Amplified Fragments of an Autosome-Borne Gene Constitute a Significant Component of the W Sex Chromosome of Eremias velox (Reptilia, Lacertidae)
Abstract
1. Introduction
2. Materials and Methods
3. Results
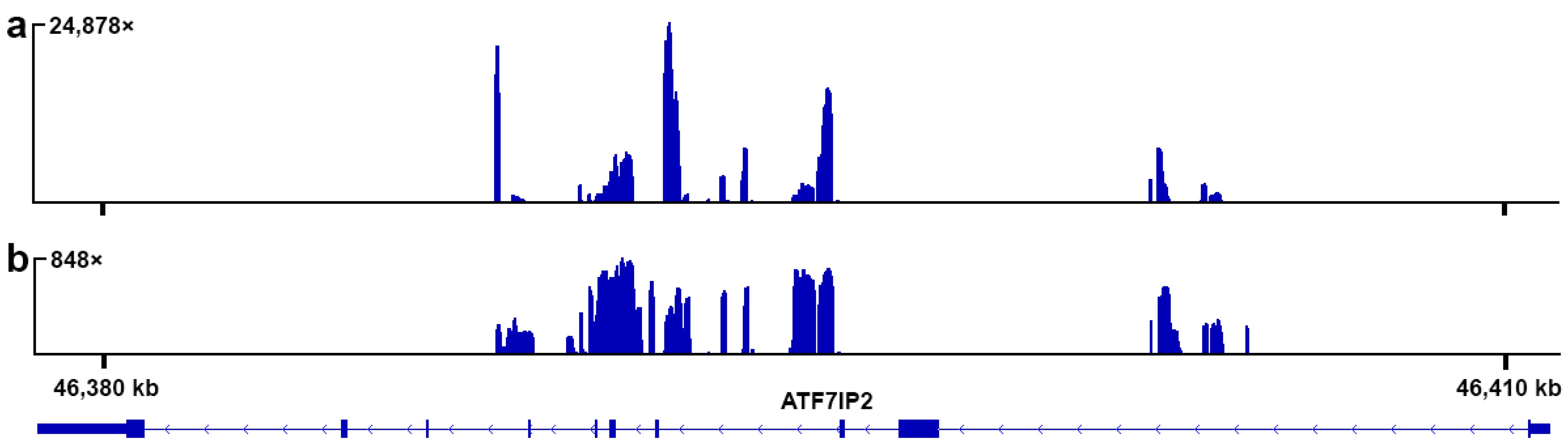
3.1. Reference-Based Assembly
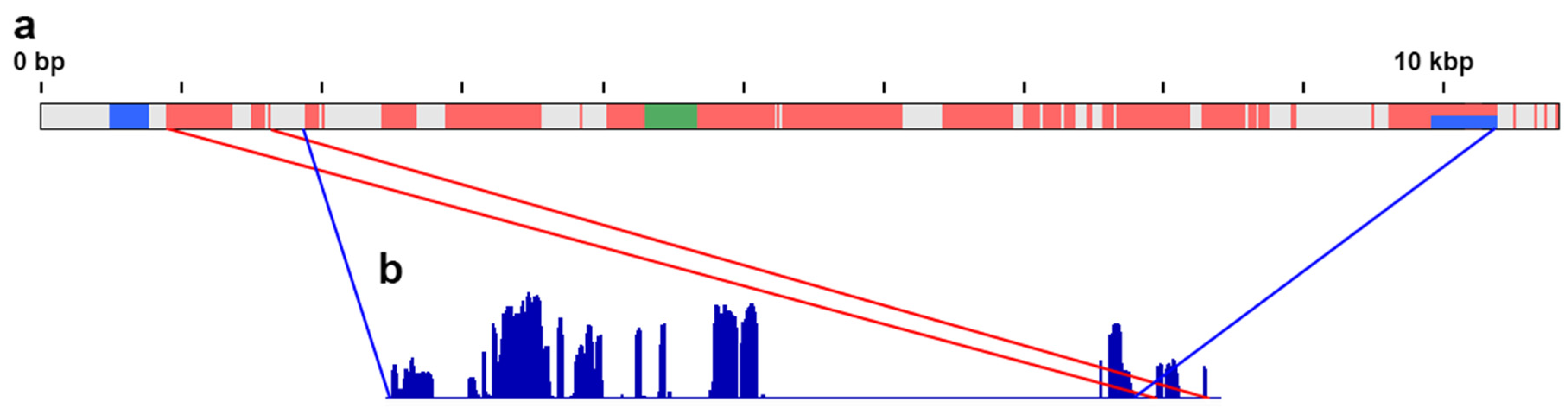
3.2. De Novo Assembly
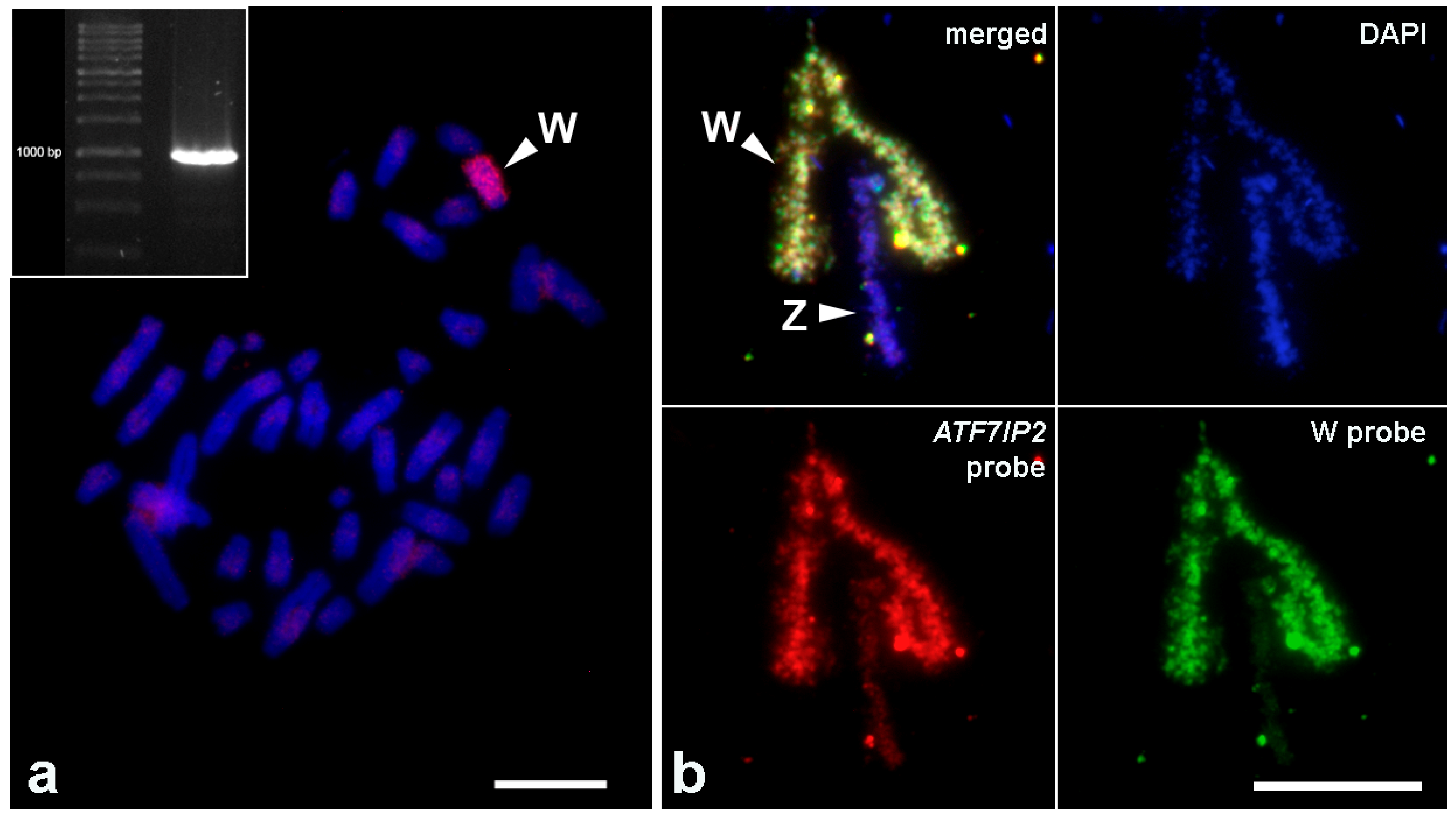
3.3. FISH with the Probe to the Fragment of ATF7IP2
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Berta, P.; Hawkins, J.R.; Sinclair, A.H.; Taylor, A.; Griffiths, B.L.; Goodfellow, P.N.; Fellous, M. Genetic evidence equating SRY and the testis-determining factor. Nature 1990, 348, 448–450. [Google Scholar] [CrossRef]
- Yoshimoto, S.; Okada, E.; Umemoto, H.; Tamura, K.; Uno, Y.; Nishida-Umehara, C.; Matsuda, Y.; Takamatsu, N.; Shiba, T.; Ito, M. A W-linked DM-domain gene, DM-W, participates in primary ovary development in Xenopus laevis. Proc. Natl. Acad. Sci. USA 2008, 105, 2469–2474. [Google Scholar] [CrossRef]
- Hirst, C.E.; Major, A.T.; Ayers, K.L.; Brown, R.J.; Mariette, M.; Sackton, T.B.; Smith, C.A. Sex Reversal and Comparative Data Undermine the W Chromosome and Support Z-linked DMRT1 as the Regulator of Gonadal Sex Differentiation in Birds. Endocrinology 2017, 158, 2970–2987. [Google Scholar] [CrossRef]
- Bachtrog, D.; Mank, J.E.; Peichel, C.L.; Kirkpatrick, M.; Otto, S.P.; Ashman, T.-L.; Hahn, M.W.; Kitano, J.; Mayrose, I.; Ming, R.; et al. Sex determination: Why so many ways of doing it? PLoS Biol. 2014, 12, e1001899. [Google Scholar] [CrossRef]
- Kostmann, A.; Kratochvíl, L.; Rovatsos, M. Poorly differentiated XX/XY sex chromosomes are widely shared across skink radiation. Proc. Biol. Sci. 2021, 288, 20202139. [Google Scholar] [CrossRef]
- Rice, W.R. The Accumulation of Sexually Antagonistic Genes as a Selective Agent Promoting the Evolution of Reduced Recombination between Primitive Sex Chromosomes. Evolution 1987, 41, 911. [Google Scholar] [CrossRef]
- Charlesworth, D. Evolution of recombination rates between sex chromosomes. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2017, 372. [Google Scholar] [CrossRef]
- Waters, P.D.; Ruiz-Herrera, A. Meiotic Executioner Genes Protect the Y from Extinction. Trends Genet. 2020, 36, 728–738. [Google Scholar] [CrossRef]
- Wright, A.E.; Darolti, I.; Bloch, N.I.; Oostra, V.; Sandkam, B.; Buechel, S.D.; Kolm, N.; Breden, F.; Vicoso, B.; Mank, J.E. Convergent recombination suppression suggests role of sexual selection in guppy sex chromosome formation. Nat. Commun. 2017, 8, 14251. [Google Scholar] [CrossRef]
