Gene Networks Driving Genetic Variation in Milk and Cheese-Making Traits of Spanish Assaf Sheep
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals and Phenotypes
2.2. Ethics Committee Approval
2.3. Design of a Custom Ovine SNP Array and Genotypes
2.4. Genetic Parameter Estimation
2.5. Genome-Wide Association Study and Pleiotropy Analysis
2.6. Stepwise Multiple Regression Analysis
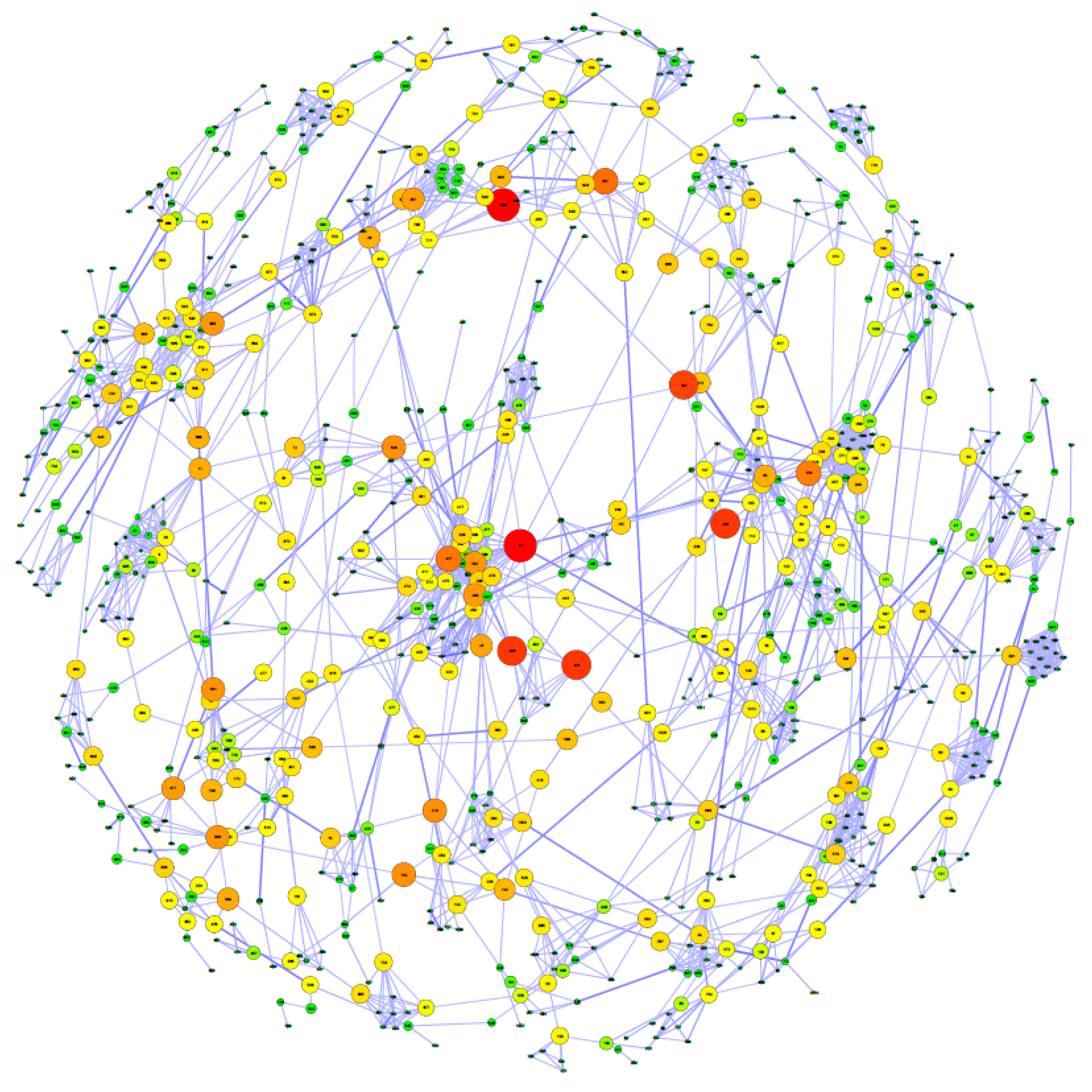
2.7. Gene Co-association Network and Functional Enrichment Analysis
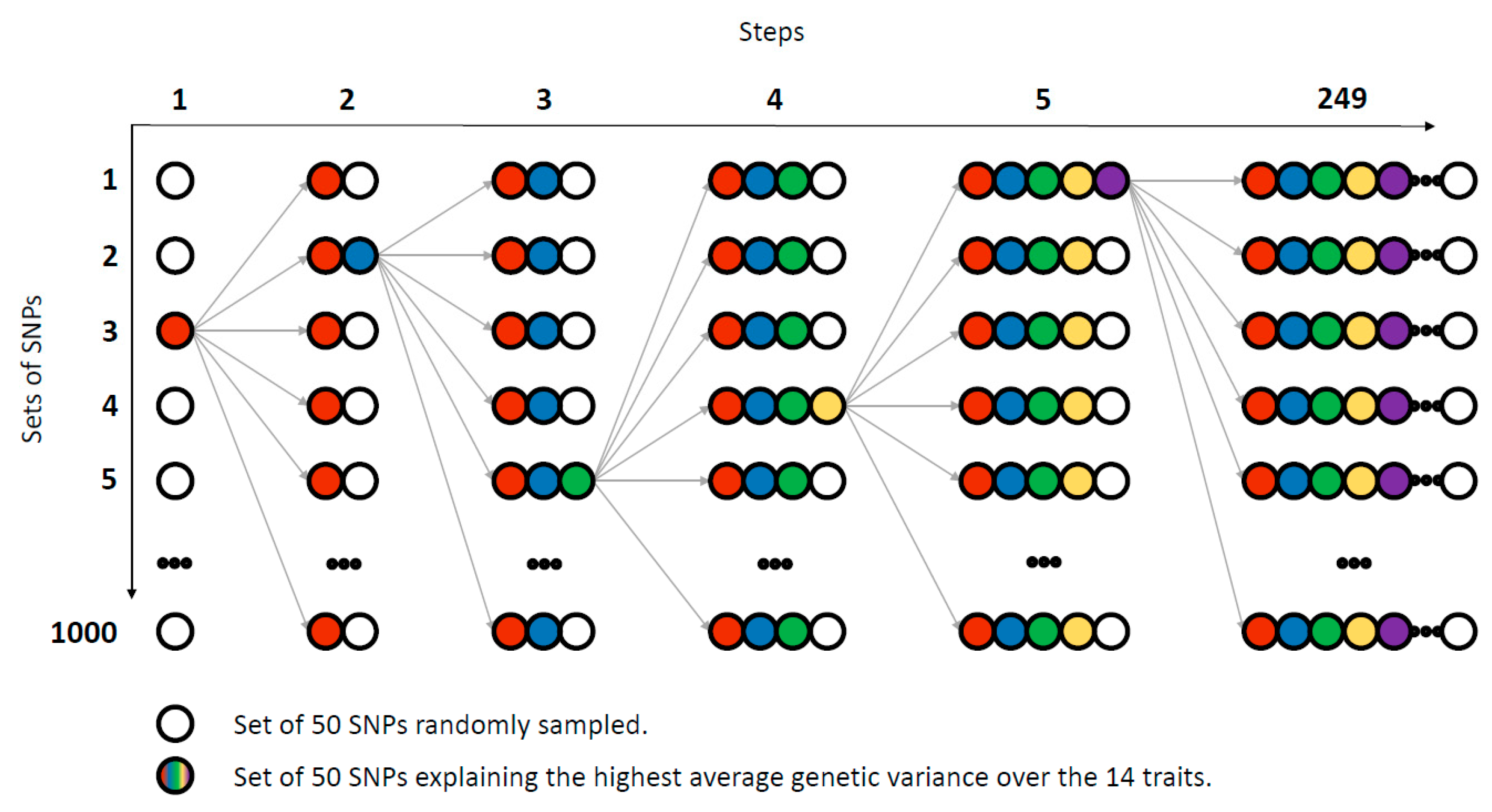
2.8. Genetic Variance Explained Randomly and Relationship Matrix
3. Results
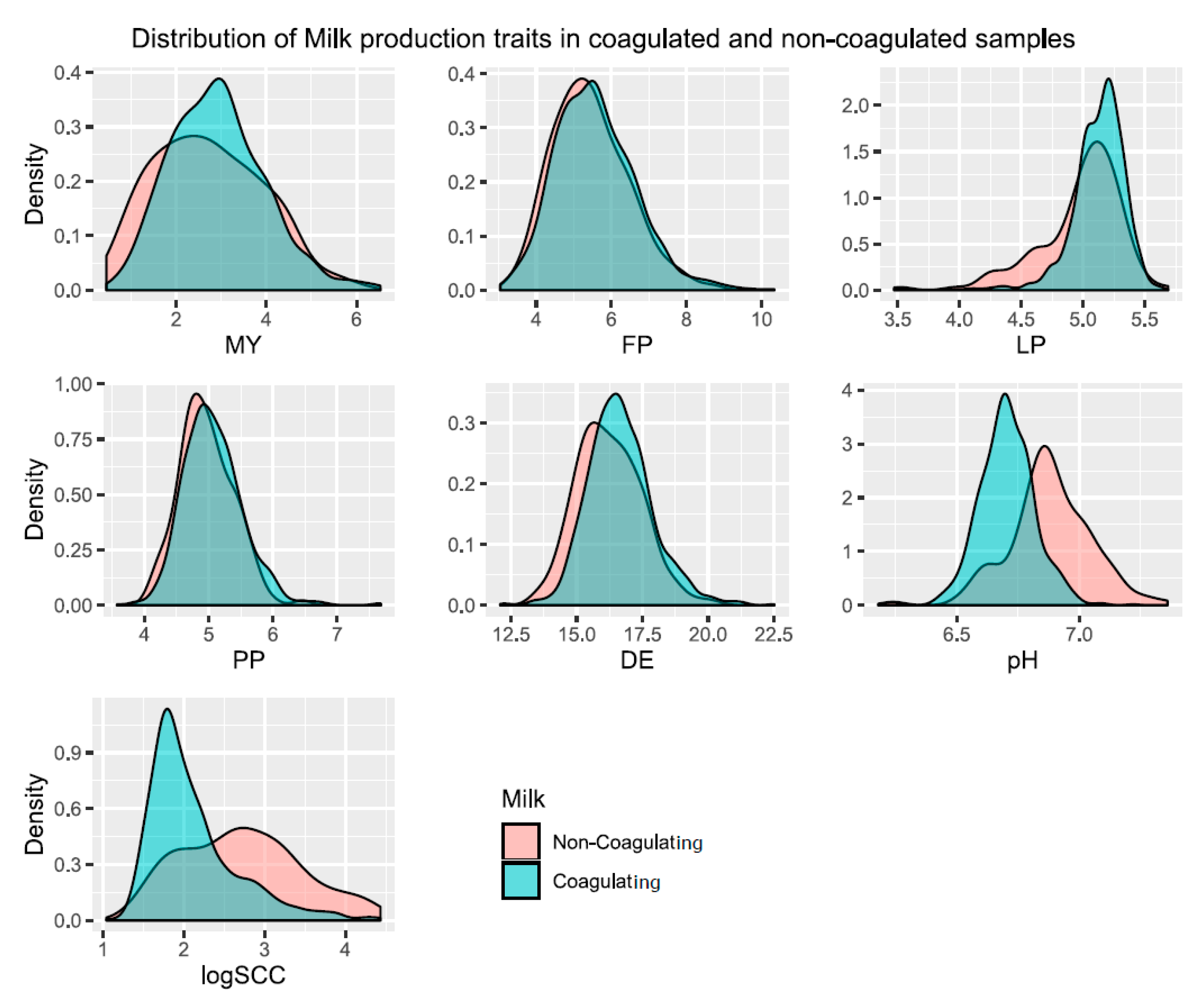
3.1. Genetic Parameters of the Analysed Phenotypes
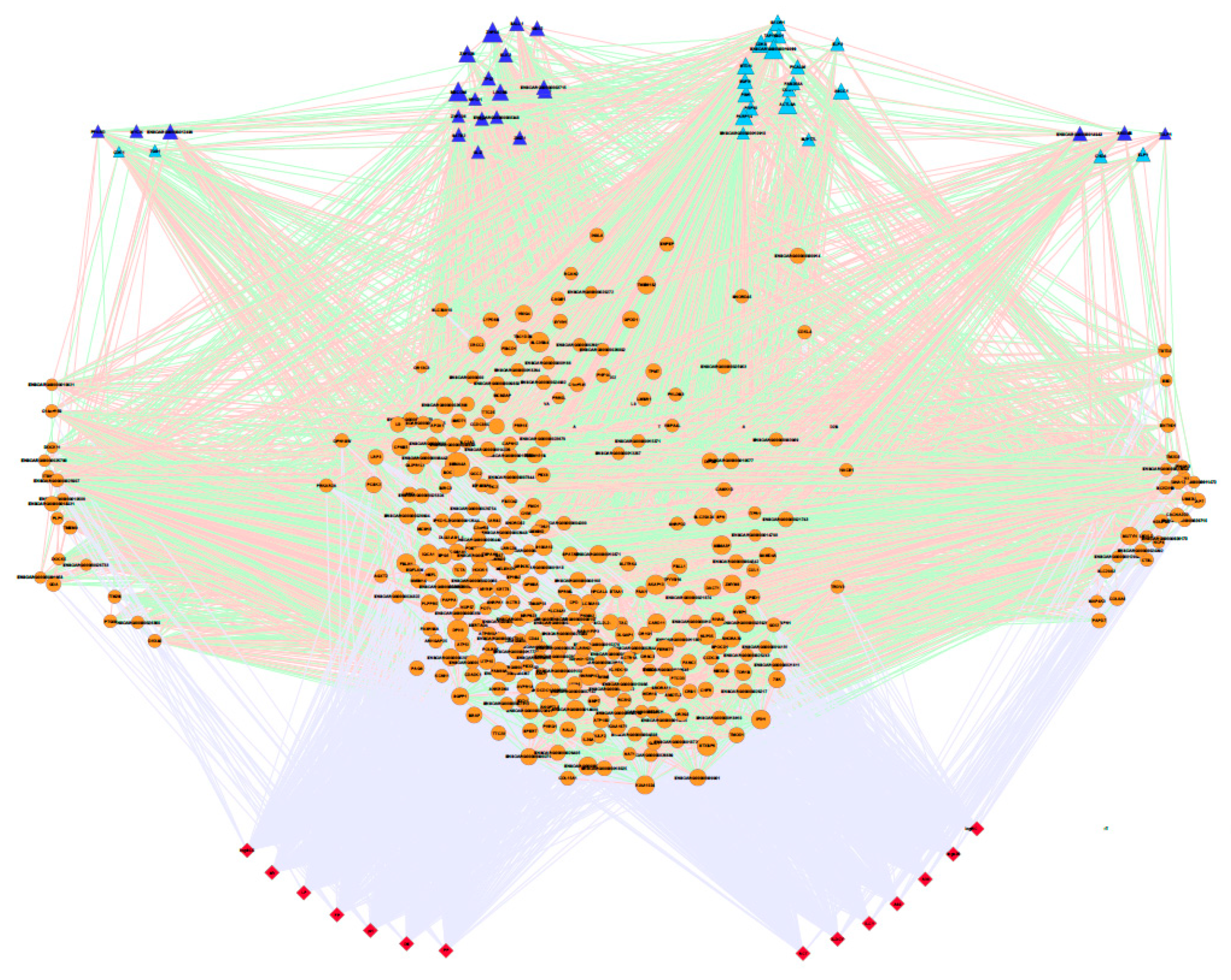
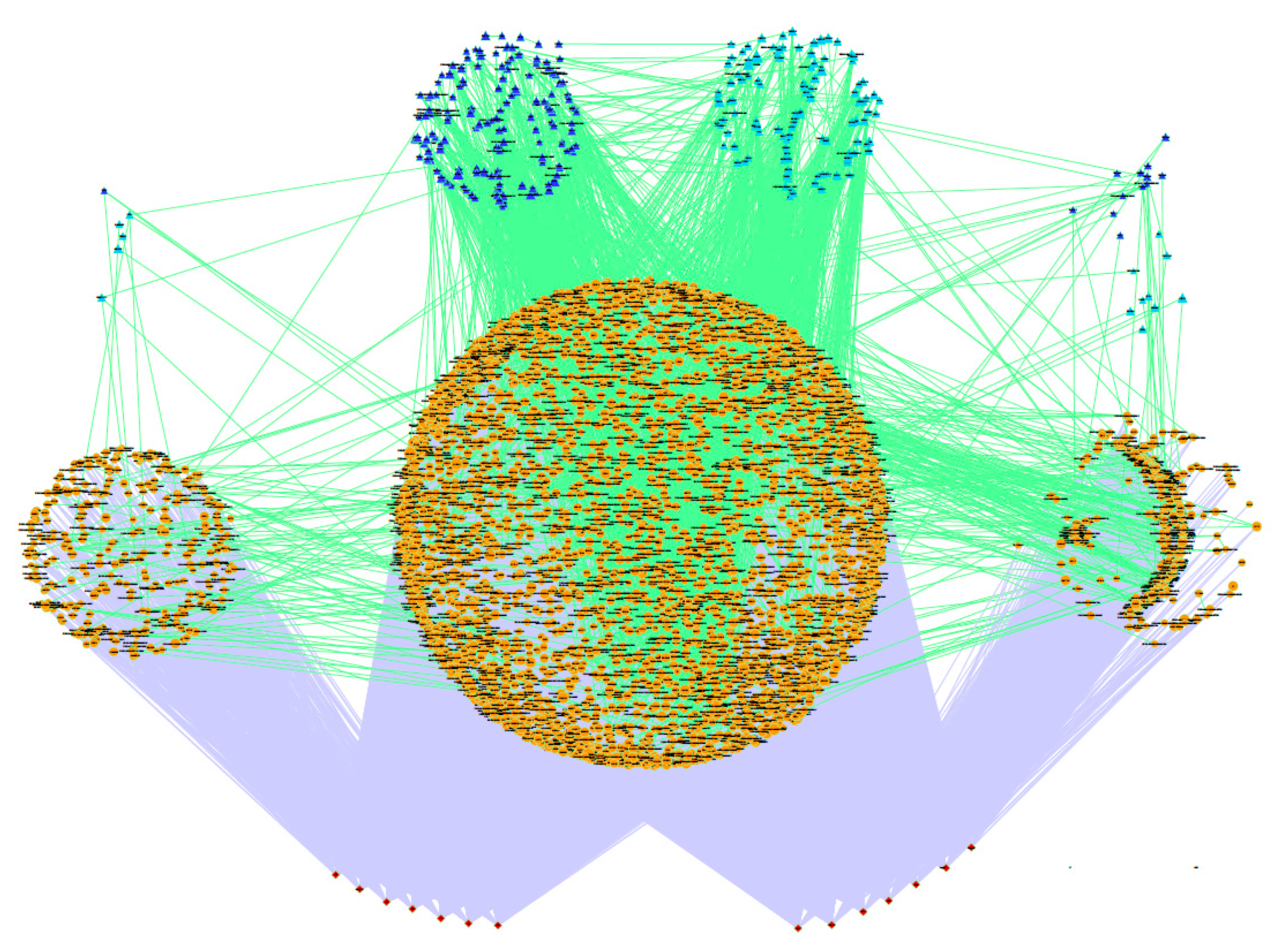
3.2. Stepwise Analysis, Gene Co-association Network and Potential Regulators
3.3. Identification of Enriched Gene Set
3.4. Pleiotropy of the Selected Genes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- FAOSTAT. Food and Agriculture Organization the United Nations Statistics Division. Available online: http://www.fao.org/faostat/ (accessed on 17 September 2019).
- Jaramillo, D.P.; Zamora, A.; Guamis, B.; Rodríguez, M.; Trujillo, A.J. Cheesemaking aptitude of two Spanish dairy ewe breeds: Changes during lactation and relationship between physico-chemical and technological properties. Small Rumin. Res. 2008, 78, 48–55. [Google Scholar] [CrossRef]
- Moioli, B.; D’Andrea, M.; Pilla, F. Candidate genes affecting sheep and goat milk quality. Small Rumin. Res. 2007, 68, 179–192. [Google Scholar] [CrossRef]
- Pappa, E.C.; Kandarakis, I.; Anifantakis, E.M.; Zerfiridis, G.K. Influence of types of milk and culture on the manufacturing practices, composition and sensory characteristics of Teleme cheese during ripening. Food Control 2006, 17, 570–581. [Google Scholar] [CrossRef]
- Gutiérrez, J.P.; Legaz, E.; Goyache, F. Genetic parameters affecting 180-days standardised milk yield, test-day milk yield and lactation length in Spanish Assaf (Assaf.E) dairy sheep. Small Rumin. Res. 2007, 70, 233–238. [Google Scholar] [CrossRef][Green Version]
- Pollott, G.E.; Gootwine, E. A genetic analysis of complete lactation milk production in Improved Awassi sheep. In Proceedings of the Livestock Production Science; Elsevier: Amsterdam, The Netherlands, 2001; Volume 71, pp. 37–47. [Google Scholar]
- Baro, J.A.; Carriedo, J.A.; San Primitivo, F. Genetic Parameters of Test Day Measures for Somatic Cell Count, Milk Yield, and Protein Percentage of Milking Ewes. J. Dairy Sci. 1994, 77, 2658–2662. [Google Scholar] [CrossRef]
