Risk of Recurrent Pregnancy Loss in the Ukrainian Population Using a Combined Effect of Genetic Variants: A Case-Control Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Sample
2.2. Blood Sample Collection, DNA Extraction and Genotyping Procedures
2.3. Association Analysis
2.4. Genetic Risk Score Calculation
2.5. Receiver Operating Characteristic Analysis
3. Results
3.1. Association Analysis
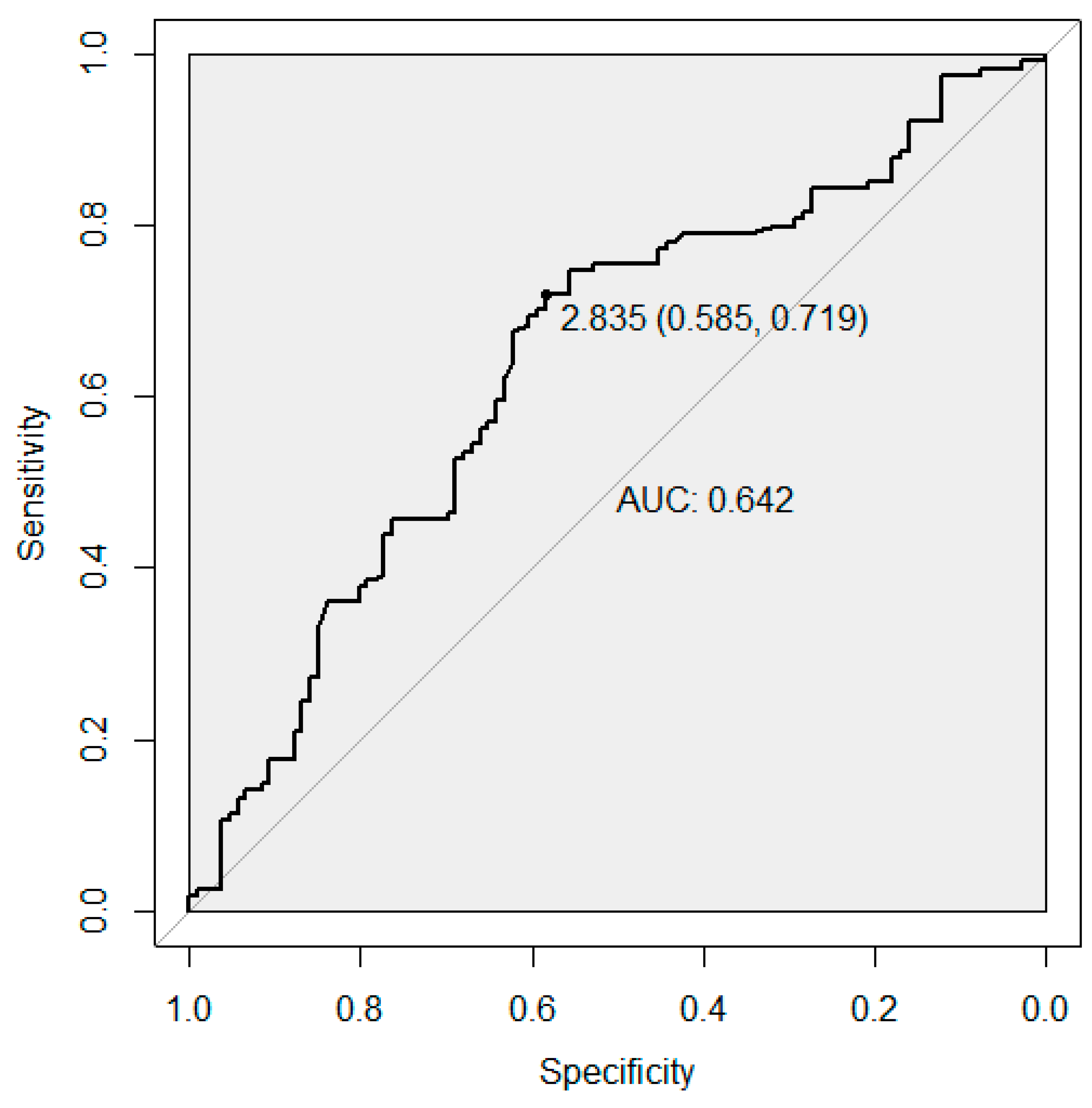
3.2. Receiver Operator Characteristic Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zegers-Hochschild, F.; Adamson, G.; De Mouzon, J.; Ishihara, O.; Mansour, R.; Nygren, K.; Sullivan, E.; Vanderpoel, S. International Committee for Monitoring Assisted Reproductive Technology (ICMART) and the World Health Organization (WHO) revised glossary of ART terminology, 2009. Fertil. Steril. 2009, 92, 1520–1524. [Google Scholar] [CrossRef]
- Branch, D.W.; Gibson, M.; Silver, R.M. Recurrent Miscarriage. N. Engl. J. Med. 2010, 363, 1740–1747. [Google Scholar] [CrossRef]
- Sugiura-Ogasawara, M.; Ozaki, Y.; Katano, K.; Suzumori, N.; Kitaori, T.; Mizutani, E. Abnormal embryonic karyotype is the most frequent cause of recurrent miscarriage. Hum. Reprod. 2012, 27, 2297–2303. [Google Scholar] [CrossRef]
- Werner, M.; Reh, A.; Grifo, J.; Perle, M.A. Characteristics of chromosomal abnormalities diagnosed after spontaneous abortions in an infertile population. J. Assist. Reprod. Genet. 2012, 29, 817–820. [Google Scholar] [CrossRef][Green Version]
- Rull, K.; Nagirnaja, L.; Laan, M. Genetics of Recurrent Miscarriage: Challenges, Current Knowledge, Future Directions. Front. Genet. 2012, 3, 34. [Google Scholar] [CrossRef]
- Li, T.C.; Makris, M.; Tomsu, M.; Tuckerman, E.; Laird, S. Recurrent miscarriage: Aetiology, management and prognosis. Hum. Reprod. Update 2002, 8, 463–481. [Google Scholar] [CrossRef]
- McQueen, D.; Perfetto, C.O.; Hazard, F.K.; Lathi, R.B. Pregnancy outcomes in women with chronic endometritis and recurrent pregnancy loss. Fertil. Steril. 2015, 104, 927–931. [Google Scholar] [CrossRef]
- Zargar, M.; Ghafourian, M.; Nikbakht, R.; Hosseini, V.M.; Choghakabodi, P.M. Evaluating Chronic Endometritis in Women with Recurrent Implantation Failure and Recurrent Pregnancy Loss by Hysteroscopy and Immunohistochemistry. J. Minim. Invasive Gynecol. 2019, 27, 116–121. [Google Scholar] [CrossRef]
- Rehman, F.; Shah, M.; Ali, A.; Rapisarda, A.; Cianci, A. Seroprevalence and risk factors of Toxoplasma gondii infection in women with recurrent fetal loss from the province of Khyber Pakhtunkhwa, Pakistan. J. Neonatal-Perinatal Med. 2020, 1–7. [Google Scholar] [CrossRef]
