Spatial and Temporal Patterns of Endopolyploidy in Mosses
Abstract
1. Introduction
2. Materials and Methods
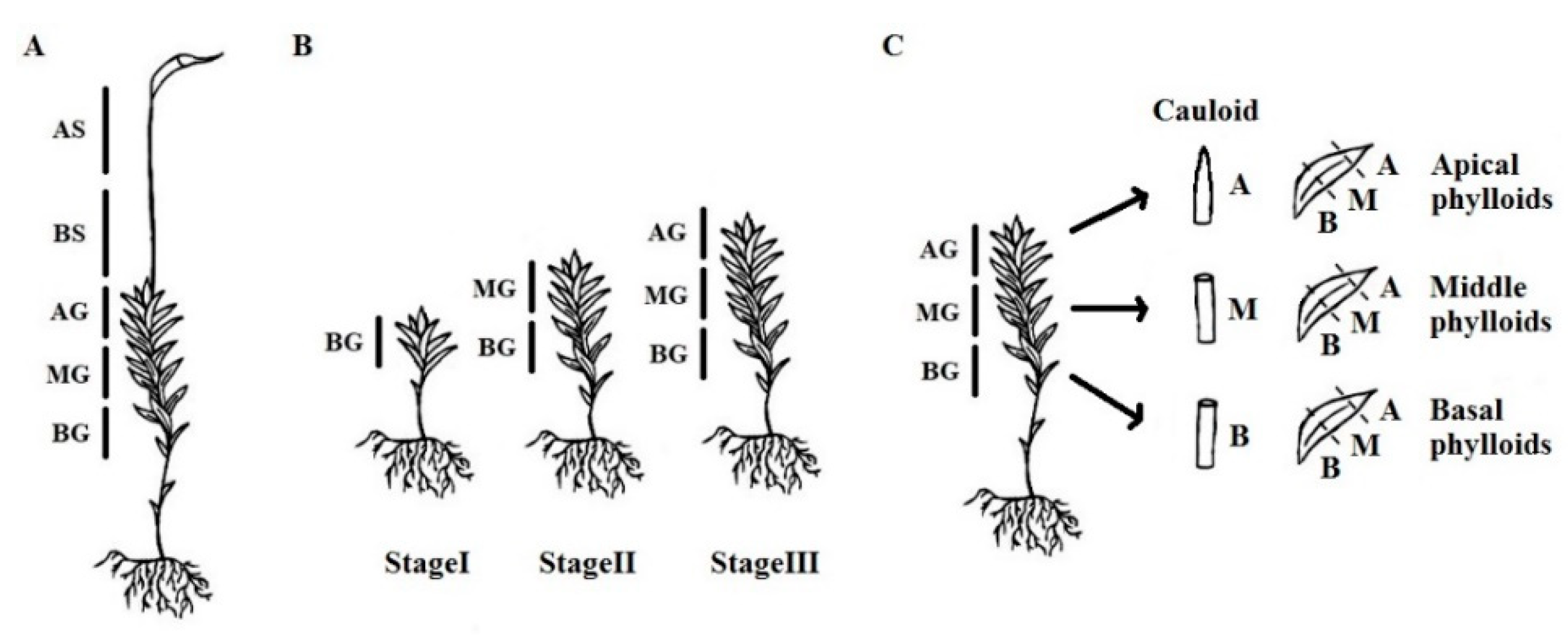
2.1. Plant Material
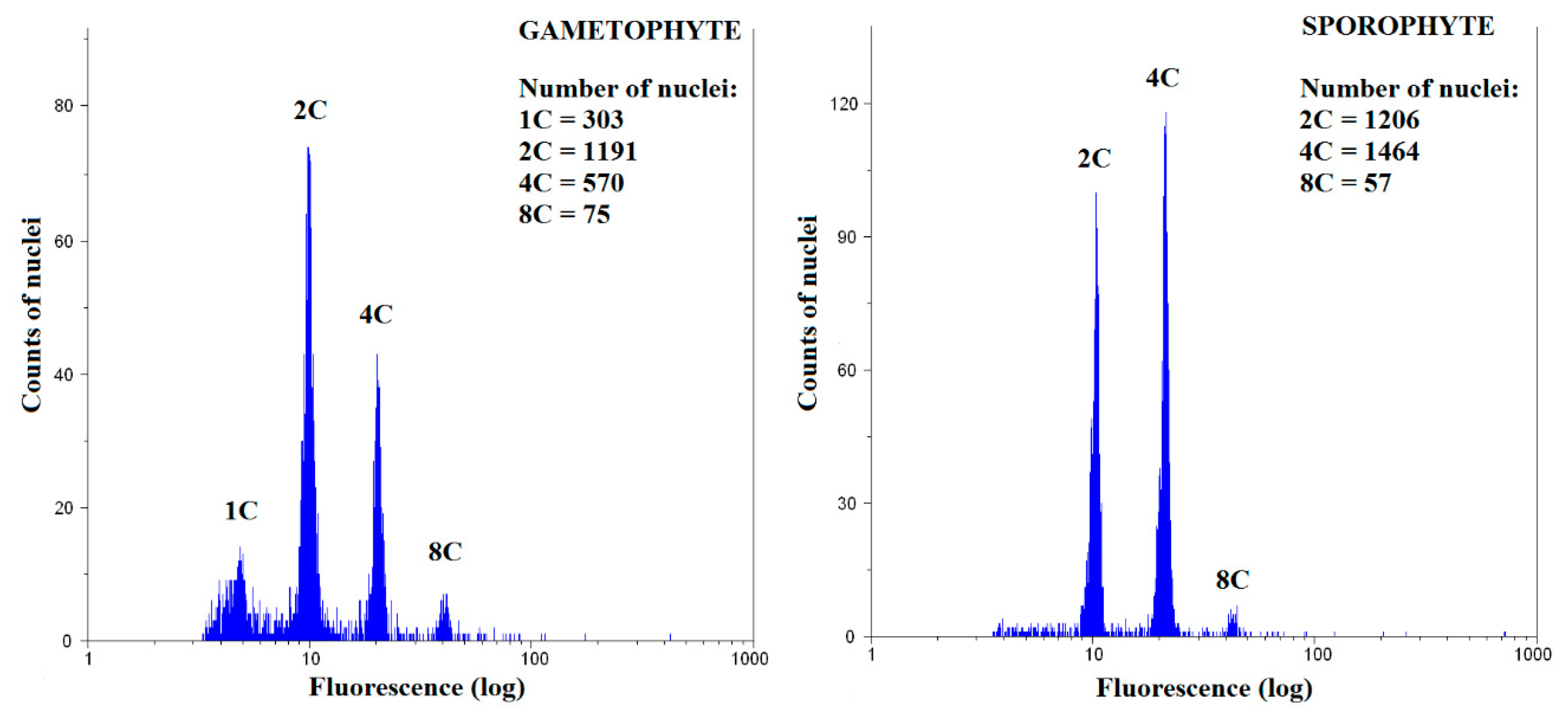
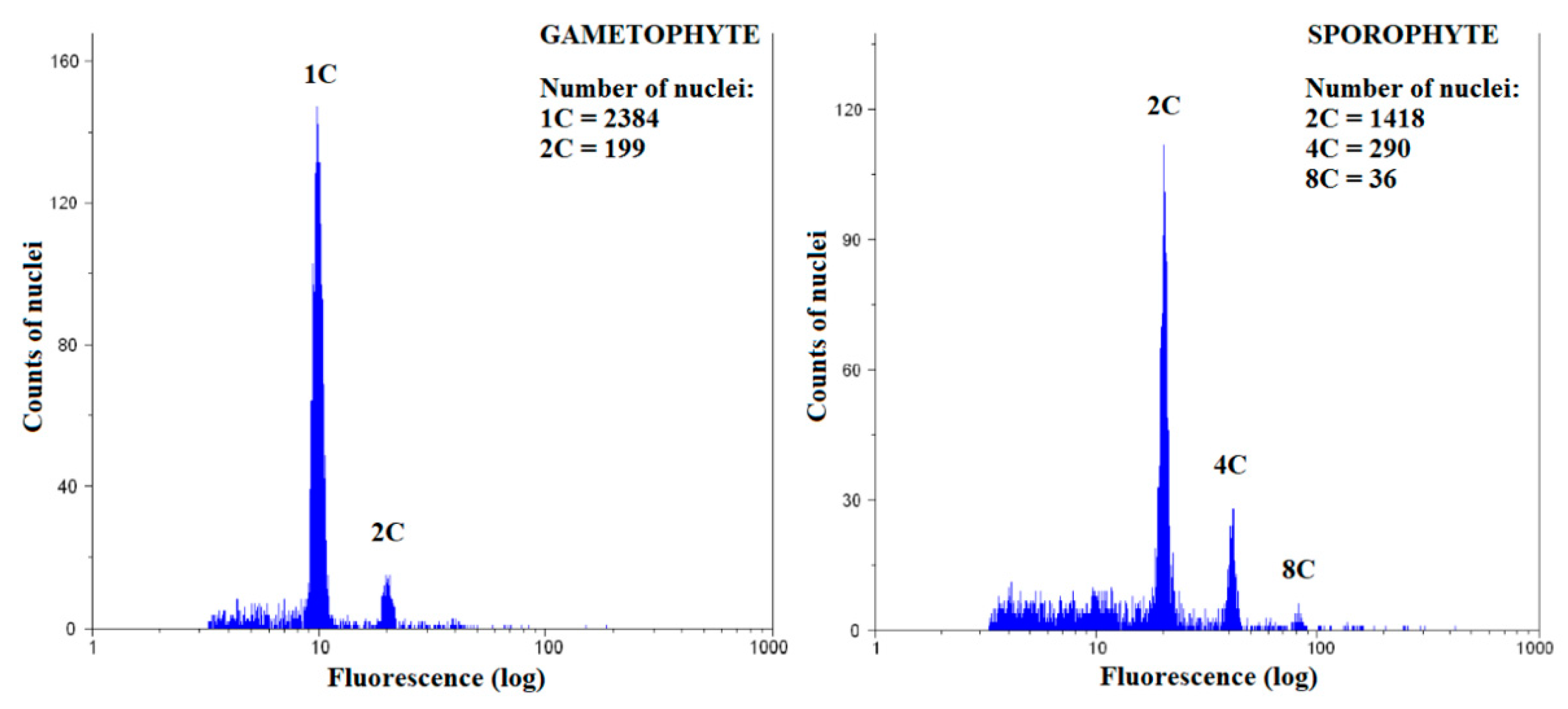
2.2. Sample Preparation for Flow Cytometry and Measurements
2.3. Calculation of Endoreduplication Index
2.4. Statistical Evaluation
3. Results
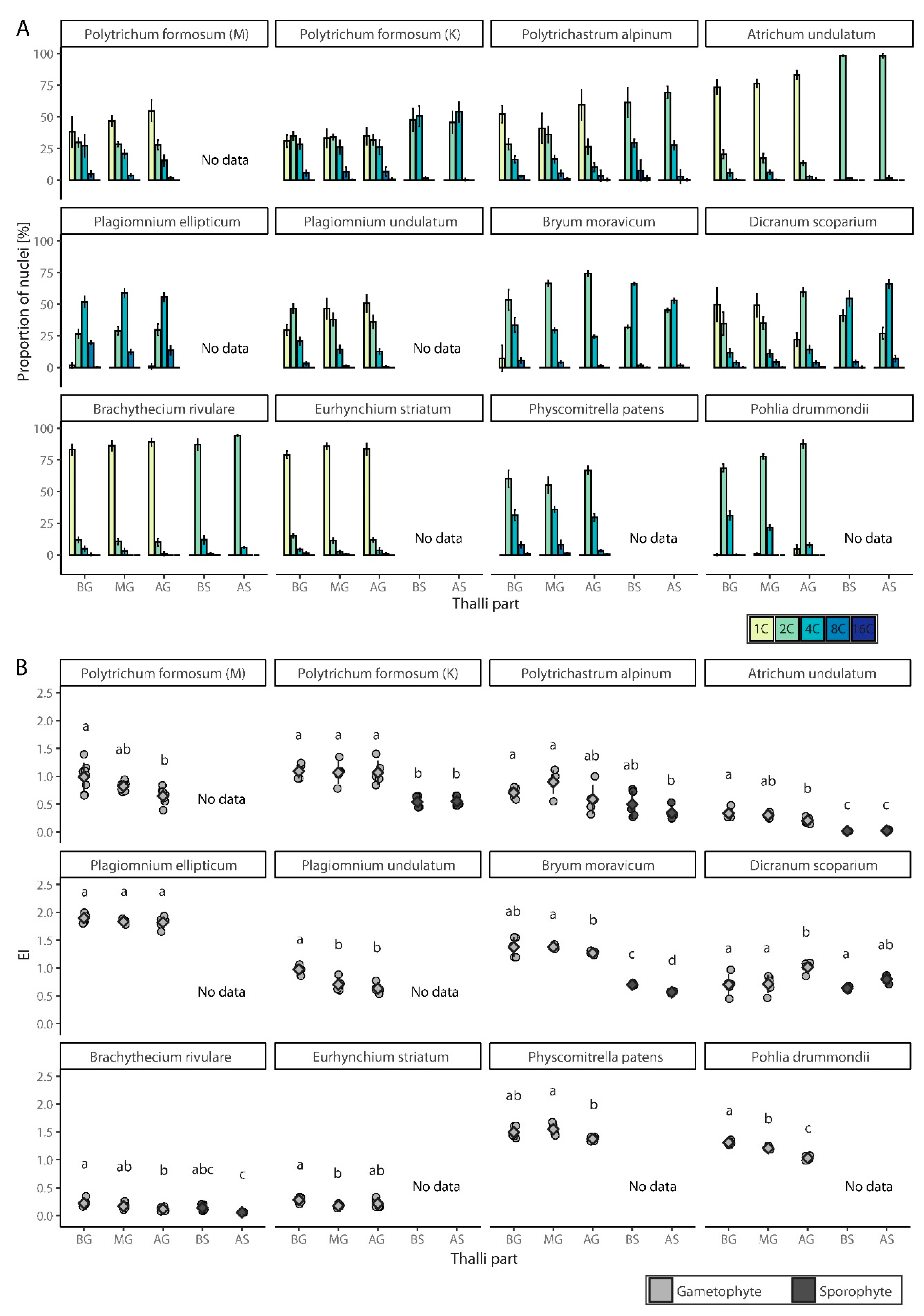
3.1. Spatial Distribution of Endopolyploidy in Moss Thalli
3.2. Spatiotemporal Analysis of Endopolyploidy in Two Moss Species Cultivated In Vitro
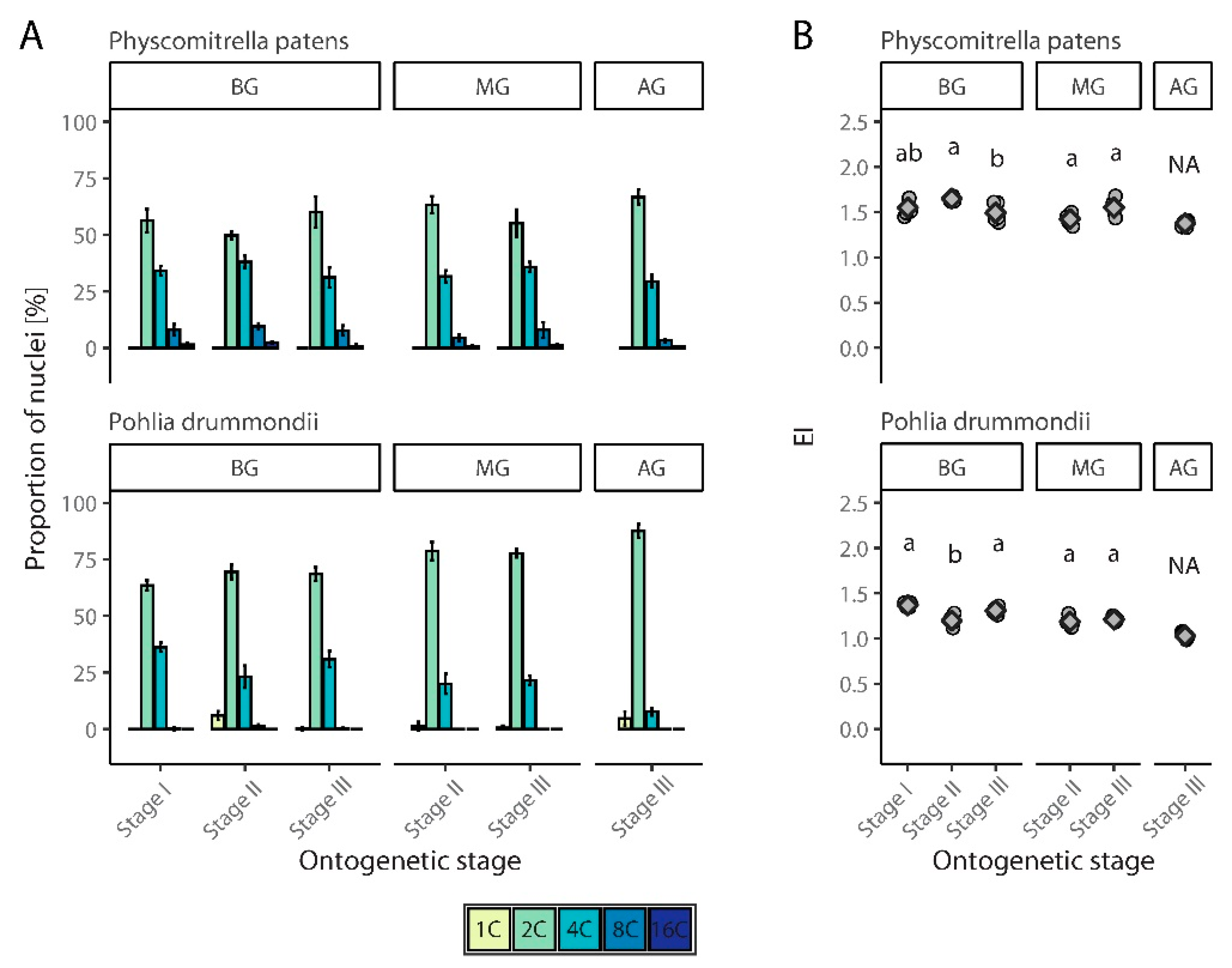
3.2.1. Physcomitrella Patens
3.2.2. Pohlia Drummondii
3.3. Spatial Distribution of Endopolyploidy in Moss Organs
3.3.1. Polytrichum Formosum
3.3.2. Plagiomnium Ellipticum
4. Discussion
4.1. Spatial Distribution of Endopolyploidy in Moss Thalli and Organs
4.2. Temporal Distribution of Endopolyploidy in Moss Thalli
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Barow, M.; Meister, A. Endopolyploidy in seed plants is differently correlated to systematics, organ, life strategy and genome size. Plant Cell Environ. 2003, 26, 571–584. [Google Scholar] [CrossRef]
- D’Amato, F. Endopolyploidy as a Factor in Plant Tissue Development. Caryologia 1964, 17, 41–52. [Google Scholar] [CrossRef][Green Version]
- Maluszynska, J.; Kolano, B.; Sas-Nowosielska, H. Endopolyploidy in Plants. In Plant Genome Diversity; Leitch, I.J., Greilhuber, J., Doležel, J., Wendel, J., Eds.; Springer: Wien, Austria, 2013; pp. 99–119. [Google Scholar]
- Lee, H.O.; Davidson, J.M.; Duronio, R.J. Endoreplication: Polyploidy with purpose. Genes Dev. 2009, 23, 2461–2477. [Google Scholar] [CrossRef] [PubMed]
- Dante, R.A.; Larkins, B.A.; Sabelli, P.A. Cell cycle control and seed development. Front. Plant Sci. 2014, 5, 14. [Google Scholar] [CrossRef]
- Joubès, J.; Chevalier, C. Endoreduplication in higher plants. Plant Mol. Biol. 2000, 43, 735–745. [Google Scholar] [CrossRef]
- Coelho, C.M.; Dante, R.A.; Sabelli, P.A.; Sun, Y.J.; Dilkes, B.P.; Gordon-Kamm, W.J.; Larkins, B.A. Cyclin-dependent kinase inhibitors in maize endosperm and their potential role in endoreduplication. Plant Physiol. 2005, 138, 2323–2336. [Google Scholar] [CrossRef]
