Sequencing and Analysis of the Sex Determination Region of Populus trichocarpa
Abstract
1. Introduction
2. Methods
2.1. Initial Genome Assembly
2.2. Variant Calling of Individuals from Natural Population
2.3. Sex-Association Analysis
2.4. Identifying the Sex-Specific Covered Region
2.5. Genetic Linkage Mapping
2.6. Gene Annotation on the SDR and X-Linked Scaffold
2.7. Estimation of the Divergence of the SDR
2.8. Identification of Recently Inserted LTR Retrotransposable Elements and Repetitive Elements
2.9. Expression of the Inverted Repeats
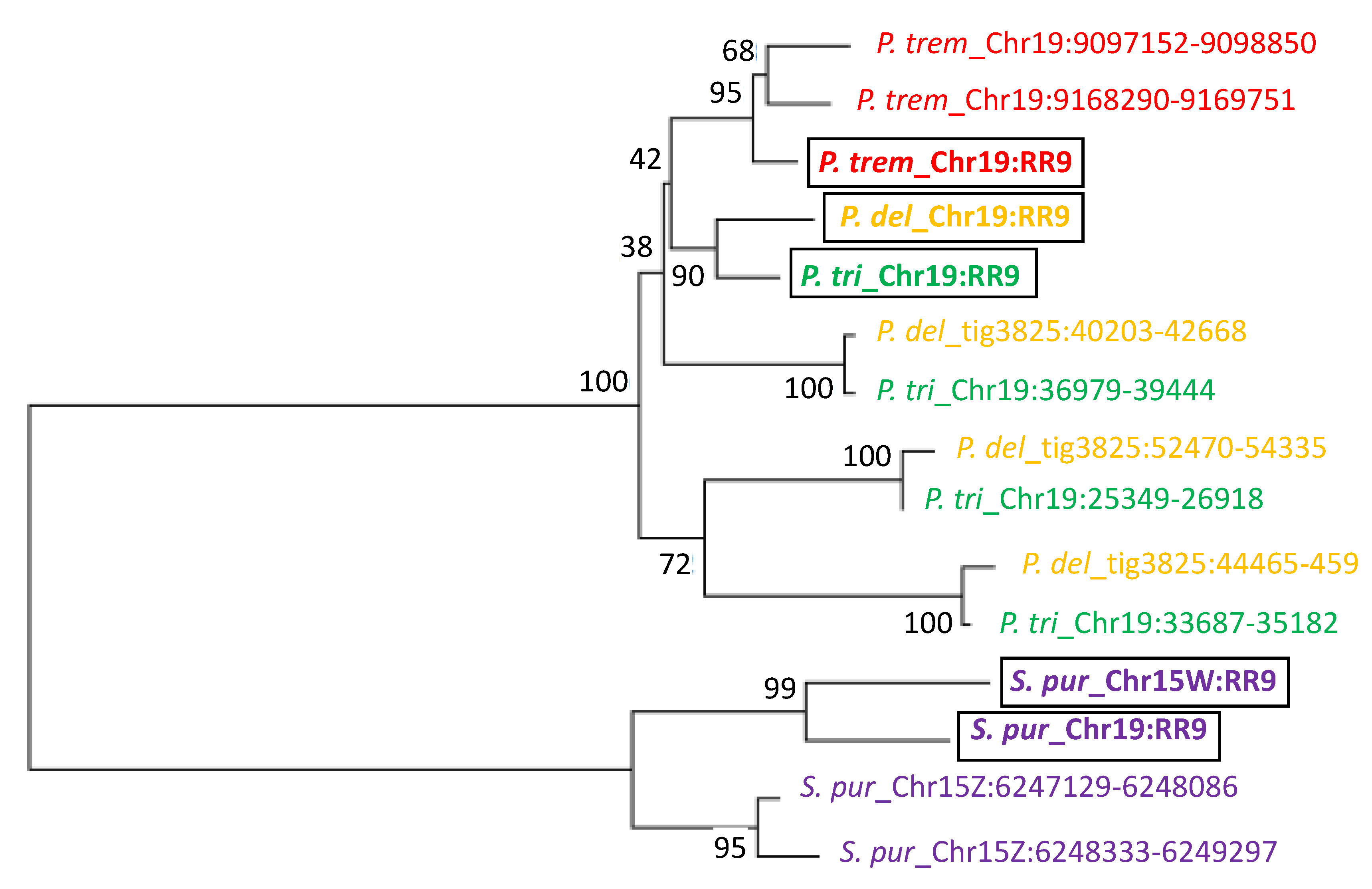
2.10. Inference of Phylogenetic Relationship of the Homologous Sequences in the SDRs
3. Results
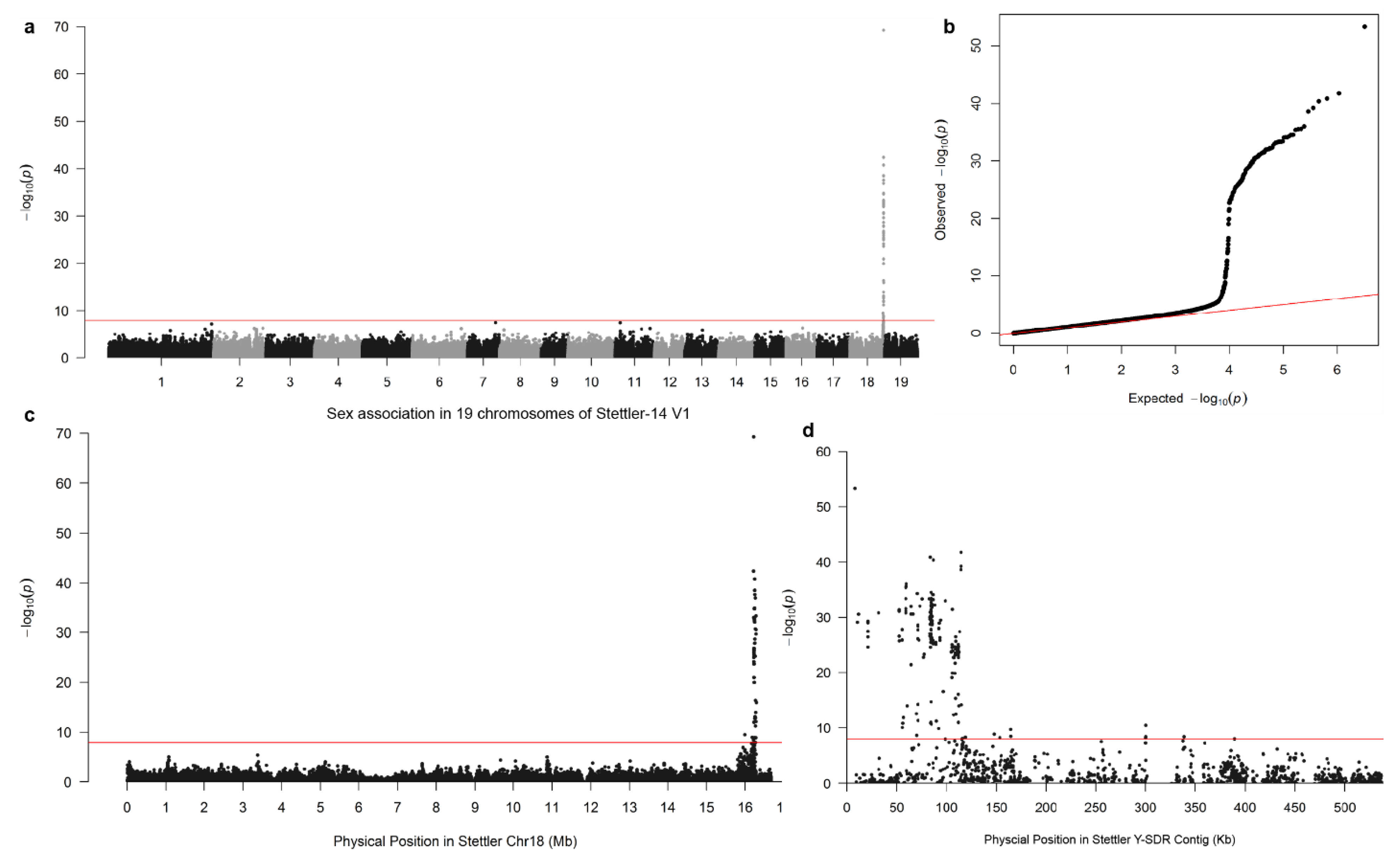
3.1. Identification of Sex-Associated Scaffolds Based on SNP Associations
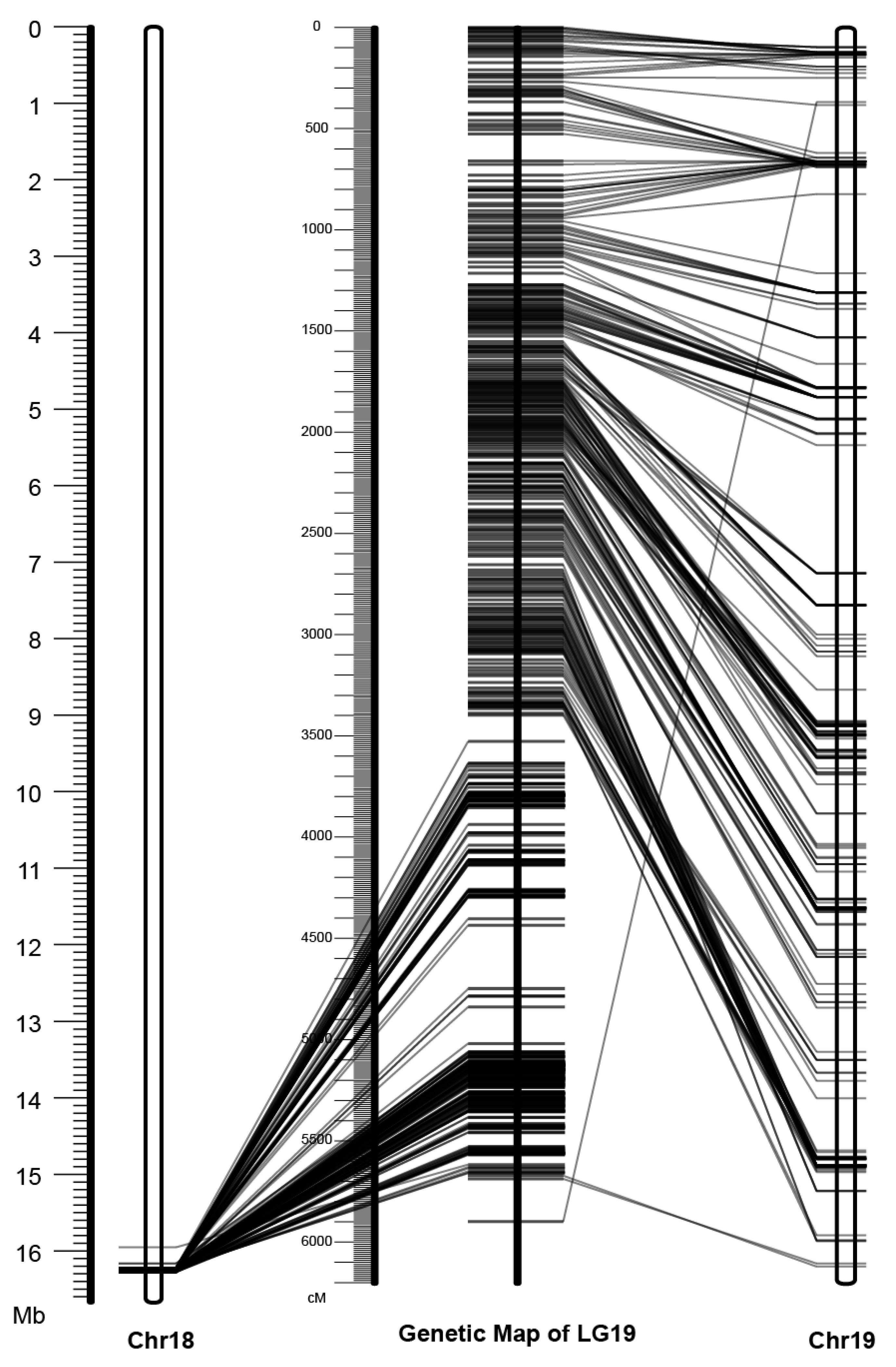
3.2. Mapping of the Y-SDR to Chr19
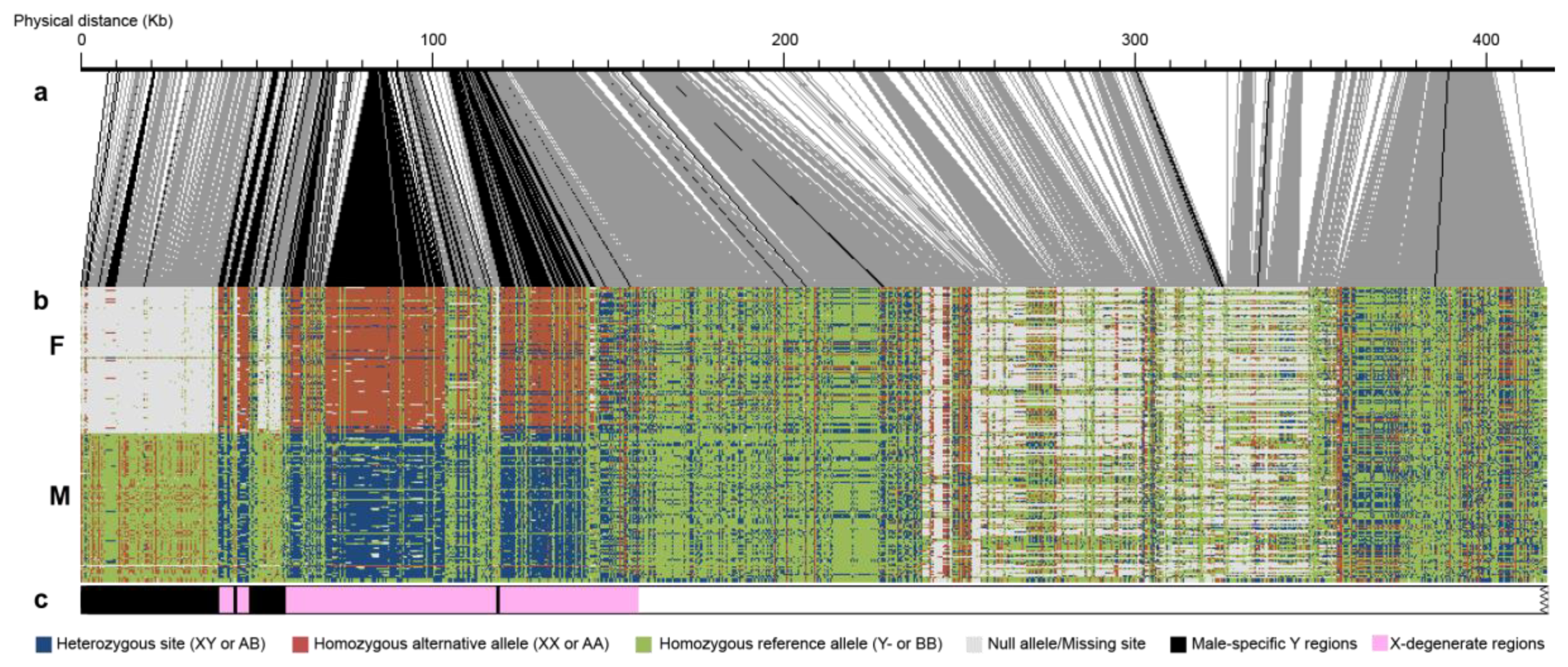
3.3. Confirmation of Male Heterogamety
3.4. Male-Specific Regions
3.5. Genomic Composition of the Y-SDR
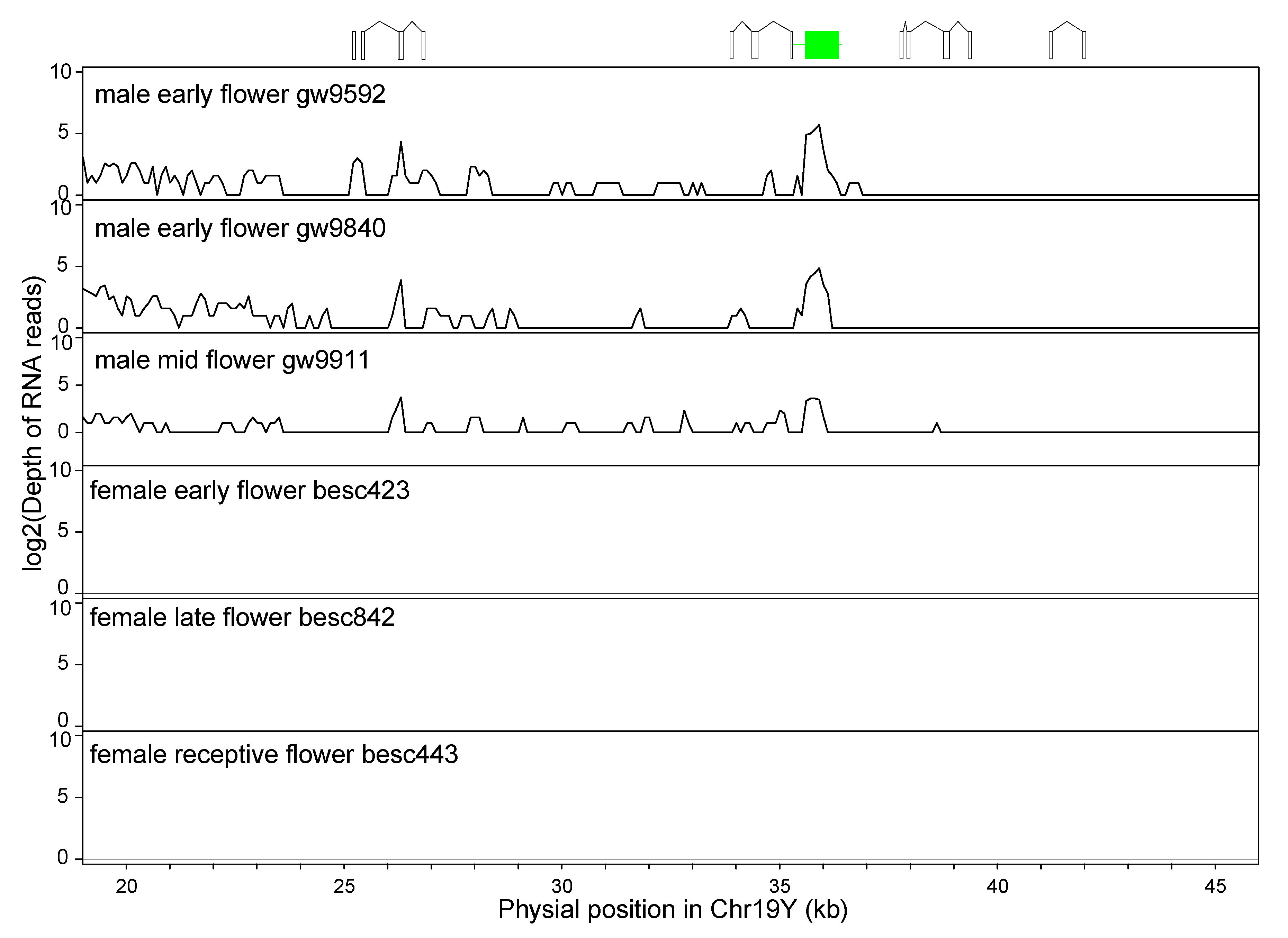
3.6. Inverted Repeats (IRs) in the Y-SDR
4. Discussion
Availability of Data
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Darwin, C. On the Origin of Species by Means of Natural Selection, or Preservation of Favoured Races in the Struggle for Life; John Murray: London, UK, 1859. [Google Scholar]
- Hosken, D.J.; House, C.M. Sexual selection. Curr. Biol. 2011, 21, R62–R65. [Google Scholar] [CrossRef] [PubMed]
