Application of Genetically Engineered Pigs in Biomedical Research
Abstract
1. Introduction
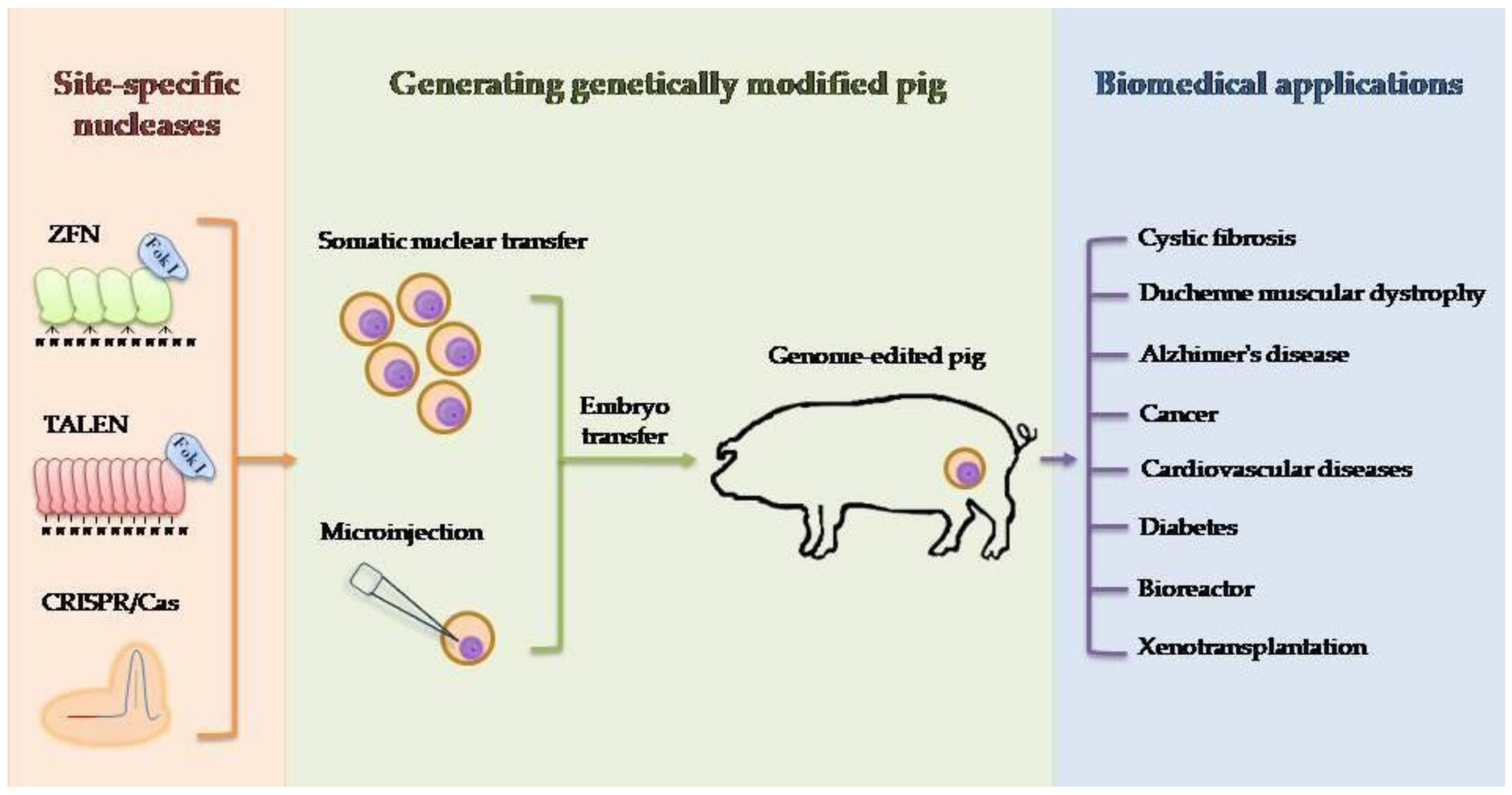
2. Techniques for Producing Genetically Engineered Pigs
3. Pig Models for Human Diseases
3.1. Cystic Fibrosis
3.2. Duchenne Muscular Dystrophy
3.3. Alzheimer’s Disease
3.4. Porcine Cancer Models
3.5. Cardiovascular Diseases
3.6. Diabetes Mellitus
4. Pigs as Bioreactors for Pharmaceutical Products
5. Pig-to-Human Xenotransplantation
5.1. Hyperacute Xenograft Rejection
5.2. Coagulation Dysregulation
5.3. Inflammatory Response
5.4. Cellular Xenograft Rejection
5.5. Porcine Endogenous Retroviruses
5.6. Preclinical Studies in Xenotransplantation
6. Conclusions and Future Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Perleberg, C.; Kind, A.; Schnieke, A. Genetically engineered pigs as models for human disease. Dis. Model. Mech. 2018, 11, dmm030783. [Google Scholar] [CrossRef]
- Gifre, L.; Arís, A.; Bach, À.; Garcia-Fruitós, E. Trends in recombinant protein use in animal production. Microb. Cell Fact. 2017, 16, 40. [Google Scholar] [CrossRef]
- Hryhorowicz, M.; Zeyland, J.; Słomski, R.; Lipiński, D. Genetically modified pigs as organ donors for xenotransplantation. Mol. Biotechnol. 2017, 59, 435–444. [Google Scholar] [CrossRef] [PubMed]
- Gordon, J.; Scangost, G.A.; Plotkin, D.J.; Barbosaf, J.A. Genetic transformation of mouse embryos by microinjection of purified DNA. Proc. Natl. Acad. Sci. USA 1980, 77, 7380–7384. [Google Scholar] [CrossRef]
- Palmiter, R.D.; Brinster, R.L.; Hammer, R.E.; Trumbauer, M.E.; Rosenfeld, M.G.; Birnberg, N.C.; Evans, R.M. Dramatic growth of mice that develop from eggs microinjected with metallothionein-growth hormone fusion genes. Nature 1982, 300, 611–615. [Google Scholar] [CrossRef] [PubMed]
- Wall, R.J. Pronuclear microinjection. Cloning Stem Cells 2001, 3, 209–220. [Google Scholar] [CrossRef] [PubMed]
- Wolf, E.; Schernthaner, W.; Zakhartchenko, V.; Prelle, K.; Stojkovic, M.; Brem, G. Transgenic technology in farm animals-progress and perspectives. Exp. Physiol. 2000, 85, 615–625. [Google Scholar] [CrossRef]
- Maga, E.A.; Sargent, R.G.; Zeng, H.; Pati, S.; Zarling, D.A.; Oppenheim, S.M.; Collette, N.M.; Moyer, A.L.; Conrad-Brink, J.S.; Rowe, J.D.; et al. Increased efficiency of transgenic livestock production. Transgenic Res. 2003, 12, 485–496. [Google Scholar] [CrossRef] [PubMed]
- Chrenek, P.; Vasicek, D.; Makarevich, A.V.; Jurcik, R.; Suvegova, K.; Parkanyi, V.; Bauer, M.; Rafay, J.; Batorova, A.; Paleyanda, R.K. Increased transgene integration efficiency upon microinjection of DNA into both pronuclei of rabbit embryos. Transgenic Res. 2005, 14, 417–428. [Google Scholar] [CrossRef]
- Sumiyama, K.; Kawakami, K.; Yagita, K. A simple and highly efficient transgenesis method in mice with the Tol2 transposon system and cytoplasmic microinjection. Genomics 2010, 95, 306–311. [Google Scholar] [CrossRef]
- Wang, Y.; Du, Y.; Shen, B.; Zhou, X.; Li, J.; Liu, Y.; Wang, J.; Zhou, J.; Hu, B.; Kang, N.; et al. Efficient generation of gene-modified pigs via injection of zygote with Cas9/sgRNA. Sci. Rep. 2015, 5, 8256. [Google Scholar] [CrossRef] [PubMed]
- Campbell, K.H.S.; McWhir, J.; Ritchie, W.A.; Wilmut, I. Sheep cloned by nuclear transfer from a cultured cell line. Nature 1996, 380, 64–66. [Google Scholar] [CrossRef] [PubMed]
- Campbell, K.H.S. A background to nuclear transfer and its applications in agriculture and human therapeutic medicine. J. Anat. 2002, 200, 267–275. [Google Scholar] [CrossRef] [PubMed]
- Rogers, C.S.; Stoltz, D.A.; Meyerholz, D.K.; Ostedgaard, L.S.; Rokhlina, T.; Taft, P.J.; Rogan, M.P.; Pezzulo, A.A.; Karp, P.H.; Itani, O.A.; et al. Disruption of the CFTR gene produces a model of cystic fibrosis in newborn pigs. Science 2008, 321, 1837–1841. [Google Scholar] [CrossRef]
- Ostedgaard, L.S.; Meyerholz, D.K.; Chen, J.H.; Pezzulo, A.A.; Karp, P.H.; Rokhlina, T.; Ernst, S.E.; Hanfland, R.A.; Reznikov, L.R.; Ludwig, P.S.; et al. The DeltaF508 mutation causes CFTR misprocessing and cystic fibrosis-like disease in pigs. Sci. Transl. Med. 2011, 3, 74ra24. [Google Scholar] [CrossRef]
- Klymiuk, N.; Mundhenk, L.; Kraehe, K.; Wuensch, A.; Plog, S.; Emrich, D.; Langenmayer, M.C.; Stehr, M.; Holzinger, A.; Kröner, C.; et al. Sequential targeting of CFTR by BAC vectors generates a novel pig model of cystic fibrosis. J. Mol. Med. 2012, 90, 597–608. [Google Scholar] [CrossRef]
- Cooney, A.L.; Abou Alaiwa, M.H.; Shah, V.S.; Bouzek, D.C.; Stroik, M.R.; Powers, L.S.; Gansemer, N.D.; Meyerholz, D.K.; Welsh, M.J.; Stoltz, D.A.; et al. Lentiviral-mediated phenotypic correction of cystic fibrosis pigs. JCI Insight 2016, 1, e88730. [Google Scholar] [CrossRef]
- Steines, B.; Dickey, D.D.; Bergen, J.; Excoffon, K.J.; Weinstein, J.R.; Li, X.; Yan, Z.; Abou Alaiwa, M.H.; Shah, V.S.; Bouzek, D.C.; et al. CFTR gene transfer with AAV improves early cystic fibrosis pig phenotypes. JCI Insight 2016, 1, e88728. [Google Scholar] [CrossRef]