- Charlesworth, D. The Guppy Sex Chromosome System and the Sexually Antagonistic Polymorphism Hypothesis for Y Chromosome Recombination Suppression. Genes 2018, 9, 264. [Google Scholar] [CrossRef]
- Lindholm, A.K.; Brooks, R.; Breden, F. Extreme polymorphism in a Y-linked sexually selected trait. Heredity 2004, 92, 156–162. [Google Scholar] [CrossRef]
- Charlesworth, B.; Charlesworth, D. The degeneration of Y chromosomes. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2000, 355, 1563–1572. [Google Scholar] [CrossRef]
- Schartl, M.; Schmid, M.; Nanda, I. Dynamics of vertebrate sex chromosome evolution: From equal size to giants and dwarfs. Chromosoma 2016, 125, 553–571. [Google Scholar] [CrossRef]
- Matsubara, K.; Nishida, C.; Matsuda, Y.; Kumazawa, Y. Sex chromosome evolution in snakes inferred from divergence patterns of two gametologous genes and chromosome distribution of sex chromosome-linked repetitive sequences. Zool. Lett. 2016, 2, 19. [Google Scholar] [CrossRef] [PubMed]
- Cortez, D.; Marin, R.; Toledo-Flores, D.; Froidevaux, L.; Liechti, A.; Waters, P.D.; Grützner, F.; Kaessmann, H. Origins and functional evolution of Y chromosomes across mammals. Nature 2014, 508, 488–493. [Google Scholar] [CrossRef] [PubMed]
- Bellott, D.W.; Skaletsky, H.; Cho, T.-J.; Brown, L.; Locke, D.; Chen, N.; Galkina, S.; Pyntikova, T.; Koutseva, N.; Graves, T.; et al. Avian W and mammalian Y chromosomes convergently retained dosage-sensitive regulators. Nat. Genet. 2017, 49, 387–394. [Google Scholar] [CrossRef]
- Zhou, Q.; Zhang, J.; Bachtrog, D.; An, N.; Huang, Q.; Jarvis, E.D.; Gilbert, M.T.P.; Zhang, G. Complex evolutionary trajectories of sex chromosomes across bird taxa. Science 2014, 346, 1246338. [Google Scholar] [CrossRef]
- Gunski, R.J.; Kretschmer, R.; Santos de Souza, M.; de Oliveira Furo, I.; Barcellos, S.A.; Costa, A.L.; Cioffi, M.B.; de Oliveira, E.H.C.; Del Valle Garnero, A. Evolution of Bird Sex Chromosomes Narrated by Repetitive Sequences: Unusual W Chromosome Enlargement in Gallinula melanops (Aves: Gruiformes: Rallidae). Cytogenet. Genome Res. 2019, 158, 152–159. [Google Scholar] [CrossRef] [PubMed]
- Augstenová, B.; Mazzoleni, S.; Kratochvíl, L.; Rovatsos, M. Evolutionary Dynamics of the W Chromosome in Caenophidian Snakes. Genes 2017, 9, 5. [Google Scholar] [CrossRef]
- Laopichienpong, N.; Muangmai, N.; Chanhome, L.; Suntrarachun, S.; Twilprawat, P.; Peyachoknagul, S.; Srikulnath, K. Evolutionary Dynamics of the Gametologous CTNNB1 Gene on the Z and W Chromosomes of Snakes. J. Hered. 2017, 108, 142–151. [Google Scholar] [CrossRef]
