Karyotypic Evolution of Sauropsid Vertebrates Illuminated by Optical and Physical Mapping of the Painted Turtle and Slider Turtle Genomes
Abstract
1. Introduction
2. Materials and Methods
2.1. BioNano Optical Mapping
2.2. Cell Culture and Chromosome Preparations
2.3. BAC-FISH Mapping
2.4. Image Analysis
2.5. Bioinformatic Analysis of Painted Turtle-Sauropsid Genome Evolution
3. Results
3.1. Improvement of the Contiguity of the Painted Turtle Genome via Optical Mapping
3.2. Improvement of Physical Mapping of the Painted Turtle Genome via Optical Mapping
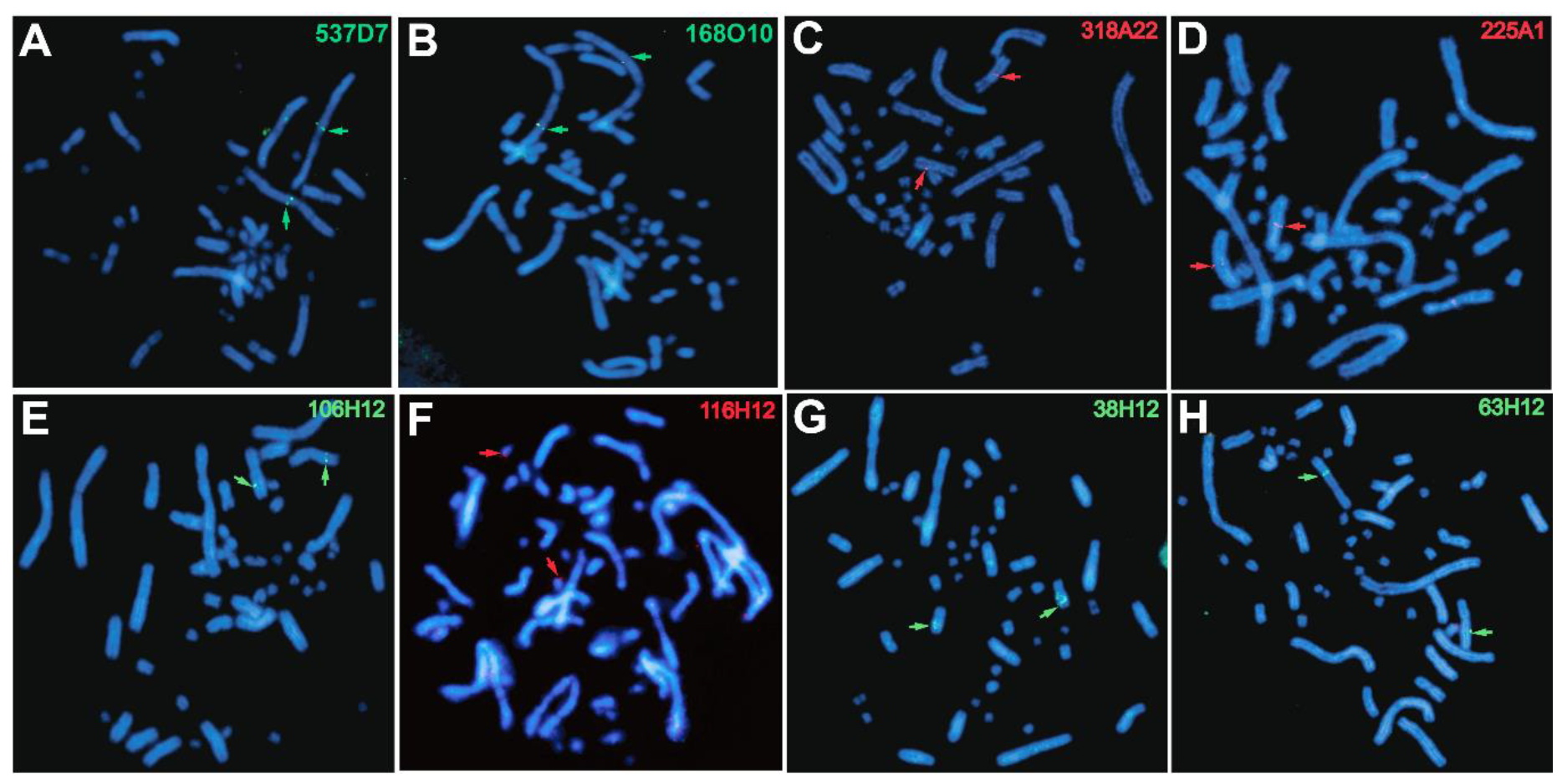
3.3. Improvement of Physical Mapping of the Painted Turtle Genome via Additional de novo BAC-FISH
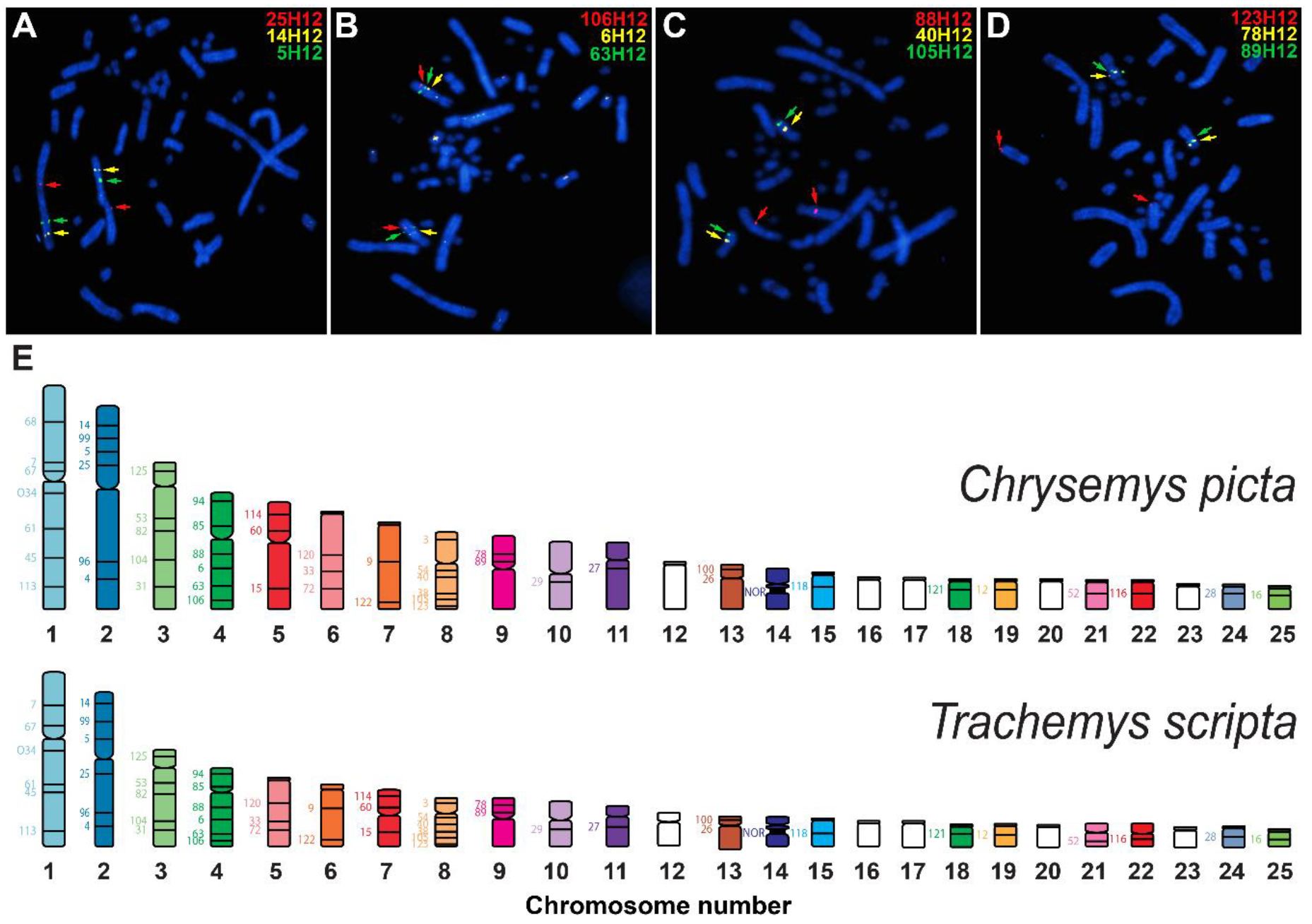
3.4. Physical Mapping of T. s. Elegans’s Genome via de Novo BAC-FISH
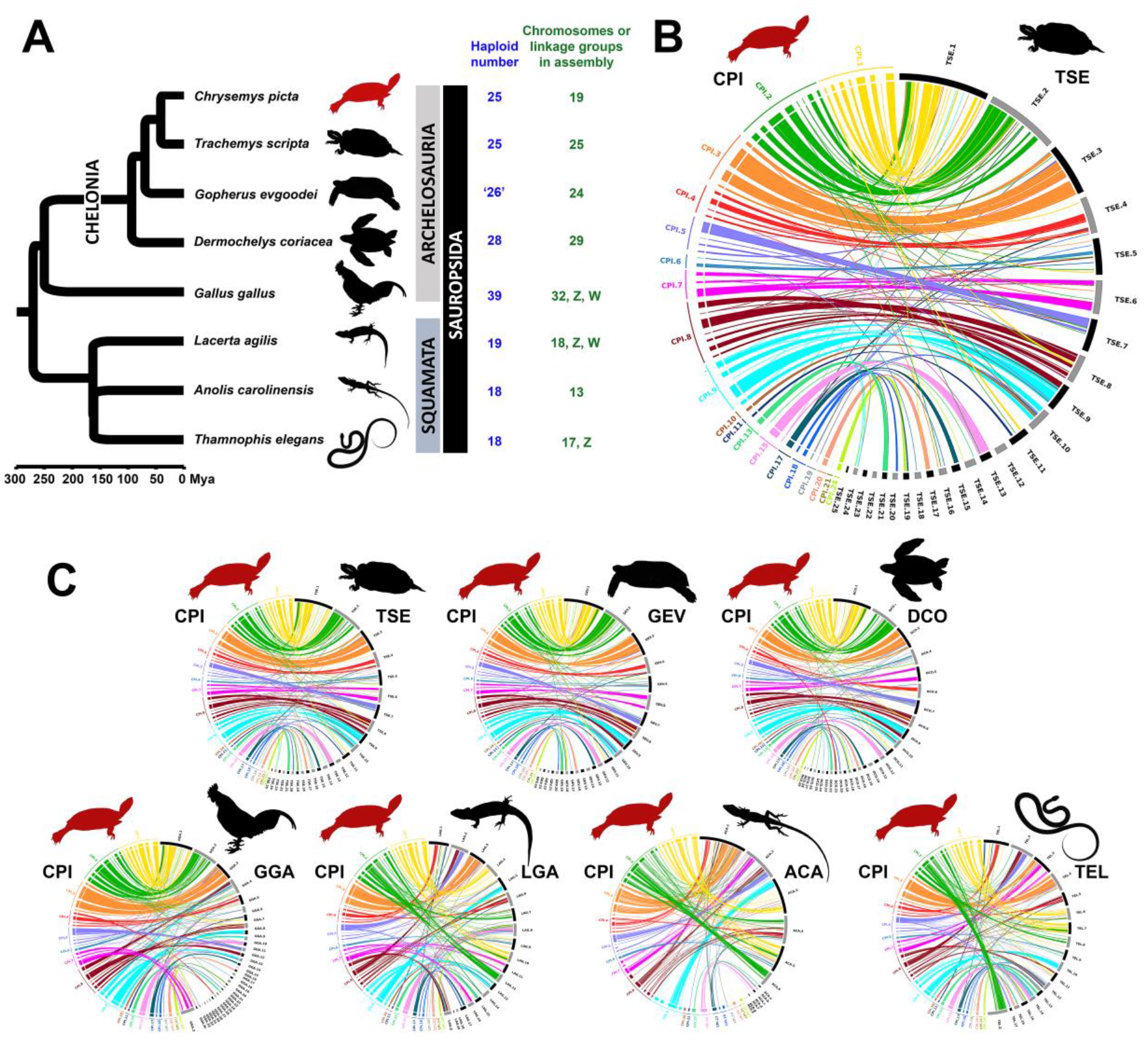
3.5. Chromosomal Homology and Synteny Decreases with Phylogenetic Distance between Painted Turtle and Other Sauropsids
4. Discussion
4.1. Contiguity of the Painted Turtle Genome Improved by Optical and Physical Mapping
4.2. Physical Mapping Uncovered Genome Assembly Errors in Slider Turtle and Highly Conserved Chromosome Homology with Painted Turtles
4.3. Karyotypic Evolution Involving Sex Chromosomes and Sexual Development Genes
4.4. Chromosomal Homology and Synteny Sheds Light on Sauropsid Karyotypic Evolution
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Van Dijk, E.L.; Jaszczyszyn, Y.; Naquin, D.; Thermes, C. The Third Revolution in Sequencing Technology. Trends Genet. 2018, 34, 666–681. [Google Scholar] [CrossRef] [PubMed]
- Putnam, N.H.; O’Connell, B.L.; Stites, J.C.; Rice, B.J.; Blanchette, M.; Calef, R.; Troll, C.J.; Fields, A.; Hartley, P.D.; Sugnet, C.W.; et al. Chromosome-scale shotgun assembly using an in vitro method for long-range linkage. Genome Res. 2016, 26, 342–350. [Google Scholar] [CrossRef] [PubMed]
- Iannucci, A.; Altmanova, M.; Ciofi, C.; Ferguson-Smith, M.; Pereira, J.C.; Rehak, I.; Stanyon, R.; Velensky, P.; Rovatsos, M.; Kratochvil, L.; et al. Isolating Chromosomes of the Komodo Dragon: New Tools for Comparative Mapping and Sequence Assembly. Cytogenet. Genome Res. 2019, 157, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Rice, E.S.; Kohno, S.; St. John, J.; Pham, S.; Howard, J.; Lareau, L.F.; O’Connell, B.L.; Hickey, G.; Armstrong, J.; Deran, A.; et al. Improved genome assembly of American alligator genome reveals conserved architecture of estrogen signaling. Genome Res. 2017, 27, 686–696. [Google Scholar] [CrossRef] [PubMed]
- Nanda, I.; Zend-Ajusch, E.; Shan, Z.; Grutzner, F.; Schartl, M.; Burt, D.W.; Koehler, M.; Fowler, V.M.; Goodwin, G.; Schneider, W.J.; et al. Conserved synteny between the chicken Z sex chromosome and human chromosome 9 includes the male regulatory gene DMRT1: A comparative (re)view on avian sex determination. Cytogenet. Cell Genet. 2000, 89, 67–78. [Google Scholar] [CrossRef]