- Ruan, J.; Hirai, H.; Yang, D.; Ma, L.; Hou, X.; Jiang, H.; Wei, H.; Rajagopalan, C.; Mou, H.; Wang, G.; et al. Efficient gene editing at major CFTR mutation loci. Mol. Ther. Nucleic Acids 2019, 16, 73–81. [Google Scholar] [CrossRef]
- Zhou, Z.P.; Yang, L.L.; Cao, H.; Chen, Z.R.; Zhang, Y.; Wen, X.Y.; Hu, J. In vitro validation of a CRISPR-mediated CFTR correction strategy for preclinical translation in pigs. Hum. Gene Ther. 2019, 30, 1101–1116. [Google Scholar] [CrossRef]
- Klymiuk, N.; Blutke, A.; Graf, A.; Krause, S.; Burkhardt, K.; Wuensch, A.; Krebs, S.; Kessler, B.; Zakhartchenko, V.; Kurome, M.; et al. Dystrophin-deficient pigs provide new insights into the hierarchy of physiological derangements of dystrophic muscle. Hum. Mol. Genet. 2013, 22, 4368–4382. [Google Scholar] [CrossRef]
- Fröhlich, T.; Kemter, E.; Flenkenthaler, F.; Klymiuk, N.; Otte, K.A.; Blutke, A.; Krause, S.; Walter, M.C.; Wanke, R.; Wolf, E.; et al. Progressive muscle proteome changes in a clinically relevant pig model of Duchenne muscular dystrophy. Sci. Rep. 2016, 6, 33362. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.H.; Zhao, H.; Qing, Y.B.; Pan, W.R.; Jia, B.Y.; Zhao, H.Y.; Huang, X.X.; Wei, H.J. Porcine zygote injection with Cas9/sgRNA results in DMD-modified pig with muscle dystrophy. Int. J. Mol. Sci. 2016, 17, 1668. [Google Scholar] [CrossRef] [PubMed]
- Moretti, A.; Fonteyne, L.; Giesert, F.; Hoppmann, P.; Meier, A.B.; Bozoglu, T.; Baehr, A.; Schneider, C.M.; Sinnecker, D.; Klett, K.; et al. Somatic gene editing ameliorates skeletal and cardiac muscle failure in pig and human models of Duchenne muscular dystrophy. Nat. Med. 2020, 26, 207–214. [Google Scholar] [CrossRef]
- Walsh, D.M.; Klyubin, I.; Shankar, G.M.; Townsend, M.; Fadeeva, J.V.; Betts, V.; Podlisny, M.B.; Cleary, J.P.; Ashe, K.H.; Rowan, M.J.; et al. The role of cell-derived oligomers of Abeta in Alzheimer’s disease and avenues for therapeutic intervention. Biochem. Soc. Trans. 2005, 33, 1087–1090. [Google Scholar] [CrossRef]
- Kragh, P.M.; Nielsen, A.L.; Li, J.; Du, Y.; Lin, L.; Schmidt, M.; Bøgh, I.B.; Holm, I.E.; Jakobsen, J.E.; Johansen, M.G.; et al. Hemizygous minipigs produced by random gene insertion and handmade cloning express the Alzheimer’s disease-causing dominant mutation APPsw. Transgenic Res. 2009, 18, 545–558. [Google Scholar] [CrossRef] [PubMed]
- Jakobsen, J.E.; Johansen, M.G.; Schmidt, M.; Dagnæs-Hansen, F.; Dam, K.; Gunnarsson, A.; Liu, Y.; Kragh, P.M.; Li, R.; Holm, I.E.; et al. Generation of minipigs with targeted transgene insertion by recombinase-mediated cassette exchange (RMCE) and somatic cell nuclear transfer (SCNT). Transgenic Res. 2013, 22, 709–723. [Google Scholar] [CrossRef] [PubMed]
- Jakobsen, J.E.; Johansen, M.G.; Schmidt, M.; Liu, Y.; Li, R.; Callesen, H.; Melnikova, M.; Habekost, M.; Matrone, C.; Bouter, Y.; et al. Expression of the Alzheimer’s disease mutations AβPP695sw and PSEN1M146I in double-transgenic Göttingen minipigs. J. Alzheimer’s Dis. 2016, 53, 1617–1630. [Google Scholar] [CrossRef]
- Lee, S.E.; Hyun, H.; Park, M.R.; Choi, Y.; Son, Y.J.; Park, Y.G.; Jeong, S.G.; Shin, M.Y.; Ha, H.J.; Hong, H.S.; et al. Production of transgenic pig as an Alzheimer’s disease model using a multi-cistronic vector system. PLoS ONE 2017, 12, e0177933. [Google Scholar] [CrossRef]
- Adam, S.J.; Rund, L.A.; Kuzmuk, K.N.; Zachary, J.F.; Schook, L.B.; Counter, C.M. Genetic induction of tumorigenesis in swine. Oncogene 2007, 26, 1038–1045. [Google Scholar] [CrossRef]
- Schook, L.B.; Collares, T.V.; Hu, W.; Liang, Y.; Rodrigues, F.M.; Rund, L.A.; Schachtschneider, K.M.; Seixas, F.K.; Singh, K.; Wells, K.D.; et al. A genetic porcine model of cancer. PLoS ONE 2015, 10, e0128864. [Google Scholar] [CrossRef] [PubMed]
- Saalfrank, A.; Janssen, K.P.; Ravon, M.; Flisikowski, K.; Eser, S.; Steiger, K.; Flisikowska, T.; Müller-Fliedner, P.; Schulze, É.; Brönner, C.; et al. A porcine model of osteosarcoma. Oncogenesis 2016, 5, e210. [Google Scholar] [CrossRef]
- Leuchs, S.; Saalfrank, A.; Merkl, C.; Flisikowska, T.; Edlinger, M.; Durkovic, M.; Rezaei, N.; Kurome, M.; Zakhartchenko, V.; Kessler, B.; et al. Inactivation and inducible oncogenic mutation of p53 in gene targeted pigs. PLoS ONE 2012, 7, e43323. [Google Scholar] [CrossRef]
- Sieren, J.C.; Meyerholz, D.K.; Wang, X.J.; Davis, B.T.; Newell, J.D., Jr.; Hammond, E.; Rohret, J.A.; Rohret, F.A.; Struzynski, J.T.; Goeken, J.A.; et al. Development and translational imaging of a TP53 porcine tumorigenesis model. J. Clin. Invest. 2014, 124, 4052–4066. [Google Scholar] [CrossRef]
- Flisikowska, T.; Merkl, C.; Landmann, M.; Eser, S.; Rezaei, N.; Cui, X.; Kurome, M.; Zakhartchenko, V.; Kessler, B.; Wieland, H.; et al. A porcine model of familial adenomatous polyposis. Gastroenterology 2012, 143, 1173–1175. [Google Scholar] [CrossRef]
