miR-143 Targeting CUX1 to Regulate Proliferation of Dermal Papilla Cells in Hu Sheep
Abstract
1. Introduction
2. Materials and Methods
2.1. Skin Tissue Collection, RNA and Protein Extraction
2.2. Cell Culture and Transfection
2.3. Cell Proliferation and Cell Cycle
2.4. Target Gene Prediction for miR-143
2.5. Quantitative RT-PCR and Western Blot
2.6. Luciferase Reporter Assay
2.7. Statistical Analysis
3. Results
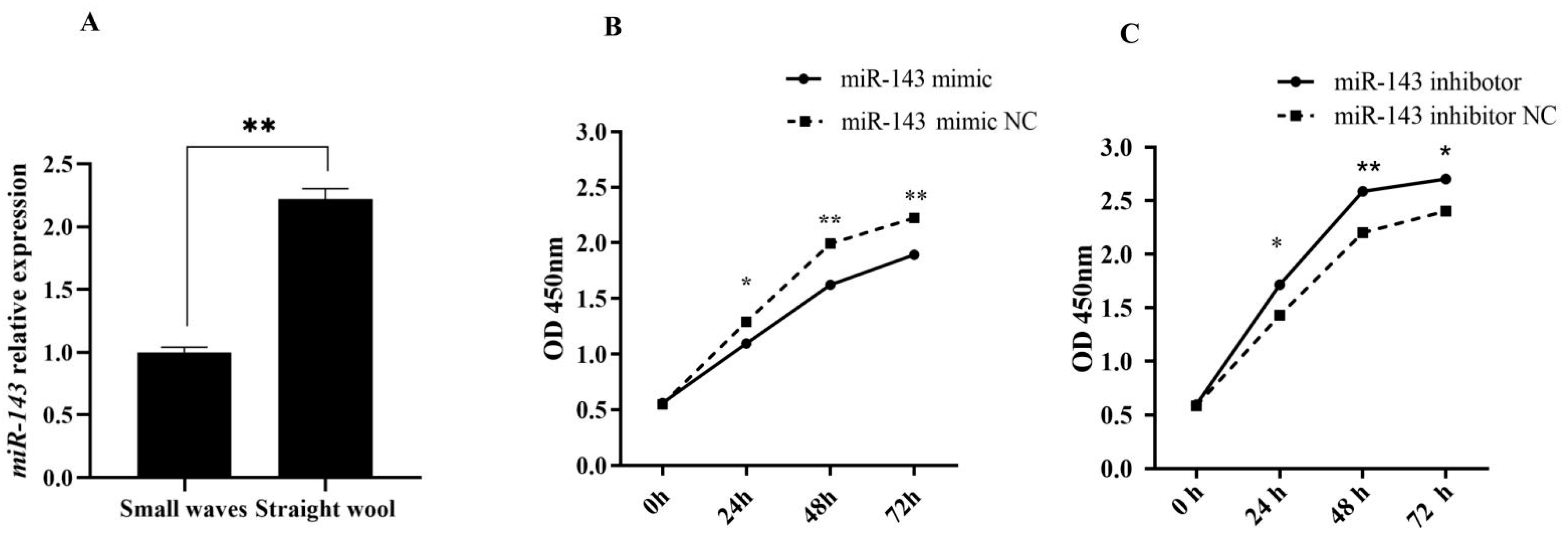
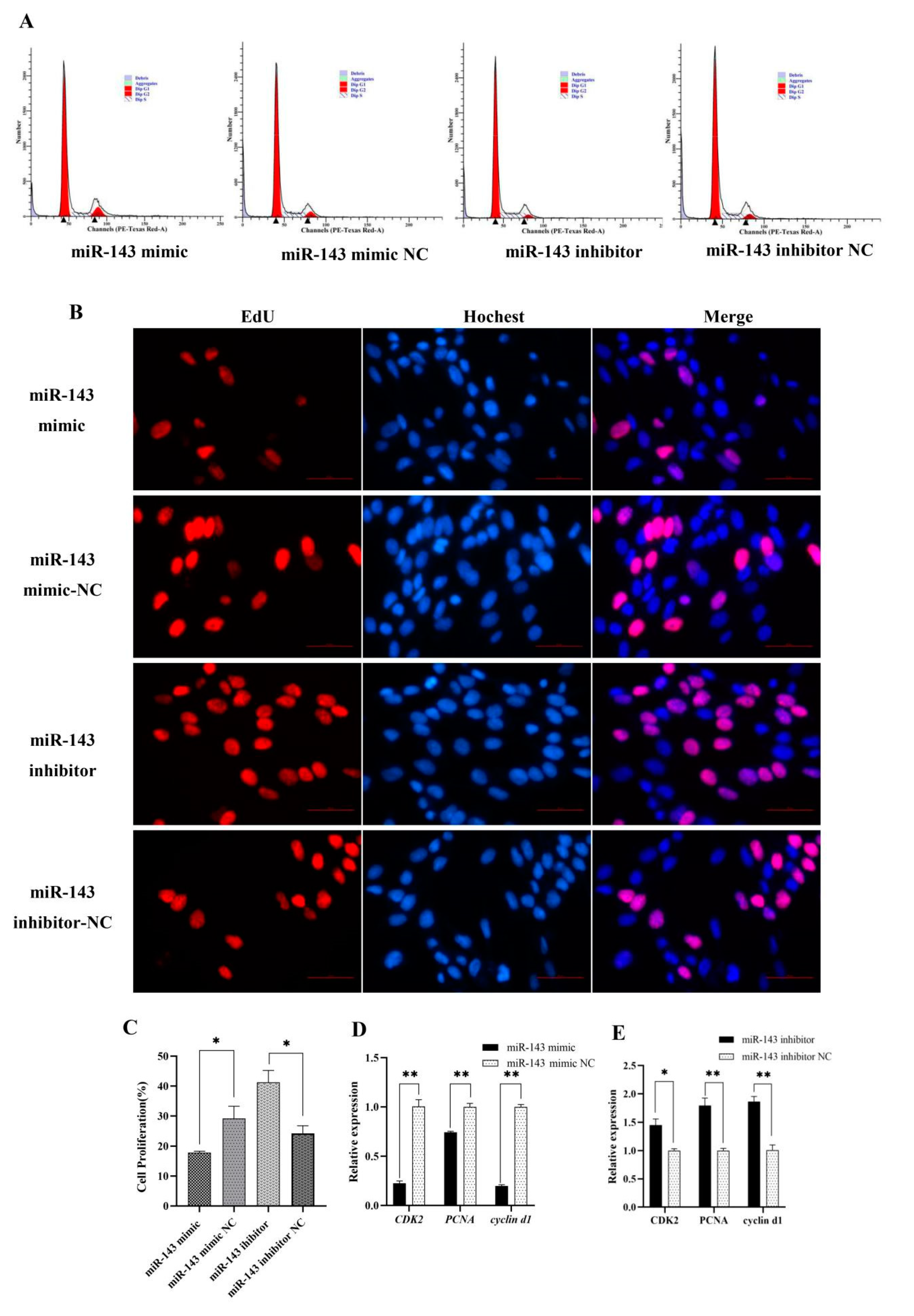
3.1. miR-143 Promoting Dermal Papilla Cells Proliferation
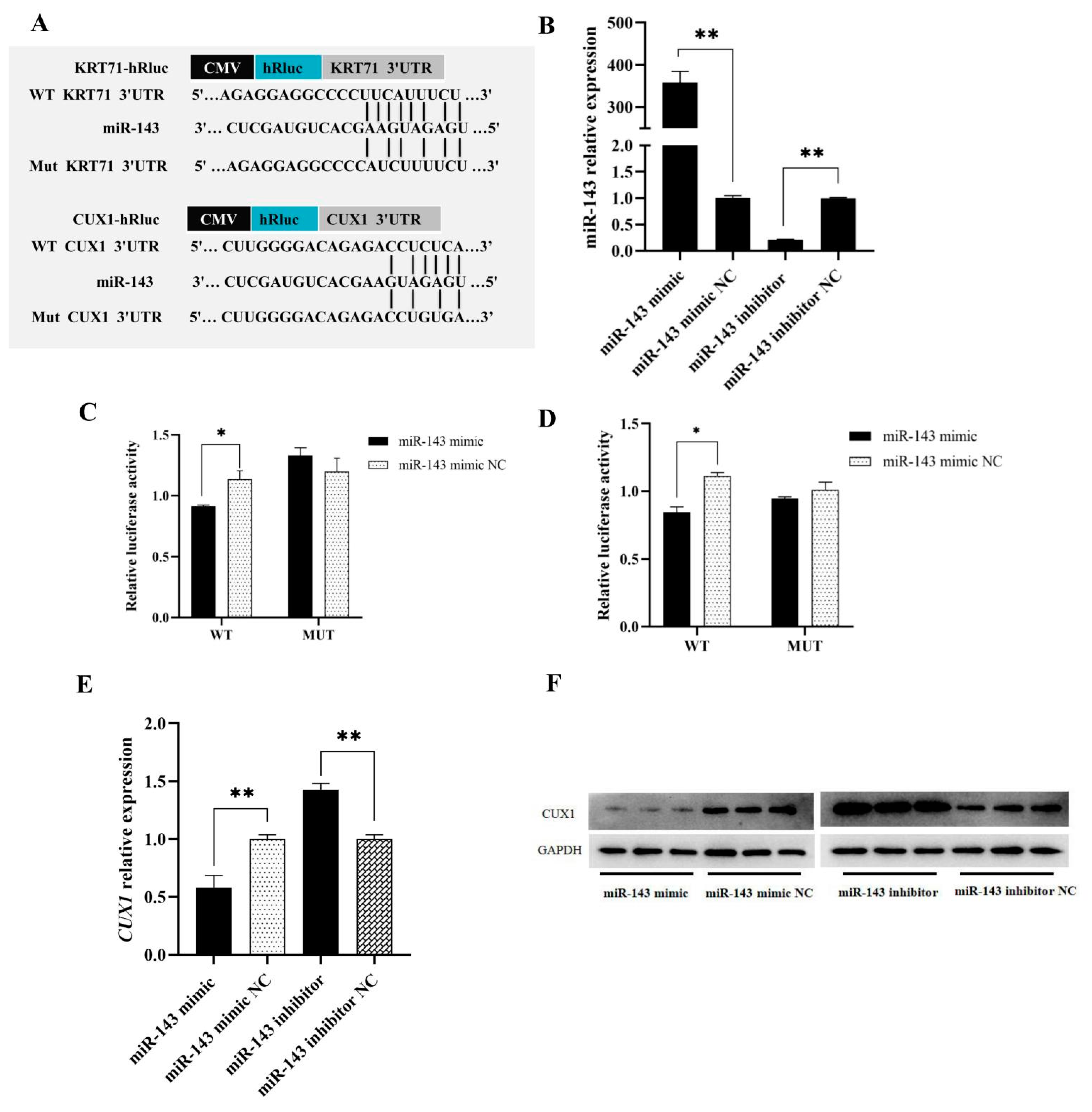
3.2. Target Genes Prediction of miR-143
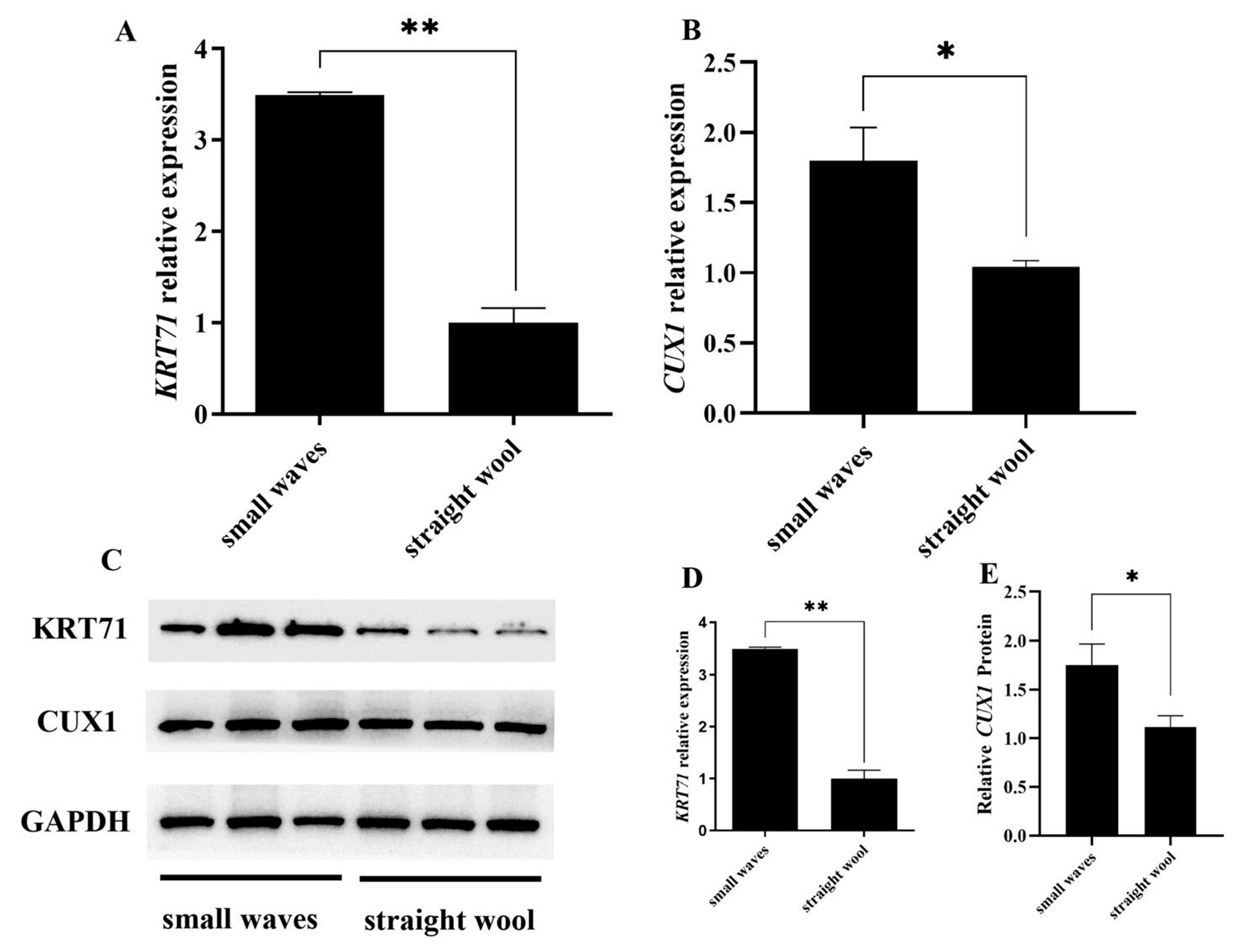
3.3. CUX1 and KRT71 Highly Expressed in Small Waves Group
3.4. CUX1 and KRT71 Targeting to miR-143
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Andl, T.; Reddy, S.T.; Gaddapara, T.; Millar, S.E. WNT signals are required for the initiation of hair follicle development. Dev. Cell. 2002, 2, 643–653. [Google Scholar] [CrossRef]
- Lyubimova, A.; Garber, J.J.; Upadhyay, G.; Sharov, A.; Anastasoaie, F.; Yajnik, V.; Cotsarelis, G.; Dotto, G.P.; Botchkarev, V.; Snapper, S.B. Neural Wiskott-Aldrich syndrome protein modulates Wnt signaling and is required for hair follicle cycling in mice. J. Clin. Investig. 2010, 120, 446–456. [Google Scholar] [CrossRef][Green Version]
- Kulessa, H.; Turk, G.; Hogan, B.L. Inhibition of Bmp signaling affects growth and differentiation in the anagen hair follicle. EMBO J. 2000, 19, 6664–6674. [Google Scholar] [CrossRef]
- Aubin-Houzelstein, G. Notch signaling and the developing hair follicle. Adv. Exp. Med. Biol. 2012, 727, 142–160. [Google Scholar] [CrossRef]
- Driskell, R.R.; Giangreco, A.; Jensen, K.B.; Mulder, K.W.; Watt, F.M. Sox2-positive dermal papilla cells specify hair follicle type in mammalian epidermis. Development 2009, 136, 2815–2823. [Google Scholar] [CrossRef]
