Comparative Genomic Analysis of 450 Strains of Salmonella enterica Isolated from Diseased Animals
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains
2.2. Genome Sequencing, Assembly and Annotation
2.3. Detection of SPIs and Virulence spvRABCD Operon
2.4. Identification of Resistance Genes, Plasmid Types, and SGI
2.5. Submission and Accession Number
3. Results
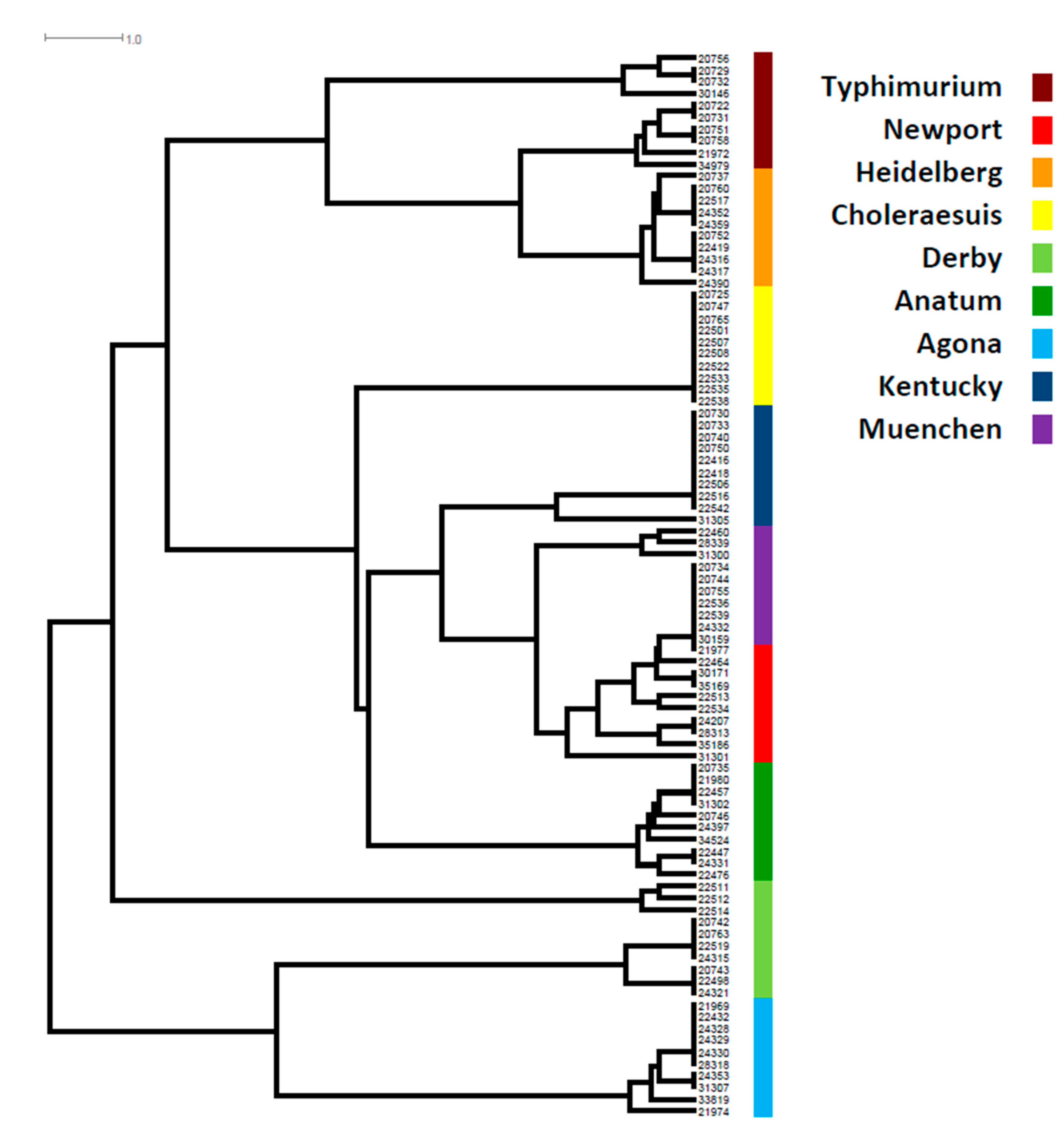
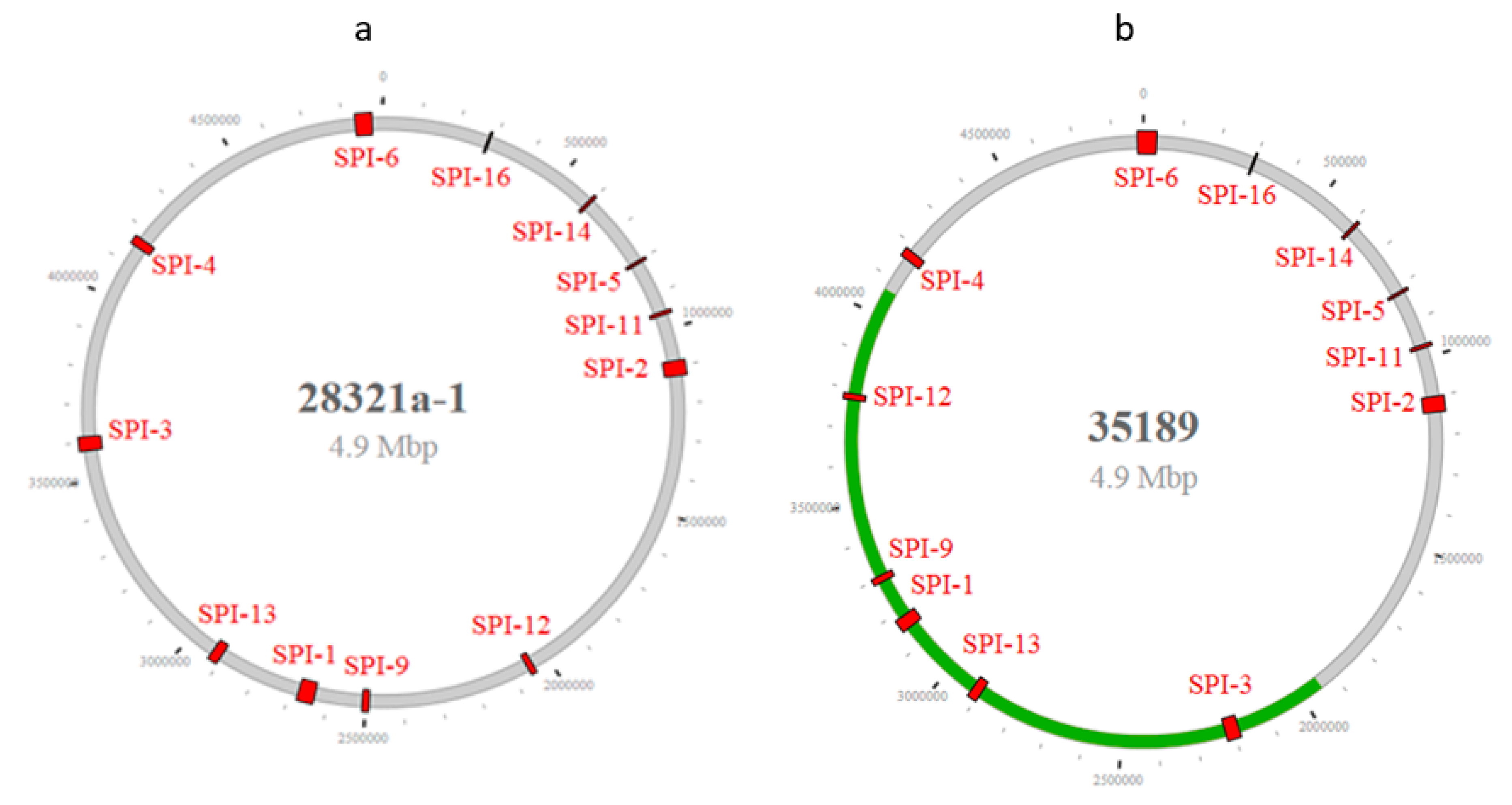
3.1. The SPI Profile, Distribution, and Diversity
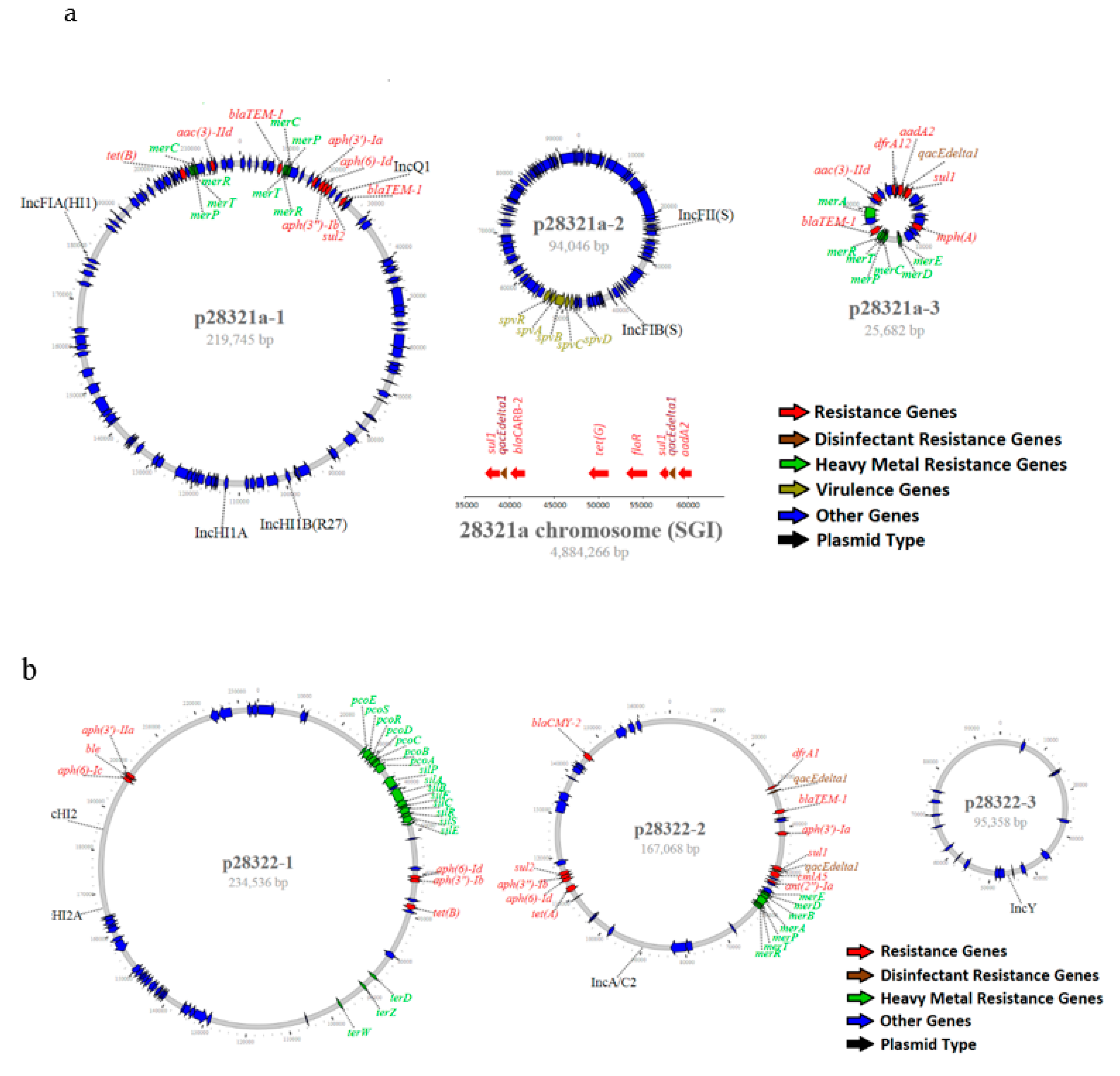
3.2. Resistance Genes
3.3. Plasmid Type, Structure and Association with Salmonella Serotypes and Source of Isolates
3.4. Prevalence of SGI and spv Virulence Operon
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Data Availability
Disclaimer
References
- Chan, K.; Baker, S.; Kim, C.C.; Detweiler, C.S.; Dougan, G.; Falkow, S. Genomic comparison of Salmonella enterica serovars and Salmonella bongori by use of an S. enterica serovar typhimurium DNA microarray. J. Bacteriol. 2003, 185, 553–563. [Google Scholar] [CrossRef]
- Kingsley, R.A.; Msefula, C.L.; Thomson, N.R.; Kariuki, S.; Holt, K.E.; Gordon, M.A.; Harris, D.; Clarke, L.; Whitehead, S.; Sangal, V.; et al. Epidemic multiple drug resistant Salmonella Typhimurium causing invasive disease in sub-Saharan Africa have a distinct genotype. Genome Res. 2009, 19, 2279–2287. [Google Scholar] [CrossRef]
- Libby, S.J.; Adams, L.G.; Ficht, T.A.; Allen, C.; Whitford, H.A.; Buchmeier, N.A.; Bossie, S.; Guiney, D.G. The spv genes on the Salmonella dublin virulence plasmid are required for severe enteritis and systemic infection in the natural host. Infect. Immun. 1997, 65, 1786–1792. [Google Scholar] [CrossRef]