- Davis, B.T.; Wang, X.J.; Rohret, J.A.; Struzynski, J.T.; Merricks, E.P.; Bellinger, D.A.; Rohret, F.A.; Nichols, T.C.; Rogers, C.S. Targeted disruption of LDLR causes hypercholesterolemia and atherosclerosis in Yucatan miniature pigs. PLoS ONE 2014, 9, e93457. [Google Scholar] [CrossRef]
- Burke, A.C.; Telford, D.E.; Sutherland, B.G.; Edwards, J.Y.; Sawyez, C.G.; Barrett, P.H.R.; Newton, R.S.; Pickering, J.G.; Huff, M.W. Bempedoic acid lowers low-density lipoprotein cholesterol and attenuates atherosclerosis in low-density lipoprotein receptor-deficient (LDLR+/− and LDLR−/−) Yucatan miniature pigs. Arterioscler. Thromb. Vasc. Biol. 2018, 38, 1178–1190. [Google Scholar] [CrossRef] [PubMed]
- Al-Mashhadi, R.H.; Sørensen, C.B.; Kragh, P.M.; Christoffersen, C.; Mortensen, M.B.; Tolbod, L.P.; Thim, T.; Du, Y.; Li, J.; Liu, Y.; et al. Familial hypercholesterolemia and atherosclerosis in cloned minipigs created by DNA transposition of a human PCSK9 gain-of-function mutant. Sci. Transl. Med. 2013, 5, 166ra1. [Google Scholar] [CrossRef]
- Wei, J.; Ouyang, H.; Wang, Y.; Pang, D.; Cong, N.X.; Wang, T.; Leng, B.; Li, D.; Li, X.; Wu, R.; et al. Characterization of a hypertriglyceridemic transgenic miniature pig model expressing human apolipoprotein CIII. FEBS J. 2012, 279, 91–99. [Google Scholar] [CrossRef]
- Renner, S.; Fehlings, C.; Herbach, N.; Hofmann, A.; von Waldthausen, D.C.; Kessler, B.; Ulrichs, K.; Chodnevskaja, I.; Moskalenko, V.; Amselgruber, W.; et al. Glucose intolerance and reduced proliferation of pancreatic beta-cells in transgenic pigs with impaired glucose-dependent insulinotropic polypeptide function. Diabetes 2010, 59, 1228–1238. [Google Scholar] [CrossRef] [PubMed]
- Renner, S.; Römisch-Margl, W.; Prehn, C.; Krebs, S.; Adamski, J.; Göke, B.; Blum, H.; Suhre, K.; Roscher, A.A.; Wolf, E. Changing metabolic signatures of amino acids and lipids during the prediabetic period in a pig model with impaired incretin function and reduced β-cell mass. Diabetes 2012, 61, 2166–2175. [Google Scholar] [CrossRef] [PubMed]
- Streckel, E.; Braun-Reichhart, C.; Herbach, N.; Dahlhoff, M.; Kessler, B.; Blutke, A.; Bähr, A.; Übel, N.; Eddicks, M.; Ritzmann, M.; et al. Effects of the glucagon-like peptide-1 receptor agonist liraglutide in juvenile transgenic pigs modeling a pre-diabetic condition. J. Transl. Med. 2015, 13, 73. [Google Scholar] [CrossRef] [PubMed]
- Umeyama, K.; Watanabe, M.; Saito, H.; Kurome, M.; Tohi, S.; Matsunari, H.; Miki, K.; Nagashima, H. Dominant-negative mutant hepatocyte nuclear factor 1α induces diabetes in transgenic-cloned pigs. Transgenic Res. 2009, 18, 697–706. [Google Scholar] [CrossRef] [PubMed]
- Umeyama, K.; Nakajima, M.; Yokoo, T.; Nagaya, M.; Nagashima, H. Diabetic phenotype of transgenic pigs introduced by dominant-negative mutant hepatocyte nuclear factor 1α. J. Diabetes Complicat. 2017, 31, 796–803. [Google Scholar] [CrossRef]
- Renner, S.; Braun-Reichhart, C.; Blutke, A.; Herbach, N.; Emrich, D.; Streckel, E.; Wünsch, A.; Kessler, B.; Kurome, M.; Bähr, A.; et al. Permanent neonatal diabetes in INS(C94Y) transgenic pigs. Diabetes 2013, 62, 1505–1511. [Google Scholar] [CrossRef]
- Kleinwort, K.J.H.; Amann, B.; Hauck, S.M.; Hirmer, S.; Blutke, A.; Renner, S.; Uhl, P.B.; Lutterberg, K.; Sekundo, W.; Wolf, E.; et al. Retinopathy with central oedema in an INS C94Y transgenic pig model of long-term diabetes. Diabetologia 2017, 60, 1541–1549. [Google Scholar] [CrossRef]
- Backman, M.; Flenkenthaler, F.; Blutke, A.; Dahlhoff, M.; Ländström, E.; Renner, S.; Philippou-Massier, J.; Krebs, S.; Rathkolb, B.; Prehn, C.; et al. Multi-omics insights into functional alterations of the liver in insulin-deficient diabetes mellitus. Mol. Metab. 2019, 26, 30–44. [Google Scholar] [CrossRef]
- Velander, W.H.; Johnson, J.L.; Page, R.L.; Russell, C.G.; Subramanian, A.; Wilkins, T.D.; Gwazdauskas, F.C.; Pittius, C.; Drohan, W.N. High-level expression of a heterologous protein in the milk of transgenic swine using the cDNA encoding human protein C. Proc. Natl. Acad. Sci. USA 1992, 89, 12003–12007. [Google Scholar] [CrossRef]
- Paleyanda, R.K.; Velander, W.H.; Lee, T.K.; Scandella, D.H.; Gwazdauskas, F.C.; Knight, J.W.; Hoyer, L.W.; Drohan, W.N.; Lubon, H. Transgenic pigs produce functional human factor VIII in milk. Nat. Biotechnol. 1997, 15, 971–975. [Google Scholar] [CrossRef]
- Lee, M.H.; Lin, Y.S.; Tu, C.F.; Yen, C.H. Recombinant human factor IX produced from transgenic porcine milk. Biomed. Res. Int. 2014, 2014, 315375. [Google Scholar] [CrossRef]
- Zhao, J.; Xu, W.; Ross, J.W.; Walters, E.M.; Butler, S.P.; Whyte, J.J.; Kelso, L.; Fatemi, M.; Vanderslice, N.C.; Giroux, K.; et al. Engineering protein processing of the mammary gland to produce abundant hemophilia B therapy in milk. Sci. Rep. 2015, 5, 14176. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.G.; Lee, H.C.; Kim, S.W.; Lee, P.; Chung, H.J.; Lee, Y.K.; Han, J.H.; Hwang, I.S.; Yoo, J.I.; Kim, Y.K.; et al. Production of recombinant human von Willebrand factor in the milk of transgenic pigs. J. Reprod. Dev. 2009, 55, 484–490. [Google Scholar] [CrossRef] [PubMed]