- Chi, W.; Wu, E.; Morgan, B.A. Dermal papilla cell number specifies hair size, shape and cycling and its reduction causes follicular decline. Development 2013, 140, 1676–1683. [Google Scholar] [CrossRef]
- Preedy, V.R. Handbook of Hair in Health and Disease. Human Health Handbooks No. 1. 2012, Volume 1, pp. 332–351. Available online: https://books.google.co.uk/books?hl=zh-TW&lr=&id=7N3nEX6eL_MC&oi=fnd&pg=PA4&dq=Preedy,+Victor,+R.+Handb.+Hair+Health+Dis&ots=GcKMCrWxsm&sig=rariLE8rfXv9eQjS07LtDlwTTf8#v=onepage&q=Preedy%2C%20Victor%2C%20R.%20Handb.%20Hair%20Health%20Dis&f=false (accessed on 20 June 2021). [CrossRef]
- Cadieu, E.; Neff, M.W.; Quignon, P.; Walsh, K.; Chase, K.; Parker, H.G.; Vonholdt, B.M.; Rhue, A.; Boyko, A.; Byers, A.; et al. Coat variation in the domestic dog is governed by variants in three genes. Science 2009, 326, 150–153. [Google Scholar] [CrossRef]
- Kang, X.; Liu, Y.; Zhang, J.; Xu, Q.; Liu, C.; Fang, M. Characteristics and expression profile of KRT71 screened by suppression subtractive hybridization cDNA library in curly fleece chinese Tan sheep. DNA Cell Biol. 2017, 36, 552–564. [Google Scholar] [CrossRef]
- Tufarelli, C.; Fujiwara, Y.; Zappulla, D.C.; Neufeld, E.J. Hair defects and pup loss in mice with targeted deletion of the first cut repeat domain of the Cux/CDP homeoprotein gene. Dev. Biol. 1998, 200, 69–81. [Google Scholar] [CrossRef]
- Nissimov, J.N.; Das, C.A. Hair curvature: A natural dialectic and review. Biol. Rev. Camb. Philos. Soc. 2014, 89, 723–766. [Google Scholar] [CrossRef]
- Huntzinger, E.; Izaurralde, E. Gene silencing by microRNAs: Contributions of translational repression and mRNA decay. Nat. Rev. Genet. 2011, 12, 99–110. [Google Scholar] [CrossRef]
- Mardaryev, A.N.; Ahmed, M.I.; Vlahov, N.V.; Fessing, M.Y.; Gill, J.H.; Sharov, A.A.; Botchkareva, N.V. Micro-RNA-31 controls hair cycle-associated changes in gene expression programs of the skin and hair follicle. FASEB J. 2010, 24, 3869–3881. [Google Scholar] [CrossRef]