- Schmid, M.; Nanda, I.; Guttenbach, M.; Steinlein, C.; Hoehn, H.; Schartl, M.; Haaf, T.; Weigend, S.; Fries, R.; Buerstedde, J.M.; et al. First report on chicken genes and chromosomes 2000. Cytogenet. Cell Genet. 2000, 90, 169–218. [Google Scholar] [CrossRef] [PubMed]
- Valenzuela, N. The painted turtle, Chrysemys picta: A model system for vertebrate evolution, ecology, and human health. Cold Spring Harb. Protoc. 2009, 4, 1–9. [Google Scholar] [CrossRef]
- Mizoguchi, B.A.; Valenzuela, N. Ecotoxicological perspectives of sex determination. Sex. Dev. 2016, 10, 45–57. [Google Scholar] [CrossRef]
- Valenzuela, N.; Adams, D.C. Chromosome number and sex determination coevolve in turtles. Evolution 2011, 65, 1808–1813. [Google Scholar] [CrossRef]
- Lee, L.S.; Montiel, E.E.; Navarro-Domínguez, B.M.; Valenzuela, N. Chromosomal Rearrangements during Turtle Evolution Altered the Synteny of Genes Involved in Vertebrate Sex Determination. Cytogenet. Genome Res. 2019, 157, 70–81. [Google Scholar] [CrossRef]
- Shaffer, H.B.; Minx, P.; Warren, D.E.; Shedlock, A.M.; Thomson, R.C.; Valenzuela, N.; Abramyan, J.; Amemiya, C.T.; Badenhorst, D.; Biggar, K.K.; et al. The western painted turtle genome, a model for the evolution of extreme physiological adaptations in a slowly evolving lineage. Genome Biol. 2013, 14, 21–22. [Google Scholar] [CrossRef]
- Badenhorst, D.; Hillier, L.W.; Literman, R.; Montiel, E.E.; Radhakrishnan, S.; Shen, Y.; Minx, P.; Janes, D.E.; Warren, W.C.; Edwards, S.V.; et al. Physical mapping and refinement of the painted turtle genome (Chrysemys picta) inform amniote genome evolution and challenge turtle-bird chromosomal conservation. Genom. Biol. Evol. 2015, 7, 2038–2050. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Simison, W.B.; Parham, J.F.; Papenfuss, T.J.; Lam, A.W.; Henderson, J.B. An Annotated Chromosome-Level Reference Genome of the Red-Eared Slider Turtle (Trachemys scripta elegans). Genome Biol. Evol. 2020, 12, 456–462. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Pascual-Anaya, J.; Zadissa, A.; Li, W.Q.; Niimura, Y.; Huang, Z.Y.; Li, C.Y.; White, S.; Xiong, Z.Q.; Fang, D.M.; et al. The draft genomes of soft-shell turtle and green sea turtle yield insights into the development and evolution of the turtle-specific body plan. Nat. Genet. 2013, 45, 701–706. [Google Scholar] [CrossRef]
- Tollis, M.; DeNardo, D.F.; Cornelius, J.A.; Dolby, G.A.; Edwards, T.; Henen, B.T.; Karl, A.E.; Murphy, R.W.; Kusumi, K. The Agassiz’s desert tortoise genome provides a resource for the conservation of a threatened species. PLoS ONE 2017, 12, e0177708. [Google Scholar] [CrossRef]
