]
- Hirsova, P.; Ibrahim, S.H.; Verma, V.K.; Morton, L.A.; Shah, V.H.; LaRusso, N.F.; Gores, G.J.; Malhi, H. Extracellular Vesicles in Liver Pathobiology: Small Particles with Big Impact. Hepatology 2016, 64, 2219–2233. [Google Scholar] [CrossRef] [PubMed]
- He, M.; Qin, H.; Poon, T.C.W.; Sze, S.C.; Ding, X.; Co, N.N.; Ngai, S.M.; Chan, T.F.; Wong, N. Hepatocellular Carcinoma-Derived Exosomes Promote Motility of Immortalized Hepatocyte through Transfer of Oncogenic Proteins and RNAs. Carcinogenesis 2015. [Google Scholar] [CrossRef] [PubMed]
- Kogure, T.; Yan, I.K.; Lin, W.L.; Patel, T. Extracellular Vesicle-Mediated Transfer of a Novel Long Noncoding RNA TUC339: A Mechanism of Intercellular Signaling in Human Hepatocellular Cancer. Genes Cancer 2013. [Google Scholar] [CrossRef] [PubMed]
- Kosaka, N.; Iguchi, H.; Yoshioka, Y.; Takeshita, F.; Matsuki, Y.; Ochiya, T. Secretory Mechanisms and Intercellular Transfer of MicroRNAs in Living Cells. J. Biol. Chem. 2010. [Google Scholar] [CrossRef]
- Meng, F.; Henson, R.; Wehbe-Janek, H.; Ghoshal, K.; Jacob, S.T.; Patel, T. MicroRNA-21 Regulates Expression of the PTEN Tumor Suppressor Gene in Human Hepatocellular Cancer. Gastroenterology 2007, 133, 647–658. [Google Scholar] [CrossRef]
- Liao, Q.; Han, P.; Huang, Y.; Wu, Z.; Chen, Q.; Li, S.; Ye, J.; Wu, X. Potential Role of Circulating MicroRNA-21 for Hepatocellular Carcinoma Diagnosis: A Meta-Analysis. PLoS ONE 2015, 10, e0130677. [Google Scholar] [CrossRef]
- Wang, H.; Hou, L.; Li, A.; Duan, Y.; Gao, H.; Song, X. Expression of Serum Exosomal MicroRNA-21 in Human Hepatocellular Carcinoma. BioMed Res. Int. 2014. [Google Scholar] [CrossRef]
- Shi, J. Considering Exosomal MiR-21 as a Biomarker for Cancer. J. Clin. Med. 2016, 42. [Google Scholar] [CrossRef]
- Feng, Y.H.; Tsao, C.J. Emerging Role of MicroRNA-21 in Cancer. Biomed. Rep. 2016, 5, 395–402. [Google Scholar] [CrossRef]
- Cao, L.Q.; Yang, X.W.; Chen, Y.B.; Zhang, D.W.; Jiang, X.F.; Xue, P. Exosomal MiR-21 Regulates the TETs/PTENp1/PTEN Pathway to Promote Hepatocellular Carcinoma Growth. Mol. Cancer. 2019, 18, 148, Erratum in 2020, 19, 59. [Google Scholar] [CrossRef]
- Bettermann, K.; Vucur, M.; Haybaeck, J.; Koppe, C.; Janssen, J.; Heymann, F.; Weber, A.; Weiskirchen, R.; Liedtke, C.; Gassler, N.; et al. TAK1 Suppresses a NEMO-Dependent but NF-KappaB-Inde-Pendent Pathway to Liver Cancer. Cancer Cell 2010, 17, 481–496. [Google Scholar] [CrossRef]
- Inokuchi, S.; Aoyama, T.; Miura, K.; Österreicher, C.H.; Kodama, Y.; Miyai, K.; Akira, S.; Brenner, D.A.; Seki, E. Disruption of TAK1 in Hepatocytes Causes Hepatic Injury, Inflammation, FIbrosis, and Carcinogenesis. Proc. Natl. Acad. Sci. USA 2010, 107, 844–849. [Google Scholar] [CrossRef]