- Scholes, D.R.; Paige, K.N. Plasticity in ploidy: A generalized response to stress. Trends Plant Sci. 2015, 20, 165–175. [Google Scholar] [CrossRef]
- Bateman, R.M.; Guy, J.J.; Rudall, P.J.; Leitch, I.J.; Pellicer, J.; Leitch, A.R. Evolutionary and functional potential of ploidy increase within individual plants: Somatic ploidy mapping of the complex labellum of sexually deceptive bee orchids. Ann. Bot. 2018, 122, 133–150. [Google Scholar] [CrossRef]
- Bainard, J.D.; Newmaster, S.G. Endopolyploidy in bryophytes: Widespread in mosses and absent in liverworts. J. Bot. 2010, 2010, 7. [Google Scholar] [CrossRef]
- Bainard, J.D.; Henry, T.A.; Bainard, L.D.; Newmaster, S.G. DNA content variation in monilophytes and lycophytes: Large genomes that are not endopolyploid. Chromosome Res. 2011, 19, 763–775. [Google Scholar] [CrossRef]
- Bhosale, R.; Boudolf, V.; Cuevas, F.; Lu, R.; Eekhout, T.; Hu, Z.B.; Van Isterdael, G.; Lambert, G.M.; Xu, F.; Nowack, M.K.; et al. A spatiotemporal DNA endoploidy map of the Arabidopsis root reveals roles for the endocycle in root development and stress adaptation. Plant Cell 2018, 30, 2330–2351. [Google Scholar] [CrossRef] [PubMed]
- Robinson, D.O.; Coate, J.E.; Singh, A.; Hong, L.L.; Bush, M.; Doyle, J.J.; Roeder, A.H.K. Ploidy and size at multiple scales in the Arabidopsis sepal. Plant Cell 2018, 30, 2308–2329. [Google Scholar] [CrossRef] [PubMed]
- Barow, M. Endopolyploidy in seed plants. Bioessays 2006, 28, 271–281. [Google Scholar] [CrossRef] [PubMed]
- Bauer, M.J.; Birchler, J.A. Organization of endoreduplicated chromosomes in the endosperm of Zea mays L. Chromosoma 2006, 115, 383–394. [Google Scholar] [CrossRef] [PubMed]
- Ceccarelli, M.; Santantonio, E.; Marmottini, F.; Amzallag, G.N.; Cionini, P.G. Chromosome endoreduplication as a factor of salt adaptation in Sorghum bicolor. Protoplasma 2006, 227, 113–118. [Google Scholar] [CrossRef] [PubMed]
- Sliwinska, E.; Lukaszewska, E. Polysomaty in growing in vitro sugar-beet (Beta vulgaris L.) seedlings of different ploidy level. Plant Sci. 2005, 168, 1067–1074. [Google Scholar] [CrossRef]