- Cao, D.N.; Wang, M.; Ge, Y.; Gong, S.P. Draft genome of the big-headed turtle Platysternon megacephalum. Sci. Data 2019, 6, 1–8. [Google Scholar] [CrossRef]
- Quesada, V.; Freitas-Rodriguez, S.; Miller, J.; Perez-Silva, J.G.; Jiang, Z.F.; Tapia, W.; Santiago-Fernandez, O.; Campos-Iglesias, D.; Kuderna, L.F.K.; Quinzin, M.; et al. Giant tortoise genomes provide insights into longevity and age-related disease. Nat. Ecol. Evol. 2019, 3, 87–95. [Google Scholar] [CrossRef]
- Lam, E.T.; Hastie, A.; Lin, C.; Ehrlich, D.; Das, S.K.; Austin, M.D.; Deshpande, P.; Cao, H.; Nagarajan, N.; Xiao, M.; et al. Genome mapping on nanochannel arrays for structural variation analysis and sequence assembly. Nat. Biotechnol. 2012, 30, 771–776. [Google Scholar] [CrossRef]
- Janes, D.E.; Organ, C.; Valenzuela, N. New resources inform study of genome size, content, and organization in nonavian reptiles. Integr. Comp. Biol. 2008, 48, 447–453. [Google Scholar] [CrossRef]
- Janes, D.E.; Valenzuela, N.; Ezaz, T.; Amemiya, C.; Edwards, S.V. Sex chromosome evolution in Amniotes: Applications for bacterial artificial chromosome libraries. J. Biomed. Biotechnol. 2011, 2011, 132975. [Google Scholar] [CrossRef]
- Montiel, E.E.; Badenhorst, D.; Lee, L.S.; Literman, R.; Trifonov, V.; Valenzuela, N. Cytogenetic insights into the evolution of chromosomes and sex determination reveal striking homology of turtle sex chromosomes to amphibian autosomes. Cytogenet. Genome Res. 2016, 148, 292–304. [Google Scholar] [CrossRef] [PubMed]
- Valenzuela, N.; Badenhorst, D.; Montiel, E.E.; Literman, R. Molecular cytogenetic search for cryptic sex chromosomes in painted turtles Chrysemys picta. Cytogenet. Genome Res. 2014, 144, 39–46. [Google Scholar] [CrossRef] [PubMed]
- Radhakrishnan, S.; Literman, R.; Neuwald, J.; Severin, A.; Valenzuela, N. Transcriptomic responses to environmental temperature by turtles with temperature-dependent and genotypic sex determination assessed by RNAseq inform the genetic architecture of embryonic gonadal development. PLoS ONE 2017, 12, e0172044. [Google Scholar] [CrossRef] [PubMed]
- Radhakrishnan, S.; Literman, R.; Neuwald, J.L.; Valenzuela, N. Thermal Response of Epigenetic Genes Informs Turtle Sex Determination with and without Sex Chromosomes. Sex. Dev. 2018, 12, 308–319. [Google Scholar] [CrossRef]
- Radhakrishnan, S.; Literman, R.; Mizoguchi, B.; Valenzuela, N. MeDIP-seq and nCpG analyses illuminate sexually dimorphic methylation of gonadal development genes with high historic methylation in turtle hatchlings with temperature-dependent sex determination. Epigenetics Chromatin 2017, 10, 28. [Google Scholar] [CrossRef]