- Cechova, M.; Vegesna, R.; Tomaszkiewicz, M.; Harris, R.S.; Chen, D.; Rangavittal, S.; Medvedev, P.; Makova, K.D. Dynamic evolution of great ape Y chromosomes. Proc. Natl. Acad. Sci. USA 2020, 117, 26273–26280. [Google Scholar] [CrossRef]
- Xu, L.; Zhou, Q. The Female-Specific W Chromosomes of Birds Have Conserved Gene Contents but Are Not Feminized. Genes 2020, 11, 1126. [Google Scholar] [CrossRef]
- Vicoso, B.; Emerson, J.J.; Zektser, Y.; Mahajan, S.; Bachtrog, D. Comparative sex chromosome genomics in snakes: Differentiation, evolutionary strata, and lack of global dosage compensation. PLoS Biol. 2013, 11, e1001643. [Google Scholar] [CrossRef] [PubMed]
- Schield, D.R.; Card, D.C.; Hales, N.R.; Perry, B.W.; Pasquesi, G.M.; Blackmon, H.; Adams, R.H.; Corbin, A.B.; Smith, C.F.; Ramesh, B.; et al. The origins and evolution of chromosomes, dosage compensation, and mechanisms underlying venom regulation in snakes. Genome Res. 2019, 29, 590–601. [Google Scholar] [CrossRef]
- Kondo, M.; Hornung, U.; Nanda, I.; Imai, S.; Sasaki, T.; Shimizu, A.; Asakawa, S.; Hori, H.; Schmid, M.; Shimizu, N.; et al. Genomic organization of the sex-determining and adjacent regions of the sex chromosomes of medaka. Genome Res. 2006, 16, 815–826. [Google Scholar] [CrossRef]
- Jeffries, D.L.; Lavanchy, G.; Sermier, R.; Sredl, M.J.; Miura, I.; Borzée, A.; Barrow, L.N.; Canestrelli, D.; Crochet, P.-A.; Dufresnes, C.; et al. A rapid rate of sex-chromosome turnover and non-random transitions in true frogs. Nat. Commun. 2018, 9, 4088. [Google Scholar] [CrossRef]
- Bewick, A.J.; Chain, F.J.J.; Zimmerman, L.B.; Sesay, A.; Gilchrist, M.J.; Owens, N.D.L.; Seifertova, E.; Krylov, V.; Macha, J.; Tlapakova, T.; et al. A large pseudoautosomal region on the sex chromosomes of the frog Silurana tropicalis. Genome Biol. Evol. 2013, 5, 1087–1098. [Google Scholar] [CrossRef]
- Stöck, M.; Horn, A.; Grossen, C.; Lindtke, D.; Sermier, R.; Betto-Colliard, C.; Dufresnes, C.; Bonjour, E.; Dumas, Z.; Luquet, E.; et al. Ever-young sex chromosomes in European tree frogs. PLoS Biol. 2011, 9, e1001062. [Google Scholar] [CrossRef]
- Tripathi, N.; Hoffmann, M.; Weigel, D.; Dreyer, C. Linkage analysis reveals the independent origin of Poeciliid sex chromosomes and a case of atypical sex inheritance in the guppy (Poecilia reticulata). Genetics 2009, 182, 365–374. [Google Scholar] [CrossRef]