- Henry, I.M.; Akagi, T.; Tao, R.; Comai, L. One Hundred Ways to Invent the Sexes: Theoretical and Observed Paths to Dioecy in Plants. Annu. Rev. Plant Biol. 2018, 69, 553–575. [Google Scholar] [CrossRef] [PubMed]
- Renner, S.S. The relative and absolute frequencies of angiosperm sexual systems: Dioecy, monoecy, gynodioecy, and an updated online database. Am. J. Bot. 2014, 101, 1588–1596. [Google Scholar] [CrossRef] [PubMed]
- Balounova, V.; Gogela, R.; Cegan, R.; Cangren, P.; Zluvova, J.; Safar, J.; Kovacova, V.; Bergero, R.; Hobza, R.; Vyskot, B.; et al. Evolution of sex determination and heterogamety changes in section Otites of the genus Silene. Sci. Rep. 2019, 9, 1045. [Google Scholar] [CrossRef] [PubMed]
- Charlesworth, D. Plant sex chromosome evolution. J. Exp. Bot. 2013, 64, 405–420. [Google Scholar] [CrossRef]
- Bachtrog, D. Y-chromosome evolution: Emerging insights into processes of Y-chromosome degeneration. Nat. Rev. Genet. 2013, 14, 113–124. [Google Scholar] [CrossRef]
- Bachtrog, D.; Mank, J.E.; Peichel, C.L.; Kirkpatrick, M.; Otto, S.P.; Ashman, T.-L.; Hahn, M.W.; Kitano, J.; Mayrose, I.; Ming, R.; et al. Sex Determination: Why So Many Ways of Doing It? PLoS Biol. 2014, 12. [Google Scholar] [CrossRef]
- Charlesworth, D. Plant contributions to our understanding of sex chromosome evolution. New Phytol. 2015, 208, 52–65. [Google Scholar] [CrossRef]
- Moore, R.C.; Harkess, A.E.; Weingartner, L.A. How to be a seXY plant model: A holistic view of sex-chromosome research. Am. J. Bot. 2016, 103, 1379–1382. [Google Scholar] [CrossRef][Green Version]