- Bourdon, M.; Pirrello, J.; Cheniclet, C.; Coriton, O.; Bourge, M.; Brown, S.; Moïse, A.; Peypelut, M.; Rouyère, V.; Renaudin, J.P.; et al. Evidence for karyoplasmic homeostasis during endoreduplication and a ploidy-dependent increase in gene transcription during tomato fruit growth. Development 2012, 139, 3817–3826. [Google Scholar] [CrossRef]
- Cookson, S.J.; Van Lijsebettens, M.; Granier, C. Correlation between leaf growth variables suggest intrinsic and early controls of leaf size in Arabidopsis thaliana. Plant Cell Environ. 2005, 28, 1355–1366. [Google Scholar] [CrossRef]
- De Veylder, L.; Larkin, J.C.; Schnittger, A. Molecular control and function of endoreplication in development and physiology. Trends Plant Sci. 2011, 16, 624–634. [Google Scholar] [CrossRef]
- Kondorosi, E.; Roudier, F.; Gendreau, E. Plant cell-size control: Growing by ploidy? Curr. Opin. Plant Biol. 2000, 3, 488–492. [Google Scholar] [CrossRef]
- Breuer, C.; Braidwood, L.; Sugimoto, K. Endocycling in the path of plant development. Curr. Opin. Plant Biol. 2014, 17, 78–85. [Google Scholar] [CrossRef] [PubMed]
- Cavallini, A.; Baroncelli, S.; Lercari, B.; Cionini, G.; Rocca, M.; D’Amato, F. Effect of light and gibberellic-acid on chromosome endoreduplication in leaf epidermis of Triticum durum Desf. Protoplasma 1995, 186, 57–62. [Google Scholar] [CrossRef]
- Kinoshita, I.; Sanbe, A.; Yokomura, E.I. Difference in light-induced increase in ploidy level and cell size between adaxial and abaxial epidermal pavement cells of Phaseolus vulgaris primary leaves. J. Exp. Bot. 2008, 59, 1419–1430. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yamasaki, S.; Shimada, E.; Kuwano, T.; Kawano, T.; Noguchi, N. Continuous UV-B irradiation induces endoreduplication and peroxidase activity in epidermal cells surrounding trichomes on cucumber cotyledons. J. Radiat. Res. 2010, 51, 187–196. [Google Scholar] [CrossRef] [PubMed]
- Gegas, V.C.; Wargent, J.J.; Pesquet, E.; Granqvist, E.; Paul, N.D.; Doonan, J.H. Endopolyploidy as a potential alternative adaptive strategy for Arabidopsis leaf size variation in response to UV-B. J. Exp. Bot. 2014, 65, 2757–2766. [Google Scholar] [CrossRef] [PubMed]
