A Novel Carboxylesterase Derived from a Compost Metagenome Exhibiting High Stability and Activity towards High Salinity
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Plasmids
2.2. Identification and Analysis of Est56 Sequence
2.3. Cloning and Expression of Est56
2.4. Purification of Recombinant Est56
2.5. Esterase Standard Assay
2.6. Characterization of Est56
2.6.1. Substrate Specificity
2.6.2. Effect of Temperature and pH
2.6.3. Effect of Salinity
2.6.4. Effect of Organic Solvents
2.6.5. Effect of Other Additives
2.7. Sequence Analysis of Halotolerant Enzymes
2.8. Homology Modeling and Putative Structure Analysis
2.9. Accession Numbers
3. Results
3.1. Identification and Sequence Analysis of a Novel Lipolytic Gene
3.2. Purification of Recombinant Est56 and Substrate Specificity
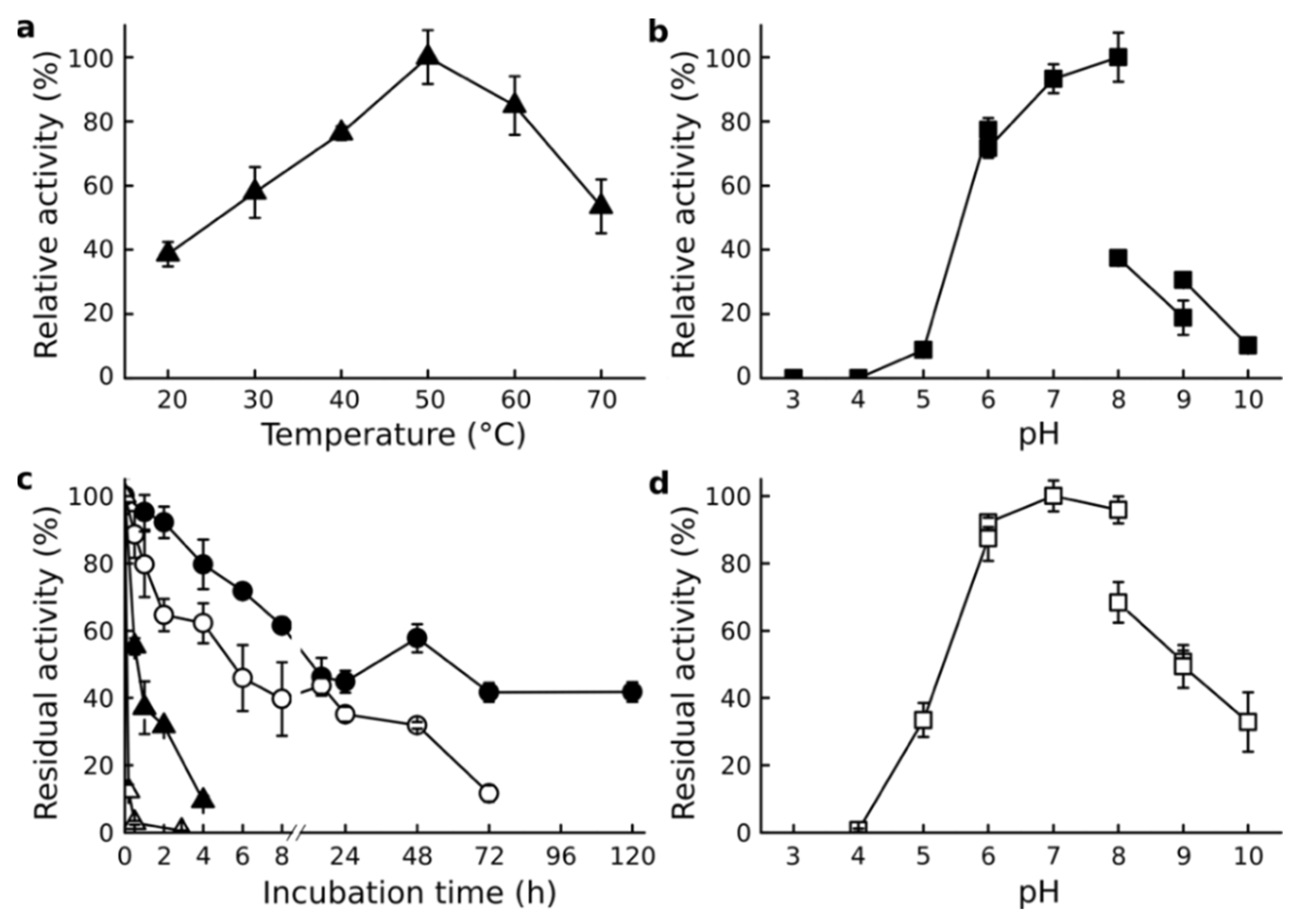
3.3. Effect of Temperature and pH
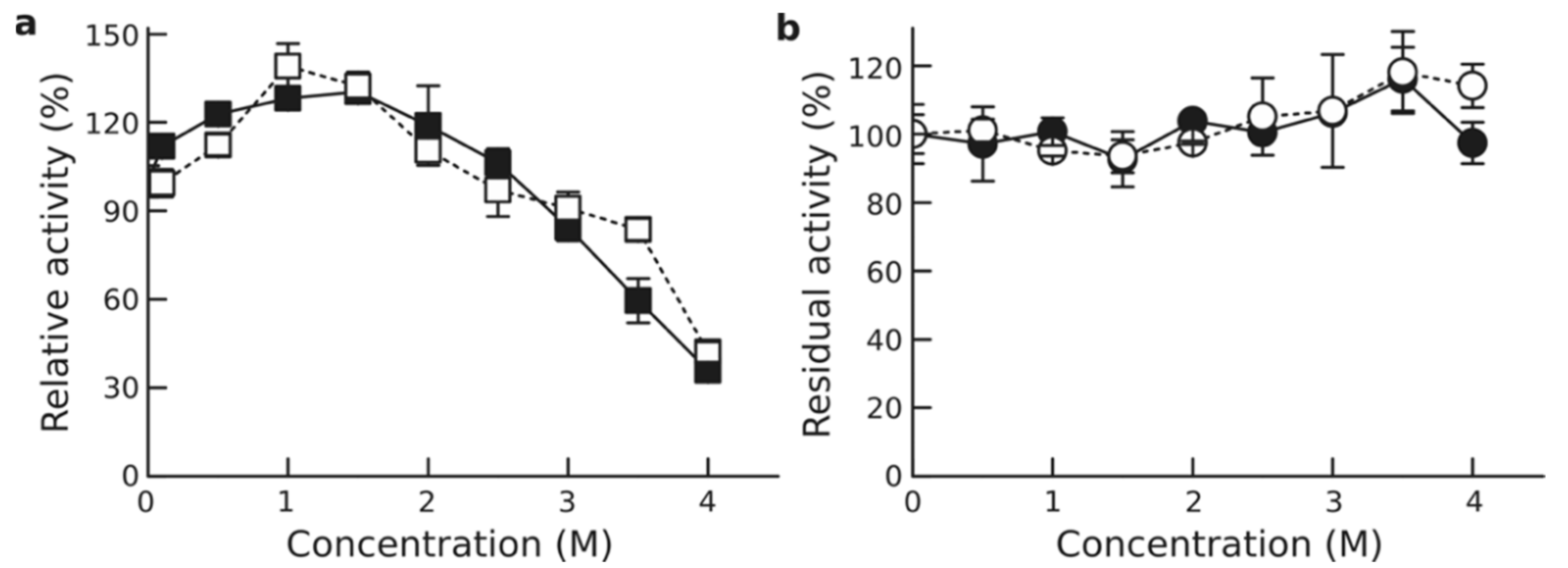
3.4. Effect of Salinity
3.5. Protective Effect of NaCl against Denaturants
3.6. Effect of Organic Solvents
3.7. Effect of Other Additives
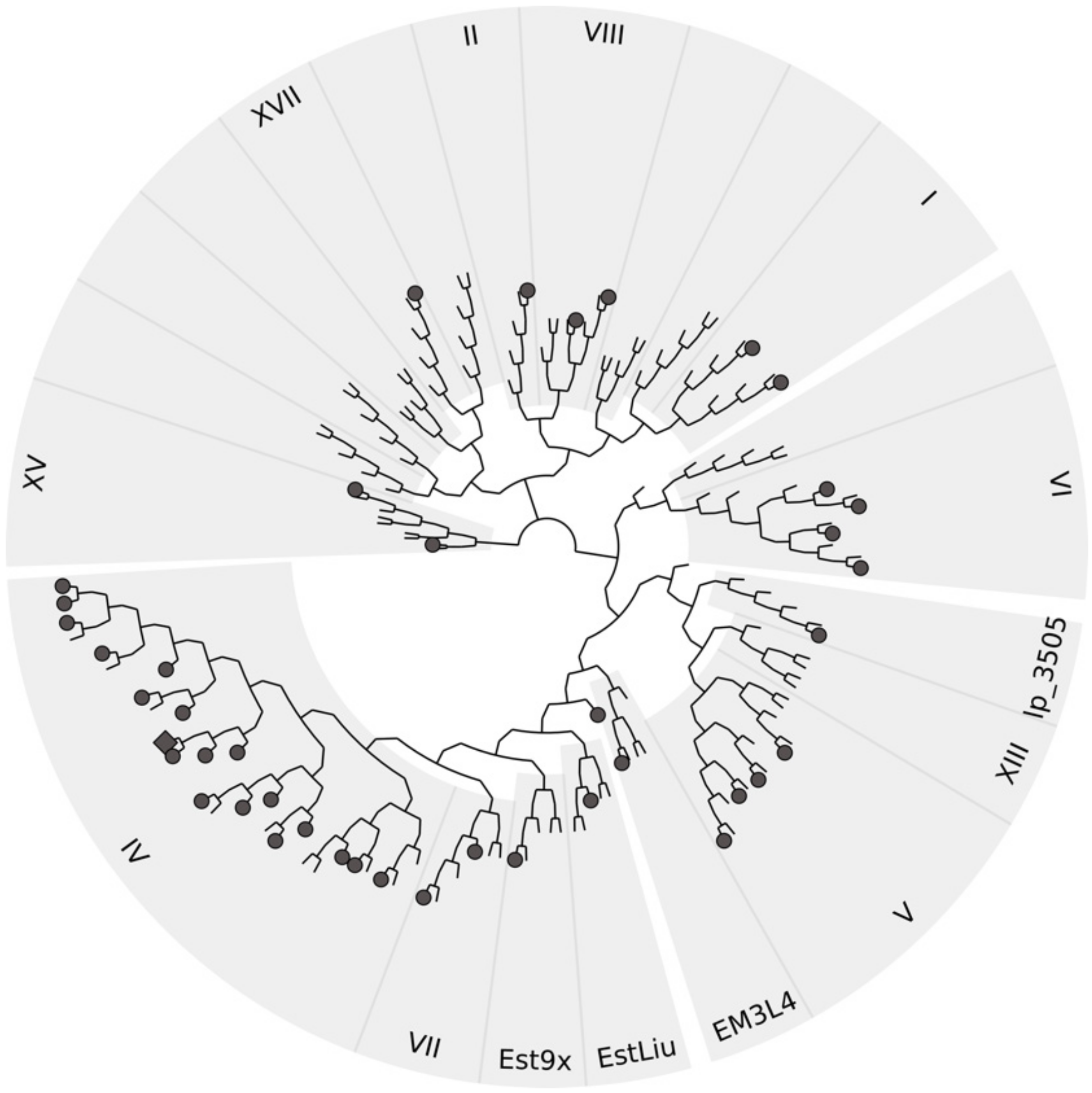
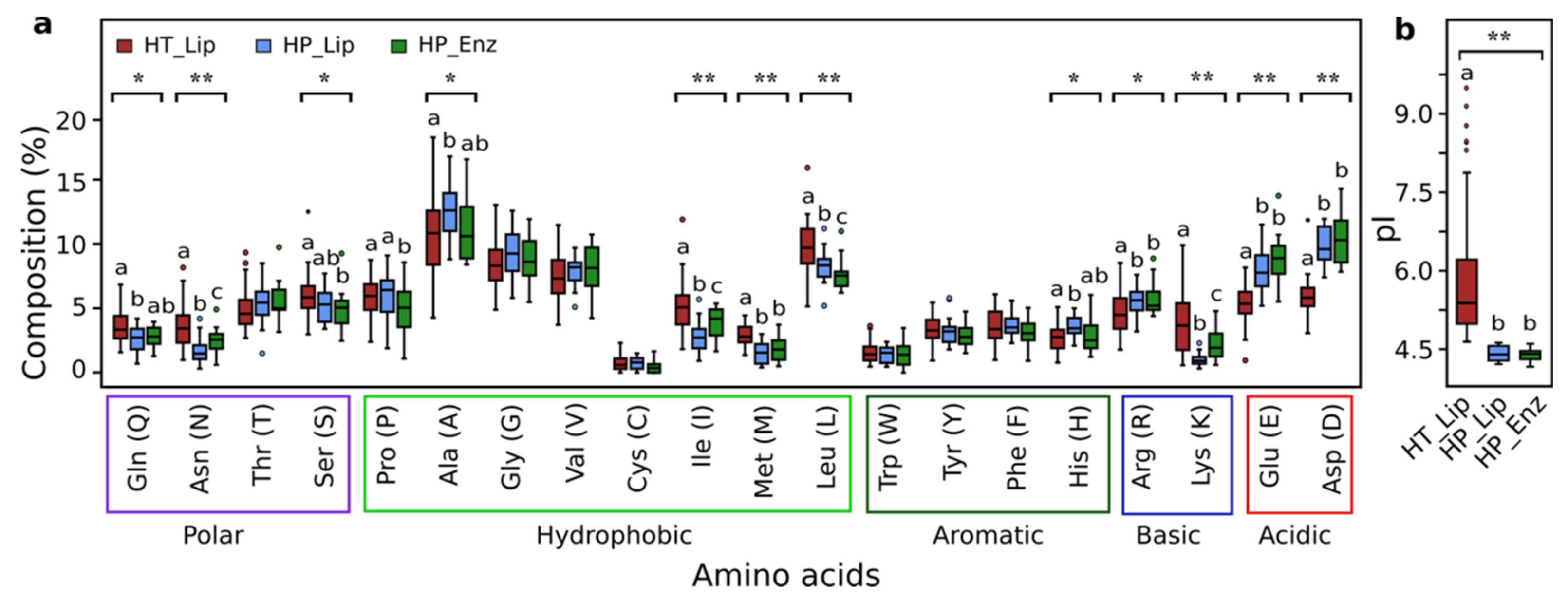
3.8. Sequence Analysis of Halotolerant Lipolytic Enzymes
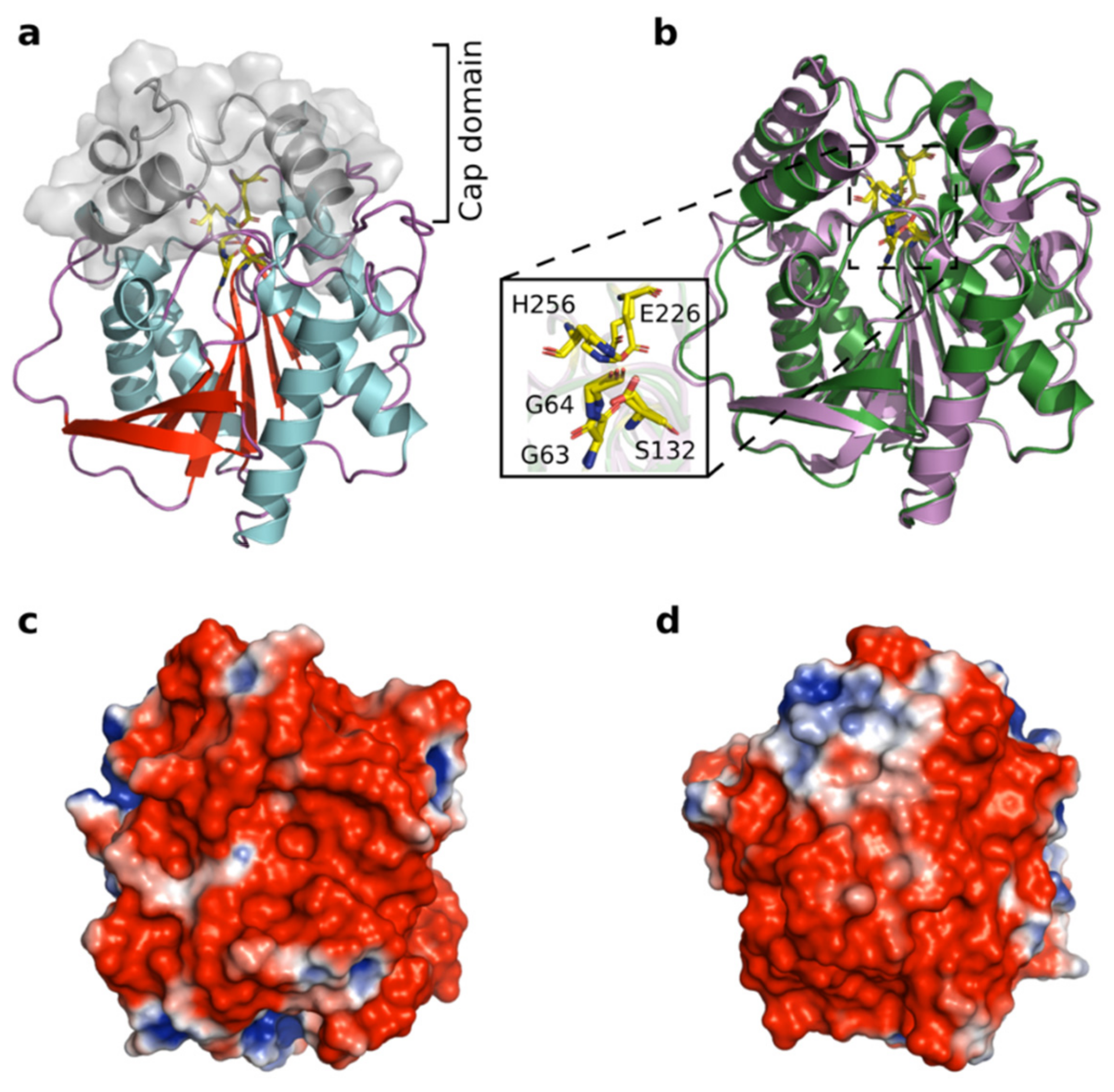
3.9. Structural Modeling of Est56
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yadav, A.N.; Saxena, A.K. Biodiversity and biotechnological applications of halophilic microbes for sustainable agriculture. J. Appl. Biol. Biotechnol. 2018, 6, 48–55. [Google Scholar] [CrossRef]
- Dassarma, S.; Dassarma, P. Halophiles and their enzymes: Negativity put to good use. Curr. Opin. Microbiol. 2015, 25, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Salma, M.; Abdulla, M.K.; Samina, M. Osmoadaptation in halophilic bacteria and archaea. Res. J. Biotechnol. 2020, 15, 154–161. [Google Scholar]
- Siglioccolo, A.; Paiardini, A.; Piscitelli, M.; Pascarella, S. Structural adaptation of extreme halophilic proteins through decrease of conserved hydrophobic contact surface. BMC Struct. Biol. 2011, 11, 50. [Google Scholar] [CrossRef] [PubMed]
- Ishibashi, M.; Hayashi, T.; Yoshida, C.; Tokunaga, M. Increase of salt dependence of halophilic nucleoside diphosphate kinase caused by a single amino acid substitution. Extremophiles 2013, 17, 585–591. [Google Scholar] [CrossRef] [PubMed]
- Moshfegh, M.; Shahverdi, A.R.; Zarrini, G.; Faramarzi, M.A. Biochemical characterization of an extracellular polyextremophilic α-amylase from the halophilic archaeon Halorubrum xinjiangense. Extremophiles 2013, 17, 677–687. [Google Scholar] [CrossRef] [PubMed]
- Cycil, L.M.; DasSarma, S.; Pecher, W.; McDonald, R.; AbdulSalam, M.; Hasan, F. Metagenomic insights into the diversity of halophilic microorganisms indigenous to the Karak Salt Mine, Pakistan. Front. Microbiol. 2020, 11, 1567. [Google Scholar] [CrossRef]
- Oren, A. Bioenergetic aspects of halophilism. Microbiol. Mol. Biol. Rev. 1999, 63, 334–348. [Google Scholar] [CrossRef]
- Ventosa, A.; Nieto, J.J.; Oren, A. Biology of moderately halophilic aerobic bacteria. Microbiol. Mol. Biol. Rev. 1998, 62, 504–544. [Google Scholar] [CrossRef]