- Besse, A.; Lamothe, B.; Campos, A.D.; Webster, W.K.; Maddineni, U.; Lin, S.C.; Wu, H.; Darnay, B.G. TAK1-Dependent Signaling Requires Functional Interaction with TAB2/TAB3. J. Biol. Chem. 2007. [Google Scholar] [CrossRef]
- Roh, Y.S.; Song, J.; Seki, E. TAK1 Regulates Hepatic Cell Survival and Carcinogenesis. J. Gastroenterol. 2014, 49, 185–194. [Google Scholar] [CrossRef]
- Lin, X.J.; Fang, J.H.; Yang, X.J.; Zhang, C.; Yuan, Y.; Zheng, L.; Zhuang, S.M. Hepatocellular Carcinoma Cell-Secreted Exosomal MicroRNA-210 Promotes Angiogenesis In Vitro and In Vivo. Mol. Ther. Nucleic Acids 2018. [Google Scholar] [CrossRef]
- Fang, J.H.; Zhang, Z.J.; Shang, L.R.; Luo, Y.W.; Lin, Y.F.; Yuan, Y.; Zhuang, S.M. Hepatoma Cell-Secreted Exosomal MicroRNA-103 Increases Vascular Permeability and Promotes Metastasis by Targeting Junction Proteins. Hepatology 2018. [Google Scholar] [CrossRef] [PubMed]
- Fang, T.; Lv, H.; Lv, G.; Li, T.; Wang, C.; Han, Q.; Yu, L.; Su, B.; Guo, L.; Huang, S.; et al. Tumor-Derived Exosomal MiR-1247-3p Induces Cancer-Associated Fibroblast Activation to Foster Lung Metastasis of Liver Cancer. Nat. Commun. 2018. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yan, I.K.; Haga, H.; Patel, T. Modulation of Hypoxia-Signaling Pathways by Extracellular Linc-RoR. J. Cell Sci. 2014. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yan, I.K.; Wood, J.; Haga, H.; Patel, T. Involvement of Extracellular Vesicle Long Noncoding RNA (Linc-VLDLR) in Tumor Cell Responses to Chemotherapy. Mol. Cancer Res. 2014. [Google Scholar] [CrossRef]
- Braconi, C.; Valeri, N.; Kogure, T.; Gasparini, P.; Huang, N.; Nuovo, G.J.; Terracciano, L.; Croce, C.M.; Patel, T. Expression and Functional Role of a Transcribed Noncoding RNA with an Ultraconserved Element in Hepatocellular Carcinoma. Proc. Natl. Acad. Sci. USA 2011, 108, 786–791. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Lei, Y.; Wu, M.; Li, N. Regulation of Macrophage Activation and Polarization by HCC-Derived Exosomal LncRNA TUC339. Int. J. Mol. Sci. 2018, 2958. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhang, A.; Ho, T.T.; Zhang, Z.; Zhou, N.; Ding, X.; Zhang, X.; Xu, M.; Mo, Y.Y. Linc-RoR Promotes c-Myc Expression through HnRNP i and AUF1. Nucleic Acids Res. 2015. [Google Scholar] [CrossRef]
- Zhang, H.; Deng, T.; Ge, S.; Liu, Y.; Bai, M.; Zhu, K.; Fan, Q.; Li, J.; Ning, T.; Tian, F.; et al. Exosome CircRNA Secreted from Adipocytes Promotes the Growth of Hepatocellular Carcinoma by Targeting Deubiquitination-Related USP7. Oncogene 2019, 38, 2844–2859. [Google Scholar] [CrossRef]
- Wang, G.; Liu, W.; Zou, Y.; Wang, G.; Deng, Y.; Luo, J.; Zhang, Y.; Li, H.; Zhang, Q.; Yang, Y.; et al. Three Isoforms of Exosomal CircPTGR1 Promote Hepatocellular Carcinoma Metastasis via the MiR449a–MET Pathway. EBioMedicine 2019. [Google Scholar] [CrossRef] [PubMed]