- Montiel, E.E.; Badenhorst, D.; Tamplin, J.; Burke, R.L.; Valenzuela, N. Discovery of the youngest sex chromosomes reveals first case of convergent co-option of ancestral autosomes in turtles. Chromosoma 2017, 126, 105–113. [Google Scholar] [CrossRef]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997v1. [Google Scholar]
- Radhakrishnan, S.; Valenzuela, N. Chromosomal context affects the molecular evolution of sex-linked genes and their autosomal counterparts in turtles and other vertebrates. J. Hered. 2017, 108, 720–730. [Google Scholar] [CrossRef]
- Delcher, A.L.; Phillippy, A.; Carlton, J.; Salzberg, S.L. Fast algorithms for large-scale genome alignment and comparison. Nucleic Acids Res. 2002, 30, 2478–2483. [Google Scholar] [CrossRef]
- Frith, M.C.; Hamada, M.; Horton, P. Parameters for accurate genome alignment. BMC Bioinform. 2010, 11, 80. [Google Scholar] [CrossRef]
- Kielbasa, S.M.; Wan, R.; Sato, K.; Horton, P.; Frith, M.C. Adaptive seeds tame genomic sequence comparison. Genome Res. 2011, 21, 487–493. [Google Scholar] [CrossRef] [PubMed]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed]
- Valenzuela, N. Co-evolution of genomic structure and selective forces underlying sexual development and reproduction. Cytogenet. Genome Res. 2009, 127, 232–241. [Google Scholar] [CrossRef] [PubMed]
- Valenzuela, N. Causes and Consequences of Evolutionary Transitions in the Level of Phenotypic Plasticity of Reptilian Sex Determination. In Transitions Between Sexual Systems; Leonard, J., Ed.; Springer: Cham, Switzerland, 2018; pp. 345–363. [Google Scholar]
- Kasai, F.; O’Brien, P.C.M.; Martin, S.; Ferguson-Smith, M.A. Extensive homology of chicken macrochromosomes in the karyotypes of Trachemys scripta elegans and Crocodylus niloticus revealed by chromosome painting despite long divergence times. Cytogenet. Genome Res. 2012, 136, 303–307. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, Y.; Nishida-Umehara, C.; Tarui, H.; Kuroiwa, A.; Yamada, K.; Isobe, T.; Ando, J.; Fujiwara, A.; Hirao, Y.; Nishimura, O.; et al. Highly conserved linkage homology between birds and turtles: Bird and turtle chromosomes are precise counterparts of each other. Chrom. Res. 2005, 13, 601–615. [Google Scholar] [CrossRef]
- Damas, J.; O’Connor, R.; Farre, M.; Lenis, V.P.E.; Martell, H.J.; Mandawala, A.; Fowler, K.; Joseph, S.; Swain, M.T.; Griffin, D.K.; et al. Upgrading short-read animal genome assemblies to chromosome level using comparative genomics and a universal probe set. Genome Res. 2017, 27, 875–884. [Google Scholar] [CrossRef]
- Shearer, L.A.; Anderson, L.K.; de Jong, H.; Smit, S.; Goicoechea, J.L.; Roe, B.A.; Hua, A.; Giovannoni, J.J.; Stack, S.M. Fluorescence In Situ Hybridization and Optical Mapping to Correct Scaffold Arrangement in the Tomato Genome. G3 2014, 4, 1395–1405. [Google Scholar] [CrossRef]