- Corral, P.; Amoozegar, M.A.; Ventosa, A. Halophiles and their biomolecules: Recent advances and future applications in biomedicine. Mar. Drugs 2020, 18, 33. [Google Scholar] [CrossRef]
- Jiang, H.; Zhang, S.; Gao, H.; Hu, N. Characterization of a cold-active esterase from Serratia sp. and improvement of thermostability by directed evolution. BMC Biotechnol. 2016, 16. [Google Scholar] [CrossRef] [PubMed]
- Bai, W.; Xue, Y.; Zhou, C.; Ma, Y. Cloning, expression, and characterization of a novel alkali-tolerant xylanase from alkaliphilic Bacillus sp. SN5. Biotechnol. Lett. 2012, 34, 2093–2099. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Paul, D.; Bhushan, B.; Wakchaure, G.C.; Meena, K.K.; Shouche, Y. Traversing the “Omic” landscape of microbial halotolerance for key molecular processes and new insights. Crit. Rev. Microbiol. 2020, 46, 631–653. [Google Scholar] [CrossRef] [PubMed]
- Delgado-García, M.; Valdivia-Urdiales, B.; Aguilar-González, C.N.; Contreras-Esquivel, J.C.; Rodríguez-Herrera, R. Halophilic hydrolases as a new tool for the biotechnological industries. J. Sci. Food Agric. 2012, 92, 2575–2580. [Google Scholar] [CrossRef] [PubMed]
- Munawar, N.; Engel, P.C. Halophilic enzymes: Characteristics, structural adaptation and potential applications for biocatalysis. Curr. Biotechnol. 2013, 2, 334–344. [Google Scholar] [CrossRef]
- Ortega, G.; Laín, A.; Tadeo, X.; López-Méndez, B.; Castaño, D.; Millet, O. Halophilic enzyme activation induced by salts. Sci. Rep. 2011, 1, 6. [Google Scholar] [CrossRef]
- Edbeib, M.F.; Aksoy, H.M.; Kaya, Y.; Wahab, R.A.; Huyop, F. Haloadaptation: Insights from comparative modeling studies between halotolerant and non-halotolerant dehalogenases. J. Biomol. Struct. Dyn. 2020, 38, 3452–3461. [Google Scholar] [CrossRef]
- Sinha, R.; Khare, S.K. Protective role of salt in catalysis and maintaining structure of halophilic proteins against denaturation. Front. Microbiol. 2014, 5, 165. [Google Scholar] [CrossRef]
- Vogler, M.; Karan, R.; Renn, D.; Vancea, A.; Vielberg, M.-T.; Grötzinger, S.W.; DasSarma, P.; DasSarma, S.; Eppinger, J.; Groll, M.; et al. Crystal structure and active site engineering of a halophilic γ-carbonic anhydrase. Front. Microbiol. 2020, 11, 742. [Google Scholar] [CrossRef]
- Bardavid, R.E.; Oren, A. The amino acid composition of proteins from anaerobic halophilic bacteria of the order Halanaerobiales. Extremophiles 2012, 16, 567–572. [Google Scholar] [CrossRef]
- Arpigny, J.L.; Jaeger, K.-E. Bacterial lipolytic enzymes: Classification and properties. Biochem. J. 1999, 343, 177–183. [Google Scholar] [CrossRef] [PubMed]
- Bornscheuer, U.T. Microbial carboxyl esterases: Classification, properties and application in biocatalysis. FEMS Microbiol. Rev. 2002, 26, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Kovacic, F.; Babic, N.; Krauss, U.; Jaeger, K. Classification of lipolytic enzymes from bacteria. In Aerobic Utilization of Hydrocarbons, Oils and Lipids. Handbook of Hydrocarbon and Lipid Microbiology; Rojo, F., Ed.; Springer: Cham, Switzerland, 2019; pp. 1–35. ISBN 9783319397825. [Google Scholar]