- Engelen-Eigles, G.; Jones, R.J.; Phillips, R.L. DNA endoreduplication in maize endosperm cells: The effect of exposure to short-term high temperature. Plant Cell Environ. 2000, 23, 657–663. [Google Scholar] [CrossRef]
- Jovtchev, G.; Barow, M.; Meister, A.; Schubert, I. Impact of environmental and endogenous factors on endopolyploidization in angiosperms. Environ. Exp. Bot. 2007, 60, 404–411. [Google Scholar] [CrossRef]
- Artlip, T.S.; Madison, J.T.; Setter, T.L. Water-deficit in developing endosperm of maize—Cell division and nuclear DNA endoreduplication. Plant Cell Environ. 1995, 18, 1034–1040. [Google Scholar] [CrossRef]
- Granier, C.; Inzé, D.; Tardieu, F. Spatial distribution of cell division rate can be deduced from that of p34(cdc2) kinase activity in maize leaves grown at contrasting temperatures and soil water conditions. Plant Physiol. 2000, 124, 1393–1402. [Google Scholar] [CrossRef]
- Elmaghrabi, A.M.; Ochatt, S.; Rogers, H.J.; Francis, D. Enhanced tolerance to salinity following cellular acclimation to increasing NaCl levels in Medicago truncatula. Plant Cell Tissue Organ Cult. 2013, 114, 61–70. [Google Scholar] [CrossRef]
- Andrea, B.; Caceres, M.E.; Cionini, G.; Cionini, P.G. A DNA cytophotometric study of salt adaptation in Allium cepa and Nicotiana bigelovii. Caryologia 2008, 61, 176–181. [Google Scholar] [CrossRef]
- Barkla, B.J.; Rhodes, T.; Tran, K.N.T.; Wijesinghege, C.; Larkin, J.C.; Dassanayake, M. Making epidermal bladder cells bigger: Developmental and salinity-induced endopolyploidy in a model halophyte. Plant Physiol. 2018, 177, 615–632. [Google Scholar] [CrossRef] [PubMed]
- Kocová, V.; Bubanová, D.; Rákai, A.; Kolarčik, V.; Mártonfi, P. Salinity has no effect on polysomatic pattern in seedlings of Trifolium pratense and T. repens. Acta Biol. Crac. Ser. Bot. 2017, 59, 55–65. [Google Scholar] [CrossRef]
- Ding, J.; Sun, Y.; Xiao, C.L.; Shi, K.; Zhou, Y.H.; Yu, J.Q. Physiological basis of different allelopathic reactions of cucumber and figleaf gourd plants to cinnamic acid. J. Exp. Bot. 2007, 58, 3765–3773. [Google Scholar] [CrossRef] [PubMed]
- Goga, M.; Ručová, D.; Kolarčik, V.; Sabovljević, M.; Bačkor, M.; Lang, I. Usnic acid, as a biotic factor, changes the ploidy level in mosses. Ecol. Evol. 2018, 8, 2781–2787. [Google Scholar] [CrossRef]