- Nanda, I.; Schories, S.; Tripathi, N.; Dreyer, C.; Haaf, T.; Schmid, M.; Schartl, M. Sex chromosome polymorphism in guppies. Chromosoma 2014, 123, 373–383. [Google Scholar] [CrossRef]
- Charlesworth, D.; Zhang, Y.; Bergero, R.; Graham, C.; Gardner, J.; Yong, L. Using GC Content to Compare Recombination Patterns on the Sex Chromosomes and Autosomes of the Guppy, Poecilia reticulata, and Its Close Outgroup Species. Mol. Biol. Evol. 2020, 37, 3550–3562. [Google Scholar] [CrossRef] [PubMed]
- Lisachov, A.P.; Zadesenets, K.S.; Rubtsov, N.B.; Borodin, P.M. Sex chromosome synapsis and recombination in male guppies. Zebrafish 2015, 12, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Marin, R.; Cortez, D.; Lamanna, F.; Pradeepa, M.M.; Leushkin, E.; Julien, P.; Liechti, A.; Halbert, J.; Brüning, T.; Mössinger, K.; et al. Convergent origination of a Drosophila-like dosage compensation mechanism in a reptile lineage. Genome Res. 2017, 27, 1974–1987. [Google Scholar] [CrossRef] [PubMed]
- Rovatsos, M.; Rehák, I.; Velenský, P.; Kratochvíl, L. Shared Ancient Sex Chromosomes in Varanids, Beaded Lizards, and Alligator Lizards. Mol. Biol. Evol. 2019, 36, 1113–1120. [Google Scholar] [CrossRef] [PubMed]
- Matsubara, K.; Sarre, S.D.; Georges, A.; Matsuda, Y.; Marshall Graves, J.A.; Ezaz, T. Highly differentiated ZW sex microchromosomes in the Australian Varanus species evolved through rapid amplification of repetitive sequences. PLoS ONE 2014, 9, e95226. [Google Scholar] [CrossRef]
- Rovatsos, M.; Vukić, J.; Kratochvíl, L. Mammalian X homolog acts as sex chromosome in lacertid lizards. Heredity 2016, 117, 8–13. [Google Scholar] [CrossRef]
- Olmo, E.; Odierna, G.; Capriglione, T. Evolution of sex-chromosomes in lacertid lizards. Chromosoma 1987, 96, 33–38. [Google Scholar] [CrossRef]
- Rovatsos, M.; Vukić, J.; Mrugała, A.; Suwala, G.; Lymberakis, P.; Kratochvíl, L. Little evidence for switches to environmental sex determination and turnover of sex chromosomes in lacertid lizards. Sci. Rep. 2019, 9, 7832. [Google Scholar] [CrossRef]
- Matsubara, K.; Uno, Y.; Srikulnath, K.; Matsuda, Y.; Miller, E.; Olsson, M. No Interstitial Telomeres on Autosomes but Remarkable Amplification of Telomeric Repeats on the W Sex Chromosome in the Sand Lizard (Lacerta agilis). J. Hered. 2015, 106, 753–757. [Google Scholar] [CrossRef]
- Giovannotti, M.; Nisi Cerioni, P.; Rojo, V.; Olmo, E.; Slimani, T.; Splendiani, A.; Caputo Barucchi, V. Characterization of a satellite DNA in the genera Lacerta and Timon (Reptilia, Lacertidae) and its role in the differentiation of the W chromosome. J. Exp. Zool. B Mol. Dev. Evol. 2018, 330, 83–95. [Google Scholar] [CrossRef]