- Tennessen, J.A.; Wei, N.; Straub, S.C.K.; Govindarajulu, R.; Liston, A.; Ashman, T.-L. Repeated translocation of a gene cassette drives sex-chromosome turnover in strawberries. PLoS Biol. 2018, 16, e2006062. [Google Scholar] [CrossRef]
- Cronk, Q.C.B.; Needham, I.; Rudall, P.J. Evolution of Catkins: Inflorescence Morphology of Selected Salicaceae in an Evolutionary and Developmental Context. Front. Plant Sci. 2015, 6, 1030. [Google Scholar] [CrossRef] [PubMed]
- Geraldes, A.; Hefer, C.A.; Capron, A.; Kolosova, N.; Martinez-Nuñez, F.; Soolanayakanahally, R.Y.; Stanton, B.; Guy, R.D.; Mansfield, S.D.; Douglas, C.J.; et al. Recent y chromosome divergence despite ancient origin of dioecy in poplars (Populus). Mol. Ecol. 2015, 24, 3243–3256. [Google Scholar] [CrossRef] [PubMed]
- Pucholt, P.; Rönnberg-Wästljung, A.; Berlin, S. Single locus sex determination and female heterogamety in the basket willow (Salix viminalis L.). Heredity (Edinb) 2015, 114, 575–583. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Ye, N.; Zhang, D.; Chen, Y.; Fang, L.; Dai, X.; Yin, T. Different autosomes evolved into sex chromosomes in the sister genera of Salix and Populus. Sci. Rep. 2015, 5, e9076. [Google Scholar] [CrossRef]
- Zhou, R.; Macaya-Sanz, D.; Rodgers-Melnick, E.; Carlson, C.H.; Gouker, F.E.; Evans, L.M.; Schmutz, J.; Jenkins, J.W.; Yan, J.; Tuskan, G.A.; et al. Characterization of a large sex determination region in Salix purpurea L. (Salicaceae). Mol. Genet. Genom. 2018, 293, 1437–1452. [Google Scholar] [CrossRef] [PubMed]