- Bittante, G.; Cipolat-Gotet, C.; Pazzola, M.; Dettori, M.L.; Vacca, G.M.; Cecchinato, A. Genetic analysis of coagulation properties, curd firming modeling, milk yield, composition, and acidity in Sarda dairy sheep. J. Dairy Sci. 2017, 100, 385–394. [Google Scholar] [CrossRef]
- Othmane, M.H.; Carriedo, J.; San Primitivo, F.; De la Fuente, L. Genetic parameters for lactation traits of milking ewes: Protein content and composition, fat, somatic cells and individual laboratory cheese yield. Genet. Sel. Evol. 2002, 34, 581. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Mayor, M.; Pong-Wong, R.; Gutiérrez-Gil, B.; Garzón, A.; de la Fuente, L.F.; Arranz, J.J. Phenotypic and genetic parameter estimates of cheese-making traits and their relationships with milk production, composition and functional traits in Spanish Assaf sheep. Livest. Sci. 2019, 228, 76–83. [Google Scholar] [CrossRef]
- García-Gámez, E.; Gutiérrez-Gil, B.; Sahana, G.; Sánchez, J.-P.; Bayón, Y.; Arranz, J.-J.; Jiang, L.; Liu, J.; Sun, D.; Ma, P.; et al. GWA Analysis for Milk Production Traits in Dairy Sheep and Genetic Support for a QTN Influencing Milk Protein Percentage in the LALBA Gene. PLoS ONE 2012, 7, e47782. [Google Scholar] [CrossRef]
- Sanz, A.; Serrano, C.; Ranera, B.; Dervishi, E.; Zaragoza, P.; Calvo, J.H.; Rodellar, C. Novel polymorphisms in the 5′UTR of FASN, GPAM, MC4R and PLIN1 ovine candidate genes: Relationship with gene expression and diet. Small Rumin. Res. 2015, 123, 70–74. [Google Scholar] [CrossRef]
- Noce, A.; Pazzola, M.; Dettori, M.L.; Amills, M.; Castelló, A.; Cecchinato, A.; Bittante, G.; Vacca, G.M. Variations at regulatory regions of the milk protein genes are associated with milk traits and coagulation properties in the Sarda sheep. Anim. Genet. 2016, 47, 717–726. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.L.; Park, C.A.; Reecy, J.M. Developmental progress and current status of the Animal QTLdb. Nucleic Acids Res. 2016, 44, D827–D833. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, M.P.; Ramayo-Caldas, Y.; Wolf, V.; Laithier, C.; El Jabri, M.; Michenet, A.; Boussaha, M.; Taussat, S.; Fritz, S.; Delacroix-Buchet, A.; et al. Sequence-based GWAS, network and pathway analyses reveal genes co-associated with milk cheese-making properties and milk composition in Montbéliarde cows. Genet. Sel. Evol. 2019, 51, 34. [Google Scholar] [CrossRef] [PubMed]
- Reverter, A.; Chan, E.K.F. Combining partial correlation and an information theory approach to the reversed engineering of gene co-expression networks. Bioinformatics 2008, 24, 2491–2497. [Google Scholar] [CrossRef]