- ElSalam, S.A.; Mansor, A.E.; Sarhan, M.H.; Shalaby, A.M.; Gobran, M.A.; Alabiad, M.A. Evaluation of Apoptosis, Proliferation, and Adhesion Molecule Expression in Trophoblastic Tissue of Women with Recurrent Spontaneous Abortion and Infected With Toxoplasma gondii. Int. J. Gynecol. Pathol. 2020. [Google Scholar] [CrossRef]
- Christiansen, O.B.; Andersen, A.-M.N.; Bosch, E.; Daya, S.; Delves, P.J.; Hviid, T.V.; Kutteh, W.H.; Laird, S.M.; Li, T.-C.; Van Der Ven, K. Evidence-based investigations and treatments of recurrent pregnancy loss. Fertil. Steril. 2005, 83, 821–839. [Google Scholar] [CrossRef]
- Kolte, A.M.; Nielsen, H.S.; Moltke, I.; Degn, B.; Pedersen, B.; Sunde, L.; Nielsen, F.C.; Christiansen, O.B. A genome-wide scan in affected sibling pairs with idiopathic recurrent miscarriage suggests genetic linkage. Mol. Hum. Reprod. 2011, 17, 379–385. [Google Scholar] [CrossRef][Green Version]
- Klarin, D.; Emdin, C.A.; Natarajan, P.; Conrad, M.F.; Kathiresan, S.; INVENT Consortium. Genetic Analysis of Venous Thromboembolism in UK Biobank Identifies the ZFPM2 Locus and Implicates Obesity as a Causal Risk Factor Derek. Circ. Cardiovasc. Genet. 2017, 10, e001643. [Google Scholar] [CrossRef]
- Germain, M.; Chasman, D.I.; De Haan, H.; Tang, W.; Lindström, S.; Weng, L.-C.; De Andrade, M.; De Visser, M.C.; Wiggins, K.L.; Suchon, P.; et al. Meta-analysis of 65,734 Individuals Identifies TSPAN15 and SLC44A2 as Two Susceptibility Loci for Venous Thromboembolism. Am. J. Hum. Genet. 2015, 96, 532–542. [Google Scholar] [CrossRef]
- Hinds, D.A.; Buil, A.; Ziemek, D.; Martinez-Perez, A.; Malik, R.; Folkersen, L.; Germain, M.; Mälarstig, A.; Brown, A.; Soria, J.M.; et al. Genome-wide association analysis of self-reported events in 6135 individuals and 252 827 controls identifies 8 loci associated with thrombosis. Hum. Mol. Genet. 2016, 25, 1867–1874. [Google Scholar] [CrossRef]
- Heit, J.A.; Armasu, S.M.; Asmann, Y.W.; Cunningham, J.M.; Matsumoto, M.E.; Petterson, T.M.; De Andrade, M. A genome-wide association study of venous thromboembolism identifies risk variants in chromosomes 1q24.2 and 9q. J. Thromb. Haemost. 2013, 10, 1521–1531. [Google Scholar] [CrossRef]
- Rai, R.; Regan, L. Recurrent miscarriage. Lancet 2006, 368, 601–611. [Google Scholar] [CrossRef]
- Kovalevsky, G.; Gracia, C.R.; Berlin, J.A.; Sammel, M.D.; Barnhart, K.T. Evaluation of the Association Between Hereditary Thrombophilias and Recurrent Pregnancy Loss. Arch. Intern. Med. 2004, 164, 558–563. [Google Scholar] [CrossRef]
- Barlik, M.; Seremak-Mrozikiewicz, A.; Drews, K.; Klejewski, A.; Kurzawińska, G.; Łowicki, Z.; Wolski, H. Correlation between factor VII and PAI-1 genetic variants and recurrent miscarriage. Ginekol. Polska 2016, 87, 504–509. [Google Scholar] [CrossRef]