- Paolucci, I.; Gaudet, M.; Jorge, V.; Beritognolo, I.; Terzoli, S.; Kuzminsky, E.; Muleo, R.; Scarascia Mugnozza, G.; Sabatti, M. Genetic linkage maps of Populus alba L. and comparative mapping analysis of sex determination across Populus species. Tree Genet. Genomes 2010, 6, 863–875. [Google Scholar] [CrossRef]
- Pakull, B.; Groppe, K.; Mecucci, F.; Gaudet, M.; Sabatti, M.; Fladung, M. Genetic mapping of linkage group XIX and identification of sex-linked SSR markers in a Populus tremula × Populus tremuloides cross. Can. J. For. Res. 2011, 41, 245–253. [Google Scholar] [CrossRef]
- Pakull, B.; Kersten, B.; Lüneburg, J.; Fladung, M. A simple PCR-based marker to determine sex in aspen. Plant Biol. 2014, 047300, 256–261. [Google Scholar] [CrossRef]
- Zhou, R.; Macaya-Sanz, D.; Carlson, C.H.; Schmutz, J.; Jenkins, J.W.; Kudrna, D.; Sharma, A.; Sandor, L.; Shu, S.; Barry, K.; et al. A willow sex chromosome reveals convergent evolution of complex palindromic repeats. Genome Biol. 2020, 21, 38. [Google Scholar] [CrossRef]
- Sanderson, B.J.; Feng, G.; Hu, N.; Grady, J.; Carlson, C.H.; Smart, L.B.; Keefover-Ring, K.; Yin, T.; Ma, T.; Liu, J.; et al. Sex determination through X-Y heterogamety in Salix nigra. bioRxiv 2020, 2020.03.23.000919. [Google Scholar] [CrossRef]
- Yin, T.; DiFazio, S.P.; Gunter, L.E.; Zhang, X.; Sewell, M.M.; Woolbright, S.A.; Allan, G.J.; Kelleher, C.T.; Douglas, C.J.; Wang, M.; et al. Genome structure and emerging evidence of an incipient sex chromosome in Populus. Genome Res. 2008, 18, 422–430. [Google Scholar] [CrossRef] [PubMed]