- Mudadu, M.A.; Porto-Neto, L.R.; Mokry, F.B.; Tizioto, P.C.; Oliveira, P.S.N.; Tullio, R.R.; Nassu, R.T.; Niciura, S.C.M.; Tholon, P.; Alencar, M.M.; et al. Genomic structure and marker-derived gene networks for growth and meat quality traits of Brazilian Nelore beef cattle. BMC Genomics 2016, 17, 235. [Google Scholar]
- Ramayo-Caldas, Y.; Mármol-Sánchez, E.; Ballester, M.; Sánchez, J.P.; González-Prendes, R.; Amills, M.; Quintanilla, R. Integrating genome-wide co-association and gene expression to identify putative regulators and predictors of feed efficiency in pigs. Genet. Sel. Evol. 2019, 51, 48. [Google Scholar] [CrossRef]
- Suárez-Vega, A.; Gutiérrez-Gil, B.; Klopp, C.; Tosser-Klopp, G.; Arranz, J.J. Variant discovery in the sheep milk transcriptome using RNA sequencing. BMC Genomics 2017, 18, 170. [Google Scholar] [CrossRef]
- Caballero-Villalobos, J.; Perea, J.M.; Angón, E.; Arias, R.; Garzón, A. Coagulation efficiency and its determinant factors: A case study for Manchega ewe milk in the region of Castilla-La Mancha, Spain. J. Dairy Sci. 2018, 101, 3878–3886. [Google Scholar] [CrossRef]
- Othmane, M.H.; De La Fuente, L.F.; Carriedo, J.A.; San Primitivo, F. Heritability and genetic correlations of test day milk yield and composition, individual laboratory cheese yield, and somatic cell count for dairy ewes. J. Dairy Sci. 2002, 85, 2692–2698. [Google Scholar] [CrossRef]
- VanRaden, P.M. Efficient Methods to Compute Genomic Predictions. J. Dairy Sci. 2008, 91, 4414–4423. [Google Scholar] [CrossRef]
- Reverter, A. Pedigromics: A network-inspired approach to visualise and analyse pedigree structures. Proc. Assoc. Advmt. Anim. Breed. Genet. 2019, 23, 540–543. [Google Scholar]
- Misztal, I.; Tsuruta, S.; Lourenco, D.; Aguilar, I.; Legarra, A.; Vitezica, Z. Manual for BLUPF90 Family of Programs. Available online: http://nce.ads.uga.edu/wiki/lib/exe/fetch.php?media=blupf90_all2.pdf (accessed on 15 October 2019).
- Garrick, D.J.; Taylor, J.F.; Fernando, R.L. Deregressing estimated breeding values and weighting information for genomic regression analyses. Genet. Sel. Evol. 2009, 41, 55. [Google Scholar] [CrossRef] [PubMed]
- Kasprzyk, A. BioMart: Driving a paradigm change in biological data management. Database 2011, 2011, bar049. [Google Scholar] [CrossRef] [PubMed]