- Suwala, G.; Altmanová, M.; Mazzoleni, S.; Karameta, E.; Pafilis, P.; Kratochvíl, L.; Rovatsos, M. Evolutionary Variability of W-Linked Repetitive Content in Lacertid Lizards. Genes 2020, 11, 531. [Google Scholar] [CrossRef] [PubMed]
- Lisachov, A.P.; Galkina, S.A.; Saifitdinova, A.F.; Svetlana, A.R.; Andreyushkova, D.A.; Trifonov, V.A.; Borodin, P.M. Identification of sex chromosomes in Eremiasvelox (Lacertidae, Reptilia) using lampbrush chromosome analysis. Comp. Cytogenet. 2019, 13, 121–132. [Google Scholar] [CrossRef]
- Morgan, G.T. Imaging the dynamics of transcription loops in living chromosomes. Chromosoma 2018, 127, 361–374. [Google Scholar] [CrossRef] [PubMed]
- Gall, J.G. Are lampbrush chromosomes unique to meiotic cells? Chromosome Res. 2012, 20, 905–909. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gaginskaya, E.; Kulikova, T.; Krasikova, A. Avian lampbrush chromosomes: A powerful tool for exploration of genome expression. Cytogenet. Genome Res. 2009, 124, 251–267. [Google Scholar] [CrossRef]
- Solovei, I.; Gaginskaya, E.; Hutchison, N.; Macgregor, H. Avian sex chromosomes in the lampbrush form: The ZW lampbrush bivalents from six species of bird. Chromosome Res. 1993, 1, 153–166. [Google Scholar] [CrossRef] [PubMed]
- Pokorná, M.; Kratochvíl, L.; Kejnovský, E. Microsatellite distribution on sex chromosomes at different stages of heteromorphism and heterochromatinization in two lizard species (Squamata: Eublepharidae: Coleonyx elegans and lacertidae: Eremias velox). BMC Genet. 2011, 12, 90. [Google Scholar] [CrossRef]
- Stanyon, R.; Galleni, L. A rapid fibroblast culture technique for high resolution karyotypes. Bolletino Zool. 1991, 58, 81–83. [Google Scholar] [CrossRef]
- Yang, F.; O’Brien, P.C.; Milne, B.S.; Graphodatsky, A.S.; Solanky, N.; Trifonov, V.; Rens, W.; Sargan, D.; Ferguson-Smith, M.A. A complete comparative chromosome map for the dog, red fox, and human and its integration with canine genetic maps. Genomics 1999, 62, 189–202. [Google Scholar] [CrossRef]
- Saifitdinova, A.; Galkina, S.; Volodkina, V.; Gaginskaya, E. Preparation of lampbrush chromosomes dissected from avian and reptilian growing oocytes. Biol. Commun. 2017, 62, 165–168. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 1989; ISBN 0879693096. [Google Scholar]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Makunin, A.I.; Kichigin, I.G.; Larkin, D.M.; O’Brien, P.C.M.; Ferguson-Smith, M.A.; Yang, F.; Proskuryakova, A.A.; Vorobieva, N.V.; Chernyaeva, E.N.; O’Brien, S.J.; et al. Contrasting origin of B chromosomes in two cervids (Siberian roe deer and grey brocket deer) unravelled by chromosome-specific DNA sequencing. BMC Genom. 2016, 17, 618. [Google Scholar] [CrossRef]