- Giusti, B.; Gori, A.M.; Marcucci, R.; Sestini, I.; Saracini, C.; Paniccia, R.; Poli, S.; Giglioli, C.; Valente, S.; Prisco, D.; et al. Role of glycoprotein Ia gene polymorphisms in determining platelet function in myocardial infarction patients undergoing percutaneous coronary intervention on dual antiplatelet treatment. Atherosclerosis 2008, 196, 341–348. [Google Scholar] [CrossRef]
- Pepe, G.J.; Albrecht, E.D. Actions of Placental and Fetal Adrenal Steroid Hormones in Primate Pregnancy. Endocr. Rev. 1995, 16, 608–648. [Google Scholar] [CrossRef]
- Sapkota, Y.; iPSYCH-SSI-Broad Group; Steinthorsdottir, V.; Morris, A.P.; Fassbender, A.; Rahmioglu, N.; De Vivo, I.; Buring, J.E.; Zhang, F.; Edwards, T.L.; et al. Meta-analysis identifies five novel loci associated with endometriosis highlighting key genes involved in hormone metabolism. Nat. Commun. 2017, 8, 15539. [Google Scholar] [CrossRef]
- Barban, N.; BIOS Consortium; Jansen, R.; De Vlaming, R.; Vaez, A.; Mandemakers, J.J.; Tropf, F.C.; Shen, X.; Wilson, J.F.; Chasman, D.I.; et al. Genome-wide analysis identifies 12 loci influencing human reproductive behavior. Nat. Genet. 2016, 48, 1462–1472. [Google Scholar] [CrossRef]
- Chen, Q.; Zhang, Y.; Peng, H.; Lei, L.; Kuang, H.; Zhang, L.; Ning, L.; Cao, Y.; Duan, E. Transient β2-Adrenoceptor Activation Confers Pregnancy Loss by Disrupting Embryo Spacing at Implantation. J. Biol. Chem. 2010, 286, 4349–4356. [Google Scholar] [CrossRef]
- Suh, Y.J.; Park, H.J.; Lee, K.A.; Lee, B.E.; Ha, E.H.; Kim, Y.J. Associations Between Genetic Polymorphisms of Beta-2 Adrenergic Receptor and Preterm Delivery in Korean Women. Am. J. Reprod. Immunol. 2013, 69, 85–91. [Google Scholar] [CrossRef]
- Peterlin, A.; Maver, A.; Jan, Z.; Lovrecic, L.; Tul, N.; Peterlin, B. Polymorphism of the ADRB2 rs1042713 gene is not associated with spontaneous preterm birth: Analyses in a Slovenian sample and meta analysis. Balk. J. Med Genet. 2017, 20, 35–41. [Google Scholar] [CrossRef]
- Ayuk, P.Y.; Theophanous, D.; D’Souza, S.W.; Sibley, C.P.; Glazier, J.D. L-Arginine Transport by the Microvillous Plasma Membrane of the Syncytiotrophoblast from Human Placenta in Relation to Nitric Oxide Production: Effects of Gestation, Preeclampsia, and Intrauterine Growth Restriction. J. Clin. Endocrinol. Metab. 2002, 87, 747–751. [Google Scholar] [CrossRef][Green Version]
- Yang, C.; FangFang, W.; Jie, L.; Yanlong, Y.; Jie, W.; Xuefei, L.; Xuerong, Z.; Hu, Y. Angiotensin-converting enzyme insertion/deletion (I/D) polymorphisms and recurrent pregnancy loss: A meta-analysis. J. Assist. Reprod. Genet. 2012, 29, 1167–1173. [Google Scholar] [CrossRef]
- Deng, C.; Tang, S.; Huang, X.; Gao, J.; Tian, J.; Zhou, X.; Xie, Y.; Liao, M.; Mo, Z.; Wang, Q. Identification of three novel loci of ALDH2 Gene for Serum Folate levels in a Male Chinese Population by Genome-Wide Association Study. Gene 2018, 674, 121–126. [Google Scholar] [CrossRef]