- Tomaszkiewicz, M.; Medvedev, P.; Makova, K.D. Y and W Chromosome Assemblies: Approaches and Discoveries. Trends Genet. 2017, 33, 266–282. [Google Scholar] [CrossRef]
- Shedlock, A.M.; Botka, C.W.; Zhao, S.Y.; Shetty, J.; Zhang, T.T.; Liu, J.S.; Deschavanne, P.J.; Edward, S.V. Phylogenomics of nonavian reptiles and the structure of the ancestral amniote genorne. Proc. Natl. Acad. Sci. USA 2007, 104, 2767–2772. [Google Scholar] [CrossRef]
- Stankova, H.; Hastie, A.R.; Chan, S.; Vrana, J.; Tulpova, Z.; Kubalakova, M.; Visendi, P.; Hayashi, S.; Luo, M.C.; Batley, J.; et al. BioNano genome mapping of individual chromosomes supports physical mapping and sequence assembly in complex plant genomes. Plant Biotechnol. J. 2016, 14, 1523–1531. [Google Scholar] [CrossRef]
- O’Connor, R.E.; Romanov, M.N.; Kiazim, L.G.; Barrett, P.M.; Farre, M.; Damas, J.; Ferguson-Smith, M.; Valenzuela, N.; Larkin, D.M.; Griffin, D.K. Reconstruction of the diapsid ancestral genome permits chromosome evolution tracing in avian and non-avian dinosaurs. Nat. Commun. 2018, 9, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Bista, B.; Valenzuela, N. Turtle insights into the evolution of the reptilian karyotype and the genomic architecture of sex determination. Genes 2020, 11, 416. [Google Scholar] [CrossRef] [PubMed]
- Kawagoshi, T.; Uno, Y.; Nishida, C.; Matsuda, Y. The Staurotypus turtles and aves share the same origin of sex chromosomes but evolved different types of heterogametic sex determination. PLoS ONE 2014, 9, e0105315. [Google Scholar] [CrossRef] [PubMed]
- Badenhorst, D.; Stanyon, R.; Engstrom, T.; Valenzuela, N. A ZZ/ZW microchromosome system in the spiny softshell turtle, Apalone spinifera, reveals an intriguing sex chromosome conservation in Trionychidae. Chrom. Res. 2013, 21, 137–147. [Google Scholar] [CrossRef] [PubMed]
- Janes, D.E.; Organ, C.L.; Stiglec, R.; O’Meally, D.; Sarre, S.D.; Georges, A.; Graves, J.A.M.; Valenzuela, N.; Literman, R.A.; Rutherford, K.; et al. Molecular evolution of Dmrt1 accompanies change of sex-determining mechanisms in reptilia. Biol. Lett. 2014, 10, 20140809. [Google Scholar] [CrossRef]
- Literman, R.; Burrett, A.; Bista, B.; Valenzuela, N. Putative Independent Evolutionary Reversals from Genotypic to Temperature-Dependent Sex Determination are Associated with Accelerated Evolution of Sex-Determining Genes in Turtles. J. Mol. Evol. 2018, 86, 11–26. [Google Scholar] [CrossRef]
- Mizoguchi, B.; Valenzuela, N. Alternative splicing and thermosensitive expression of Dmrt1 during urogenital development in the painted turtle, Chrysemys picta. PeerJ 2020, 8, e8639. [Google Scholar] [CrossRef] [PubMed]