- Maróti, G.; Kondorosi, E. Nitrogen-fixing Rhizobium-legume symbiosis: Are polyploidy and host peptide-governed symbiont differentiation general principles of endosymbiosis? Front. Microbiol. 2014, 5, 6. [Google Scholar] [CrossRef]
- Bainard, L.D.; Bainard, J.D.; Newmaster, S.G.; Klironomos, J.N. Mycorrhizal symbiosis stimulates endoreduplication in angiosperms. Plant Cell Environ. 2011, 34, 1577–1585. [Google Scholar] [CrossRef]
- Ishida, T.; Adachi, S.; Yoshimura, M.; Shimizu, K.; Umeda, M.; Sugimoto, K. Auxin modulates the transition from the mitotic cycle to the endocycle in Arabidopsis. Development 2010, 137, 63–71. [Google Scholar] [CrossRef]
- Scholes, D.R.; Paige, K.N. Plasticity in ploidy underlies plant fitness compensation to herbivore damage. Mol. Ecol. 2014, 23, 4862–4870. [Google Scholar] [CrossRef]
- Scholes, D.R. Ploidy in plant tolerance to apical meristem damage: A test of relative costs and benefits. Int. J. Plant Sci. 2020, 181, 509–517. [Google Scholar] [CrossRef]
- Paige, K.N. Overcompensation, environmental stress, and the role of endoreduplication. Am. J. Bot. 2018, 105, 1105–1108. [Google Scholar] [CrossRef] [PubMed]
- Vieira, P.; De Clercq, A.; Stals, H.; Van Leene, J.; Van de Slijke, E.; Van Isterdael, G.; Eeckhout, D.; Persiau, G.; Van Damme, D.; Verkest, A.; et al. The cyclin-dependent kinase inhibitor KRP6 induces mitosis and impairs cytokinesis in giant cells induced by plant-parasitic nematodes in Arabidopsis. Plant Cell 2014, 26, 2633–2647. [Google Scholar] [CrossRef] [PubMed]
- Vieira, P.; Engler, J.D. Plant cyclin-dependent kinase inhibitors of the KRP family: Potent inhibitors of root-knot nematode feeding sites in plant roots. Front. Plant Sci. 2017, 8, 9. [Google Scholar] [CrossRef] [PubMed]
- Siddique, S.; Radakovic, Z.S.; De La Torre, C.M.; Chronis, D.; Novák, O.; Ramireddy, E.; Holbein, J.; Matera, C.; Hütten, M.; Gutbrod, P.; et al. A parasitic nematode releases cytokinin that controls cell division and orchestrates feeding site formation in host plants. Proc. Natl. Acad. Sci. USA 2015, 112, 12669–12674. [Google Scholar] [CrossRef]