- Paré, G.; Chasman, D.I.; Parker, A.N.; Zee, R.R.; Mälarstig, A.; Seedorf, U.; Collins, R.; Watkins, H.; Hamsten, A.; Miletich, J.P.; et al. Novel Associations of CPS1, MUT, NOX4 and DPEP1 with Plasma Homocysteine in a Healthy Population: A Genome Wide Evaluation of 13,974 Participants in the Women’s Genome Health Study Guillaumé. Circ. Cardiovasc. Genet. 2009, 2, 142–150. [Google Scholar] [CrossRef]
- Van Meurs, J.B.J.; Pare, G.; Schwartz, S.M.; Hazra, A.; Tanaka, T.; Vermeulen, S.H.; Cotlarciuc, I.; Yuan, X.; Mälarstig, A.; Bandinelli, S.; et al. Common genetic loci influencing plasma homocysteine concentrations and their effect on risk of coronary artery disease. Am. J. Clin. Nutr. 2013, 98, 668–676. [Google Scholar] [CrossRef]
- Obwegeser, R.; Hohlagschwandtner, M.; Sinzinger, H. Homocysteine—A pathophysiological cornerstone in obstetrical and gynaecological disorders? Hum. Reprod. Update 1999, 5, 64–72. [Google Scholar] [CrossRef] [PubMed]
- Steegers-Theunissen, R.; Boers, G.; Blom, H.; Trijbels, F.; Eskes, T. Hyperhomocysteinaemia and recurrent spontaneous abortion or abruptio placentae. Lancet 1992, 339, 1122–1123. [Google Scholar] [CrossRef]
- Wouters, M.G.; Boers, G.H.; Blom, H.J.; Trijbels, F.J.; Thomas, C.M.; Borm, G.F.; Steegers-Theunissen, R.P.; Eskes, T.K. Hyperhomocysteinemia: A risk factor in women with unexplained recurrent early pregnancy loss. Fertil. Steril. 1993, 60, 820–825. [Google Scholar] [CrossRef]
- Quéré, I.; Wutschert, R.; Zittoun, J.; Bellet, H.; Reber, G.; Gris, J.C.; Janbon, C.; Schved, J.F.; De Moerloose, P. Association of red-blood methylfolate but not plasma folate with C677T MTHFR polymorphism in venous thromboembolic disease. Thromb. Haemost. 1998, 80, 707–709. [Google Scholar]
- Coumans, A.; Huijgens, P.; Jakobs, C.; Schats, R.; De Vries, J.; Van Pampus, M.; Dekker, G. Haemostatic and metabolic abnormalities in women with unexplained recurrent abortion. Hum. Reprod. 1999, 14, 211–214. [Google Scholar] [CrossRef]
- Nelen, W.L.; Steegers, E.A.; Eskes, T.K.; Blom, H.J. Genetic risk factor for unexplained recurrent early pregnancy loss Fetuin protects the fetus from TNF Differences between RhD-negative Africans and RhD-negative Europeans. Lancet 1997, 350, 861–862. [Google Scholar] [CrossRef]
- Cao, Y.; Xu, J.; Zhang, Z.; Huang, X.; Zhang, A.; Wang, J.; Zheng, Q.; Fu, L.; Du, J. Association study between methylenetetrahydrofolate reductase polymorphisms and unexplained recurrent pregnancy loss: A meta-analysis. Gene 2013, 514, 105–111. [Google Scholar] [CrossRef]
- Kucherenko, A.; Gulkovskyi, R.; Khazhylenko, K.; Vorobiova, I.; Nakvasiuk, T.; Livshits, L. Recurrent pregnancy loss association with allelic variants of IL8 and IL10 genes. ScienceRise 2014, 2, 7–10. [Google Scholar] [CrossRef][Green Version]