- Sabbagh, S.C.; Forest, C.G.; Lepage, C.; Leclerc, J.M.; Daigle, F. So similar, yet so different: Uncovering distinctive features in the genomes of Salmonella enterica serovars Typhimurium and Typhi. FEMS Microbiol. Lett. 2010, 305, 1–13. [Google Scholar] [CrossRef]
- Stevens, M.P.; Humphrey, T.J.; Maskell, D.J. Molecular insights into farm animal and zoonotic Salmonella infections. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2009, 364, 2709–2723. [Google Scholar] [CrossRef]
- Gerlach, R.G.; Hensel, M. Salmonella pathogenicity islands in host specificity, host pathogen-interactions and antibiotics resistance of Salmonella enterica. Berl. Munch. Tierarztl. Wochenschr. 2007, 120, 317–327. [Google Scholar]
- Hensel, M. Evolution of pathogenicity islands of Salmonella enterica. Int. J. Med. Microbiol. 2004, 294, 95–102. [Google Scholar] [CrossRef]
- Rychlik, I.; Karasova, D.; Sebkova, A.; Volf, J.; Sisak, F.; Havlickova, H.; Kummer, V.; Imre, A.; Szmolka, A.; Nagy, B. Virulence potential of five major pathogenicity islands (SPI-1 to SPI-5) of Salmonella enterica serovar Enteritidis for chickens. BMC Microbiol. 2009, 9, 268. [Google Scholar] [CrossRef]
- Suez, J.; Porwollik, S.; Dagan, A.; Marzel, A.; Schorr, Y.I.; Desai, P.T.; Agmon, V.; McClelland, M.; Rahav, G.; Gal-Mor, O. Virulence gene profiling and pathogenicity characterization of non-typhoidal Salmonella accounted for invasive disease in humans. PLoS ONE 2013, 8, e58449. [Google Scholar] [CrossRef]