- Park, J.K.; Lee, Y.K.; Lee, P.; Chung, H.J.; Kim, S.; Lee, H.G.; Seo, M.K.; Han, J.H.; Park, C.G.; Kim, H.T.; et al. Recombinant human erythropoietin produced in milk of transgenic pigs. J. Biotechnol. 2006, 122, 362–371. [Google Scholar] [CrossRef]
- Lu, D.; Liu, S.; Shang, S.; Wu, F.; Wen, X.; Li, Z.; Li, Y.; Hu, X.; Zhao, Y.; Li, Q.; et al. Production of transgenic-cloned pigs expressing large quantities of recombinant human lysozyme in milk. PLoS ONE 2015, 10, e0123551. [Google Scholar] [CrossRef]
- Swanson, M.E.; Martin, M.J.; O’Donnell, J.K.; Hoover, K.; Lago, W.; Huntress, V.; Parsons, C.T.; Pinkert, C.A.; Pilder, S.; Logan, J.S. Production of functional human hemoglobin in transgenic swine. Biotechnology (N. Y.) 1992, 10, 557–559. [Google Scholar] [CrossRef]
- Sharma, A.; Martin, M.J.; Okabe, J.F.; Truglio, R.A.; Dhanjal, N.K.; Logan, J.S.; Kumar, R. An isologous porcine promoter permits high level expression of human hemoglobin in transgenic swine. Biotechnology (N. Y.) 1994, 12, 55–59. [Google Scholar] [CrossRef] [PubMed]
- Houdebine, L.M. Production of pharmaceutical proteins by transgenic animals. Rev. Sci. Tech. 2018, 37, 131–139. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhao, S.; Bai, L.; Fan, J.; Liu, E. Expression systems and species used for transgenic animal bioreactors. Biomed. Res. Int. 2013, 2013, 580463. [Google Scholar] [CrossRef] [PubMed]
- Lai, L.; Kolber-Simonds, D.; Park, K.W.; Cheong, H.T.; Greenstein, J.L.; Im, G.S.; Samuel, M.; Bonk, A.; Rieke, A.; Day, B.N.; et al. Production of alpha-1,3-galactosyltransferase knockout pigs by nuclear transfer cloning. Science 2002, 295, 1089–1092. [Google Scholar] [CrossRef]
- Phelps, C.J.; Koike, C.; Vaught, T.D.; Boone, J.; Wells, K.D.; Chen, S.H.; Ball, S.; Specht, S.M.; Polejaeva, I.A.; Monahan, J.A.; et al. Production of alpha 1,3-galactosyltransferase-deficient pigs. Science 2003, 299, 411–414. [Google Scholar] [CrossRef]
- Lipiński, D.; Nowak-Terpiłowska, A.; Hryhorowicz, M.; Jura, J.; Korcz, A.; Słomski, R.; Juzwa, W.; Mazurkiewicz, N.; Smorąg, Z.; Zeyland, J. Production of ZFN-mediated GGTA1 knock-out pigs by microinjection of gene constructs into pronuclei of zygotes. Pol. J. Vet. Sci. 2019, 22, 91–100. [Google Scholar] [PubMed]
- Kang, J.T.; Kwon, D.K.; Park, A.R.; Lee, E.J.; Yun, Y.J.; Ji, D.Y.; Lee, K.; Park, K.W. Production of α1,3-galactosyltransferase targeted pigs using transcription activator-like effector nuclease-mediated genome editing technology. J. Vet. Sci. 2016, 17, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Chuang, C.K.; Chen, C.H.; Huang, C.L.; Su, Y.H.; Peng, S.H.; Lin, T.Y.; Tai, H.C.; Yang, T.S.; Tu, C.F. Generation of GGTA1 mutant pigs by direct pronuclear microinjection of CRISPR/Cas9 plasmid vectors. Animal Biotechnol. 2017, 28, 174–181. [Google Scholar] [CrossRef]
- Estrada, J.L.; Martens, G.; Li, P.; Adams, A.; Newell, K.A.; Ford, M.L.; Butler, J.R.; Sidner, R.; Tector, M.; Tector, J. Evaluation of human and non-human primate antibody binding to pig cells lacking GGTA1/CMAH/β4GalNT2 genes. Xenotransplantation 2015, 22, 194–202. [Google Scholar] [CrossRef]
- Fodor, W.L.; Williams, B.L.; Matis, L.A.; Madri, J.A.; Rollins, S.A.; Knight, J.W.; Velander, W.; Squinto, S.P. Expression of a functional human complement inhibitor in a transgenic pig as a model for the prevention of xenogeneic hyperacute organ rejection. Proc. Natl. Acad. Sci. USA 1994, 91, 11153–11157. [Google Scholar] [CrossRef] [PubMed]
- McGregor, C.G.; Davies, W.R.; Oi, K.; Teotia, S.S.; Schirmer, J.M.; Risdahl, J.M.; Tazelaar, H.D.; Kremers, W.K.; Walker, R.C.; Byrne, G.W.; et al. Cardiac xenotransplantation: Recent preclinical progress with 3-month median survival. J. Thorac. Cardiovasc. Surg. 2005, 130, 844–851. [Google Scholar] [CrossRef]
- Niemann, H.; Verhoeyen, E.; Wonigeit, K.; Lorenz, R.; Hecker, J.; Schwinzer, R.; Hauser, H.; Kues, W.A.; Halter, R.; Lemme, E.; et al. Cytomegalovirus early promoter induced expression of hCD59 in porcine organs provides protection against hyperacute rejection. Transplantation 2001, 72, 1898–1906. [Google Scholar] [CrossRef]
- Cozzi, E.; Bhatti, F.; Schmoeckel, M.; Chavez, G.; Smith, K.G.; Zaidi, A.; Bradley, J.R.; Thiru, S.; Goddard, M.; Vial, C.; et al. Long-term survival of nonhuman primates receiving life-supporting transgenic porcine kidney xenografts. Transplantation 2000, 70, 15–21. [Google Scholar]
- Liu, F.; Liu, J.; Yuan, Z.; Qing, Y.; Li, H.; Xu, K.; Zhu, W.; Zhao, H.; Jia, B.; Pan, W.; et al. Generation of GTKO diannan miniature pig expressing human complementary regulator proteins hCD55 and hCD59 via T2A peptide-based bicistronic vectors and SCNT. Mol. Biotechnol. 2018, 60, 550–562. [Google Scholar] [CrossRef]