- Takaki, Y.; Saito, Y.; Takasugi, A.; Toshimitsu, K.; Yamada, S.; Muramatsu, T.; Kimura, M.; Sugiyama, K.; Suzuki, H.; Arai, E.; et al. Silencing of MicroRNA-122 Is an Early Event during Hepatocarcinogenesis from Non-Alcoholic Steatohepatitis. Cancer Sci. 2014, 105, 1254–1260. [Google Scholar] [CrossRef]
- Esau, C.; Davis, S.; Murray, S.F.; Yu, X.X.; Pandey, S.K.; Pear, M.; Watts, L.; Booten, S.L.; Graham, M.; McKay, R.; et al. MiR-122 Regulation of Lipid Metabolism Revealed by in vivo Antisense Targeting. Cell Metab. 2006. [Google Scholar] [CrossRef]
- Povero, D.; Eguchi, A.; Li, H.; Johnson, C.D.; Papouchado, B.G.; Wree, A.; Messer, K.; Feldstein, A.E. Circulating Extracellular Vesicles with Specific Proteome and Liver MicroRNAs Are Potential Biomarkers for Liver Injury in Experimental Fatty Liver Disease. PLoS ONE 2014. [Google Scholar] [CrossRef] [PubMed]
- Povero, D.; Eguchi, A.; Niesman, I.R.; Andronikou, N.; de Mollerat Du Jeu, X.; Mulya, A.; Berk, M.; Lazic, M.; Thapaliya, S.; Parola, M.; et al. Lipid-Induced Toxicity Stimulates Hepatocytes to Release Angiogenic Microparticles That Require Vanin-1 for Uptake by Endothelial Cells. Sci. Signal. 2013, 6, Ra88. [Google Scholar] [CrossRef]
- Csak, T.; Bala, S.; Lippai, D.; Satishchandran, A.; Catalano, D.; Kodys, K.; Szabo, G. MicroRNA-122 Regulates Hypoxia-Inducible Factor-1 and Vimentin in Hepatocytes and Correlates with Fibrosis in Diet-Induced Steatohepatitis. Liver Int. 2015, 35, 532–541. [Google Scholar] [CrossRef]
- Basu, S.; Bhattacharyya, S.N. Insulin-like Growth Factor-1 Prevents MiR-122 Production in Neighbouring Cells to Curtail Its Intercellular Transfer to Ensure Proliferation of Human Hepatoma Cells. Nucleic Acids Res. 2014. [Google Scholar] [CrossRef] [PubMed]
- Yamada, H.; Ohashi, K.; Suzuki, K.; Munetsuna, E.; Ando, Y.; Yamazaki, M.; Ishikawa, H.; Ichino, N.; Teradaira, R.; Hashimoto, S. Longitudinal Study of Circulating MiR-122 in a Rat Model of Non-Alcoholic Fatty Liver Disease. Clin. Chim. Acta Int. J. Clin. Chem. 2015, 446, 267–271. [Google Scholar] [CrossRef]
- Wei, J.X.; Lv, L.H.; Wan, Y.L.; Cao, Y.; Li, G.L.; Lin, H.M.; Zhou, R.; Shang, C.Z.; Cao, J.; He, H.; et al. Vps4A Functions as a Tumor Suppressor by Regulating the Secretion and Uptake of Exosomal MicroRNAs in Human Hepatoma Cells. Hepatology 2015, 61, 1284–1294. [Google Scholar] [CrossRef]
- Mann, J.; Reeves, H.L.; Feldstein, A.E. Liquid Biopsy for Liver Diseases. Gut 2018. [Google Scholar] [CrossRef]
- De Rubis, G.; Rajeev Krishnan, S.; Bebawy, M. Liquid Biopsies in Cancer Diagnosis, Monitoring, and Prognosis. Trends Pharmacol. Sci. 2019, 40, 172–186. [Google Scholar] [CrossRef]