- Müller, N.A.; Kersten, B.; Leite Montalvão, A.P.; Mähler, N.; Bernhardsson, C.; Bräutigam, K.; Carracedo Lorenzo, Z.; Hoenicka, H.; Kumar, V.; Mader, M.; et al. A single gene underlies the dynamic evolution of poplar sex determination. Nat. Plants 2020, 6, 630–637. [Google Scholar] [CrossRef] [PubMed]
- Hofmeister, B.T.; Denkena, J.; Colomé-Tatché, M.; Shahryary, Y.; Hazarika, R.; Grimwood, J.; Mamidi, S.; Jenkins, J.; Grabowski, P.P.; Sreedasyam, A.; et al. The somatic genetic and epigenetic mutation rate in a wild long-lived perennial Populus trichocarpa. bioRxiv 2019, 862623. [Google Scholar] [CrossRef]
- Koren, S.; Walenz, B.P.; Berlin, K.; Miller, J.R.; Bergman, N.H.; Phillippy, A.M. Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017, 27, 722–736. [Google Scholar] [CrossRef] [PubMed]
- Evans, L.M.; Slavov, G.T.; Rodgers-Melnick, E.; Martin, J.; Ranjan, P.; Muchero, W.; Brunner, A.M.; Schackwitz, W.; Gunter, L.; Chen, J.-G.; et al. Population genomics of Populus trichocarpa identifies signatures of selection and adaptive trait associations. Nat. Genet. 2014, 46, 1089–1096. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Picard Software. Available online: http://broadinstitute.github.io/picard/ (accessed on 25 October 2019).
- Van der Auwera, G.A.; Carneiro, M.O.; Hartl, C.; Poplin, R.; del Angel, G.; Levy-Moonshine, A.; Jordan, T.; Shakir, K.; Roazen, D.; Thibault, J.; et al. From FastQ Data to High-Confidence Variant Calls: The Genome Analysis Toolkit Best Practices Pipeline. Curr. Protoc. Bioinform. 