- Bolormaa, S.; Pryce, J.E.; Reverter, A.; Zhang, Y.; Barendse, W.; Kemper, K.; Tier, B.; Savin, K.; Hayes, B.J.; Goddard, M.E. A Multi-Trait, Meta-analysis for Detecting Pleiotropic Polymorphisms for Stature, Fatness and Reproduction in Beef Cattle. PLoS Genet. 2014, 10, e1004198. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software Environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Miao, Y.R.; Jia, L.H.; Yu, Q.Y.; Zhang, Q.; Guo, A.Y. AnimalTFDB 3.0: A comprehensive resource for annotation and prediction of animal transcription factors. Nucleic Acids Res. 2019, 47, D33–D38. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Wang, J.; Jaehnig, E.J.; Shi, Z.; Zhang, B. WebGestalt 2019: Gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 2019, 47, W199–W205. [Google Scholar] [CrossRef]
- Mi, H.; Muruganujan, A.; Huang, X.; Ebert, D.; Mills, C.; Guo, X.; Thomas, P.D. Protocol Update for large-scale genome and gene function analysis with the PANTHER classification system (v.14.0). Nat. Protoc. 2019, 14, 703–721. [Google Scholar] [CrossRef]
- Leitner, G.; Silanikove, N.; Merin, U. Estimate of milk and curd yield loss of sheep and goats with intrammamary infection and its relation to somatic cell count. Small Rumin. Res. 2008, 74, 221–225. [Google Scholar] [CrossRef]
- Beerli, R.R.; Barbas, C.F. Engineering polydactyl zinc-finger transcription factors. Nat. Biotechnol. 2002, 20, 135–141. [Google Scholar] [CrossRef]
- Gehring, W.J. The homeobox in perspective. Trends Biochem. Sci. 1992, 17, 277–280. [Google Scholar] [CrossRef]
- Sharrocks, A.D. The ETS-domain transcription factor family. Nat. Rev. Mol. Cell Biol. 2001, 2, 827–837. [Google Scholar] [CrossRef] [PubMed]
- Scatà, M.C.; Napolitano, F.; Casu, S.; Carta, A.; De Matteis, G.; Signorelli, F.; Annicchiarico, G.; Catillo, G.; Moioli, B. Ovine acyl CoA: diacylglycerol acyltransferase 1—molecular characterization, polymorphisms and association with milk traits. Anim. Genet. 2009, 40, 737–742. [Google Scholar] [CrossRef] [PubMed]
- Rajeswari, M.R.; Jain, A. High-mobility-group chromosomal proteins, HMGA1 as potential tumour markers. Curr. Sci. 2002, 82, 838–844. [Google Scholar]
- Carbon, S.; Ireland, A.; Mungall, C.J.; Shu, S.; Marshall, B.; Lewis, S.; Hub, A.; Presence, W.; Group, W. AmiGO: Online access to ontology and annotation data. Bioinforma. Appl. NOTE 2009, 25, 288–289. [Google Scholar] [CrossRef]
- Blum, J.W.; Kunz, P.; Leuenberger, H.; Gautschi, K.; Keller, M. Thyroid hormones, blood plasma metabolites and haematological parameters in relationship to milk yield in dairy cows. Anim. Prod. 1983, 36, 93–104. [Google Scholar] [CrossRef]