- Cochery-Nouvellon, E.; Nguyen, P.; Attaoua, R.; Cornillet-Lefebvre, P.; Mercier, E.; Vitry, F.; Gris, J.-C. Interleukin 10 Gene Promoter Polymorphisms in Women with Pregnancy Loss: Preferential Association with Embryonic Wastage1. Biol. Reprod. 2009, 80, 1115–1120. [Google Scholar] [CrossRef][Green Version]
- Daher, S.; Shulzhenko, N.; Morgun, A.; Mattar, R.; Rampim, G.F.; Camano, L.; DeLima, M.G. Associations between cytokine gene polymorphisms and recurrent pregnancy loss. J. Reprod. Immunol. 2003, 58, 69–77. [Google Scholar] [CrossRef]
- Lee, Y.; Choi, S.J.; Ji, J.D. Association between IL-6 −174 G/C, IL-6 −634 G/C, and IFN-γ +874 A/T polymorphisms and susceptibility to recurrent pregnancy loss: A meta-analysis. J. Assist. Reprod. Genet. 2015, 32, 1421–1427. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Weimar, C.H.; Macklon, N.; Uiterweer, E.D.P.; Brosens, J.J.; Gellersen, B. The motile and invasive capacity of human endometrial stromal cells: Implications for normal and impaired reproductive function. Hum. Reprod. Update 2013, 19, 542–557. [Google Scholar] [CrossRef] [PubMed]
- The Practice Committee of the American Society for Reproductive Medicine. Evaluation and treatment of recurrent pregnancy loss: A committee opinion. Fertil. Steril. 2012, 98, 1103–1111. [Google Scholar] [CrossRef] [PubMed]
- Koksal, V.; Barıs, I.; Etlik, O.; Baris, I. Primer-engineered multiplex PCR–RFLP for detection of MTHFR C677T, prothrombin G20210A and factor V Leiden mutations. Exp. Mol. Pathol. 2007, 83, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Saha, N.; Liu, Y.; Heng, C.-K.; Hong, S.; Low, P.S.; Tay, J.S. Association of factor VII genotype with plasma factor VII activity and antigen levels in healthy Indian adults and interaction with triglycerides. Arter. Thromb. A J. Vasc. Biol. 1994, 14, 1923–1927. [Google Scholar] [CrossRef] [PubMed]
- Fechter, A.; Gardemann, A.; Kroll, H. The role of the glycoprotein IIb fibrinogen receptor subunit T2622G gene polymorphism (HPA-3) on coronary artery disease and acute myocardial infarction. Thromb. Haemost. 2001, 85, 182–183. [Google Scholar] [CrossRef]
- Su, M.-T.; Lin, S.-H.; Chen, Y.-C. Association of sex hormone receptor gene polymorphisms with recurrent pregnancy loss: A systematic review and meta-analysis. Fertil. Steril. 2011, 96, 1435–1444.e1. [Google Scholar] [CrossRef] [PubMed]
- Livshyts, G.B.; Kucherenko, A.M.; Podlesna, S.S.; Kravchenko, S.A.; Livshits, L. Analysis of allelic polymorphism in the ESR1 gene in the Ukraine’s population. Cytol. Genet. 2012, 46, 220–226. [Google Scholar] [CrossRef][Green Version]