- Rahman, M.A.; Culsum, U.; Tang, W.; Zhang, S.W.; Wu, G.; Liu, Z. Characterization of a novel cold active and salt tolerant esterase from Zunongwangia profunda. Enzyme Microb. Technol. 2016, 85, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Jeon, J.H.; Kim, J.T.; Lee, H.S.; Kim, S.J.; Kang, S.G.; Choi, S.H. Novel lipolytic enzymes identified from metagenomic library of deep-sea sediment. Evid. Based Complement. Altern. Med. 2011, 2011. [Google Scholar] [CrossRef]
- Hu, Y.; Fu, C.; Huang, Y.; Yin, Y.; Cheng, G.; Lei, F.; Lu, N.; Li, J.; Ashforth, E.J.; Zhang, L.; et al. Novel lipolytic genes from the microbial metagenomic library of the South China Sea marine sediment. FEMS Microbiol. Ecol. 2010, 72, 228–237. [Google Scholar] [CrossRef] [PubMed]
- Fang, Z.; Li, J.; Wang, Q.; Fang, W.; Peng, H.; Zhang, X.; Xiao, Y. A novel esterase from a marine metagenomic library exhibiting salt tolerance ability. J. Microbiol. Biotechnol. 2014, 24, 771–780. [Google Scholar] [CrossRef] [PubMed]
- Asyari, M.; Aditiawati, P.; Akhmaloka; Hertadi, R. Cloning and sequence analysis of lipase gene of halophilic bacteria isolated from mud crater of Bledug Kuwu, Central Java, Indonesia. Biosci. Biotechnol. Res. Asia 2015, 12, 1903–1912. [Google Scholar] [CrossRef][Green Version]
- Jeon, J.H.; Lee, H.S.; Kim, J.T.; Kim, S.J.; Choi, S.H.; Kang, S.G.; Lee, J.H. Identification of a new subfamily of salt-tolerant esterases from a metagenomic library of tidal flat sediment. Appl. Microbiol. Biotechnol. 2012, 93, 623–631. [Google Scholar] [CrossRef]
- Jayanath, G.; Mohandas, S.P.; Kachiprath, B.; Solomon, S.; Sajeevan, T.P.; Singh, I.S.B.; Philip, R. A novel solvent tolerant esterase of GDSGG motif subfamily from solar saltern through metagenomic approach: Recombinant expression and characterization. Int. J. Biol. Macromol. 2018, 119, 393–401. [Google Scholar] [CrossRef]
- Li, P.-Y.; Zhang, Y.; Xie, B.-B.; Zhang, Y.-Q.; Hao, J.; Wang, Y.; Wang, P.; Li, C.-Y.; Qin, Q.-L.; Zhang, X.-Y.; et al. Structural and mechanistic insights into the improvement of the halotolerance of a marine microbial esterase by increasing intra- and interdomain hydrophobic interactions. Appl. Environ. Microbiol. 2017, 83, e01286-17. [Google Scholar] [CrossRef]
- Mandrich, L.; de Pascale, D. An overview on thermal adaptation of esterases and lipases belonging to the HSL family: New insight on the computational analysis. Curr. Chem. Biol. 2011, 5, 17–28. [Google Scholar] [CrossRef]
- Mandrich, L.; Menchise, V.; Alterio, V.; De Simone, G.; Pedone, C.; Rossi, M.; Manco, G. Functional and structural features of the oxyanion hole in a thermophilic esterase from Alicyclobacillus acidocaldarius. Proteins Struct. Funct. Genet. 2008, 71, 1721–1731. [Google Scholar] [CrossRef]
- Li, P.Y.; Ji, P.; Li, C.Y.; Zhang, Y.; Wang, G.L.; Zhang, X.Y.; Xie, B.B.; Qin, Q.L.; Chen, X.L.; Zhou, B.C.; et al. Structural basis for dimerization and catalysis of a novel sterase from the GTSAG motif subfamily of the bacterial hormone-sensitive lipase family. J. Biol. Chem. 2014, 289, 19031–19041. [Google Scholar] [CrossRef] [PubMed]