- Staiger, E.A.; Thonney, M.L.; Buchanan, J.W.; Rogers, E.R.; Oltenacu, P.A.; Mateescu, R.G. Effect of prolactin, β-lactoglobulin, and κ-casein genotype on milk yield in East Friesian sheep. J. Dairy Sci. 2010, 93, 1736–1742. [Google Scholar] [CrossRef] [PubMed]
- Wegner, T.N.; Schuh, J.D.; Nelson, F.E.; Stott, G.H. Effect of Streess on Blood Leucocyte and Milk Somatic Cell Counts in Dairy Cows. J. Dairy Sci. 1976, 59, 949–956. [Google Scholar] [CrossRef]
- Almeida, O.F.X.; Yassouridis, A.; Forgas-Moya, I. Reduced Availability of Milk after Central Injections of Corticotropin-Releasing Hormone in Lactating Rats. Neuroendocrinology 1994, 59, 72–77. [Google Scholar] [CrossRef]
- Bionaz, M.; Loor, J.J. Gene networks driving bovine milk fat synthesis during the lactation cycle. BMC Genomics 2008, 9, 366. [Google Scholar] [CrossRef]
- Russell, T.D.; Palmer, C.A.; Orlicky, D.J.; Bales, E.S.; Chang, B.H.-J.; Chan, L.; McManaman, J.L. Mammary glands of adipophilin-null mice produce an amino-terminally truncated form of adipophilin that mediates milk lipid droplet formation and secretion. J. Lipid Res. 2008, 49, 206–216. [Google Scholar] [CrossRef]
- Sanchez, M.-P.; Govignon-Gion, A.; Croiseau, P.; Fritz, S.; Hozé, C.; Miranda, G.; Martin, P.; Barbat-Leterrier, A.; Letaïef, R.; Rocha, D.; et al. Within-breed and multi-breed GWAS on imputed whole-genome sequence variants reveal candidate mutations affecting milk protein composition in dairy cattle. Genet. Sel. Evol. 2017, 49, 68. [Google Scholar] [CrossRef] [PubMed]
- Knutsen, T.M.; Olsen, H.G.; Tafintseva, V.; Svendsen, M.; Kohler, A.; Kent, M.P.; Lien, S. Unravelling genetic variation underlying de novo-synthesis of bovine milk fatty acids. Sci. Rep. 2018, 8, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Crisà, A.; Marchitelli, C.; Pariset, L.; Contarini, G.; Signorelli, F.; Napolitano, F.; Catillo, G.; Valentini, A.; Moioli, B. Exploring polymorphisms and effects of candidate genes on milk fat quality in dairy sheep. J. Dairy Sci. 2010, 93, 3834–3845. [Google Scholar] [CrossRef] [PubMed]
- Cánovas, A.; Rincón, G.; Islas-Trejo, A.; Jimenez-Flores, R.; Laubscher, A.; Medrano, J.F. RNA sequencing to study gene expression and single nucleotide polymorphism variation associated with citrate content in cow milk. J. Dairy Sci. 2013, 96, 2637–2648. [Google Scholar] [CrossRef]
- Watanabe, K.; Stringer, S.; Frei, O.; Mirkov, M.U.; De Leeuw, C.; Polderman, T.J.C.; Van Der Sluis, S.; Andreassen, O.A.; Neale, B.M.; Posthuma, D. A global overview of pleiotropy and genetic architecture in complex traits. Nat. Genet. 2019, 51, 1339–1348. [Google Scholar] [CrossRef]
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef]