- Flynn, J.M.; Hubley, R.; Goubert, C.; Rosen, J.; Clark, A.G.; Feschotte, C.; Smit, A.F. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA 2020, 117, 9451–9457. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 2006, 22, 1658–1659. [Google Scholar] [CrossRef] [PubMed]
- Liehr, T. Fluorescence In situ Hybridization (FISH): Application Guide, 2nd ed.; Springer: New York, NY, USA, 2017; ISBN 9783662529591. [Google Scholar]
- Waters, P.D.; Duffy, B.; Frost, C.J.; Delbridge, M.L.; Graves, J.A. The human Y chromosome derives largely from a single autosomal region added to the sex chromosomes 80–130 million years ago. Cytogenet. Cell Genet. 2001, 92, 74–79. [Google Scholar] [CrossRef] [PubMed]
- Pokorná, M.; Rábová, M.; Ráb, P.; Ferguson-Smith, M.A.; Rens, W.; Kratochvíl, L. Differentiation of sex chromosomes and karyotypic evolution in the eye-lid geckos (Squamata: Gekkota: Eublepharidae), a group with different modes of sex determination. Chromosome Res. 2010, 18, 809–820. [Google Scholar] [CrossRef]
- Rovatsos, M.; Johnson Pokorná, M.; Altmanová, M.; Kratochvíl, L. Mixed-Up Sex Chromosomes: Identification of Sex Chromosomes in the X1X1X2X2/X1X2Y System of the Legless Lizards of the Genus Lialis (Squamata: Gekkota: Pygopodidae). Cytogenet. Genome Res. 2016, 149, 282–289. [Google Scholar] [CrossRef]
- Giovannotti, M.; Trifonov, V.A.; Paoletti, A.; Kichigin, I.G.; O’Brien, P.C.M.; Kasai, F.; Giovagnoli, G.; Ng, B.L.; Ruggeri, P.; Cerioni, P.N.; et al. New insights into sex chromosome evolution in anole lizards (Reptilia, Dactyloidae). Chromosoma 2017, 126, 245–260. [Google Scholar] [CrossRef]
- Lisachov, A.P.; Tishakova, K.V.; Romanenko, S.A.; Molodtseva, A.S.; Prokopov, D.Y.; Pereira, J.C.; Ferguson-Smith, M.A.; Borodin, P.M.; Trifonov, V.A. Whole-Chromosome Fusions in the Karyotype Evolution of Sceloporus (Iguania, Reptilia) are More Intense in Sex Chromosomes Than Autosomes. bioRxiv 2020. [Google Scholar]
- Kupriyanova, L.; Kuksin, A.; Odierna, G. Karyotype, chromosome structure, reproductive modalities of three Southern Eurasian populations of the common lacertid lizard, Zootoca vivipara Jacquin, 1787. Acta Herpetol. 2008, 2, 99–106. [Google Scholar]