- Kawai, A.; Nishida-Umehara, C.; Ishijima, J.; Tsuda, Y.; Ota, H.; Matsuda, Y. Different origins of bird and reptile sex chromosomes inferred from comparative mapping of chicken Z-linked genes. Cytogenet. Genome Res. 2007, 117, 92–102. [Google Scholar] [CrossRef]
- Ezaz, T.; Sarre, S.D.; O’Meally, D.; Graves, J.A.M.; Georges, A. Sex chromosome evolution in lizards: Independent origins and rapid transitions. Cytogenet. Genome Res. 2009, 127, 249–260. [Google Scholar] [CrossRef]
- Uno, Y.; Nishida, C.; Tarui, H.; Ishishita, S.; Takagi, C.; Nishimura, O.; Ishijima, J.; Ota, H.; Kosaka, A.; Matsubara, K.; et al. Inference of the Protokaryotypes of Amniotes and Tetrapods and the Evolutionary Processes of Microchromosomes from Comparative Gene Mapping. PLoS ONE 2012, 7, e0053027. [Google Scholar] [CrossRef]
- Matsubara, K.; Tarui, H.; Toriba, M.; Yamada, K.; Nishida-Umehara, C.; Agata, K.; Matsuda, Y. Evidence for different origin of sex chromosomes in snakes, birds, and mammals and step-wise differentiation of snake sex chromosomes. Proc. Natl. Acad. Sci. USA 2006, 103, 18190–18195. [Google Scholar] [CrossRef] [PubMed]
- Alam, S.M.I.; Sarre, S.D.; Gleeson, D.; Georges, A.; Ezaz, T. Did Lizards Follow Unique Pathways in Sex Chromosome Evolution? Genes 2018, 9, 239. [Google Scholar] [CrossRef] [PubMed]
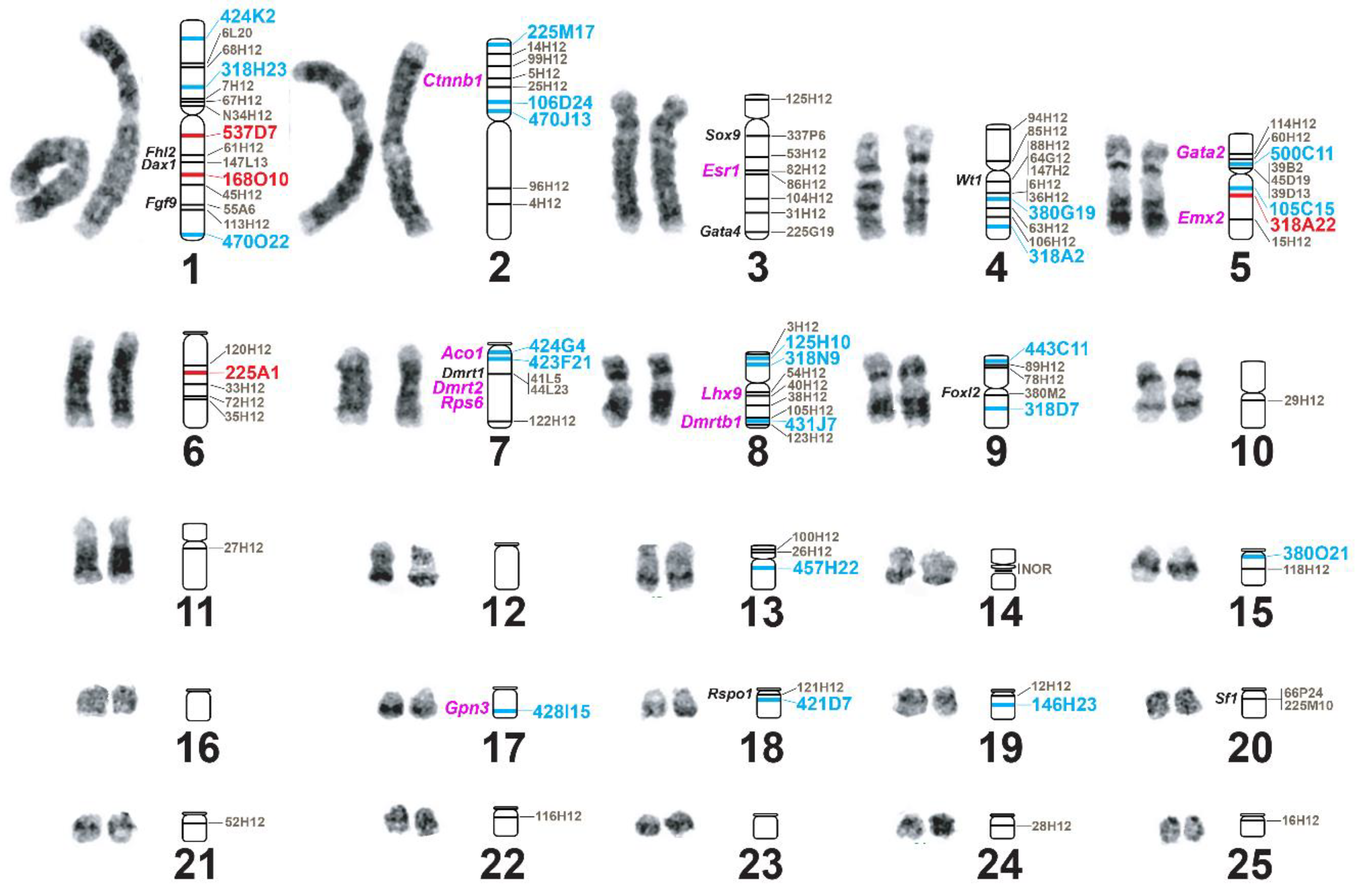
| BAC ID | BAC Size | ID BioNano Scaffold | ID NCBI Scaffold | Scaffold Size (bp) | CPI Chromosome |
|---|---|---|---|---|---|
| 424K2 | 163,122 | 203 | ML621420.1 | 15,883,268 | 1p |
| 318H23 | 145,864 | 217 | ML621429.1 | 8,721,052 | 1p |
| 537D7 | 154,142 | 60 | ML621305.1 | 21,088,589 | 1q |
| 168O10 | 162,311 | 153 | ML621386.1 | 8,440,838 | 1q |
| 470O22 | 158,668 | 331 | ML621495.1 | 5,731,011 | 1q |
| 225M17 | 149,235 | 179 | ML621405.1 | 57,448,544 | 2p |
| 106D24 | 132,359 | 23 | ML621269.1 | 31,217,720 | 2p |
| 470J13 | 136,853 | 304 | ML621480.1 | 3,056,537 | 2p |
| 380G19 | 160,060 | 141 | ML621375.1 | 5,625,148 | 4q |
| 318A2 | 138,653 | 60 | ML621305.1 | 21,088,589 | 4q |
| 500C11 | 176,356 | 29,527 | ML621534.1 | 1,293,745 | 5p |
| 105C15 | 151,304 | 87 | ML621330.1 | 5,648,650 | 5q |
| 318A22 | 134,050 | 163 | ML621393.1 | 21,409,697 | 5q |
| 225A1 | 149,859 | 178 | ML621404.1 | 40,323,178 | 6 |
| 424G4 | 146,986 | 75 | ML621319.1 | 13,166,690 | 7 |
| 423F21 | 152,254 | 75 | ML621319.1 | 13,166,690 | 7 |
| 125H10 | 146,876 | 18 | ML621264.1 | 1,390,666 | 8p |
| 318N9 | 153,773 | 76 | ML621320.1 | 21,927,555 | 8p |
| 431J7 | 125,453 | 309 | ML621483.1 | 5,421,227 | 8q |
| 443C11 | 136,412 | 194 | ML621413.1 | 18,639,614 | 9p |
| 318D7 | 137,214 | 14 | ML621260.1 | 13,251,999 | 9q |