- Mohiuddin, M.M.; Corcoran, P.C.; Singh, A.K.; Azimzadeh, A.; Hoyt, R.F., Jr.; Thomas, M.L.; Eckhaus, M.A.; Seavey, C.; Ayares, D.; Pierson, R.N., 3rd; et al. B-cell depletion extends the survival of GTKO.hCD46Tg pig heart xenografts in baboons for up to 8 months. Am. J. Transplant. 2012, 12, 763–771. [Google Scholar] [CrossRef]
- Miwa, Y.; Yamamoto, K.; Onishi, A.; Iwamoto, M.; Yazaki, S.; Haneda, M.; Iwasaki, K.; Liu, D.; Ogawa, H.; Nagasaka, T.; et al. Potential value of human thrombomodulin and DAF expression for coagulation control in pig-to-human xenotransplantation. Xenotransplantation 2010, 17, 26–37. [Google Scholar] [CrossRef]
- Bongoni, A.K.; Kiermeir, D.; Jenni, H.; Bähr, A.; Ayares, D.; Klymiuk, N.; Wolf, E.; Voegelin, E.; Constantinescu, M.A.; Seebach, J.D.; et al. Complement dependent early immunological responses during ex vivo xenoperfusion of hCD46/HLA-E double transgenic pig forelimbs with human blood. Xenotransplantation 2014, 21, 230–243. [Google Scholar] [CrossRef] [PubMed]
- Nowak-Terpiłowska, A.; Lipiński, D.; Hryhorowicz, M.; Juzwa, W.; Jura, J.; Słomski, R.; Mazurkiewicz, N.; Gawrońska, B.; Zeyland, J. Production of ULBP1-KO pigs with human CD55 expression using CRISPR technology. J. Appl. Animal Res. 2020, 48, 93–101. [Google Scholar] [CrossRef]
- Iwase, H.; Ekser, B.; Hara, H.; Phelps, C.; Ayares, D.; Cooper, D.K.; Ezzelarab, M.B. Regulation of human platelet aggregation by genetically modified pig endothelial cells and thrombin inhibition. Xenotransplantation 2014, 21, 72–83. [Google Scholar] [CrossRef] [PubMed]
- Harris, D.G.; Quinn, K.J.; French, B.M.; Schwartz, E.; Kang, E.; Dahi, S.; Phelps, C.J.; Ayares, D.L.; Burdorf, L.; Azimzadeh, A.M.; et al. Meta-analysis of the independent and cumulative effects of multiple genetic modifications on pig lung xenograft performance during ex vivo perfusion with human blood. Xenotransplantation 2015, 22, 102–111. [Google Scholar] [CrossRef]
- Iwase, H.; Hara, H.; Ezzelarab, M.; Li, T.; Zhang, Z.; Gao, B.; Liu, H.; Long, C.; Wang, Y.; Cassano, A.; et al. Immunological and physiological observations in baboons with life-supporting genetically engineered pig kidney grafts. Xenotransplantation 2017, 24. [Google Scholar] [CrossRef]
- Lin, C.C.; Ezzelarab, M.; Hara, H.; Long, C.; Lin, C.W.; Dorling, A.; Cooper, D.K. Atorvastatin or transgenic expression of TFPI inhibits coagulation initiated by anti-nonGal IgG binding to porcine aortic endothelial cells. J. Thromb. Haemost. 2010, 8, 2001–2010. [Google Scholar] [CrossRef]
- Kwon, D.J.; Kim, D.H.; Hwang, I.S.; Kim, D.E.; Kim, H.J.; Kim, J.S.; Lee, K.; Im, G.S.; Lee, J.W.; Hwang, S. Generation of α-1,3-galactosyltransferase knocked-out transgenic cloned pigs with knocked-in five human genes. Transgenic Res. 2017, 26, 153–163. [Google Scholar] [CrossRef]
- Wheeler, D.G.; Joseph, M.E.; Mahamud, S.D.; Aurand, W.L.; Mohler, P.J.; Pompili, V.J.; Dwyer, K.M.; Nottle, M.B.; Harrison, S.J.; d’Apice, A.J.F.; et al. Transgenic swine: Expression of human CD39 protects against myocardial injury. J. Mol. Cell. Cardiol. 2012, 52, 958–961. [Google Scholar] [CrossRef]
- Petersen, B.; Ramackers, W.; Lucas-Hahn, A.; Lemme, E.; Hassel, P.; Queisser, A.L.; Herrmann, D.; Barg-Kues, B.; Carnwath, J.W.; Klose, J.; et al. Transgenic expression of human heme oxygenase-1 in pigs confers resistance against xenograft rejection during ex vivo perfusion of porcine kidneys. Xenotransplantation 2011, 18, 355–368. [Google Scholar] [CrossRef]
- Oropeza, M.; Petersen, B.; Carnwath, J.W.; Lucas-Hahn, A.; Lemme, E.; Hassel, P.; Herrmann, D.; Barg-Kues, B.; Holler, S.; Queisser, A.L.; et al. Transgenic expression of the human A20 gene in cloned pigs provides protection against apoptotic and inflammatory stimuli. Xenotransplantation 2009, 16, 522–534. [Google Scholar] [CrossRef]
- Ahrens, H.E.; Petersen, B.; Ramackers, W.; Petkov, S.; Herrmann, D.; Hauschild-Quintern, J.; Lucas-Hahn, A.; Hassel, P.; Ziegler, M.; Baars, W.; et al. Kidneys from α1,3-galactosyltransferase knockout/human heme oxygenase-1/human A20 transgenic pigs are protected from rejection during ex vivo perfusion with human blood. Transplant. Direct 2015, 1, e23. [Google Scholar] [CrossRef]
- Weiss, E.H.; Lilienfeld, B.G.; Müller, S.; Müller, E.; Herbach, N.; Kessler, B.; Wanke, R.; Schwinzer, R.; Seebach, J.D.; Wolf, E.; et al. HLA-E/human beta2-microglobulin transgenic pigs: Protection against xenogeneic human anti-pig natural killer cell cytotoxicity. Transplantation 2009, 87, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Hryhorowicz, M.; Zeyland, J.; Nowak-Terpiłowska, A.; Jura, J.; Juzwa, W.; Słomski, R.; Bocianowski, J.; Smorąg, Z.; Woźniak, A.; Lipiński, D. Characterization of three generations of transgenic pigs expressing the HLA-E gene. Ann. Animal Sci. 2018, 18, 919–935. [Google Scholar] [CrossRef]
- Maeda, A.; Kawamura, T.; Ueno, T.; Usui, N.; Eguchi, H.; Miyagawa, S. The suppression of inflammatory macrophage-mediated cytotoxicity and proinflammatory cytokine production by transgenic expression of HLA-E. Transpl. Immunol. 2013, 29, 76–81. [Google Scholar] [CrossRef]