- Xu, L.; Auer, G.; Peona, V.; Suh, A.; Deng, Y.; Feng, S.; Zhang, G.; Blom, M.P.K.; Christidis, L.; Prost, S.; et al. Dynamic evolutionary history and gene content of sex chromosomes across diverse songbirds. Nat. Ecol. Evol. 2019, 3, 834–844. [Google Scholar] [CrossRef] [PubMed]
- Saxena, R.; Brown, L.G.; Hawkins, T.; Alagappan, R.K.; Skaletsky, H.; Reeve, M.P.; Reijo, R.; Rozen, S.; Dinulos, M.B.; Disteche, C.M.; et al. The DAZ gene cluster on the human Y chromosome arose from an autosomal gene that was transposed, repeatedly amplified and pruned. Nat. Genet. 1996, 14, 292–299. [Google Scholar] [CrossRef] [PubMed]
- Tsai, K.L.; Evans, J.M.; Noorai, R.E.; Starr-Moss, A.N.; Clark, L.A. Novel Y Chromosome Retrocopies in Canids Revealed through a Genome-Wide Association Study for Sex. Genes 2019, 10, 320. [Google Scholar] [CrossRef] [PubMed]
- Lisachov, A.P.; Makunin, A.I.; Giovannotti, M.; Pereira, J.C.; Druzhkova, A.S.; Caputo Barucchi, V.; Ferguson-Smith, M.A.; Trifonov, V.A. Genetic Content of the Neo-Sex Chromosomes in Ctenonotus and Norops (Squamata, Dactyloidae) and Degeneration of the Y Chromosome as Revealed by High-Throughput Sequencing of Individual Chromosomes. Cytogenet. Genome Res. 2019, 157, 115–122. [Google Scholar] [CrossRef]
- Giovannotti, M.; Rojo, V.; Nisi Cerioni, P.; González-Tizón, A.; Martínez-Lage, A.; Splendiani, A.; Naveira, H.; Ruggeri, P.; Arribas, Ó.; Olmo, E.; et al. Isolation and characterization of two satellite DNAs in some Iberian rock lizards (Squamata, Lacertidae). J. Exp. Zool. B Mol. Dev. Evol. 2014, 322, 13–26. [Google Scholar] [CrossRef] [PubMed]
- Rojo, V.; Martínez-Lage, A.; Giovannotti, M.; González-Tizón, A.M.; Nisi Cerioni, P.; Caputo Barucchi, V.; Galán, P.; Olmo, E.; Naveira, H. Evolutionary dynamics of two satellite DNA families in rock lizards of the genus Iberolacerta (Squamata, Lacertidae): Different histories but common traits. Chromosome Res. 2015, 23, 441–461. [Google Scholar] [CrossRef]
- Peona, V.; Blom, M.P.K.; Xu, L.; Burri, R.; Sullivan, S.; Bunikis, I.; Liachko, I.; Haryoko, T.; Jønsson, K.A.; Zhou, Q.; et al. Identifying the causes and consequences of assembly gaps using a multiplatform genome assembly of a bird-of-paradise. Mol. Ecol. Resour. 2021, 21, 263–286. [Google Scholar] [CrossRef]
- Ichimura, T.; Watanabe, S.; Sakamoto, Y.; Aoto, T.; Fujita, N.; Nakao, M. Transcriptional repression and heterochromatin formation by MBD1 and MCAF/AM family proteins. J. Biol. Chem. 2005, 280, 13928–13935. [Google Scholar] [CrossRef]