- Bainard, J.D.; Newmaster, S.G.; Budke, J.M. Genome size and endopolyploidy evolution across the moss phylogeny. Ann. Bot. 2020, 125, 543–555. [Google Scholar] [CrossRef]
- Buck, W.; Goffinet, B. Morphology and classification of the Bryophyta. In Bryophyte Biology, 2nd ed.; Cambridge University Press: Cambridge, UK, 2009; pp. 55–138. [Google Scholar]
- Crandall-Stotler, B.J.; Bartholomew-Began, S.E. Morphology of mosses (phylum Bryophyta). Flora N. Am. N. Mex. 2007, 27, 3–13. [Google Scholar]
- Slack, N.G. The ecological value of bryophytes as indicators of climate change. In Bryophyte Ecology and Climate Change, 1st ed.; Tuba, Z., Slack, N.G., Stark, L.R., Eds.; Cambridge University Press: Cambridge, UK, 2011; p. 528. [Google Scholar]
- Magill, R.E. Moss diversity: New look at old numbers. Phytotaxa 2010, 9, 167–174. [Google Scholar] [CrossRef][Green Version]
- Ennos, R.; Sheffield, L. Life on the ground. In Plant Life; John Wiley and Sons Ltd.: Hoboken, NJ, USA; Blackwell Science Inc.: Oxford, UK, 2000; p. 232. [Google Scholar]
- Nehira, K. Spore germination, protonema development and soreling development. In New Manual of Bryology; Schuster, R.M., Ed.; The Hattori Botanical Laboratory: Nichinan, Japan, 1983; pp. 343–379. [Google Scholar]
- Bainard, J.D.; Forrest, L.L.; Goffinet, B.; Newmaster, S.G. Nuclear DNA content variation and evolution in liverworts. Mol. Phylogenet. Evol. 2013, 68, 619–627. [Google Scholar] [CrossRef]
- Bainard, J.D.; Villarreal, J.C. Genome size increases in recently diverged hornwort clades. Genome 2013, 56, 431–435. [Google Scholar] [CrossRef]
- Gang, Y.Y.; Du, G.S.; Shi, D.J.; Wang, M.Z.; Li, X.D.; Hua, Z.L. Establishment of in vitro regeneration system of the Atrichum mosses. Acta Bot. Sin. 2003, 45, 1475–1480. [Google Scholar]
- Loureiro, J.; Rodriguez, E.; Doležel, J.; Santos, C. Two new nuclear isolation buffers for plant DNA flow cytometry: A test with 37 species. Ann. Bot. 2007, 100, 875–888. [Google Scholar] [CrossRef] [PubMed]
- Doležel, J.; Greilhuber, J.; Suda, J. Estimation of nuclear DNA content in plants using flow cytometry. Nat. Protoc. 2007, 2, 2233–2244. [Google Scholar] [CrossRef] [PubMed]
- Givan, A.L. Principles of flow cytometry: An overview. Methods Cell Biol. 2001, 63, 19–50. [Google Scholar] [CrossRef] [PubMed]