- Dong, C.; Wang, H.; Xue, L.; Dong, Y.; Yang, L.; Fan, R.; Yu, X.; Tian, X.; Ma, S.; Smith, G.W. Coat color determination by miR-137 mediated down-regulation of microphthalmia-associated transcription factor in a mouse model. RNA 2012, 18, 1679–1686. [Google Scholar] [CrossRef]
- Sonkoly, E.; Wei, T.; Janson, P.C.; Saaf, A.; Lundeberg, L.; Tengvall-Linder, M.; Norstedt, G.; Alenius, H.; Homey, B.; Scheynius, A.; et al. MicroRNAs: Novel regulators involved in the pathogenesis of psoriasis? PLoS ONE 2007, 2, e0000610. [Google Scholar] [CrossRef]
- Wang, D.; Zhang, Z.; O’Loughlin, E.; Wang, L.; Fan, X.; Lai, E.C.; Yi, R. MicroRNA-205 controls neonatal expansion of skin stem cells by modulating the PI(3)K pathway. Nat. Cell Biol. 2013, 15, 1153–1163. [Google Scholar] [CrossRef]
- Yuan, S.; Li, F.; Meng, Q.; Zhao, Y.; Chen, L.; Zhang, H.; Xue, L.; Zhang, X.; Lengner, C.; Yu, Z. Post-transcriptional regulation of keratinocyte progenitor cell expansion, differentiation and hair follicle regression by miR-22. PLoS Genet. 2015, 11, e1005253. [Google Scholar] [CrossRef]
- Zhang, L.; Stokes, N.; Polak, L.; Fuchs, E. Specific microRNAs are preferentially expressed by skin stem cells to balance self-renewal and early lineage commitment. Cell Stem Cell 2011, 8, 294–308. [Google Scholar] [CrossRef]
- Yi, R.; Poy, M.N.; Stoffel, M.; Fuchs, E. A skin microRNA promotes differentiation by repressing ‘stemness’. Nature 2008, 452, 225–229. [Google Scholar] [CrossRef]
- Wanbao Yang, Q.L.B.S. MicroRNA-148b Promotes Proliferation of Hair Follicle Cells by Targeting NFAT5. Frontier of Agricultural Science and Engineering (English Version); Higher Education Press: Wuhan, China, 2016. [Google Scholar] [CrossRef]
- Gao, W.; Sun, W.; Yin, J.; Lv, X.; Bao, J.; Yu, J.; Wang, L.; Jin, C.; Hu, L. Screening candidate microRNAs (miRNAs) in different lambskin hair follicles in Hu sheep. PLoS ONE 2017, 12, e0176532. [Google Scholar] [CrossRef]