- Dávalos, A.; Goedeke, L.; Smibert, P.; Ramírez, C.M.; Warrier, N.P.; Andreo, U.; Cirera-Salinas, D.; Rayner, K.; Suresh, U.; Pastor-Pareja, J.C.; et al. miR-33a/b contribute to the regulation of fatty acid metabolism and insulin signaling. Proc. Natl. Acad. Sci. USA 2011, 108, 9232–9237. [Google Scholar] [CrossRef]
- Li, R.; Dudemaine, P.-L.; Zhao, X.; Lei, C.; Ibeagha-Awemu, E.M. Comparative Analysis of the miRNome of Bovine Milk Fat, Whey and Cells. PLoS ONE 2016, 11, e0154129. [Google Scholar] [CrossRef]
- Ramos, A.M.; Matos, C.A.P.; Russo-Almeida, P.A.; Bettencourt, C.M.V.; Matos, J.; Martins, A.; Pinheiro, C.; Rangel-Figueiredo, T. Candidate genes for milk production traits in Portuguese dairy sheep. Small Rumin. Res. 2009, 82, 117–121. [Google Scholar] [CrossRef]

| Traits 1 | Fixed factors 2 | Coagulation 3 | R-square (%) 4 | |||
|---|---|---|---|---|---|---|
| AGE_NB | NBL | DIM | FTD | |||
| Milk production traits | ||||||
| MY | *** | * | * | *** | *** | 34.52 |
| PP | ns | ns | ns | *** | ns | 28.42 |
| FP | ns | ns | ns | *** | ns | 20.57 |
| LP | ns | ns | * | *** | *** | 21.11 |
| DE | ns | ns | ns | *** | ns | 22.17 |
| pH | ns | ns | *** | *** | *** | 47.04 |
| SCC (log) | ns | ns | ns | *** | *** | 18.80 |
| Cheese-making traits | ||||||
| A30 | * | ns | ns | *** | na | 26.81 |
| A60 | * | ns | ns | *** | na | 11.96 |
| RCT | ns | ns | ns | *** | na | 10.61 |
| RCT/A60 (log) | ns | ns | * | *** | na | 18.60 |
| K20 (log) | ns | ns | ns | *** | na | 6.13 |
| ILCY | ns | ns | ns | *** | na | 8.17 |
| ILDCY | * | ns | * | *** | na | 11.52 |
| Traits 1 | Days in Milk | ||
|---|---|---|---|
| Effect | SD | p-value | |
| Milk production traits | |||
| MY | 0.0027 | ±0.0015 | 0.0401 |
| PP | −0.0005 | ±0.0006 | 0.3998 |
| FP | 0.0024 | ±0.0014 | 0.0901 |
| LP | −0.0007 | ±0.0003 | 0.0425 |
| TS | 0.0012 | ±0.0016 | 0.4655 |
| pH | −0.0007 | ±0.0001 | <0.0001 |
| SCC (log) | −0.0011 | ±0.0008 | 0.1755 |
| Cheese-making traits | |||
| A30 | 0.0462 | ±0.0235 | 0.0495 |
| A60 | 0.0288 | ±0.0247 | 0.2447 |
| RCT | −0.0331 | ±0.0200 | 0.0986 |
| RCT/A60 (log) | −0.0008 | ±0.0003 | 0.0289 |
| K20 (log) | −0.0006 | ±0.0004 | 0.1724 |
| ILCY | 0.0018 | ±0.0013 | 0.1846 |
| ILDCY | 0.0013 | ±0.0005 | 0.0165 |
| Traits 1 | Mean | SD 2 | σ2 | SE3 | h2 | SE 3 |
|---|---|---|---|---|---|---|
| Milk production traits | ||||||
| MY | 2.8885 | 1.0730 | 0.1999 | 0.0617 | 0.2501 | 0.0739 |
| PP | 5.0535 | 0.4639 | 0.0546 | 0.0128 | 0.3433 | 0.0746 |
| FP | 5.5558 | 1.0539 | 0.0366 | 0.0528 | 0.0402 | 0.0582 |