2013, 43, 11.10.1–11.10.33. [Google Scholar]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.R.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Harman-Ware, A.E.; Macaya Sanz, D.; Abeyratne, C.R.; Doeppke, C.; Haiby, K.; Tuskan, G.A.; Stanton, B.; DiFazio, S.P.; Davis, M.F. Accurate Determination of Genotypic Variance of Cell Wall Characteristics of a Populus trichocarpa Pedigree Using High-Throughput Pyrolysis-Molecular Beam Mass Spectrometry. Biotechnol. Biofuels 2020. [Google Scholar] [CrossRef]
- Margarido, G.R.; Souza, A.P.; Garcia, A.A. OneMap: Software for genetic mapping in outcrossing species. Hereditas 2007, 144, 78–79. [Google Scholar] [CrossRef] [PubMed]
- Muchero, W.; Guo, J.; DiFazio, S.P.; Chen, J.-G.; Ranjan, P.; Slavov, G.T.; Gunter, L.E.; Jawdy, S.; Bryan, A.C.; Sykes, R.; et al. High-resolution genetic mapping of allelic variants associated with cell wall chemistry in Populus. BMC Genom. 2015, 16, 24. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Zhang, X.; Miao, C.; Zhang, J.; Ming, R.; Schnable, J.C.; Schnable, P.S.; Lyons, E.; Lu, J. ALLMAPS: Robust scaffold ordering based on multiple maps. Genome Biol. 2015, 16, 3. [Google Scholar] [CrossRef]
- Solovyev, V.; Kosarev, P.; Seledsov, I.; Vorobyev, D. Automatic annotation of eukaryotic genes, pseudogenes and promoters. Genome Biol. 2006, 7, S10. [Google Scholar] [CrossRef] [PubMed]
- FGENESH Software. Available online: http://www.softberry.com/berry.phtml?topic=fgenesh (accessed on 1 August 2019).