- Lv, X.; Sun, W.; Yin, J.; Ni, R.; Su, R.; Wang, Q.; Gao, W.; Bao, J.; Yu, J.; Wang, L.; et al. An integrated analysis of microRNA and mRNA expression profiles to identify RNA expression signatures in lambskin hair follicles in Hu Sheep. PLoS ONE 2016, 11, e0157463. [Google Scholar] [CrossRef]
- Sun, W.; Ni, R.; Yin, J.F.; Musa, H.H.; Ding, T.; Chen, L. Genome array of hair follicle genes in lambskin with different patterns. PLoS ONE 2013, 8, e0068840. [Google Scholar] [CrossRef]
- Lv, X.; Gao, W.; Jin, C.; Wang, L.; Wang, Y.; Chen, W.; Zou, S.; Huang, S.; Li, Z.; Wang, J.; et al. Preliminary study on microR-148a and microR-10a in dermal papilla cells of Hu sheep. BMC Genet. 2019, 20, 70. [Google Scholar] [CrossRef]
- Rehmsmeier, M.; Steffen, P.; Hochsmann, M.; Giegerich, R. Fast and effective prediction of microRNA/target duplexes. RNA 2004, 10, 1507–1517. [Google Scholar] [CrossRef]
- Griffiths-Jones, S. miRBase: The microRNA sequence database. Methods Mol. Biol. 2006, 342, 129–138. [Google Scholar] [CrossRef]
- Huang, J.C.; Morris, Q.D.; Frey, B.J. Bayesian inference of microRNA targets from sequence and expression data. J. Comput. Biol. 2007, 14, 550–563. [Google Scholar] [CrossRef] [PubMed]
- Trick, A.Y.; Chen, F.E.; Schares, J.A.; Freml, B.E.; Lor, P.; Yun, Y.; Wang, T.H. High resolution estimates of relative gene abundance with quantitative ratiometric regression PCR (qRR-PCR). Analyst 2021, 146, 6463–6469. [Google Scholar] [CrossRef]
- Park, E.; Cho, M.; Ki, C.S. Correct use of repeated measures analysis of variance. Korean J. Lab. Med. 2009, 29, 1–9. [Google Scholar] [CrossRef]
- Cloete, E.; Khumalo, N.P.; Ngoepe, M.N. The what, why and how of curly hair: A review. Proc. Math. Phys. Eng. Sci. 2019, 475, 20190516. [Google Scholar] [CrossRef] [PubMed]
- Westgate, G.E.; Ginger, R.S.; Green, M.R. The biology and genetics of curly hair. Exp. Dermatol. 2017, 26, 483–490. [Google Scholar] [CrossRef]