- Chahinian, H.; Ali, Y.B.; Abousalham, A.; Petry, S.; Mandrich, L.; Manco, G.; Canaan, S.; Sarda, L. Substrate specificity and kinetic properties of enzymes belonging to the hormone-sensitive lipase family: Comparison with non-lipolytic and lipolytic carboxylesterases. Biochim. Biophys. Acta 2005, 1738, 29–36. [Google Scholar] [CrossRef]
- Rao, L.; Zhao, X.; Li, Y.; Xue, Y.; Ma, Y.; Lu, J.R. Solution behavior and activity of a halophilic esterase under high salt concentration. PLoS ONE 2009, 4, e6980. [Google Scholar] [CrossRef] [PubMed]
- Alcaide, M.; Stogios, P.J.; Lafraya, Á.; Tchigvintsev, A.; Flick, R.; Bargiela, R.; Chernikova, T.N.; Reva, O.N.; Hai, T.; Leggewie, C.C.; et al. Pressure adaptation is linked to thermal adaptation in salt-saturated marine habitats. Environ. Microbiol. 2015, 17, 332–345. [Google Scholar] [CrossRef] [PubMed]
- Ai, L.; Huang, Y.; Wang, C. Purification and characterization of halophilic lipase of Chromohalobacter sp. from ancient salt well. J. Basic Microbiol. 2018, 58, 647–657. [Google Scholar] [CrossRef] [PubMed]
- Castilla, A.; Panizza, P.; Rodríguez, D.; Bonino, L.; Díaz, P.; Irazoqui, G.; Giordano, S.R. A novel thermophilic and halophilic esterase from Janibacter sp. R02, the first member of a new lipase family (Family XVII). Enzyme Microb. Technol. 2017, 98, 86–95. [Google Scholar] [CrossRef] [PubMed]
- Glogauer, A.; Martini, V.P.; Faoro, H.; Couto, G.H.; Müller-Santos, M.; Monteiro, R.A.; Mitchell, D.A.; de Souza, E.M.; Pedrosa, F.O.; Krieger, N. Identification and characterization of a new true lipase isolated through metagenomic approach. Microb. Cell Fact. 2011, 10, 54. [Google Scholar] [CrossRef]
- Mohamed, Y.M.; Ghazy, M.A.; Sayed, A.; Ouf, A.; El-Dorry, H.; Siam, R. Isolation and characterization of a heavy metal-resistant, thermophilic esterase from a Red Sea Brine Pool. Sci. Rep. 2013, 3, 3358. [Google Scholar] [CrossRef]
- Borchert, E.; Selvin, J.; Kiran, S.G.; Jackson, S.A.; O’Gara, F.; Dobson, A.D.W. A novel cold active esterase from a deep sea sponge Stelletta normani metagenomic library. Front. Mar. Sci. 2017, 4, 1–13. [Google Scholar] [CrossRef]
- Marhuenda-Egea, F.C.; Bonete, M.J. Extreme halophilic enzymes in organic solvents. Curr. Opin. Biotechnol. 2002, 13, 385–389. [Google Scholar] [CrossRef]
- Pérez, D.; Martín, S.; Fernández-Lorente, G.; Filice, M.; Guisán, J.M.; Ventosa, A.; García, M.T.; Mellado, E. A novel halophilic lipase, LipBL, showing high efficiency in the production of eicosapentaenoic acid (EPA). PLoS ONE 2011, 6, e23325. [Google Scholar] [CrossRef]
- Oren, A. Industrial and environmental applications of halophilic microorganisms. Environ. Technol. 2010, 31, 825–834. [Google Scholar] [CrossRef] [PubMed]
- Ryckeboer, J.; Mergaert, J.; Vaes, K.; Klammer, S.; Clercq, D.; Coosemans, J.; Insam, H.; Swings, J. A survey of bacteria and fungi occurring during composting and self-heating processes. Ann. Microbiol. 2003, 53, 349–410. [Google Scholar]