- Pardini, B.; Sabo, A.A.; Birolo, G.; Calin, G.A. Noncoding RNAs in Extracellular Fluids as Cancer Biomarkers: The New Frontier of Liquid Biopsies. Cancers 2019, 11, 1170. [Google Scholar] [CrossRef]
- Lee, Y.R.; Kim, G.; Tak, W.Y.; Jang, S.Y.; Kweon, Y.O.; Park, J.G.; Lee, H.W.; Han, Y.S.; Chun, J.M.; Park, S.Y.; et al. Circulating Exosomal Noncoding RNAs as Prognostic Biomarkers in Human Hepatocellular Carcinoma. Int. J. Cancer 2019. [Google Scholar] [CrossRef]
- Afonso, M.; Rodrigues, P.; Simão, A.; Castro, R. Circulating MicroRNAs as Potential Biomarkers in Non-Alcoholic Fatty Liver Disease and Hepatocellular Carcinoma. J. Clin. Med. 2016, 5, 30. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Chen, S.; Liu, B. Diagnostic and Prognostic Values of Serum Exosomal MicroRNA-21 in Children with Hepatoblastoma: A Chinese Population-Based Study. Pediatr. Surg. Int. 2016. [Google Scholar] [CrossRef]
- Li, N.; Miao, Y.; Shan, Y.; Liu, B.; Li, Y.; Zhao, L.; Jia, L. MiR-106b and MiR-93 Regulate Cell Progression by Suppression of PTEN via PI3K/Akt Pathway in Breast Cancer. Cell Death Dis. 2017, 8, E2796. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Guo, Y.; Ma, Z.; Ma, G.; Xue, Q.; Li, F.; Liu, L. Long Non-Coding RNA PTENP1 Functions as a CeRNA to Modulate PTEN Level by Decoying MiR-106b and MiR-93 in Gastric Cancer. Oncotarget 2017, 8, 26079–26089. [Google Scholar] [CrossRef] [PubMed]
- Xue, X.; Wang, X.; Zhao, Y.; Hu, R.; Qin, L. Exosomal MiR-93 Promotes Proliferation and Invasion in Hepatocellular Carcinoma by Directly Inhibiting TIMP2/TP53INP1/CDKN1A. Biochem. Biophys. Res. Commun. 2018, 502, 515–521. [Google Scholar] [CrossRef]
- Mjelle, R.; Dima, S.O.; Bacalbasa, N.; Chawla, K.; Sorop, A.; Cucu, D.; Herlea, V.; Sætrom, P.; Popescu, I. Comprehensive Transcriptomic Analyses of Tissue, Serum, and Serum Exosomes from Hepatocellular Carcinoma Patients. BMC Cancer 2019. [Google Scholar] [CrossRef]
- Tian, X.P.; Wang, C.Y.; Jin, X.H.; Li, M.; Wang, F.W.; Huang, W.J.; Yun, J.P.; Xu, R.H.; Cai, Q.Q.; Xie, D. Acidic Microenvironment Up-Regulates Exosomal MiR-21 and MiR-10b in Early-Stage Hepatocellular Carcinoma to Promote Cancer Cell Proliferation and Metastasis. Theranostics 2019, 9, 1965–1979. [Google Scholar] [CrossRef]
- Sohn, W.; Kim, J.; Kang, S.H.; Yang, S.R.; Cho, J.Y.; Cho, H.C.; Shim, S.G.; Paik, Y.H. Serum Exosomal MicroRNAs as Novel Biomarkers for Hepatocellular Carcinoma. Exp. Mol. Med. 2015. [Google Scholar] [CrossRef]
- Tang, J.; Li, Y.; Liu, K.; Zhu, Q.; Yang, W.H.; Xiong, L.K.; Guo, D.L. Exosomal MiR- 9-3p Suppresses HBGF-5 Expression and Is a Functional Biomarker in Hepatocellular Carcinoma. Minerva Med. 2018, 109, 15–23. [Google Scholar] [PubMed]