- Amavisit, P.; Lightfoot, D.; Browning, G.F.; Markham, P.F. Variation between pathogenic serovars within Salmonella pathogenicity islands. J. Bacteriol. 2003, 185, 3624–3635. [Google Scholar] [CrossRef]
- Groisman, E.A. Bacterial responses to host-defense peptides. Trends Microbiol. 1996, 4, 127–128; discussion 128–129. [Google Scholar] [CrossRef]
- Marcus, S.L.; Brumell, J.H.; Pfeifer, C.G.; Finlay, B.B. Salmonella pathogenicity islands: Big virulence in small packages. Microbes Infect. 2000, 2, 145–156. [Google Scholar] [CrossRef]
- Blondel, C.J.; Jimenez, J.C.; Contreras, I.; Santiviago, C.A. Comparative genomic analysis uncovers 3 novel loci encoding type six secretion systems differentially distributed in Salmonella serotypes. BMC Genom. 2009, 10, 354. [Google Scholar] [CrossRef]
- Hayward, M.R.; AbuOun, M.; La Ragione, R.M.; Tchorzewska, M.A.; Cooley, W.A.; Everest, D.J.; Petrovska, L.; Jansen, V.A.; Woodward, M.J. SPI-23 of S. Derby: Role in adherence and invasion of porcine tissues. PLoS ONE 2014, 9, e107857. [Google Scholar] [CrossRef] [PubMed]
- Fookes, M.; Schroeder, G.N.; Langridge, G.C.; Blondel, C.J.; Mammina, C.; Connor, T.R.; Seth-Smith, H.; Vernikos, G.S.; Robinson, K.S.; Sanders, M.; et al. Salmonella bongori provides insights into the evolution of the Salmonellae. PLoS Pathog. 2011, 7, e1002191. [Google Scholar] [CrossRef]
- Hsu, C.H.; Li, C.; Hoffmann, M.; McDermott, P.; Abbott, J.; Ayers, S.; Tyson, G.H.; Tate, H.; Yao, K.; Allard, M.; et al. Comparative Genomic Analysis of Virulence, Antimicrobial Resistance, and Plasmid Profiles of Salmonella Dublin Isolated from Sick Cattle, Retail Beef, and Humans in the United States. Microb. Drug Resist. 2019, 25, 1238–1249. [Google Scholar] [CrossRef] [PubMed]