- Mou, C.; Thomason, H.A.; Willan, P.M.; Clowes, C.; Harris, W.E.; Drew, C.F.; Dixon, J.; Dixon, M.J.; Headon, D.J. Enhanced ectodysplasin-A receptor (EDAR) signaling alters multiple fiber characteristics to produce the East Asian hair form. Hum. Mutat. 2008, 29, 1405–1411. [Google Scholar] [CrossRef]
- Wu, S.; Tan, J.; Yang, Y.; Peng, Q.; Zhang, M.; Li, J.; Lu, D.; Liu, Y.; Lou, H.; Feng, Q.; et al. Genome-wide scans reveal variants at EDAR predominantly affecting hair straightness in Han Chinese and Uyghur populations. Hum. Genet. 2016, 135, 1279–1286. [Google Scholar] [CrossRef]
- Pospiech, E.; Lee, S.D.; Kukla-Bartoszek, M.; Karlowska-Pik, J.; Wozniak, A.; Boron, M.; Zubanska, M.; Bronikowska, A.; Hong, S.R.; Lee, J.H.; et al. Variation in the RPTN gene may facilitate straight hair formation in Europeans and East Asians. J. Dermatol. Sci. 2018, 91, 331–334. [Google Scholar] [CrossRef]
- Demay, M.B. The hair cycle and Vitamin D receptor. Arch. Biochem. Biophys. 2012, 523, 19–21. [Google Scholar] [CrossRef]
- Liu, F.; Chen, Y.; Zhu, G.; Hysi, P.G.; Wu, S.; Adhikari, K.; Breslin, K.; Pospiech, E.; Hamer, M.A.; Peng, F.; et al. Meta-analysis of genome-wide association studies identifies 8 novel loci involved in shape variation of human head hair. Hum. Mol. Genet. 2018, 27, 559–575. [Google Scholar] [CrossRef]
- Rong, E.G.; Yang, H.; Zhang, Z.W.; Wang, Z.P.; Yan, X.H.; Li, H.; Wang, N. Association of methionine synthase gene polymorphisms with wool production and quality traits in Chinese Merino population. J. Anim. Sci. 2015, 93, 4601–4609. [Google Scholar] [CrossRef]
- Salmela, E.; Niskanen, J.; Arumilli, M.; Donner, J.; Lohi, H.; Hytonen, M.K. A novel KRT71 variant in curly-coated dogs. Anim. Genet. 2019, 50, 101–104. [Google Scholar] [CrossRef]
- Bauer, A.; Hadji, R.S.; Brunner, M.T.; Jagannathan, V.; Bucher, I.; Bannoehr, J.; Varjonen, K.; Bond, R.; Bergvall, K.; Welle, M.M.; et al. A second KRT71 allele in curly coated dogs. Anim. Genet. 2019, 50, 97–100. [Google Scholar] [CrossRef]