- Littlejohn, M.D.; Taylor, D.R.; Miller, A.; Kennedy, M.A. Determination of ?2-adrenergic receptor (ADRB2) haplotypes by a multiplexed polymerase chain reaction assay. Hum. Mutat. 2002, 20, 479. [Google Scholar] [CrossRef] [PubMed]
- Su, M.-T.; Lin, S.-H.; Chen, Y.-C. Genetic association studies of angiogenesis- and vasoconstriction-related genes in women with recurrent pregnancy loss: A systematic review and meta-analysis. Hum. Reprod. Update 2011, 17, 803–812. [Google Scholar] [CrossRef] [PubMed]
- Safarinejad, M.R.; Khoshdel, A.; Shekarchi, B.; Taghva, A.; Safarinejad, S. Association of the T-786C, G894T and 4a/4b polymorphisms of the endothelial nitric oxide synthase gene with vasculogenic erectile dysfunction in Iranian subjects. BJU Int. 2010, 107, 1994–2001. [Google Scholar] [CrossRef] [PubMed]
- Lindpaintner, K.; Pfeffer, M.A.; Kreutz, R.; Stampfer, M.J.; Grodstein, F.; LaMotte, F.; Buring, J.; Hennekens, C.H. A Prospective Evaluation of an Angiotensin-Converting–Enzyme Gene Polymorphism and the Risk of Ischemic Heart Disease. N. Engl. J. Med. 1995, 332, 706–712. [Google Scholar] [CrossRef] [PubMed]
- Shazia, M.; Raheel, Q.; Farah, A.; Wajid, A.K.; Asifa, A. C677T polymorphism in the methylenetetrahydrofolate reductase gene is associated with primary closed angle glaucoma. Mol. Vis. 2008, 14, 661–665. [Google Scholar]
- Fishman, D.; Faulds, G.; Jeffery, R.; Mohamed-Ali, V.; Yudkin, J.S.; Humphries, S.; Woo, P. The effect of novel polymorphisms in the interleukin-6 (IL-6) gene on IL-6 transcription and plasma IL-6 levels, and an association with systemic-onset juvenile chronic arthritis. J. Clin. Investig. 1998, 102, 1369–1376. [Google Scholar] [CrossRef]
- Heinzmann, A.; Ahlert, I.; Kurz, T.; Berner, R.; Deichmann, K.A. Association study suggests opposite effects of polymorphisms within IL8 on bronchial asthma and respiratory syncytial virus bronchiolitis. J. Allergy Clin. Immunol. 2004, 114, 671–676. [Google Scholar] [CrossRef]
- Costa, G.C.; Rocha, M.O.D.C.; Moreira, P.R.; Menezes, C.A.S.; Silva, M.R.; Gollob, K.J.; Dutra, W.O. Functional IL-10 Gene Polymorphism Is Associated with Chagas Disease Cardiomyopathy. J. Infect. Dis. 2009, 199, 451–454. [Google Scholar] [CrossRef]
- Temple, S.E.L.; Lim, E.; Cheong, K.; Almeida, C.-A.M.; Proce, P.; Ardlie, K.G.; Waterer, G.W. Alleles carried at positions -819 and -592 of the IL10 promoter affect transcription following stimulation of peripheral blood cells with Streptococcus pneumoniae. Immunogenetics 2003, 55, 629–632. [Google Scholar] [CrossRef]
- Belsky, D.W.; Moffitt, T.E.; Sugden, K.; Williams, B.; Houts, R.; McCarthy, J.J.; Caspi, A. Development and Evaluation of a Genetic Risk Score for Obesity. Biodemography Soc. Biol. 2013, 59, 85–100. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018; Available online: https://www.R-project.org/ (accessed on 10 November 2020).