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 2012, 40, 1178–1186. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Wilm, A.; Higgins, D.G.; Valentin, F.; Blackshields, G.; McWilliam, H.; Wallace, I.M.; Thompson, J.D.; Larkin, M.A.; Brown, N.P.; McGettigan, P.A.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar]
- Yang, Z. PAML 4: Phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 2007, 24, 1586–1591. [Google Scholar] [CrossRef]
- Ellinghaus, D.; Kurtz, S.; Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinform. 2008, 9, 18. [Google Scholar] [CrossRef]
- Steinbiss, S.; Willhoeft, U.; Gremme, G.; Kurtz, S. Fine-grained annotation and classification of de novo predicted LTR retrotransposons. Nucleic Acids Res. 2009, 37, 7002–7013. [Google Scholar] [CrossRef]
- Benson, G. Tandem repeats finder: A program to analyze DNA sequences. Nucleic Acids Res. 1999, 27, 573–580. [Google Scholar] [CrossRef] [PubMed]
- Richards, E.J.; Ausubel, F.M. Isolation of a higher eukaryotic telomere from Arabidopsis thaliana. Cell 1988, 53, 127–136. [Google Scholar] [CrossRef]
- Repeatmasker and Repeatmodeler software. Available online: http://www.repeatmasker.org (accessed on 30 November 2019).
- JGI Portal. Available online: https://genome.jgi.doe.gov/portal/ (accessed on 26 April 2019).
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef]
- Integrative Genomics Viewer. Available online: https://software.broadinstitute.org/software/igv/ (accessed on 2 August 2019).
- Tamura, K.; Peterson, D.; Stecher, G.; Peterson, N.; Kumar, S.; Nei, M. MEGA5: Molecular Evolutionary Genetics Analysis Using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- Ingvarsson, P.K. Multilocus patterns of nucleotide polymorphism and the demographic history of Populus tremula. Genetics 2008, 180, 329–340. [Google Scholar] [CrossRef] [PubMed]
- Charlesworth, D. Plant Sex Chromosomes. Annu. Rev. Plant Biol. 2016, 67, 397–420. [Google Scholar] [CrossRef] [PubMed]
- Chefdor, F.; Héricourt, F.; Koudounas, K.; Carqueijeiro, I.; Courdavault, V.; Mascagni, F.; Bertheau, L.; Larcher, M.; Depierreux, C.; Lamblin, F.; et al. Highlighting type A RRs as potential regulators of the dkHK1 multi-step phosphorelay pathway in Populus. Plant Sci. 2018, 277, 68–78. [Google Scholar] [CrossRef]
- Melnikova, N.V.; Kudryavtseva, A.V.; Borkhert, E.V.; Pushkova, E.N.; Fedorova, M.S.; Snezhkina, A.V.; Krasnov, G.S.; Dmitriev, A.A. Sex-specific polymorphism of MET1 and ARR17 genes in Populus × sibirica. Biochimie 2019, 162, 26–32. [Google Scholar] [CrossRef]
- Matzke, M.A.; Mosher, R.A. RNA-directed DNA methylation: An epigenetic pathway of increasing complexity. Nat. Rev. Genet. 2014, 15, 394–408. [Google Scholar] [CrossRef]


| seqID | Start | End | Ori | Size (bp) |
|---|---|---|---|---|
| scaffold_1208 | 1 | 85,028 | - | 85,028 |
| scaffold_1534 | 95,029 | 169,954 | - | 74,926 |
| scaffold_43 | 179,955 | 543,096 | + | 363,142 |
| LTR-ID | SuperFamily | SITE Count | Substitution Rate (SE) | Element Start | Element End | l/rLTR Length | TSD Motif | Pfam |
|---|---|---|---|---|---|---|---|---|
| Ltr-y-a | Gypsy | 162 | 0.078(0.025) | 64,125 | 69,169 | 175/162 | aaat | Retrotrans_gag-223..315;RVP_2-479..564;RVT_1-724..861 |
| Ltr-y-b | Gypsy | 409 | 0.007(0.004) | 90,484 | 95,727 | 409/409 | tattt | Retrotrans_gag-262..351;RVP_2-512..601;RVT_1-746..906;rve-1255..1363;Chromo-1552..1599 |