- Guiney, D.G.; Fang, F.C.; Krause, M.; Libby, S. Plasmid-mediated virulence genes in non-typhoid Salmonella serovars. FEMS Microbiol. Lett. 1994, 124, 1–9. [Google Scholar] [CrossRef][Green Version]
- Guiney, D.G.; Fang, F.C.; Krause, M.; Libby, S.; Buchmeier, N.A.; Fierer, J. Biology and clinical significance of virulence plasmids in Salmonella serovars. Clin. Infect Dis. 1995, 21 (Suppl. 2), S146–S151. [Google Scholar] [CrossRef]
- Gulig, P.A.; Doyle, T.J. The Salmonella typhimurium virulence plasmid increases the growth rate of salmonellae in mice. Infect. Immun. 1993, 61, 504–511. [Google Scholar] [CrossRef]
- Haneda, T.; Okada, N.; Nakazawa, N.; Kawakami, T.; Danbara, H. Complete DNA sequence and comparative analysis of the 50-kilobase virulence plasmid of Salmonella enterica serovar Choleraesuis. Infect. Immun. 2001, 69, 2612–2620. [Google Scholar] [CrossRef]
- Rotger, R.; Casadesus, J. The virulence plasmids of Salmonella. Int. Microbiol. 1999, 2, 177–184. [Google Scholar] [PubMed]
- Zhao, S.; McDermott, P.F.; White, D.G.; Qaiyumi, S.; Friedman, S.L.; Abbott, J.W.; Glenn, A.; Ayers, S.L.; Post, K.W.; Fales, W.H.; et al. Characterization of multidrug resistant Salmonella recovered from diseased animals. Vet. Microbiol. 2007, 123, 122–132. [Google Scholar] [CrossRef] [PubMed]
- FDA. National Antimicrobial Resistance Monitoring System (NARMS): 2017 Report; U.S. Department of Health and Human Services, Food and Drug Administration: Rockville, MD, USA, 2017.