- Kang, C.H.; Oh, K.H.; Lee, M.H.; Oh, T.K.; Kim, B.H.; Yoon, J.H. A novel family VII esterase with industrial potential from compost metagenomic library. Microb. Cell Fact. 2011, 10, 41. [Google Scholar] [CrossRef]
- Ilmberger, N.; Meske, D.; Juergensen, J.; Schulte, M.; Barthen, P.; Rabausch, U.; Angelov, A.; Mientus, M.; Liebl, W.; Schmitz, R.A.; et al. Metagenomic cellulases highly tolerant towards the presence of ionic liquids—Linking thermostability and halotolerance. Appl. Microbiol. Biotechnol. 2012, 95, 135–146. [Google Scholar] [CrossRef] [PubMed]
- Dougherty, M.J.; D’haeseleer, P.; Hazen, T.C.; Simmons, B.A.; Adams, P.D.; Hadi, M.Z. Glycoside hydrolases from a targeted compost metagenome, activity-screening and functional characterization. BMC Biotechnol. 2012, 12, 38. [Google Scholar] [CrossRef]
- Lu, M.; Dukunde, A.; Daniel, R. Biochemical profiles of two thermostable and organic solvent–tolerant esterases derived from a compost metagenome. Appl. Microbiol. Biotechnol. 2019, 103, 3421–3437. [Google Scholar] [CrossRef]
- Ishikawa, J.; Hotta, K. FramePlot: A new implementation of the frame analysis for predicting protein-coding regions in bacterial DNA with a high G + C content. FEMS Microbiol. Lett. 1999, 174, 251–263. [Google Scholar] [CrossRef]
- Bendtsen, J.D.; Nielsen, H.; Von Heijne, G.; Brunak, S. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 2004, 340, 783–795. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; McGinnis, S.; Madden, T.L. BLAST: Improvements for better sequence analysis. Nucleic Acids Res. 2006, 34, W6–W9. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinform. 2008, 9, 40. [Google Scholar] [CrossRef] [PubMed]
- Robert, X.; Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 2014, 42, 320–324. [Google Scholar] [CrossRef] [PubMed]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Lineweaver, H.; Burk, D. The determination of enzyme dissociation constants. J. Am. Chem. Soc. 1934, 56, 658–666. [Google Scholar] [CrossRef]
- Fitter, J.; Haber-Pohlmeier, S. Structural stability and unfolding properties of thermostable bacterial α-amylases: A comparative study of homologous enzymes. Biochemistry 2004, 43, 9589–9599. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Asnicar, F.; Weingart, G.; Tickle, T.L.; Huttenhower, C.; Segata, N. Compact graphical representation of phylogenetic data and metadata with GraPhlAn. PeerJ 2015, 3, e1029. [Google Scholar] [CrossRef]
- Loukas, A.; Kappas, I.; Abatzopoulos, T.J. HaloDom: A new database of halophiles across all life domains. J. Biol. Res. 2018, 25, 2. [Google Scholar] [CrossRef] [PubMed]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein identification and analysis tools on the ExPASy server. In The Proteomics Protocols Handbook; Humana Press: Totowa, NJ, USA, 2005; pp. 571–607. [Google Scholar]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; Mcglinn, D.; Minchin, P.R.; O’hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. Available online: https://cran.r-project.org/web/packages/vegan/vegan.pdf (accessed on 30 October 2018).