- Pierard-Franchimont, C.; Paquet, P.; Quatresooz, P.; Pierard, G.E. Mechanobiology and cell tensegrity: The root of ethnic hair curling? J. Cosmet. Dermatol. 2011, 10, 163–167. [Google Scholar] [CrossRef]
- Ji, K.; Zhang, P.; Zhang, J.; Fan, R.; Liu, Y.; Yang, S.; Hu, S.; Liu, X.; Dong, C. MicroRNA 143-5p regulates alpaca melanocyte migration, proliferation and melanogenesis. Exp. Dermatol. 2018, 27, 166–171. [Google Scholar] [CrossRef]
- Qi, S.; Liu, B.; Zhang, J.; Liu, X.; Dong, C.; Fan, R. Knockdown of microRNA1435p by STTM technology affects eumelanin and pheomelanin production in melanocytes. Mol. Med. Rep. 2019, 20, 2649–2656. [Google Scholar] [CrossRef]
- Han, W.; Yang, F.; Wu, Z.; Guo, F.; Zhang, J.; Hai, E.; Shang, F.; Su, R.; Wang, R.; Wang, Z.; et al. Inner mongolian cashmere goat secondary follicle development regulation research based on mRNA-miRNA co-analysis. Sci. Rep. 2020, 10, 4519. [Google Scholar] [CrossRef]
- Guoping, M.; Ran, L.; Yanru, Q. miR-143 inhibits cell proliferation of gastric cancer cells through targeting GATA6. Oncol. Res. 2018, 26, 1023–1029. [Google Scholar] [CrossRef]
- Sun, X.; Dai, G.; Yu, L.; Hu, Q.; Chen, J.; Guo, W. miR-143-3p inhibits the proliferation, migration and invasion in osteosarcoma by targeting FOSL2. Sci. Rep. 2018, 8, 606. [Google Scholar] [CrossRef]
- Cordes, K.R.; Sheehy, N.T.; White, M.P.; Berry, E.C.; Morton, S.U.; Muth, A.N.; Lee, T.H.; Miano, J.M.; Ivey, K.N.; Srivastava, D. miR-145 and miR-143 regulate smooth muscle cell fate and plasticity. Nature 2009, 460, 705–710. [Google Scholar] [CrossRef]
- Papanota, A.M.; Tsiakanikas, P.; Kontos, C.K.; Malandrakis, P.; Liacos, C.I.; Ntanasis-Stathopoulos, I.; Kanellias, N.; Gavriatopoulou, M.; Kastritis, E.; Avgeris, M.; et al. A molecular signature of circulating microRNA can predict osteolytic bone disease in multiple myeloma. Cancers 2021, 13, 3877. [Google Scholar] [CrossRef]