- Abicht, J.M.; Sfriso, R.; Reichart, B.; Längin, M.; Gahle, K.; Puga Yung, G.L.; Seebach, J.D.; Rieben, R.; Ayares, D.; Wolf, E.; et al. Multiple genetically modified GTKO/hCD46/HLA-E/hβ2-mg porcine hearts are protected from complement activation and natural killer cell infiltration during ex vivo perfusion with human blood. Xenotransplantation 2018, 25, e12390. [Google Scholar] [CrossRef]
- Zeyland, J.; Hryhorowicz, M.; Nowak-Terpiłowska, A.; Jura, J.; Słomski, R.; Smorąg, Z.; Gajda, B.; Lipiński, D. The production of UL16-binding protein 1 targeted pigs using CRISPR technology. 3 Biotech 2018, 8, 70. [Google Scholar] [CrossRef] [PubMed]
- Ide, K.; Wang, H.; Tahara, H.; Liu, J.; Wang, X.; Asahara, T.; Sykes, M.; Yang, Y.G.; Ohdan, H. Role for CD47-SIRPalpha signaling in xenograft rejection by macrophages. Proc. Natl. Acad. Sci. USA 2007, 104, 5062–5066. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.J.; Koo, T.Y.; Lee, H.S.; Lee, W.B.; Kang, B.; Lee, J.G.; Jang, J.Y.; Fang, T.; Ryu, J.H.; Ahn, C.; et al. Role of human CD200 overexpression in pig-to-human xenogeneic immune response compared with human CD47 overexpression. Transplantation 2018, 102, 406–416. [Google Scholar] [CrossRef] [PubMed]
- Tena, A.; Kurtz, J.; Leonard, D.A.; Dobrinsky, J.R.; Terlouw, S.L.; Mtango, N.; Verstegen, J.; Germana, S.; Mallard, C.; Arn, J.S.; et al. Transgenic expression of human CD47 markedly increases engraftment in a murine model of pig-to-human hematopoietic cell transplantation. Am. J. Transplant. 2014, 14, 2713–2722. [Google Scholar] [CrossRef] [PubMed]
- Tena, A.A.; Sachs, D.H.; Mallard, C.; Yang, Y.G.; Tasaki, M.; Farkash, E.; Rosales, I.A.; Colvin, R.B.; Leonard, D.A.; Hawley, R.J. Prolonged survival of pig skin on baboons after administration of pig cells expressing human CD47. Transplantation 2017, 101, 316–321. [Google Scholar] [CrossRef]
- Martin, C.; Plat, M.; Nerriére-Daguin, V.; Coulon, F.; Uzbekova, S.; Venturi, E.; Condé, F.; Hermel, J.M.; Hantraye, P.; Tesson, L.; et al. Transgenic expression of CTLA4-Ig by fetal pig neurons for xenotransplantation. Transgenic Res. 2005, 14, 373–384. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yang, H.Q.; Jiang, W.; Fan, N.N.; Zhao, B.T.; Ou-Yang, Z.; Liu, Z.M.; Zhao, Y.; Yang, D.S.; Zhou, X.Y.; et al. Transgenic expression of human cytoxic T-lymphocyte associated antigen4-immunoglobulin (hCTLA4Ig) by porcine skin for xenogeneic skin grafting. Transgenic Res. 2015, 24, 199–211. [Google Scholar] [CrossRef] [PubMed]
- Reyes, L.M.; Estrada, J.L.; Wang, Z.Y.; Blosser, R.J.; Smith, R.F.; Sidner, R.A.; Paris, L.L.; Blankenship, R.L.; Ray, C.N.; Miner, A.C.; et al. Creating class I MHC-null pigs using guide RNA and the Cas9 endonuclease. J. Immunol. 2014, 193, 5751–5757. [Google Scholar] [CrossRef]
- Fischer, K.; Rieblinger, B.; Hein, R.; Sfriso, R.; Zuber, J.; Fischer, A.; Klinger, B.; Liang, W.; Flisikowski, K.; Kurome, M.; et al. Viable pigs after simultaneous inactivation of porcine MHC class I and three xenoreactive antigen genes GGTA1, CMAH and B4GALNT2. Xenotransplantation 2020, 27, e12560. [Google Scholar] [CrossRef]
- Niu, D.; Wei, H.J.; Lin, L.; George, H.; Wang, T.; Lee, I.H.; Zhao, H.Y.; Wang, Y.; Kan, Y.; Shrock, E.; et al. Inactivation of porcine endogenous retrovirus in pigs using CRISPR-Cas9. Science 2017, 357, 1303–1307. [Google Scholar] [CrossRef]
- Tseng, Y.L.; Kuwaki, K.; Dor, F.J.; Shimizu, A.; Houser, S.; Hisashi, Y.; Yamada, K.; Robson, S.C.; Awwad, M.; Schuurman, H.J.; et al. α1,3-Galactosyltransferase gene-knockout pig heart transplantation in baboons with survival approaching 6 months. Transplantation 2005, 80, 1493–1500. [Google Scholar] [CrossRef]
- Simon, P.M.; Neethling, F.A.; Taniguchi, S.; Goode, P.L.; Zopf, D.; Hancock, W.W.; Cooper, D.K. Intravenous infusion of Galα-3Gal oligosaccharides in baboons delays hyperacute rejection of porcine heart xenografts. Transplantation 1998, 65, 346–353. [Google Scholar] [CrossRef]
- Mohiuddin, M.M.; Singh, A.K.; Corcoran, P.C.; Thomas Iii, M.L.; Clark, T.; Lewis, B.G.; Hoyt, R.F.; Eckhaus, M.; Pierson, R.N., III; Belli, A.J.; et al. Chimeric 2C10R4 anti-CD40 antibody therapy is critical for long-term survival of GTKO.hCD46.hTBM pig-to-primate cardiac xenograft. Nat. Commun. 2016, 7, 11138. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.K.; Chan, J.L.; DiChiacchio, L.; Hardy, N.L.; Corcoran, P.C.; Lewis, B.G.T.; Thomas, M.L.; Burke, A.P.; Ayares, D.; Horvath, K.A.; et al. Cardiac xenografts show reduced survival in the absence of transgenic human thrombomodulin expression in donor pigs. Xenotransplantation 2019, 26, e12465. [Google Scholar] [CrossRef]
- Längin, M.; Mayr, T.; Reichart, B.; Michel, S.; Buchholz, S.; Guethoff, S.; Dashkevich, A.; Baehr, A.; Egerer, S.; Bauer, A.; et al. Consistent success in life-supporting porcine cardiac xenotransplantation. Nature 2018, 564, 430–433. [Google Scholar] [CrossRef]