- Harbottle, H.; Thakur, S.; Zhao, S.; White, D.G. Genetics of antimicrobial resistance. Anim. Biotechnol. 2006, 17, 111–124. [Google Scholar] [CrossRef]
- McMillan, E.A.; Gupta, S.K.; Williams, L.E.; Jove, T.; Hiott, L.M.; Woodley, T.A.; Barrett, J.B.; Jackson, C.R.; Wasilenko, J.L.; Simmons, M.; et al. Antimicrobial Resistance Genes, Cassettes, and Plasmids Present in Salmonella enterica Associated With United States Food Animals. Front. Microbiol. 2019, 10, 832. [Google Scholar] [CrossRef] [PubMed]
- McDermott, P.F.; Zhao, S.; Tate, H. Antimicrobial Resistance in nontyphoidal Salmonella. In Antimicrobial Resistance in Bacteria from Livestock and Companion Animals; Aarestrup, F.M., Schwarz, S., Shen, J., Cavaco, L., Eds.; ASM Press: Washington, DC, USA, 2018. [Google Scholar]
- Folster, J.P.; Katz, L.; McCullough, A.; Parsons, M.B.; Knipe, K.; Sammons, S.A.; Boncy, J.; Tarr, C.L.; Whichard, J.M. Multidrug-resistant IncA/C plasmid in Vibrio cholerae from Haiti. Emerg Infect. Dis. 2014, 20, 1951–1953. [Google Scholar] [CrossRef] [PubMed]
- Tatusova, T.; DiCuccio, M.; Badretdin, A.; Chetvernin, V.; Nawrocki, E.P.; Zaslavsky, L.; Lomsadze, A.; Pruitt, K.D.; Borodovsky, M.; Ostell, J. NCBI prokaryotic genome annotation pipeline. Nucleic. Acids Res. 2016, 44, 6614–6624. [Google Scholar] [CrossRef]
- Hunt, M.; Silva, N.D.; Otto, T.D.; Parkhill, J.; Keane, J.A.; Harris, S.R. Circlator: Automated circularization of genome assemblies using long sequencing reads. Genome Biol. 2015, 16, 294. [Google Scholar] [CrossRef]
- Hall, R.M. Salmonella genomic islands and antibiotic resistance in Salmonella enterica. Future Microbiol. 2010, 5, 1525–1538. [Google Scholar] [CrossRef]
- Siebor, E.; Neuwirth, C. Proteus genomic island 1 (PGI1), a new resistance genomic island from two Proteus mirabilis French clinical isolates. J. Antimicrob. Chemother. 2014, 69, 3216–3220. [Google Scholar] [CrossRef]
- Monte, D.F.; Lincopan, N.; Berman, H.; Cerdeira, L.; Keelara, S.; Thakur, S.; Fedorka-Cray, P.J.; Landgraf, M. Genomic Features of High-Priority Salmonella enterica Serovars Circulating in the Food Production Chain, Brazil, 2000-2016. Sci. Rep. 2019, 9, 11058. [Google Scholar] [CrossRef]
- Silva, C.; Puente, J.L.; Calva, E. Salmonella virulence plasmid: Pathogenesis and ecology. Pathog. Dis. 2017. [Google Scholar] [CrossRef] [PubMed]
- Libby, S.J.; Lesnick, M.; Hasegawa, P.; Kurth, M.; Belcher, C.; Fierer, J.; Guiney, D.G. Characterization of the spv locus in Salmonella enterica serovar Arizona. Infect. Immun. 2002, 70, 3290–3294. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Mohammed, M.; Le Hello, S.; Leekitcharoenphon, P.; Hendriksen, R. The invasome of Salmonella Dublin as revealed by whole genome sequencing. BMC Infect. Dis. 2017, 17, 544. [Google Scholar] [CrossRef] [PubMed]
- Mangat, C.S.; Bekal, S.; Irwin, R.J.; Mulvey, M.R. A Novel Hybrid Plasmid Carrying Multiple Antimicrobial Resistance and Virulence Genes in Salmonella enterica Serovar Dublin. Antimicrob. Agents Chemother. 2017, 61. [Google Scholar] [CrossRef]
- Feng, Y.; Liu, J.; Li, Y.G.; Cao, F.L.; Johnston, R.N.; Zhou, J.; Liu, G.R.; Liu, S.L. Inheritance of the Salmonella virulence plasmids: Mostly vertical and rarely horizontal. Infect. Genet Evol. 2012, 12, 1058–1063. [Google Scholar] [CrossRef]
- Zhao, S.; Qaiyumi, S.; Friedman, S.; Singh, R.; Foley, S.L.; White, D.G.; McDermott, P.F.; Donkar, T.; Bolin, C.; Munro, S.; et al. Characterization of Salmonella enterica serotype newport isolated from humans and food animals. J. Clin. Microbiol. 2003, 41, 5366–5371. [Google Scholar] [CrossRef]