- GWAS Results of the UK Biobank Data. Available online: http://www.nealelab.is/uk-biobank (accessed on 9 November 2018).
- Kirshenbaum, M.; Orvieto, R. Should We Offer In Vitro Fertilization to Couples with Unexplained Recurrent Pregnancy Loss? J. Clin. Med. 2019, 8, 2001. [Google Scholar] [CrossRef]
- Khera, A.V.; Chaffin, M.; Aragam, K.G.; Haas, M.E.; Roselli, C.; Choi, S.H.; Natarajan, P.; Lander, E.S.; Lubitz, S.; Ellinor, P.T.; et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 2018, 50, 1219–1224. [Google Scholar] [CrossRef] [PubMed]
- Christiansen, O.B.; Steffensen, R.; Nielsen, H.S.; Varming, K. Multifactorial Etiology of Recurrent Miscarriage and Its Scientific and Clinical Implications. Gynecol. Obstet. Investig. 2008, 66, 257–267. [Google Scholar] [CrossRef] [PubMed]
- Prokopenko, I.; McCarthy, M.I.; Lindgren, C.M. Type 2 diabetes: New genes, new understanding. Trends Genet. 2008, 24, 613–621. [Google Scholar] [CrossRef] [PubMed]
| Locus Name | SNP rsID | EA/NEA | EAF (1000 G *) | OR (95% CI) | p-Value | Case/Control Sample Size | Outcome | Reference | Genotyping Method | Reference |
|---|---|---|---|---|---|---|---|---|---|---|
| Blood coagulation | ||||||||||
| F2 | rs1799963 | A/G | 0.008 | 2.00 (1.00, 4.00) | <0.03 | 342/123 | RPL | [18] | RFLP | [45] |
| F5 | rs6025 $ | T/C | 0.012 | 2.5 (1.80, 3.40) | <10−3 | 342/123 | RPL | [18] | RFLP | [45] |
| F7 | rs6046 | G/A | 0.89 | - | - | - | - | - | RFLP | [46] |
| GP1A | rs1126643 | T/C | 0.40 | - | - | - | - | - | RFLP | [47] |
| Hormonal regulation | ||||||||||
| ESR1 | rs2234693 ^ | T/C | 0.58 | 1.10 (0.57, 2.13) | >0.05 | 350/646 | RPL | [48] | RFLP | [49] |
| ADRB2 | rs1042714 | G/C | 0.41 | - | - | - | - | - | RFLP | [50] |
| Endometrium and placental function | ||||||||||
| ENOS | Intron-4 ǂ VNTR ^ | B/A | - | 1.005 (0.74, 1.37) | >0.05 | 410/357 | RPL | [51] | Allele-specific PCR | [52] |
| ACE | rs1799752 †,$ | D/I | - | 2.06 (1.46, 2.91) | NA | 740/329 | RPL | [28] | Allele-specific PCR | [53] |
| Folate metabolism | ||||||||||
| MTHFR | rs1801133 $ | T/C | 0.36 | 1.25 (0.93, 1.67) | 0.14 | 1830/3037 | RPL | [38] | RFLP | [54] |
| Inflammatory response | ||||||||||
| IL6 | 14718 $ | G/C | - | 1.214 (0.88, 1.67) | 0.24 | 230/188 | RPL | [42] | RFLP | [55] |
| IL8 | rs2227306 | C/T | 0.61 | - | - | - | - | - | RFLP | [56] |
| IL10 | rs1800896 ^ | G/A | 0.45 | 1.27 (0.95, 1.70) | >0.05 | 635/571 | RPL | [41] | RFLP | [57] |
| IL10 | rs1800872 | T/G | 0.76 | 3.01 (1.92, 4.72) | <10−4 | 342/123 | Early PL | [40] | RFLP | [58] |
| Locus Name | Chr: Position | Variant ID | EA/NEA | EAF Cases/Controls | N Cases/N Controls | EAE | OR (95% CI) | p-Value |
|---|---|---|---|---|---|---|---|---|