- Kim, S.C.; Mathews, D.V.; Breeden, C.P.; Higginbotham, L.B.; Ladowski, J.; Martens, G.; Stephenson, A.; Farris, A.B.; Strobert, E.A.; Jenkins, J.; et al. Long-term survival of pig-to-rhesus macaque renal xenografts is dependent on CD4 T cell depletion. Am. J. Transplant. 2019, 19, 2174–2185. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Cooper, D.K.C.; Burdorf, L.; Wang, Y.; Iwase, H. Overcoming coagulation dysregulation in pig solid organ transplantation in nonhuman primates: Recent progress. Transplantation 2018, 102, 1050–1058. [Google Scholar] [CrossRef] [PubMed]
- Shah, J.A.; Patel, M.S.; Elias, N.; Navarro-Alvarez, N.; Rosales, I.; Wilkinson, R.A.; Louras, N.J.; Hertl, M.; Fishman, J.A.; Colvin, R.B.; et al. Prolonged survival following pig-to-primate liver xenotransplantation utilizing exogenous coagulation factors and costimulation blockade. Am. J. Transplant. 2017, 17, 2178–2185. [Google Scholar] [CrossRef]
- Watanabe, H.; Ariyoshi, Y.; Pomposelli, T.; Takeuchi, K.; Ekanayake-Alper, D.K.; Boyd, L.K.; Arn, S.J.; Sahara, H.; Shimizu, A.; Ayares, D.; et al. Intra-bone bone marrow transplantation from hCD47 transgenic pigs to baboons prolongs chimerism to >60 days and promotes increased porcine lung transplant survival. Xenotransplantation 2020, 27, e12552. [Google Scholar] [CrossRef] [PubMed]
- Kwon, D.N.; Lee, K.; Kang, M.J.; Choi, Y.J.; Park, C.; Whyte, J.J.; Brown, A.N.; Kim, J.H.; Samuel, M.; Mao, J.; et al. Production of biallelic CMP-Neu5Ac hydroxylase knock-out pigs. Sci. Rep. 2013, 3, 1981. [Google Scholar] [CrossRef]
- Cozzi, E.; White, D.J. The generation of transgenic pigs as potential organ donors for humans. Nat. Med. 1995, 1, 964–966. [Google Scholar] [CrossRef] [PubMed]
- Diamond, L.E.; Quinn, C.M.; Martin, M.J.; Lawson, J.; Platt, J.L.; Logan, J.S. A human CD46 transgenic pig model system for the study of discordant xenotransplantation. Transplantation 2001, 71, 132–142. [Google Scholar] [CrossRef]
- Petersen, B.; Ramackers, W.; Tiede, A.; Lucas-Hahn, A.; Herrmann, D.; Barg-Kues, B.; Schuettler, W.; Friedrich, L.; Schwinzer, R.; Winkler, M.; et al. Pigs transgenic for human thrombomodulin have elevated production of activated protein C. Xenotransplantation 2009, 16, 486–495. [Google Scholar] [CrossRef]
- Aigner, B.; Renner, S.; Kessler, B.; Klymiuk, N.; Kurome, M.; Wünsch, A.; Wolf, E. Transgenic pigs as models for translational biomedical research. J. Mol. Med. (Berl.) 2010, 88, 653–664. [Google Scholar] [CrossRef]
- Ran, F.A.; Hsu, P.D.; Lin, C.Y.; Gootenberg, J.S.; Konermann, S.; Trevino, A.E.; Scott, D.A.; Inoue, A.; Matoba, S.; Zhang, Y.; et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 2013, 154, 1380–1389. [Google Scholar] [CrossRef] [PubMed]
- Kleinstiver, B.P.; Pattanayak, V.; Prew, M.S.; Tsai, S.Q.; Nguyen, N.T.; Zheng, Z.; Joung, J.K. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 2016, 529, 490–495. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.; Staahl, B.T.; Alla, R.K.; Doudna, J.A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. eLife 2014, 3, e04766. [Google Scholar] [CrossRef] [PubMed]
- Nishiyama, J.; Mikuni, T.; Yasuda, R. Virus-mediated genome editing via homology-directed repair in mitotic and postmitotic cells in mammalian brain. Neuron 2017, 96, 755–768. [Google Scholar] [CrossRef] [PubMed]
- Hryhorowicz, M.; Grześkowiak, B.; Mazurkiewicz, N.; Śledziński, P.; Lipiński, D.; Słomski, R. Improved delivery of CRISPR/Cas9 system using magnetic nanoparticles into porcine fibroblast. Mol. Biotechnol. 2019, 61, 173–180. [Google Scholar] [CrossRef]
- Gün, G.; Kues, W.A. Current progress of genetically engineered pig models for biomedical research. Biores. Open Access 2014, 3, 255–264. [Google Scholar] [CrossRef]
- Yang, H.; Wu, Z. Genome editing of pigs for agriculture and biomedicine. Front. Genet. 2018, 9, 360. [Google Scholar] [CrossRef]
- Kalla, D.; Kind, A.; Schnieke, A. Genetically engineered pigs to study cancer. Int. J. Mol. Sci. 2020, 21, 488. [Google Scholar] [CrossRef]
- Iwatsuki-Horimoto, K.; Nakajima, N.; Shibata, M.; Takahashi, K.; Sato, Y.; Kiso, M.; Yamayoshi, S.; Ito, M.; Enya, S.; Otake, M.; et al. The microminipig as an animal model for influenza a virus infection. J. Virol. 2017, 91, e01716-16. [Google Scholar] [CrossRef]
- Whitney, J.B.; Brad Jones, R. In vitro and in vivo models of HIV latency. Adv. Exp. Med. Biol. 2018, 1075, 241–263. [Google Scholar]
- Siragam, V.; Wong, G.; Qiu, X.G. Animal models for filovirus infections. Zool. Res. 2018, 39, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Xu, W. Production of Recombinant Human Coagulation Factor IX by Transgenic Pig. Ph.D. Thesis, University of Nebraska, Lincoln, NE, USA, 2014. Available online: https://digitalcommons.unl.edu/dissertations/AAI3632487 (accessed on 10 August 2014).