| LP | 5.1072 | 0.2396 | 0.0091 | 0.0035 | 0.1802 | 0.0668 |
| DE | 16.6060 | 1.2547 | 0.2101 | 0.0898 | 0.1656 | 0.0693 |
| pH | 6.7276 | 0.1336 | 0.0049 | 0.0011 | 0.3706 | 0.0751 |
| SCC (log) | 2.1896 | 0.6082 | 0.0170 | 0.0217 | 0.0479 | 0.0614 |
| Cheese-making traits | ||||||
| A30 | 29.7650 | 13.9750 | 48.5430 | 21.6810 | 0.2845 | 0.1230 |
| A60 | 40.9590 | 11.2960 | 19.0450 | 8.1594 | 0.1658 | 0.0697 |
| RCT | 29.1540 | 10.5250 | 19.1840 | 6.5753 | 0.2255 | 0.0747 |
| RCT/A60 (log) | -0.1535 | 0.2839 | 0.0130 | 0.0049 | 0.1956 | 0.0710 |
| K20 (log) | 0.5714 | 0.2227 | 0.0139 | 0.0040 | 0.3347 | 0.0905 |
| ILCY | 2.4938 | 0.4136 | 0.0519 | 0.0142 | 0.3366 | 0.0864 |
| ILDCY | 0.9724 | 0.1758 | 0.0065 | 0.0023 | 0.2593 | 0.0866 |
| Traits | pH | RCT | logk20 | A30 | A60 | ILCY | ILCDY | FP | PP | LP | DE | MY | logSCC | logRCT/A60 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| pH | 0.37 (0.07) | 0.47 (0.02) | 0.37 (0.03) | −0.24 (0.04) | −0.44 (0.02) | 0.03 (0.03) | −0.17 (0.03) | −0.09 (0.03) | −0.09 (0.03) | −0.16 (0.03) | −0.14 (0.03) | −0.11 (0.03) | 0.30 (0.02) | 0.50 (0.02) |
| RCT | 0.63 (0.02) | 0.22 (0.07) | 0.68 (0.02) | −0.85 (0.02) | −0.61 (0.02) | 0.03 (0.03) | −0.12 (0.03) | 0.02 (0.03) | 0.04 (0.03) | −0.04 (0.03) | 0.02 (0.03) | 0.03 (0.03) | 0.15 (0.03) | 0.91 (0.01) |
| logk20 | 0.57 (0.02) | 0.92 (0.01) | 0.31 (0.08) | −0.68 (0.03) | −0.59 (0.02) | −0.05 (0.03) | −0.21 (0.03) | −0.02 (0.03) | −0.12 (0.03) | −0.01 (0.03) | −0.07 (0.03) | −0.05 (0.03) | 0.12 (0.03) | 0.74 (0.02) |
| A30 | −0.44 (0.03) | −0.97 (0.01) | −0.86 (0.02) | 0.28 (0.09) | 0.34 (0.04) | 0.05 (0.04) | 0.14 (0.04) | −0.02 (0.04) | 0.05 (0.04) | −0.02 (0.04) | 0.02 (0.04) | 0.05 (0.04) | −0.08 (0.04) | −0.75 (0.02) |
| A60 | −0.76 (0.02) | −0.79 (0.02) | −0.95 (0.01) | 0.91 (0.01) | 0.16 (0.06) | 0.01 (0.03) | 0.22 (0.03) | 0.07 (0.03) | 0.13 (0.03) | 0.02 (0.03) | 0.19 (0.03) | −0.01 (0.03) | −0.07 (0.03) | −0.85 (0.01) |
| ILCY | −0.89 (0.01) | 0.37 (0.03) | 0.14 (0.03) | 0.11 (0.04) | 0.11 (0.03) | 0.33 (0.08) | 0.83 (0.01) | 0.17 (0.03) | 0.27 (0.03) | −0.06 (0.03) | 0.23 (0.03) | −0.13 (0.03) | 0.08 (0.03) | 0.01 (0.03) |
| ILCDY | −0.36 (0.03) | 0.23 (0.03) | −0.51 (0.02) | 0.71 (0.03) | 0.14 (0.03) | 0.87 (0.01) | 0.26 (0.08) | 0.29 (0.03) | 0.30 (0.03) | −0.14 (0.03) | 0.27 (0.03) | −0.08 (0.03) | 0.03 (0.03) | −0.19 (0.03) |
| FP | −0.50 (0.02) | 0.36 (0.03) | 0.66 (0.02) | −0.52 (0.03) | −0.99 (0.00) | 0.99 (0.00) | 0.99 (0.00) | 0.04 (0.05) | 0.27 (0.02) | −0.22 (0.03) | 0.89 (0.01) | −0.03 (0.03) | 0.04 (0.03) | −0.01 (0.03) |