- Boyd, D.A.; Peters, G.A.; Ng, L.; Mulvey, M.R. Partial characterization of a genomic island associated with the multidrug resistance region of Salmonella enterica Typhymurium DT104. FEMS Microbiol. Lett. 2000, 189, 285–291. [Google Scholar] [CrossRef]
- de Curraize, C.; Amoureux, L.; Bador, J.; Chapuis, A.; Siebor, E.; Clement, C.; Sauge, J.; Aho-Glele, L.S.; Neuwirth, C. “Does the Salmonella Genomic Island 1 (SGI1) confer invasiveness properties to human isolates?”. BMC Infect. Dis. 2017, 17, 741. [Google Scholar] [CrossRef]
- Yang, S.; Deng, W.; Liu, S.; Yu, X.; Mustafa, G.R.; Chen, S.; He, L.; Ao, X.; Yang, Y.; Zhou, K.; et al. Presence of Heavy Metal Resistance Genes in Escherichia coli and Salmonella, and Analysis of Resistance Gene Structure in E. coli E308. J. Glob. Antimicrob. Resist. 2020. [Google Scholar] [CrossRef]
| Serotype | SPI 1 | SPI 2 | SPI 3 | SPI 4 | SPI 5 | SPI 6 | SPI 7 | SPI 8 | SPI 9 | SPI 10 | SPI 11 | SPI 12 | SPI 13 | SPI 14 | SPI 15 | SPI 16 | SPI 17 | SPI 18 | SPI 19 | SPI 20 | SPI 21 | SPI 22 | SPI 23 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Typhimurium (136) 2 | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Newport (43) | + | + | + | + | + | + | - | - | + | 5% | + | + | + | + | - | + | - | 2% | - | - | - | - | - |
| Heidelberg (38) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Choleraesuis (35) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Derby (29) | + | + | + | + | + | + | - | 17% | + | - | + | + | - | - | - | + | 17% | - | - | - | - | - | 83% |
| Anatum (16) | + | + | + | + | + | + | - | - | + | 87% | + | + | + | + | - | - | - | - | - | - | - | - | + |
| Agona (14) | + | + | + | + | + | + | - | + | + | - | + | + | - | - | - | + | - | - | + | - | - | - | + |
| Kentucky (13) | + | + | + | + | + | + | - | + | + | - | + | + | - | - | - | + | - | - | - | - | - | - | - |
| Muenchen (11) | + | + | + | + | + | + | 20% | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | 27% |
| Worthington (9) | + | + | + | + | + | + | - | + | + | - | + | + | - | - | - | + | - | - | - | - | - | - | - |
| Mbandaka (9) | + | + | + | + | + | + | - | - | + | - | + | + | - | - | - | + | - | - | - | - | - | - | - |
| Uganda (9) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | - | - | - | - | - | - | - | - |
| Senftenberg (8) | + | + | + | + | + | + | - | + | + | - | + | + | - | - | - | + | - | - | - | - | - | - | - |
| Oranienburg (7) | + | + | + | + | + | + | - | - | + | - | + | - | + | + | - | + | - | + | - | - | - | - | 86% |
| I 4,5,12: i: -(3) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Schwarzengrund (6) | + | + | + | + | + | + | - | - | + | - | + | - | + | + | - | + | - | + | - | - | - | - | - |
| Muenster (5) | + | + | + | + | + | + | - | - | + | + | + | - | + | + | - | - | - | + | - | - | - | - | + |
| Reading (5) | + | + | + | + | + | + | - | - | + | - | + | 80% | + | + | - | + | - | 20% | - | - | - | - | - |
| Montevideo (4) 3 | + | + | + | + | + | + | - | - | + | - | + | - | + | + | - | + | - | + | - | - | - | - | - |
| Enteritidis (4) | + | + | + | + | + | + | - | - | + | + | + | + | + | + | - | + | + | - | + | - | - | - | - |
| Saintpaul (4) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Give (4) | + | + | + | + | + | + | - | - | + | - | + | - | + | + | - | - | - | + | - | - | - | - | - |
| Bredeney (3) | + | + | + | + | + | + | - | - | + | - | + | - | + | + | - | + | - | + | - | - | - | - | - |
| Hadar (3) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Albany (2) | + | + | + | + | + | + | - | + | + | - | + | + | - | - | - | + | - | - | - | - | - | - | - |