| F2 | 11:46761055 | rs1799963 | A/G | 0.018/0.011 | 110/46 | 0.49 | 1.64 (0.23, 32.48) | 0.66 |
| F5 | 1:169519049 | rs6025 | A/G | 0.018/0.022 | 114/46 | −0.22 | 0.80 (0.15, 5.92) | 0.80 |
| F7 | 13:113773159 | rs6046 | G/A | 0.89/0.85 | 75/46 | 0.34 | 1.40 (0.65, 3.04) | 0.38 |
| GP1A | 5:52347369 | rs1126643 | C/T | 0.63/0.54 | 81/46 | 0.35 | 1.43 (0.85, 2.44) | 0.18 |
| ESR1 | 6:152163335 | rs2234693 | T/C | 0.49/0.55 | 110/106 | −0.18 | 0.84 (0.57, 1.23) | 0.37 |
| ADRB2 | 5:148206473 | rs1042714 | C/G | 0.60/0.51 | 81/46 | 0.38 | 1.47 (0.86, 2.56) | 0.16 |
| ENOS | 15:35147732-35262040 | Intron-4 ǂ VNTR | B/A | 0.83/0.80 | 102/46 | 0.15 | 1.16 (0.63, 2.08) | 0.63 |
| ACE | 17:61565890 | rs1799752 † | D/I | 0.52/0.53 | 100/46 | −0.048 | 0.95 (0.59, 1.54) | 0.85 |
| MTHFR | 1:11856378 | rs1801133 | T/C | 0.30/0.24 | 114/46 | 0.33 | 1.40 (0.80, 2.51) | 0.25 |
| IL6 | 7:22766840 | 14718 | G/C | 0.59/0.56 | 106/106 | 0.11 | 1.10 (0.77, 1.64) | 0.57 |
| IL8 | 4:74607055 | rs2227306 | T/C | 0.40/0.37 | 114/106 | 0.11 | 1.10 (0.76, 1.64) | 0.60 |
| IL10 | 1:206946897 | rs1800896 | A/G | 0.59/0.49 | 114/106 | 0.47 | 1.60 (1.07, 2.42) | 0.025 |
| GRS | 114/106 | 1.56 (1.21, 2.04) | 8.7 × 10−4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Loizidou, E.M.; Kucherenko, A.; Tatarskyy, P.; Chernushyn, S.; Livshyts, G.; Gulkovskyi, R.; Vorobiova, I.; Antipkin, Y.; Gorodna, O.; Kaakinen, M.A.; et al. Risk of Recurrent Pregnancy Loss in the Ukrainian Population Using a Combined Effect of Genetic Variants: A Case-Control Study. Genes 2021, 12, 64. https://doi.org/10.3390/genes12010064
Loizidou EM, Kucherenko A, Tatarskyy P, Chernushyn S, Livshyts G, Gulkovskyi R, Vorobiova I, Antipkin Y, Gorodna O, Kaakinen MA, et al. Risk of Recurrent Pregnancy Loss in the Ukrainian Population Using a Combined Effect of Genetic Variants: A Case-Control Study. Genes. 2021; 12(1):64. https://doi.org/10.3390/genes12010064
Chicago/Turabian StyleLoizidou, Eleni M., Anastasia Kucherenko, Pavlo Tatarskyy, Sergey Chernushyn, Ganna Livshyts, Roman Gulkovskyi, Iryna Vorobiova, Yurii Antipkin, Oleksandra Gorodna, Marika A. Kaakinen, and et al. 2021. "Risk of Recurrent Pregnancy Loss in the Ukrainian Population Using a Combined Effect of Genetic Variants: A Case-Control Study" Genes 12, no. 1: 64. https://doi.org/10.3390/genes12010064
APA StyleLoizidou, E. M., Kucherenko, A., Tatarskyy, P., Chernushyn, S., Livshyts, G., Gulkovskyi, R., Vorobiova, I., Antipkin, Y., Gorodna, O., Kaakinen, M. A., Prokopenko, I., & Livshits, L. (2021). Risk of Recurrent Pregnancy Loss in the Ukrainian Population Using a Combined Effect of Genetic Variants: A Case-Control Study. Genes, 12(1), 64. https://doi.org/10.3390/genes12010064