- Timms, R.T.; Tchasovnikarova, I.A.; Antrobus, R.; Dougan, G.; Lehner, P.J. ATF7IP-Mediated Stabilization of the Histone Methyltransferase SETDB1 Is Essential for Heterochromatin Formation by the HUSH Complex. Cell Rep. 2016, 17, 653–659. [Google Scholar] [CrossRef]
- Gardner, E.J.; Nizami, Z.F.; Talbot, C.C.; Gall, J.G. Stable intronic sequence RNA (sisRNA), a new class of noncoding RNA from the oocyte nucleus of Xenopus tropicalis. Genes Dev. 2012, 26, 2550–2559. [Google Scholar] [CrossRef]
- Liu, J.; Guo, X.; Chen, D.; Li, J.; Yue, B.; Zeng, X. Diversification and historical demography of the rapid racerunner (Eremias velox) in relation to geological history and Pleistocene climatic oscillations in arid Central Asia. Mol. Phylogenetics Evol. 2019, 130, 244–258. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Tang, X.; Xin, Y.; Yue, F.; Yan, X.; Liu, B.; An, B.; Wang, X.; Chen, Q. Identification of Sex Chromosomes by Means of Comparative Genomic Hybridization in a Lizard, Eremias multiocellata. Zool. Sci. 2015, 32, 151–156. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lisachov, A.; Andreyushkova, D.; Davletshina, G.; Prokopov, D.; Romanenko, S.; Galkina, S.; Saifitdinova, A.; Simonov, E.; Borodin, P.; Trifonov, V. Amplified Fragments of an Autosome-Borne Gene Constitute a Significant Component of the W Sex Chromosome of Eremias velox (Reptilia, Lacertidae). Genes 2021, 12, 779. https://doi.org/10.3390/genes12050779
Lisachov A, Andreyushkova D, Davletshina G, Prokopov D, Romanenko S, Galkina S, Saifitdinova A, Simonov E, Borodin P, Trifonov V. Amplified Fragments of an Autosome-Borne Gene Constitute a Significant Component of the W Sex Chromosome of Eremias velox (Reptilia, Lacertidae). Genes. 2021; 12(5):779. https://doi.org/10.3390/genes12050779
Chicago/Turabian StyleLisachov, Artem, Daria Andreyushkova, Guzel Davletshina, Dmitry Prokopov, Svetlana Romanenko, Svetlana Galkina, Alsu Saifitdinova, Evgeniy Simonov, Pavel Borodin, and Vladimir Trifonov. 2021. "Amplified Fragments of an Autosome-Borne Gene Constitute a Significant Component of the W Sex Chromosome of Eremias velox (Reptilia, Lacertidae)" Genes 12, no. 5: 779. https://doi.org/10.3390/genes12050779
APA StyleLisachov, A., Andreyushkova, D., Davletshina, G., Prokopov, D., Romanenko, S., Galkina, S., Saifitdinova, A., Simonov, E., Borodin, P., & Trifonov, V. (2021). Amplified Fragments of an Autosome-Borne Gene Constitute a Significant Component of the W Sex Chromosome of Eremias velox (Reptilia, Lacertidae). Genes, 12(5), 779. https://doi.org/10.3390/genes12050779