| Ltr-y-c | Copia | 166 | 0.045(0.017) | 96,610 | 98,445 | 166/166 | tttc | UBN2_3-153..247;RVT_2-244..308 |
| Ltr-y-d | Copia | 294 | 0.065(0.016) | 99,790 | 104,250 | 303/295 | ttca | DUF4219-126..152;UBN2-204..281;gag_pre-integrs-514..572;rve-587..665;RVT_2-992..1139 |
| GeneID | Start | End | Size | Strand | Description | Annotation in V3 Genome | Nisqually V4 | dS(S.E.) | dN(S.E.) |
|---|---|---|---|---|---|---|---|---|---|
| Po14v11g055363m | 52,354 | 56,656 | 4303 | + | T-complex protein 1 subunit γ (TCP-1,CCT3, TRIC5) | Potri.018G138200; Potri.T046300 | Potriv41g055126m; | 0.0737(0.0136) | 0.0016(0.0012) |
| Potriv41g057391m | 0.0600(0.012) | 0.0008(0.0008) | |||||||
| Po14v11g055362m | 59,327 | 69,212 | 9886 | + | Chloride channel protein CLC-C | Potri.018G138100; Potri.T046200 | Potriv41g055125m; | 0.0117(0.0044) | 0.0206(0.0078) |
| Potriv41g057390m | 0.033(0.0075) | 0.0105(0.0025) | |||||||
| Po14v11g055360m | 73,031 | 82,422 | 9392 | + | similar to DNA (cytosine-5)-methyltransferase AthI (EC 2.1.1.37) (MET1) | Potri.018G138000; Potri.T046100 | Potriv41g055122m; | 0.0194(0.0041) | 0.006(0.0013) |
| Potriv41g057386m | 0.0158(0.0036) | 0.0057(0.0013) | |||||||
| Po14v11g055357m | 96,006 | 105,907 | 9902 | + | Archaeal ATPase (Arch_ATPase)//Leucine rich repeat (LRR_8) | Potri.018G137900 | NA | NA | NA |
| Po14v11g055355m | 116,621 | 118,021 | 1401 | - | hypothetical protein | Potri.018G137700 | Potriv41g055119m; | 0(0) | 0.0206(0.0078) |
| Potriv41g057380m | 0(0) | 0.0236(0.0084) |
| Start (bp) | End (bp) | Size (bp) | ARMs |
|---|---|---|---|
| 23,726 | 25,349 | 1624 | ARM-1 |
| 25,381 | 29,199 | 3819 | ARM-2 |
| 29,200 | 31,389 | 2190 | Spacer-1 |
| 31,390 | 35,225 | 3836 | ARM-3 |
| 35,226 | 37,885 | 2660 | Spacer-2 |
| 37,886 | 40,646 | 2761 | ARM-4a |
| 40,531 | 41,958 | 1428 | ARM-4b |
| Po14v11g057342m (Chr19) | Size (bp) | ARM-1 | ARM-2 | ARM-3 | ARM-4a | ARM-4b |
|---|---|---|---|---|---|---|
| exon1 (5′-UTR) | 76 | + | - | + | - | + |
| exon2 | 139 | Absent | - | + | - | Absent |
| exon3 | 74 | Absent | - | +(truncated) | - | + |
| exon4 | 78 | Absent | Absent | Absent | -(Spacer) | Absent |
| exon5 | 71 | Absent | Absent | Absent | Absent | Absent |
| exon6 (3′-UTR) | 373 | Absent | Absent | Absent | Absent | Absent |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhou, R.; Macaya-Sanz, D.; Schmutz, J.; Jenkins, J.W.; Tuskan, G.A.; DiFazio, S.P. Sequencing and Analysis of the Sex Determination Region of Populus trichocarpa. Genes 2020, 11, 843. https://doi.org/10.3390/genes11080843
Zhou R, Macaya-Sanz D, Schmutz J, Jenkins JW, Tuskan GA, DiFazio SP. Sequencing and Analysis of the Sex Determination Region of Populus trichocarpa. Genes. 2020; 11(8):843. https://doi.org/10.3390/genes11080843
Chicago/Turabian StyleZhou, Ran, David Macaya-Sanz, Jeremy Schmutz, Jerry W. Jenkins, Gerald A. Tuskan, and Stephen P. DiFazio. 2020. "Sequencing and Analysis of the Sex Determination Region of Populus trichocarpa" Genes 11, no. 8: 843. https://doi.org/10.3390/genes11080843
APA StyleZhou, R., Macaya-Sanz, D., Schmutz, J., Jenkins, J. W., Tuskan, G. A., & DiFazio, S. P. (2020). Sequencing and Analysis of the Sex Determination Region of Populus trichocarpa. Genes, 11(8), 843. https://doi.org/10.3390/genes11080843