- Baker, N.A.; Sept, D.; Joseph, S.; Holst, M.J.; McCammon, J.A. Electrostatics of nanosystems: Application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 2001, 98, 10037–10041. [Google Scholar] [CrossRef]
- Lee, S.W.; Won, K.; Lim, H.K.; Kim, J.C.; Choi, G.J.; Cho, K.Y. Screening for novel lipolytic enzymes from uncultured soil microorganisms. Appl. Microbiol. Biotechnol. 2004, 65, 720–726. [Google Scholar] [CrossRef]
- Jin, P.; Pei, X.; Du, P.; Yin, X.; Xiong, X.; Wu, H.; Zhou, X.; Wang, Q. Overexpression and characterization of a new organic solvent-tolerant esterase derived from soil metagenomic DNA. Bioresour. Technol. 2012, 116, 234–240. [Google Scholar] [CrossRef] [PubMed]
- Dukunde, A.; Schneider, D.; Lu, M.; Brady, S.; Daniel, R. A novel, versatile family IV carboxylesterase exhibits high stability and activity in a broad pH spectrum. Biotechnol. Lett. 2017, 39, 577–587. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Zhang, K.; Han, W. Cloning and biochemical characterization of a novel lipolytic gene from activated sludge metagenome, and its gene product. Microb. Cell Fact. 2010, 9, 83. [Google Scholar] [CrossRef]
- Lindgren, M.; Westlund, P.-O. On the stability of chymotrypsin inhibitor 2 in a 10 M urea solution. The role of interaction energies for urea-induced protein denaturation. Phys. Chem. Chem. Phys. 2010, 12, 9358. [Google Scholar] [CrossRef]
- Peng, Q.; Zhang, X.; Shang, M.; Wang, X.; Wang, G.; Li, B.; Guan, G.; Li, Y.; Wang, Y. A novel esterase gene cloned from a metagenomic library from neritic sediments of the South China Sea. Microb. Cell Fact. 2011, 10, 95. [Google Scholar] [CrossRef]
- Esteban-Torres, M.; Santamaría, L.; de las Rivas, B.; Muñoz, R. Characterisation of a cold-active and salt-tolerant esterase from Lactobacillus plantarum with potential application during cheese ripening. Int. Dairy J. 2014, 39, 312–315. [Google Scholar] [CrossRef]
- Li, P.; Chen, X.; Ji, P.; Li, C.; Wang, P.; Zhang, Y.; Xie, B.; Qin, Q.; Su, H.; Zhou, B.; et al. Interdomain hydrophobic interactions modulate the thermostability of microbial esterases from the hormone-sensitive lipase family. J. Biol. Chem. 2015, 290, 11188–11198. [Google Scholar] [CrossRef]
- Khan, M.; Sathya, T.A. Extremozymes from metagenome: Potential applications in food processing. Crit. Rev. Food Sci. Nutr. 2018, 58, 2017–2025. [Google Scholar] [CrossRef] [PubMed]
- Simon, C.; Daniel, R. Metagenomic analyses: Past and future trends. Appl. Environ. Microbiol. 2011, 77, 1153–1161. [Google Scholar] [CrossRef] [PubMed]
- Huo, Y.Y.; Rong, Z.; Jian, S.L.; Xu, C.D.; Li, J.; Xu, X.W. A novel halotolerant thermoalkaliphilic esterase from marine bacterium Erythrobacter seohaensis SW-135. Front. Microbiol. 2017, 8, 2315. [Google Scholar] [CrossRef]
- Wu, G.; Zhang, S.; Zhang, H.; Zhang, S.; Liu, Z. A novel esterase from a psychrotrophic bacterium Psychrobacter celer 3Pb1 showed cold-adaptation and salt-tolerance. J. Mol. Catal. B Enzym. 2013, 98, 119–126. [Google Scholar] [CrossRef]
- Wang, G.; Wang, Q.; Lin, X.; Ng, T.B.; Yan, R.; Lin, J.; Ye, X. A novel cold-adapted and highly salt-tolerant esterase from Alkalibacterium sp. SL3 from the sediment of a soda lake. Sci. Rep. 2016, 6, 19494. [Google Scholar] [CrossRef] [PubMed]
- Tchigvintsev, A.; Tran, H.; Popovic, A.; Kovacic, F.; Brown, G.; Flick, R.; Hajighasemi, M.; Egorova, O.; Somody, J.C.; Tchigvintsev, D.; et al. The environment shapes microbial enzymes: Five cold-active and salt-resistant carboxylesterases from marine metagenomes. Appl. Microbiol. Biotechnol. 2015, 99, 2165–2178. [Google Scholar] [CrossRef] [PubMed]
- Selvin, J.; Kennedy, J.; Lejon, D.P.H.; Kiran, G.S.; Dobson, A.D.W. Isolation identification and biochemical characterization of a novel halo-tolerant lipase from the metagenome of the marine sponge Haliclona simulans. Microb. Cell Fact. 2012, 11, 72. [Google Scholar] [CrossRef]
- Zhang, Y.; Hao, J.; Zhang, Y.Q.; Chen, X.L.; Xie, B.B.; Shi, M.; Zhou, B.C.; Zhang, Y.Z.; Li, P.Y. Identification and characterization of a novel salt-tolerant esterase from the deep-sea sediment of the South China Sea. Front. Microbiol. 2017, 8, 441. [Google Scholar] [CrossRef]
- Alcaide, M.; Tchigvintsev, A.; Martínez-Martínez, M.; Popovic, A.; Reva, O.N.; Lafraya, Á.; Bargiela, R.; Nechitaylo, T.Y.; Matesanz, R.; Cambon-Bonavita, M.-A.; et al. Identification and characterization of carboxyl esterases of gill chamber-associated microbiota in the deep-sea shrimp Rimicaris exoculata by using functional metagenomics. Appl. Environ. Microbiol. 2015, 81, 2125–2136. [Google Scholar] [CrossRef]
- Oliveira, L.C.G.; Ramos, P.L.; Marem, A.; Kondo, M.Y.; Rocha, R.C.S.; Bertolini, T.; Silveira, M.A.; da Cruz, J.B.; de Vasconcellos, S.P.; Juliano, L.; et al. Halotolerant bacteria in the São Paulo Zoo composting process and their hydrolases and bioproducts. Braz. J. Microbiol. 2015, 46, 347–354. [Google Scholar] [CrossRef]
- Chandna, P.; Mayilraj, S.; Kuhad, R.C. Bacillus pseudoflexus sp. nov., a moderately halophilic bacterium isolated from compost. Ann. Microbiol. 2016, 66, 895–905. [Google Scholar] [CrossRef]