- Kolarčik, V.; Fraková, V.; Kocová, V.; Koprivý, L.; Mártonfi, P. Endopolyploidy pattern in Corydalis early spring geophytes. Flora 2020, 270. [Google Scholar] [CrossRef]
- Bainard, J.D.; Bainard, L.D.; Henry, T.A.; Fazekas, A.J.; Newmaster, S.G. A multivariate analysis of variation in genome size and endoreduplication in angiosperms reveals strong phylogenetic signal and association with phenotypic traits. New Phytol. 2012, 196, 1240–1250. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Core Team: Wien, Austria, 2019. [Google Scholar]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2009. [Google Scholar] [CrossRef]
- Schween, G.; Gorr, G.; Hohe, A.; Reski, R. Unique tissue-specific cell cycle in Physcomitrella. Plant. Biol. 2003, 5, 50–58. [Google Scholar] [CrossRef]
- Reski, R. Development, genetics and molecular biology of mosses. Bot. Acta 1998, 111, 1–15. [Google Scholar] [CrossRef]
- Lim, W.L.; Loh, C.S. Endopolyploidy in Vanda Miss Joaquim (Orchidaceae). New Phytol. 2003, 159, 279–287. [Google Scholar] [CrossRef]
- Qiu, Y.L.; Taylor, A.B.; McManus, H.A. Evolution of the life cycle in land plants. J. Syst. Evol. 2012, 50, 171–194. [Google Scholar] [CrossRef]
- Pacey, E.K.; Maherali, H.; Husband, B.C. The influence of experimentally induced polyploidy on the relationships between endopolyploidy and plant function in Arabidopsis thaliana. Ecol. Evol. 2020, 10, 198–216. [Google Scholar] [CrossRef]
- Straková, N.; Kocová, V.; Kolarčik, V.; Mártonfi, P. Endopolyploidy in organs of Trifolium pratense L. in different ontogenetic stages. Caryologia 2014, 67, 116–123. [Google Scholar] [CrossRef]
- Kolano, B.; Siwinska, D.; Maluszynska, J. Endopolyploidy patterns during development of Chenopodium quinoa. Acta Biol. Crac. Ser. Bot. 2009, 51, 85–92. [Google Scholar]
- Turetsky, M.R.; Bond-Lamberty, B.; Euskirchen, E.; Talbot, J.; Frolking, S.; McGuire, A.D.; Tuittila, E.S. The resilience and functional role of moss in boreal and arctic ecosystems. New Phytol. 2012, 196, 49–67. [Google Scholar] [CrossRef] [PubMed]