| 457H22 | 146,712 | 126 | ML621362.1 | 19,176,785 | 13 |
| 380O21 | 149,406 | 120 | ML621356.1 | 1,852,121 | 15 |
| 428I15 | 135,794 | 145 | ML621378.1 | 23,325,489 | 17 |
| 421D7 | 172,285 | 56 | ML621301.1 | 6,383,505 | 18 |
| 146H23 | 170,310 | 177 | ML621403.1 | 1,209,793 | 19 |
| ID BioNano Scaffold | ID NCBI Scaffold | Scaffold Size (bp) | BAC ID | Chromosome |
|---|---|---|---|---|
| 60 | ML621305.1 | 21,088,589 | 537D7 | 1q |
| 106H12 | 4 | |||
| 153 | ML621386.1 | 8,440,838 | 168O10 | 1q |
| 116H12 | 22 | |||
| 163 | ML621393.1 | 21,409,697 | 318A22 | 5q |
| 38H12 | 8 | |||
| 178 | ML621404.1 | 40,323,178 | 225A1 | 6 |
| 63H12 | 4 | |||
| 15 | ML621261.1 | 59,628,869 | 15H12 | 5 |
| 72H12, 35H18 | 6 | |||
| 70 | ML621314.1 | 11,495,585 | 147L13 | 1 |
| 27H12 | 11 | |||
| 207 | ML621424.1 | 27,887,944 | 45H12, 55A6 | 1 |
| 26H12 | 13 |
| ID BioNano Scaffold | ID NCBI Scaffold | Sex-Related Genes | CPI Chromosome | GGA Chromosome | Gene Order |
|---|---|---|---|---|---|
| 289 | ML621472.1 | Fhl2 | 1 | 1 | CPI-1:Fhl2-Dax1-Fgf9 GGA-1:Dax1-Fhl2-Fgf9 |
| 70 | ML621314.1 | Dax1 (Nr0b1) | 1 | 1 | |
| 207 | ML621424.1 | Fgf9 | 1 | 1 | |
| 339 | ML621502.1 | Ctnnb1 | 2 | 2 | |
| 65 | ML621310.1 | Sox9 | 3 | 18 | CPI-3:Sox9-Esr1-Gata4 GGA-3:Esr1-Gata4 GGA-18:Sox9 |
| 65 | ML621310.1 | Esr1 | 3 | 3 | |
| 135 | ML621369.1 | Gata4 | 3 | 3 | |
| 330 | ML621494.1 | Wt1 | 4 | 5 | |
| 59 | ML621304.1 | Gata2 | 5 | 12 | CPI-5:Gata2-Emx2 GGA-6:Emx2 GGA-12:Gata2 |
| 15 | ML621261.1 | Emx2 | 5 | 6 | |
| 75 | ML621319.1 | Aco1 | 7 | Z | CPI-7:Aco1-Dmrt1-Dmrt2-Rps6 GGA-Z:Dmrt1-Dmrt2-Rps6-Aco1 |
| 128 | ML621363.1 | Dmrt1 Dmrt2 Rps6 | 7 | Z | |
| 163 | ML621393.1 | Lhx9 | 8 | 8 | CPI-8:Lhx9-Dmrtb1 GGA-8:Lhx9-Dmrtb1 |
| 309 | ML621483.1 | Dmrtb1 | 8 | 8 | |
| 4 | ML621250.1 | Foxl2 | 9 | 9 | |
| 145 | ML621378.1 | Gpn3 | 17 | 15 | |
| 198 | ML621416.1 | Rspo1 | 18 | 23 | |
| 83 | ML621327.1 | Sf1 (Nr5a1) | 20 | 17 |
| BAC ID | TSE Assembly Scaffold | Assembly TSE Chromosome Assignment | BAC-FISH-Based TSC Chromosome (True Location) |
|---|---|---|---|
| 68H12 | CM023056.1 | TSE 1 | TSC 1 |
| 7H12 | CM023056.1 | TSE 1 | TSC 1 |
| 67H12 | CM023058.1 | TSE 3 | TSC 1 |
| O34H12 | CM023056.1 | TSE 1 | TSC 1 |
| 61H12 | CM023056.1 | TSE 1 | TSC 1 |
| 45H12 | CM023056.1 | TSE 1 | TSC 1 |
| 113H12 | CM023056.1 | TSE 1 | TSC 1 |
| 14H12 | CM023057.1 | TSE 2 | TSC 2 |
| 99H12 | CM023057.1 | TSE 2 | TSC 2 |
| 5H12 | CM023057.1 | TSE 2 | TSC 2 |
| 25H12 | CM023057.1 | TSE 2 | TSC 2 |
| 96H12 | CM023057.1 | TSE 2 | TSC 2 |
| 4H12 | CM023057.1 | TSE 2 | TSC 2 |
| 125H12 | CM023058.1 | TSE 3 | TSC 3 |
| 53H12 | CM023058.1 | TSE 3 | TSC 3 |
| 82H12 | CM023058.1 | TSE 3 | TSC 3 |
| 104H12 | CM023058.1 | TSE 3 | TSC 3 |