- Cui, Y.; Xu, H.-F.; Liu, M.-Y.; Xu, Y.-J.; He, J.-C.; Zhou, Y.; Cang, S.-D. Mechanism of Exosomal MicroRNA-224 in Development of Hepatocellular Carcinoma and Its Diagnostic and Prognostic Value. World J. Gastroenterol. 2019, 25, 1890–1898. [Google Scholar] [CrossRef]
- Cho, H.J.; Eun, J.W.; Baek, G.O.; Seoù, C.W.; Ahn, H.R.; Kim, S.S.; Cho, S.W.; Cheong, J.Y. Serum Exosomal MicroRNA, MiR-10b-5p, as a Potential Diagnostic Biomarker for Early-Stage Hepatocellular Carcinoma. J. Clin. Med. 2020, 9, 281. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Tang, J.; Zhang, B.; Yang, W.; LiuGao, M.; Wang, R.; Tan, Y.; Fan, J.; Chang, Y.; Fu, J.; et al. Epigenetic Modification of MiR-429 Promotes Liver Tumour-Initiating Cell Properties by Targeting Rb Binding Protein 4. Gut 2015. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Hu, J.; Zhou, K.; Chen, F.; Wang, Z.; Liao, B.; Dai, Z.; Cao, Y.; Fan, J.; Zhou, J. Serum Exosomal MiR-125b Is a Novel Prognostic Marker for Hepatocellular Carcinoma. OncoTargets Ther. 2017. [Google Scholar] [CrossRef]
- Sugimachi, K.; Matsumura, T.; Hirata, H.; Uchi, R.; Ueda, M.; Ueo, H.; Shinden, Y.; Iguchi, T.; Eguchi, H.; Shirabe, K.; et al. Identification of a Bona Fide MicroRNA Biomarker in Serum Exosomes That Predicts Hepatocellular Carcinoma Recurrence after Liver Transplantation. Br. J. Cancer 2015. [Google Scholar] [CrossRef] [PubMed]
- Suehiro, T.; Miyaaki, H.; Shibata, H.; Honda, T.; Ozawa, E.; Miuma, S.; Taura, N.; Nakao, K. Significance of Serum Exosomal MiR-122 and MiR-21 as a Predictive Biomarker in Hepatocellular Carcinoma Patients Who Underwent Transarterial Chemoembolization. J. Hepatol. 2017. [Google Scholar] [CrossRef]
- Shi, M.; Jiang, Y.; Yang, L.; Yan, S.; Wang, Y.-G.; Lu, X.-J. Decreased Levels of Serum Exosomal MiR-638 Predict Poor Prognosis in Hepatocellular Carcinoma. J. Cell. Biochem. 2018, 119, 4711–4716. [Google Scholar] [CrossRef]
- Li, Y.; Zhao, J.; Yu, S.; Wang, Z.; He, X.; Su, Y.; Guo, T.; Sheng, H.; Chen, J.; Zheng, Q.; et al. Extracellular Vesicles Long RNA Sequencing Reveals Abundant MRNA, CircRNA, and LncRNA in Human Blood as Potential Biomarkers for Cancer Diagnosis. Clin. Chem. 2019, 65, 798–808. [Google Scholar] [CrossRef]
- Ma, X.; Yuan, T.; Yang, C.; Wang, Z.; Zang, Y.; Wu, L.; Zhuang, L. X-Inactive-Specific Transcript of Peripheral Blood Cells Is Regulated by Exosomal Jpx and Acts as a Biomarker for Female Patients with Hepatocellular Carcinoma. Ther. Adv. Med. Oncol. 2017, 9, 665–677. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Su, Y.; Liu, X.; Xu, M.; Chen, X.; Zhu, Y.; Guo, Z.; Bai, T.; Dong, L.; Wei, C.; et al. Serum and Exosome Long Non Coding RNAs as Potential Biomarkers for Hepatocellular Carcinoma. J. Cancer 2018, 9, 2631–2639. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Chen, Y.; Dong, X.; Wang, X. Serum Exosomal Long Noncoding RNAs ENSG00000258332.1 and LINC00635 for the Diagnosis and Prognosis of Hepatocellular Carcinoma. Cancer Epidemiol. Biomark. Prev. 2018, 27, 710–716. [Google Scholar] [CrossRef]