- Rensing, S.A.; Lang, D.; Zimmer, A.D.; Terry, A.; Salamov, A.; Shapiro, H.; Nishiyama, T.; Perroud, P.F.; Lindquist, E.A.; Kamisugi, Y.; et al. The Physcomitrella genome reveals evolutionary insights into the conquest of land by plants. Science 2008, 319, 64–69. [Google Scholar] [CrossRef] [PubMed]

| Species | Collection Details | Sample Size | ||
|---|---|---|---|---|
| Exp1 | Exp2 | Exp3 | ||
| G/S | StI/StII/StIII | C/PBG/PMG/PAG | ||
| Naturally growing plants | ||||
| Atrichum undulatum | ES, Milpoš, 49.193448° N, 21.015777° E | 5/5 | -/-/- | -/-/-/- |
| Brachythecium rivulare | ES, Milpoš, 49.193448° N, 21.015777° E | 5/5 | -/-/- | -/-/-/- |
| Bryum moravicum | ES, Milpoš, 49.193448° N, 21.015777° E | 5/5 | -/-/- | -/-/-/- |
| Dicranum scoparium | ES, Vysoké Tatry, 49.211943° N, 20.308818° E | 5/5 | -/-/- | -/-/-/- |
| Eurhynchium striatum | CS, Stankovany, 49.154203° N, 19.151767° E | 5/- | -/-/- | -/-/-/- |
| Plagiomnium ellipticum | ES, Drahňov, 48.569892° N, 21.955160° E | 5/- | -/-/- | 3/3/3/3 |
| Plagiomnium undulatum | ES, Rovné, 49.2736989° N, 21.5209615° E | 5/- | -/-/- | -/-/-/- |
| Polytrichastrum alpinum | ES, Vysoké Tatry, 49.151811° N, 20.080580° E | 5/5 | -/-/- | -/-/-/- |
| Polytrichum formosum | ES, Milpoš, 49.193448° N, 21.015777° E (M code) | 8/- | -/-/- | 3/3/3/3 |
| ES, Kamenica, 49.212205° N, 20.991363° E (K code) | 5/5 | -/-/- | -/-/-/- | |
| Laboratory culture | ||||
| Physcomitrella patens | - | 5/- | 5/5/5 | -/-/-/- |
| Pohlia drummondii | - | 5/- | 5/5/5 | -/-/-/- |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Paľová, M.; Ručová, D.; Goga, M.; Kolarčik, V. Spatial and Temporal Patterns of Endopolyploidy in Mosses. Genes 2021, 12, 27. https://doi.org/10.3390/genes12010027
Paľová M, Ručová D, Goga M, Kolarčik V. Spatial and Temporal Patterns of Endopolyploidy in Mosses. Genes. 2021; 12(1):27. https://doi.org/10.3390/genes12010027
Chicago/Turabian StylePaľová, Marianna, Dajana Ručová, Michal Goga, and Vladislav Kolarčik. 2021. "Spatial and Temporal Patterns of Endopolyploidy in Mosses" Genes 12, no. 1: 27. https://doi.org/10.3390/genes12010027
APA StylePaľová, M., Ručová, D., Goga, M., & Kolarčik, V. (2021). Spatial and Temporal Patterns of Endopolyploidy in Mosses. Genes, 12(1), 27. https://doi.org/10.3390/genes12010027