| PP | −0.11 (0.03) | 0.30 (0.03) | −0.11 (0.03) | 0.70 (0.03) | 0.16 (0.03) | 0.60 (0.02) | 0.74 (0.02) | 0.65 (0.02) | 0.34 (0.06) | −0.22 (0.03) | 0.60 (0.02) | −0.30 (0.02) | 0.18 (0.03) | −0.03 (0.03) |
| LP | −0.17 (0.03) | −0.23 (0.03) | −0.47 (0.03) | 0.19 (0.04) | 0.18 (0.03) | −0.40 (0.03) | −0.65 (0.02) | −0.41 (0.02) | −0.24 (0.03) | 0.18 (0.06) | −0.08 (0.03) | 0.21 (0.03) | −0.45 (0.02) | −0.04 (0.03) |
| DE | −0.16 (0.03) | 0.22 (0.03) | 0.16 (0.03) | −0.20 (0.04) | 0.13 (0.03) | 0.79 (0.02) | 0.99 (0.00) | 0.89 (0.01) | 0.84 (0.01) | −0.11 (0.03) | 0.16 (0.06) | −0.12 (0.03) | 0.01 (0.03) | −0.03 (0.03) |
| MY | −0.27 (0.02) | −0.28 (0.03) | −0.22 (0.03) | −0.32 (0.04) | 0.26 (0.03) | −0.34 (0.03) | −0.30 (0.03) | −0.42 (0.02) | −0.52 (0.02) | 0.92 (0.01) | −0.50 (0.02) | 0.24 (0.07) | −0.31 (0.02) | 0.02 (0.03) |
| logSCC | 0.58 (0.02) | 0.12 (0.03) | 0.26 (0.03) | 0.32 (0.04) | 0.83 (0.01) | 0.48 (0.02) | 0.36 (0.03) | −0.75 (0.02) | 0.60 (0.02) | −0.73 (0.02) | −0.25 (0.03) | −0.97 (0.00) | 0.05 (0.05) | 0.13 (0.03) |
| logRCT/A60 | 0.70 (0.02) | 0.97 (0.00) | 0.96 (0.00) | −0.99 (0.00) | −0.92 (0.01) | 0.19 (0.03) | 0.70 (0.02) | 0.13 (0.03) | 0.15 (0.03) | −0.20 (0.03) | 0.34 (0.03) | −0.25 (0.03) | 0.32 (0.03) | 0.20 (0.06) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marina, H.; Reverter, A.; Gutiérrez-Gil, B.; Alexandre, P.A.; Porto-Neto, L.R.; Suárez-Vega, A.; Li, Y.; Esteban-Blanco, C.; Arranz, J.-J. Gene Networks Driving Genetic Variation in Milk and Cheese-Making Traits of Spanish Assaf Sheep. Genes 2020, 11, 715. https://doi.org/10.3390/genes11070715
Marina H, Reverter A, Gutiérrez-Gil B, Alexandre PA, Porto-Neto LR, Suárez-Vega A, Li Y, Esteban-Blanco C, Arranz J-J. Gene Networks Driving Genetic Variation in Milk and Cheese-Making Traits of Spanish Assaf Sheep. Genes. 2020; 11(7):715. https://doi.org/10.3390/genes11070715
Chicago/Turabian StyleMarina, Héctor, Antonio Reverter, Beatriz Gutiérrez-Gil, Pâmela Almeida Alexandre, Laercio R. Porto-Neto, Aroa Suárez-Vega, Yutao Li, Cristina Esteban-Blanco, and Juan-José Arranz. 2020. "Gene Networks Driving Genetic Variation in Milk and Cheese-Making Traits of Spanish Assaf Sheep" Genes 11, no. 7: 715. https://doi.org/10.3390/genes11070715
APA StyleMarina, H., Reverter, A., Gutiérrez-Gil, B., Alexandre, P. A., Porto-Neto, L. R., Suárez-Vega, A., Li, Y., Esteban-Blanco, C., & Arranz, J.-J. (2020). Gene Networks Driving Genetic Variation in Milk and Cheese-Making Traits of Spanish Assaf Sheep. Genes, 11(7), 715. https://doi.org/10.3390/genes11070715