| 31H12 | CM023058.1 | TSE 3 | TSC 3 |
| 94H12 | CM023059.1 | TSE 4 | TSC 4 |
| 85H12 | CM023059.1 | TSE 4 | TSC 4 |
| 88H12 | CM023059.1 | TSE 4 | TSC 4 |
| 6H12 | CM023059.1 | TSE 4 | TSC 4 |
| 63H12 | CM023059.1 | TSE 4 | TSC 4 |
| 106H12 | CM023059.1 | TSE 4 | TSC 4 |
| 120H12 | CM023060.1 | TSE 5 | TSC 5 |
| 33H12 | CM023060.1 | TSE 5 | TSC 5 |
| 72H12 | CM023060.1 | TSE 5 | TSC 5 |
| 9H12 | CM023058.1 | TSE 3 | TSC 6 |
| 122H12 | CM023061.1 | TSE 6 | TSC 6 |
| 114H12 | CM023062.1 | TSE 7 | TSC 7 |
| 60H12 | CM023062.1 | TSE 7 | TSC 7 |
| 15H12 | CM023062.1 | TSE 7 | TSC 7 |
| 3H12 | CM023063.1 | TSE 8 | TSC 8 |
| 54H12 | CM023063.1 | TSE 8 | TSC 8 |
| 40H12 | CM023063.1 | TSE 8 | TSC 8 |
| 38H12 | CM023063.1 | TSE 8 | TSC 8 |
| 105H12 | CM023063.1 | TSE 8 | TSC 8 |
| 123H12 | CM023063.1 | TSE 8 | TSC 8 |
| 78H12 | CM023064.1 | TSE 9 | TSC 9 |
| 89H12 | CM023064.1 | TSE 9 | TSC 9 |
| 29H12 | CM023065.1 | TSE 10 | TSC 10 |
| 27H12 | CM023066.1 | TSE 11 | TSC 11 |
| 100H12 | CM023069.1 | TSE 14 | TSC 13 |
| 26H12 | CM023069.1 | TSE 14 | TSC 13 |
| 118H12 | CM023068.1 | TSE 13 | TSC 15 |
| 121H12 | CM023075.1 | TSE 20 | TSC 18 |
| 12H12 | CM023058.1 | TSE 3 | TSC 19 |
| 28H12 | CM023074.1 | TSE 19 | TSC 24 |
| 52H12 | CM023071.1 | TSE 16 | TSC 21 |
| 116H12 | CM023077.1 | TSE 22 | TSC 22 |
| 16H12 | CM023071.1 | TSE 16 | TSC 25 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, L.S.; Navarro-Domínguez, B.M.; Wu, Z.; Montiel, E.E.; Badenhorst, D.; Bista, B.; Gessler, T.B.; Valenzuela, N. Karyotypic Evolution of Sauropsid Vertebrates Illuminated by Optical and Physical Mapping of the Painted Turtle and Slider Turtle Genomes. Genes 2020, 11, 928. https://doi.org/10.3390/genes11080928
Lee LS, Navarro-Domínguez BM, Wu Z, Montiel EE, Badenhorst D, Bista B, Gessler TB, Valenzuela N. Karyotypic Evolution of Sauropsid Vertebrates Illuminated by Optical and Physical Mapping of the Painted Turtle and Slider Turtle Genomes. Genes. 2020; 11(8):928. https://doi.org/10.3390/genes11080928
Chicago/Turabian StyleLee, Ling Sze, Beatriz M. Navarro-Domínguez, Zhiqiang Wu, Eugenia E. Montiel, Daleen Badenhorst, Basanta Bista, Thea B. Gessler, and Nicole Valenzuela. 2020. "Karyotypic Evolution of Sauropsid Vertebrates Illuminated by Optical and Physical Mapping of the Painted Turtle and Slider Turtle Genomes" Genes 11, no. 8: 928. https://doi.org/10.3390/genes11080928
APA StyleLee, L. S., Navarro-Domínguez, B. M., Wu, Z., Montiel, E. E., Badenhorst, D., Bista, B., Gessler, T. B., & Valenzuela, N. (2020). Karyotypic Evolution of Sauropsid Vertebrates Illuminated by Optical and Physical Mapping of the Painted Turtle and Slider Turtle Genomes. Genes, 11(8), 928. https://doi.org/10.3390/genes11080928