- De Santi, C.; Leiros, H.K.S.; Di Scala, A.; de Pascale, D.; Altermark, B.; Willassen, N.P. Biochemical characterization and structural analysis of a new cold-active and salt-tolerant esterase from the marine bacterium Thalassospira sp. Extremophiles 2016, 20, 323–336. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.; Huo, Y.; Cheng, H.; Zhang, X.; Zhu, X.; Wu, M. Cloning, expression and characterization of a halotolerant esterase from a marine bacterium Pelagibacterium halotolerans B2T. Extremophiles 2012, 16, 427–435. [Google Scholar] [CrossRef] [PubMed]
- Leite, A.E.T.; Briganti, L.; de Araújo, E.A.; Pellegrini, V.D.; Camilo, C.M.; Polikarpov, I. Low-resolution molecular shape, biochemical characterization and emulsification properties of a halotolerant esterase from Bacillus licheniformis. Eur. Biophys. J. 2020, 49, 435–447. [Google Scholar] [CrossRef]
- Wu, G.; Wu, G.; Zhan, T.; Shao, Z.; Liu, Z. Characterization of a cold-adapted and salt-tolerant esterase from a psychrotrophic bacterium Psychrobacter pacificensis. Extremophiles 2013, 17, 809–819. [Google Scholar] [CrossRef]
- Adıgüzel, A.O. Production and characterization of thermo-, halo- and solvent-stable esterase from Bacillus mojavensis TH309. Biocatal. Biotransform. 2020, 38, 210–226. [Google Scholar] [CrossRef]
- De Santi, C.; Altermark, B.; Pierechod, M.M.; Ambrosino, L.; de Pascale, D.; Willassen, N.-P. Characterization of a cold-active and salt tolerant esterase identified by functional screening of Arctic metagenomic libraries. BMC Biochem. 2016, 17. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Xu, H.; Wu, Y.; Zeng, J.; Guo, Z.; Wang, L.; Shen, C.; Qiao, D.; Cao, Y. A new cold-adapted, alkali-stable and highly salt-tolerant esterase from Bacillus licheniformis. Int. J. Biol. Macromol. 2018, 111, 1183–1193. [Google Scholar] [CrossRef]
- Hang, Y.; Ran, S.; Wang, X.; Jiao, J.; Wang, S.; Liu, Z. Mutational analysis and stability characterization of a novel esterase of lipolytic enzyme family VI from Shewanella sp. Int. J. Biol. Macromol. 2016, 93, 655–664. [Google Scholar] [CrossRef]
- Lee, C.W.; Kwon, S.; Park, S.H.; Kim, B.Y.; Yoo, W.; Ryu, B.H.; Kim, H.W.; Shin, S.C.; Kim, S.; Park, H.; et al. Crystal structure and functional characterization of an esterase (EaEST) from Exiguobacterium antarcticum. PLoS ONE 2017, 12, e0169540. [Google Scholar] [CrossRef]
- Yang, X.; Wu, L.; Xu, Y.; Ke, C.; Hu, F.; Xiao, X.; Huang, J. Identification and characterization of a novel alkalistable and salt-tolerant esterase from the deep-sea hydrothermal vent of the East Pacific Rise. Microbiology 2018, 7, e00601. [Google Scholar] [CrossRef]
- Dennis, P.P.; Shimmin, L.C. Evolutionary divergence and salinity-mediated selection in halophilic archaea. Microbiol. Mol. Biol. Rev. 1997, 61, 90–104. [Google Scholar] [CrossRef] [PubMed]
- Margesin, R.; Schinner, F. Potential of halotolerant and halophilic microorganisms for biotechnology. Extremophiles 2001, 5, 73–83. [Google Scholar] [CrossRef] [PubMed]
- Lanyi, J.K. Salt-dependent properties of proteins from extremely halophilic bacteria. Bacteriol. Rev. 1974, 38, 272–290. [Google Scholar] [CrossRef]
- Madern, D.; Ebel, C.; Zaccai, G. Halophilic adaptation of enzymes. Extremophiles 2000, 4, 91–98. [Google Scholar] [CrossRef]
- Hutcheon, G.W.; Vasisht, N.; Bolhuis, A. Characterisation of a highly stable α-amylase from the halophilic archaeon Haloarcula hispanica. Extremophiles 2005, 9, 487–495. [Google Scholar] [CrossRef] [PubMed]
- Kastritis, P.L.; Papandreou, N.C.; Hamodrakas, S.J. Haloadaptation: Insights from comparative modeling studies of halophilic archaeal DHFRs. Int. J. Biol. Macromol. 2007, 41, 447–453. [Google Scholar] [CrossRef]
- Madern, D.; Pfister, C.; Zaccai, G. Mutation at a single acidic amino acid enhances the halophilic behaviour of malate dehydrogenase from Haloarcula marismortui in physiological salts. Eur. J. Biochem. 1995, 230, 1088–1095. [Google Scholar] [CrossRef]
- Paul, S.; Bag, S.K.; Das, S.; Harvill, E.T.; Dutta, C. Molecular signature of hypersaline adaptation: Insights from genome and proteome composition of halophilic prokaryotes. Genome Biol. 2008, 9, R70. [Google Scholar] [CrossRef]
- Winter, J.A.; Christofi, P.; Morroll, S.; Bunting, K.A. The crystal structure of Haloferax volcanii proliferating cell nuclear antigen reveals unique surface charge characteristics due to halophilic adaptation. BMC Struct. Biol. 2009, 9, 1–15. [Google Scholar] [CrossRef]
- Tadeo, X.; Ló Pez-Mé Ndez, B.; Trigueros, T.; Laín, A.; Castañ, D.; Millet, O. Structural basis for the amino acid composition of proteins from halophilic Archea. PLoS Biol. 2009, 7, e1000257. [Google Scholar] [CrossRef] [PubMed]
- Munawar, N.; Engel, P.C. Overexpression in a non-native halophilic host and biotechnological potential of NAD+-dependent glutamate dehydrogenase from Halobacterium salinarum strain NRC-36014. Extremophiles 2012, 16, 463–476. [Google Scholar] [CrossRef] [PubMed]
- Coquelle, N.; Talon, R.; Juers, D.H.; Girard, É.; Kahn, R.; Madern, D. Gradual adaptive changes of a protein facing high salt concentrations. J. Mol. Biol. 2010, 404, 493–505. [Google Scholar] [CrossRef] [PubMed]