- Cooper, D.K.; Ekser, B.; Ramsoondar, J.; Phelps, C.; Ayares, D. The role of genetically engineered pigs in xenotransplantation research. J. Pathol. 2016, 238, 288–299. [Google Scholar] [CrossRef] [PubMed]
- Wolf, E.; Kemter, E.; Klymiuk, N.; Reichart, B. Genetically modified pigs as donors of cells, tissues, and organs for xenotransplantation. Animal Front. 2019, 9, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Lu, T.; Yang, B.; Wang, R.; Qin, C. Xenotransplantation: Current status in preclinical research. Front. Immunol. 2020, 10, 3060. [Google Scholar] [CrossRef]
- Wu, J.; Platero-Luengo, A.; Sakurai, M.; Sugawara, A.; Gil, M.A.; Yamauchi, T.; Suzuki, K.; Bogliotti, Y.S.; Cuello, C.; Morales Valencia, M.; et al. Interspecies chimerism with mammalian pluripotent stem cells. Cell 2017, 168, 473–486. [Google Scholar] [CrossRef]
| Microinjection | SCNT | |
|---|---|---|
| Advantages | increased efficiency of the transgene integration | precise transformation and selection of modified cells used in cloning |
| the number of damaged zygotes do not exceed 10% | the obtained animals do not exhibit mosaicism | |
| modification is also revealed in germ cells—transgenic offspring | ||
| Disadvantages | low process efficiency (2%–3% in pigs) | very low efficiency |
| the possibility of random integration of the transgene | early fetal mortality | |
| high process invasiveness | the possibility of genetic defects |
| Human Disease | Genetic Modification | Reference |
|---|---|---|
| Cystic fibrosis | targeted disruption of CFTR gene | [14,15,16] |
| Duchenne muscular dystrophy | targeted deletion of DMD exon 52 targeted knock-out of DMD gene | [21] [23] |
| Alzheimer’s disease | expression of human APPsw and PSEN1 M146I genes expression of human APP (K670N/M671L, I716V, V717I), Tau (P301L), and PSEN1 (M146V, L286P) genes | [28] [29] |
| Osteosarcoma | targeted knock-out of TP53 gene targeted homozygous TP53 R167H mutation | [32] [34] |
| Colorectal cancer | targeted heterozygous APC 1311 mutation | [35] |
| Cardiovascular Diseases | targeted disruption of LDLR gene expression of human PCSK9 D374Y gene expression of human ApoCIII gene | [36] [38] [39] |
| Diabetes mellitus | expression of human GIPRdn gene expression of human HNF1α P291fsinsC gene expression of porcine INS C94Y gene | [40] [43] [45] |
| Protein | Production System | Yield | Reference |
|---|---|---|---|
| Human protein C | milk | up to 1 g/L | [48] |
| Human factor VIII | milk | up to 2.7 μg/mL | [49] |
| Human factor IX | milk | up to 0.25 mg/mL | [50] |
| Human von Willebrand factor | milk | mean 280 μg/mL | [52] |
| Human erythropoietin | milk | mean 877.9 IU/1 mL | [53] |
| Human lysozyme | milk | up to 2759.6 mg/L | [54] |
| Human hemoglobin | blood | up to 32 g/L | [56] |
| Genetic Modification | Function | Reference |
|---|---|---|
| GGTA1 knock-out | deletion of Gal xenoantigen | [59,60] |
| CMAH knock-out | deletion of Neu5Gc xenoantigen | [106] |
| β4GALNT2 knock-out | deletion of SDa xenoantigen | [64] |
| expression of human CD55 gene | complement regulation | [107] |
| expression of human CD46 gene | complement regulation | [108] |
| expression of human CD59 gene | complement regulation | [65] |
| expression of human TBM gene | coagulation regulation | [109] |
| expression of human EPCR gene | coagulation regulation | [74] |
| expression of human TFPI gene | coagulation regulation | [77] |
| expression of human CD39 gene | coagulation regulation | [79] |
| expression of human HO-1 gene | anti-inflammatory/antiapoptotic | [80] |
| expression of human A20 gene | anti-inflammatory/antiapoptotic | [81] |
| expression of HLA-E | regulation of NK-cells-mediated responses | [83,84] |
| ULBP1 knock-out | regulation of NK-cells-mediated responses | [87] |
| expression of human CD47 gene | regulation of macrophage-mediated responses | [90] |
| expression of human CTLA4-Ig | regulation of T cells-mediated responses | [92] |
| SLA class I knock-out | regulation of T cells-mediated responses | [94] |
| PERV inactivation | xenozoonosis | [96] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hryhorowicz, M.; Lipiński, D.; Hryhorowicz, S.; Nowak-Terpiłowska, A.; Ryczek, N.; Zeyland, J. Application of Genetically Engineered Pigs in Biomedical Research. Genes 2020, 11, 670. https://doi.org/10.3390/genes11060670
Hryhorowicz M, Lipiński D, Hryhorowicz S, Nowak-Terpiłowska A, Ryczek N, Zeyland J. Application of Genetically Engineered Pigs in Biomedical Research. Genes. 2020; 11(6):670. https://doi.org/10.3390/genes11060670
Chicago/Turabian StyleHryhorowicz, Magdalena, Daniel Lipiński, Szymon Hryhorowicz, Agnieszka Nowak-Terpiłowska, Natalia Ryczek, and Joanna Zeyland. 2020. "Application of Genetically Engineered Pigs in Biomedical Research" Genes 11, no. 6: 670. https://doi.org/10.3390/genes11060670
APA StyleHryhorowicz, M., Lipiński, D., Hryhorowicz, S., Nowak-Terpiłowska, A., Ryczek, N., & Zeyland, J. (2020). Application of Genetically Engineered Pigs in Biomedical Research. Genes, 11(6), 670. https://doi.org/10.3390/genes11060670