| Braenderup (2) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Cerro (2) | + | + | + | + | + | + | - | - | + | - | + | + | - | - | - | + | - | - | - | - | - | - | - |
| Hartford (2) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Infantis (2) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Java (2) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Tennessee (2) | + | + | + | + | + | + | - | + | + | - | + | + | - | - | - | + | - | - | - | - | - | - | - |
| Bovismorbificans (1) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Cubana (1) | + | + | + | + | + | + | - | + | + | - | + | + | - | - | - | + | - | - | - | - | - | - | - |
| Gaminara (1) | + | + | + | + | + | + | - | - | + | - | + | - | + | + | - | - | - | + | - | - | - | - | - |
| Javiana (1) | + | + | + | + | + | + | - | - | + | - | + | - | + | + | - | + | - | + | - | - | - | - | - |
| Johannesburg (1) | + | + | + | + | + | + | - | - | + | - | + | - | + | + | - | - | - | + | - | - | - | - | - |
| Krefeld (1) | + | + | + | + | + | + | - | + | + | - | + | - | - | - | - | + | + | - | + | - | - | - | - |
| Litchfield (1) | + | + | + | + | + | + | - | - | + | + | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Lille (1) | + | + | + | + | + | + | - | + | + | - | + | + | - | - | - | + | - | - | - | - | - | - | - |
| London (1) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | - | - | - | - | - | - | - | - |
| Meleagridis (1) | + | + | + | + | + | + | - | + | + | - | + | + | - | - | - | - | - | - | - | - | - | - | - |
| Miami (1) | + | + | + | + | + | + | - | - | + | - | + | - | + | + | - | + | - | + | - | - | - | - | - |
| Norwich (1) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | + | - | - | - | - | - | - | - |
| Panama (1) | + | + | + | + | + | + | - | - | + | - | + | - | + | + | - | + | - | + | - | - | - | - | - |
| Poona (1) | + | + | + | + | + | + | - | - | + | + | + | - | + | + | - | + | - | + | - | - | - | - | + |
| Havana (1) | + | + | + | + | + | + | - | - | + | - | + | - | - | - | - | + | - | - | + | - | - | - | - |
| Arizona (IIIa 56:z4) (1) | + | + | + | - | + | + | - | - | + | - | + | + | + | + | - | - | - | - | - | + | + | - | - |
| Arizona(IIIa 41:z4) (1) | + | + | + | - | + | + | - | - | + | - | + | + | + | + | - | - | - | - | - | + | - | - | - |
| Houtenae (IV 43:z4) (2) | + | + | + | + | + | + | - | - | + | - | + | + | + | + | - | - | - | - | - | - | - | - | - |
| Resistance Class | Genes |
|---|---|
| Aminoglycoside | aadA1, aadA2, aadA3, aadA5, aadA6, aadA7, aadA12, aadA13, aadA15, aadA22, aadA25, aph(3′)-Ia, aph(3′)-IIa, aph(3′′)-Ib, aph(4)-Ia, aph(6)-Id, aph(6)-Ic, ant(2′′)-Ia, aac(3)-VIa, aac(3)-IId, aac(3)-IV, aac(3)-IIa, aac(6′)-Ib, aac(6′)-Ib4 |
| β-lactam | blaCARB-2, blaCMY, blaCMY-2, blaHER-3, blaTEM-1, blaTEM, blaOXA-9 |
| Fosfomycin | Fos, fosA7 |
| Macrolide | erm(B), mph(A) |
| Polymyxins | mcr-9.1 |
| Phenicol | catA1, cmlA1, cmlA5, cmlA6, floR |
| Quinolone | oqxA, oqxB, qnrB19 |
| Sulfonamide | sul1, sul2, sul3 |
| Tetracycline | tet(A), tet(B), tet(C), tet(G), tet(M) |
| Trimethoprim | dfrA1, dfrA5, dfrA12, dfrA14 |
| Bleomycin | ble, bleO |
| Lincosamide | lnu(G) |
| Streptothricn | sat2 |
| Biocide resistance | qacEdelta1, qacE, qacL, qacG2 |
| Heavy Metal Resistance | golS, golT, merA, merB, merC, merD, merE, merF, merP, merT, merR, pcoA, pcoB, pcoC, pcoD, pcoE, pcoR, pcoS, silA, silB, silC, silE, silF, silP, silR, silS, terD, terE, terW, terZ, arsA, arsB-mob, arsD |
| Serotype 1 | Source | Plasmid Type | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| IncA/C | IncI | Col | IncN | IncHI | IncFII | IncFIB | IncFIA | IncQ | IncX | IncY | ||