- Oren, A.; Gavrieli, I.; Gavrieli, J.; Kohen, M.; Lati, J.; Aharoni, M. Microbial communities in the Dead Sea—Past, present and future. In Adaptation to Life at High Salt Concentrations in Archaea, Bacteria, and Eukarya. Cellular Origin, Life in Extreme Habitats and Astrobiology; Gunde-Cimerman, N., Oren, A., Plemenitaš, A., Eds.; Springer: Berlin/Heidelberg, Germany, 2005; Volume 9, pp. 27–39. ISBN 1402036337. [Google Scholar]
- Bieger, B.; Essen, L.-O.; Oesterhelt, D. Crystal structure of halophilic dodecin: A novel, dodecameric flavin binding protein from Halobacterium salinarum. Structure 2003, 11, 375–385. [Google Scholar] [CrossRef]
- Altermark, B.; Helland, R.; Moe, E.; Willassen, N.P.; Smalås, A.O. Structural adaptation of endonuclease I from the cold-adapted and halophilic bacterium Vibrio salmonicida. Acta Crystallogr. Sect. D Biol. Crystallogr. 2008, 64, 368–376. [Google Scholar] [CrossRef] [PubMed]
- Sivakumar, N.; Li, N.; Tang, J.W.; Patel, B.K.C.; Swaminathan, K. Crystal structure of AmyA lacks acidic surface and provide insights into protein stability at poly-extreme condition. FEBS Lett. 2006, 580, 2646–2652. [Google Scholar] [CrossRef]
- Kim, T.D. Bacterial hormone-sensitive lipases (bHSLs): Emerging enzymes for biotechnological applications. J. Microbiol. Biotechnol. 2017, 27, 1907–1915. [Google Scholar] [CrossRef] [PubMed]
- Noby, N.; Hussein, A.; Saeed, H.; Embaby, A.M. Recombinant cold-adapted halotolerant, organic solvent-stable esterase (estHIJ) from Bacillus halodurans. Anal. Biochem. 2020, 591, 113554. [Google Scholar] [CrossRef]
- Panda, T.; Gowrishankar, B.S. Production and applications of esterases. Appl. Microbiol. Biotechnol. 2005, 67, 160–169. [Google Scholar] [CrossRef]
- Hasan, F.; Shah, A.A.; Hameed, A. Industrial applications of microbial lipases. Enzyme Microb. Technol. 2006, 39, 235–251. [Google Scholar] [CrossRef]
- Karan, R.; Capes, M.D.; DasSarma, S. Function and biotechnology of extremophilic enzymes in low water activity. Aquat. Biosyst. 2012, 8, 4. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Lopez, O.; Cerdan, M.; Siso, M. New extremophilic lipases and esterases from metagenomics. Curr. Protein Pept. Sci. 2014, 15, 445–455. [Google Scholar] [CrossRef] [PubMed]
- Rathi, P.; Bradoo, S.; Saxena, R.K.; Gupta, R. A hyper-thermostable, alkaline lipase from Pseudomonas sp. with the property of thermal activation. Biotechnol. Lett. 2000, 22, 495–498. [Google Scholar] [CrossRef]
- Castro-Ochoa, L.D.; Rodríguez-Gómez, C.; Valerio-Alfaro, G.; Oliart Ros, R. Screening, purification and characterization of the thermoalkalophilic lipase produced by Bacillus thermoleovorans CCR11. Enzyme Microb. Technol. 2005, 37, 648–654. [Google Scholar] [CrossRef]
- Kamarudin, N.H.A.; Rahman, R.N.Z.R.A.; Ali, M.S.M.; Leow, T.C.; Basri, M.; Salleh, A.B. A new cold-adapted, organic solvent stable lipase from mesophilic Staphylococcus epidermidis AT2. Protein J. 2014, 33, 296–307. [Google Scholar] [CrossRef] [PubMed]
| Residual Relative Activity (%) a | |||||
|---|---|---|---|---|---|
| 0 M | 1 M | 2 M | 3 M | 4 M | |
| Temperature b | |||||
| 50 °C | 54.9 ± 2.4 | 71.3 ± 8.1 * | 82.8 ± 7.4 ** | 72.9 ± 4.2 * | 38.9 ± 3.1 ** |
| 60 °C | 1.4 ± 0.7 | 12.7 ± 2.2 ** | 11.8 ± 1.6 ** | 8.0 ± 0.7 ** | 10.6 ± 0.6 ** |
| Urea c | |||||
| 2 M | 36.3 ± 4.7 | 36.2 ± 4.0 | 43.0 ± 4.7 | 54.0 ± 2.4 ** | 41.5 ± 9.5 |
| 4 M | 19.5 ± 3.5 | 31.6 ± 2.5 ** | 33.6 ± 2.2 ** | 29.3 ± 4.0 * | ND d |
| 6 M | 17.2 ± 2.4 | 28.6 ± 2.2 ** | ND d | ND d | ND d |
| Organic Solvent | Residual Activity (%) a | |
|---|---|---|
| 15% (v/v) | 30% (v/v) | |
| DMSO | 135.0 ± 11.8 | 135.4 ± 4.7 |
| Methanol | 116.1 ± 6.7 | 96.6 ± 7.6 |
| Ethanol | 116.2 ± 8.7 | 107.3 ± 4.2 |
| Acetone | 138.3 ± 8.9 | 115.3 ± 14.8 |
| Isopropanol | 136.3 ± 10.1 | 286.6 ± 9.0 |
| 1-Propanol | 114.0 ± 3.7 | 23.0 ± 1.0 |
| Organic Solvent | Residual Activity (%) a | |
|---|---|---|
| 15% (v/v) | 30% (v/v) | |
| Ethyl acetate | 55.3 ± 8.1 | 50.2 ± 3.5 |
| Diethyl ether | 110.8 ± 5.7 | 106.7 ± 10.8 |
| Chloroform | 46.5 ± 0.5 | 6.3 ± 2.5 |
| Toluene | 50.0 ± 2.2 | 54.9 ± 0.7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, M.; Daniel, R. A Novel Carboxylesterase Derived from a Compost Metagenome Exhibiting High Stability and Activity towards High Salinity. Genes 2021, 12, 122. https://doi.org/10.3390/genes12010122
Lu M, Daniel R. A Novel Carboxylesterase Derived from a Compost Metagenome Exhibiting High Stability and Activity towards High Salinity. Genes. 2021; 12(1):122. https://doi.org/10.3390/genes12010122
Chicago/Turabian StyleLu, Mingji, and Rolf Daniel. 2021. "A Novel Carboxylesterase Derived from a Compost Metagenome Exhibiting High Stability and Activity towards High Salinity" Genes 12, no. 1: 122. https://doi.org/10.3390/genes12010122
APA StyleLu, M., & Daniel, R. (2021). A Novel Carboxylesterase Derived from a Compost Metagenome Exhibiting High Stability and Activity towards High Salinity. Genes, 12(1), 122. https://doi.org/10.3390/genes12010122