- Tewari, R.S.; Ala, U.; Accornero, P.; Baratta, M.; Miretti, S. Circulating skeletal muscle related microRNAs profile in Piedmontese cattle during different age. Sci. Rep. 2021, 11, 15815. [Google Scholar] [CrossRef]
- Rong, Z.W.; Soucie, E.; Sung, M.N.; Martin-Soudant, N.; Berube, G.; Leduy, L.; Nepveu, A. Exon/intron structure and alternative transcripts of the CUTL1 gene. Gene 2000, 241, 75–85. [Google Scholar] [CrossRef]
- Alcalay, N.I.; Vanden, H.G. Regulation of cell proliferation and differentiation in the kidney. Front. Biosci. (Landmark Ed.) 2009, 14, 4978–4991. [Google Scholar] [CrossRef]
- Luong, M.X.; van der Meijden, C.M.; Xing, D.; Hesselton, R.; Monuki, E.S.; Jones, S.N.; Lian, J.B.; Stein, J.L.; Stein, G.S.; Neufeld, E.J.; et al. Genetic ablation of the CDP/Cux protein C terminus results in hair cycle defects and reduced male fertility. Mol. Cell. Biol. 2002, 22, 1424–1437. [Google Scholar] [CrossRef]
- Ellis, T.; Gambardella, L.; Horcher, M.; Tschanz, S.; Capol, J.; Bertram, P.; Jochum, W.; Barrandon, Y.; Busslinger, M. The transcriptional repressor CDP (Cutl1) is essential for epithelial cell differentiation of the lung and the hair follicle. Genes Dev. 2001, 15, 2307–2319. [Google Scholar] [CrossRef]
- Fujimoto, A.; Farooq, M.; Fujikawa, H.; Inoue, A.; Ohyama, M.; Ehama, R.; Nakanishi, J.; Hagihara, M.; Iwabuchi, T.; Aoki, J.; et al. A missense mutation within the helix initiation motif of the keratin K71 gene underlies autosomal dominant woolly hair/hypotrichosis. J. Investig. Dermatol. 2012, 132, 2342–2349. [Google Scholar] [CrossRef]
- Harel, S.; Christiano, A.M. Keratin 71 mutations: From water dogs to woolly hair. J. Investig. Dermatol. 2012, 132, 2315–2317. [Google Scholar] [CrossRef]
| Gene/miRNA Name | Forward Primer (5′-3′) | Reverse Primer (5′-3′) | Product Length (bp) |
|---|---|---|---|
| CUX1 | GCACGACATTGAGACGGAG | AGCTATGGTCTCAGCCTGGT | 160 |