| Typhimurium 2 (n = 136) | Bovine (32) | 6 | 6 | 1 | 0 | 0 | 25 | 22 | 1 | 0 | 0 | 0 |
| Swine (84) | 1 | 11 | 40 | 0 | 5 | 75 | 76 | 16 | 3 | 3 | 2 | |
| Chicken (5) | 0 | 2 | 2 | 0 | 0 | 5 | 5 | 0 | 0 | 1 | 0 | |
| Turkey (5) | 1 | 3 | 2 | 0 | 0 | 5 | 5 | 1 | 0 | 0 | 0 | |
| Equine (9) | 0 | 0 | 0 | 0 | 0 | 9 | 9 | 0 | 0 | 0 | 0 | |
| Caprine (1) | 0 | 1 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | |
| Total | 8 | 23 | 45 | 0 | 5 | 120 | 118 | 18 | 3 | 4 | 0 | |
| Newport (n = 43) | Bovine (30) | 25 | 5 | 4 | 1 | 0 | 5 | 2 | 0 | 0 | 2 | 0 |
| Swine (5) | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Equine (7) | 0 | 0 | 0 | 0 | 0 | 2 | 1 | 0 | 0 | 0 | 0 | |
| Caprine (1) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Total | 26 | 5 | 4 | 1 | 0 | 7 | 3 | 0 | 0 | 2 | 0 | |
| Heidelberg (n = 38) | Swine (20) | 0 | 10 | 3 | 0 | 19 | 0 | 0 | 0 | 0 | 0 | 3 |
| Turkey (13) | 2 | 7 | 0 | 1 | 7 | 0 | 0 | 0 | 0 | 3 | 0 | |
| Chicken (5) | 0 | 2 | 4 | 0 | 0 | 0 | 0 | 0 | 0 | 5 | 0 | |
| Total | 2 | 19 | 7 | 1 | 27 | 0 | 0 | 0 | 0 | 8 | 0 | |
| Choleraesuis (n = 35) | Swine (35) | 0 | 0 | 1 | 0 | 25 | 35 | 35 | 0 | 33 | 1 | 0 |
| Derby (n = 29) | Swine (29) | 0 | 13 | 14 | 0 | 3 | 1 | 1 | 4 | 6 | 5 | 4 |
| Anatum (n = 16) | Swine (9) | 0 | 6 | 9 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 |
| Bovine (5) | 2 | 1 | 1 | 0 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | |
| Equine (1) | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 1 | 0 | 0 | |
| Caprine (1) | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Total | 2 | 8 | 10 | 1 | 2 | 1 | 1 | 2 | 1 | 0 | 0 | |
| Agona (n = 14) | Bovine (5) | 5 | 3 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Swine (8) | 6 | 1 | 2 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Turkey (1) | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Total | 11 | 5 | 3 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Kentucky 3 (n = 13) | Bovine (2) | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Chicken (10) | 0 | 6 | 0 | 0 | 3 | 6 | 5 | 0 | 0 | 10 | 0 | |
| Swine (1) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Total | 0 | 7 | 0 | 0 | 3 | 6 | 5 | 0 | 0 | 10 | 0 | |
| Muenchen (n = 11) | Bovine (2) | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 |
| Swine (9) | 0 | 1 | 6 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Total | 0 | 1 | 6 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, S.; Li, C.; Hsu, C.-H.; Tyson, G.H.; Strain, E.; Tate, H.; Tran, T.-T.; Abbott, J.; McDermott, P.F. Comparative Genomic Analysis of 450 Strains of Salmonella enterica Isolated from Diseased Animals. Genes 2020, 11, 1025. https://doi.org/10.3390/genes11091025
Zhao S, Li C, Hsu C-H, Tyson GH, Strain E, Tate H, Tran T-T, Abbott J, McDermott PF. Comparative Genomic Analysis of 450 Strains of Salmonella enterica Isolated from Diseased Animals. Genes. 2020; 11(9):1025. https://doi.org/10.3390/genes11091025
Chicago/Turabian StyleZhao, Shaohua, Cong Li, Chih-Hao Hsu, Gregory H. Tyson, Errol Strain, Heather Tate, Thu-Thuy Tran, Jason Abbott, and Patrick F. McDermott. 2020. "Comparative Genomic Analysis of 450 Strains of Salmonella enterica Isolated from Diseased Animals" Genes 11, no. 9: 1025. https://doi.org/10.3390/genes11091025
APA StyleZhao, S., Li, C., Hsu, C.-H., Tyson, G. H., Strain, E., Tate, H., Tran, T.-T., Abbott, J., & McDermott, P. F. (2020). Comparative Genomic Analysis of 450 Strains of Salmonella enterica Isolated from Diseased Animals. Genes, 11(9), 1025. https://doi.org/10.3390/genes11091025