- Zhang, C.; Yang, X.; Qi, Q.; Gao, Y.; Wei, Q.; Han, S. LncRNA-HEIH in Serum and Exosomes as a Potential Biomarker in the HCV-Related Hepatocellular Carcinoma. Cancer Biomark. 2018, 21, 651–659. [Google Scholar] [CrossRef]
- Kim, S.S.; Baek, G.O.; Ahn, H.R.; Sung, S.; Seo, C.W.; Cho, H.J.; Nam, S.W.; Cheong, J.Y.; Eun, J.W. Serum Small ExtracellularVesicle-derived LINC00853 as a Novel Diagnostic Marker for Early Hepatocellular Carcinoma. Mol. Oncol. 2020, 14, 2646–2659. [Google Scholar] [CrossRef]
- Lin, H.; Zhang, Z. Diagnostic Value of a MicroRNAsignature Panel in Exosomes for Patients with Hepatocellular Carcinoma. Int. J. Clin. Exp. Pathol. 2019, 12, 1478–1487. [Google Scholar] [PubMed]
- Wang, Y.; Zhang, C.; Zhang, P.; Guo, G.; Jiang, T.; Zhao, X.; Jiang, J.; Huang, X.; Tong, H.; Tian, Y. Serum Exosomal MicroRNAs Combined with Alpha-Fetoprotein as Diagnostic Markers of Hepatocellular Carcinoma. Cancer Med. 2018, 7, 1670–1679. [Google Scholar] [CrossRef]
- Wang, G.; Zhao, W.; Wang, H.; Qiu, G.; Jiang, Z.; Wei, G.; Li, X. Exosomal MiR-744 Inhibits Proliferation and Sorafenib Chemoresistance in Hepatocellular Carcinoma by Targeting PAX2. Med. Sci. Monit. 2019, 25, 7209–7217. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Costanzi, E.; Simioni, C.; Varano, G.; Brenna, C.; Conti, I.; Neri, L.M. The Role of Extracellular Vesicles as Shuttles of RNA and Their Clinical Significance as Biomarkers in Hepatocellular Carcinoma. Genes 2021, 12, 902. https://doi.org/10.3390/genes12060902
Costanzi E, Simioni C, Varano G, Brenna C, Conti I, Neri LM. The Role of Extracellular Vesicles as Shuttles of RNA and Their Clinical Significance as Biomarkers in Hepatocellular Carcinoma. Genes. 2021; 12(6):902. https://doi.org/10.3390/genes12060902
Chicago/Turabian StyleCostanzi, Eva, Carolina Simioni, Gabriele Varano, Cinzia Brenna, Ilaria Conti, and Luca Maria Neri. 2021. "The Role of Extracellular Vesicles as Shuttles of RNA and Their Clinical Significance as Biomarkers in Hepatocellular Carcinoma" Genes 12, no. 6: 902. https://doi.org/10.3390/genes12060902
APA StyleCostanzi, E., Simioni, C., Varano, G., Brenna, C., Conti, I., & Neri, L. M. (2021). The Role of Extracellular Vesicles as Shuttles of RNA and Their Clinical Significance as Biomarkers in Hepatocellular Carcinoma. Genes, 12(6), 902. https://doi.org/10.3390/genes12060902