| KRT71 | GGCTCATCCAGAGAATCCGC | GAGCATTGTCACCCCTCTGT | 102 |
| GAPDH | TCTCAAGGGCATTCTAGGCTAC | GCCGAATTCATTGTCGTACCAG | 151 |
| CDK2 | AGAAGTGGCTGCATCACAAG | TCTCAGAATCTCCAGGGAATAG | 92 |
| PCNA | CGAGGGCTTCGACACTTAC | GTCTTCATTGCCAGCACATT | 97 |
| Cyclin d1 | CCGAGGAGAACAAGCAGATC | GAGGGTGGGTTGGAAATG | 91 |
| miR-143 | CGCGTGAGATGAAGCACTG | AGTGCAGGGTCCGAGGTATT | variable |
| U6 | CTCGCTTCGGCAGCACA | AACGCTTCACGAATTTGCGT | 95 |
| Gene Name | Primer | Forward Primer (5′-3′) |
|---|---|---|
| CUX1 | CUX1-WT-F | CCctcgagCCATCTCCCTTTAAAGAGGG |
| CUX1-WI-R | TTgcggccgcGACTTAGAGCTCGCTGTCC | |
| KRT71 | KRT71-WT-F | CCctcgagTTGTCCCAGCTCCTGCTT |
| KRT71-WT-R | TTgcggccgcTATTGAGGATGCAG | |
| CUX1 | CUX1-MUT1-F | CTTGGGGACAGAGACCTGTGAGCTGTGACACTG |
| CUX1-MUT1-R | CAGTGTCACAGCTCACAGGTCTCTGTCCCCAAG | |
| KRT71 | KRT71-MUT1-F | GGGAAGAGGAGGCCCCATCTTTTCTCCTTTC |
| KRT71-MUT1-R | GAAAGGAGAAAAGATGGGGCCTCCTCTTCCC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hu, T.; Huang, S.; Lv, X.; Wang, S.; Getachew, T.; Mwacharo, J.M.; Haile, A.; Sun, W. miR-143 Targeting CUX1 to Regulate Proliferation of Dermal Papilla Cells in Hu Sheep. Genes 2021, 12, 2017. https://doi.org/10.3390/genes12122017
Hu T, Huang S, Lv X, Wang S, Getachew T, Mwacharo JM, Haile A, Sun W. miR-143 Targeting CUX1 to Regulate Proliferation of Dermal Papilla Cells in Hu Sheep. Genes. 2021; 12(12):2017. https://doi.org/10.3390/genes12122017
Chicago/Turabian StyleHu, Tingyan, Sainan Huang, Xiaoyang Lv, Shanhe Wang, Tesfaye Getachew, Joram M. Mwacharo, Aynalem Haile, and Wei Sun. 2021. "miR-143 Targeting CUX1 to Regulate Proliferation of Dermal Papilla Cells in Hu Sheep" Genes 12, no. 12: 2017. https://doi.org/10.3390/genes12122017
APA StyleHu, T., Huang, S., Lv, X., Wang, S., Getachew, T., Mwacharo, J. M., Haile, A., & Sun, W. (2021). miR-143 Targeting CUX1 to Regulate Proliferation of Dermal Papilla Cells in Hu Sheep. Genes, 12(12), 2017. https://doi.org/10.3390/genes12122017





