Trans-Atlantic Distribution and Introgression as Inferred from Single Nucleotide Polymorphism: Mussels Mytilus and Environmental Factors
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and DNA Preparation
2.2. SNP Genotyping
2.3. Data Analysis
2.3.1. Genetic Diversity
2.3.2. Population Genetic Differentiation and Structure
2.3.3. Environmental Variables
2.3.4. Relationships between Environment and Allele Frequencies
3. Results
3.1. SNP Validation
3.2. Genetic Diversity and Hardy–Weinberg Equilibrium
3.3. Genetic Variation and Differentiation among Populations
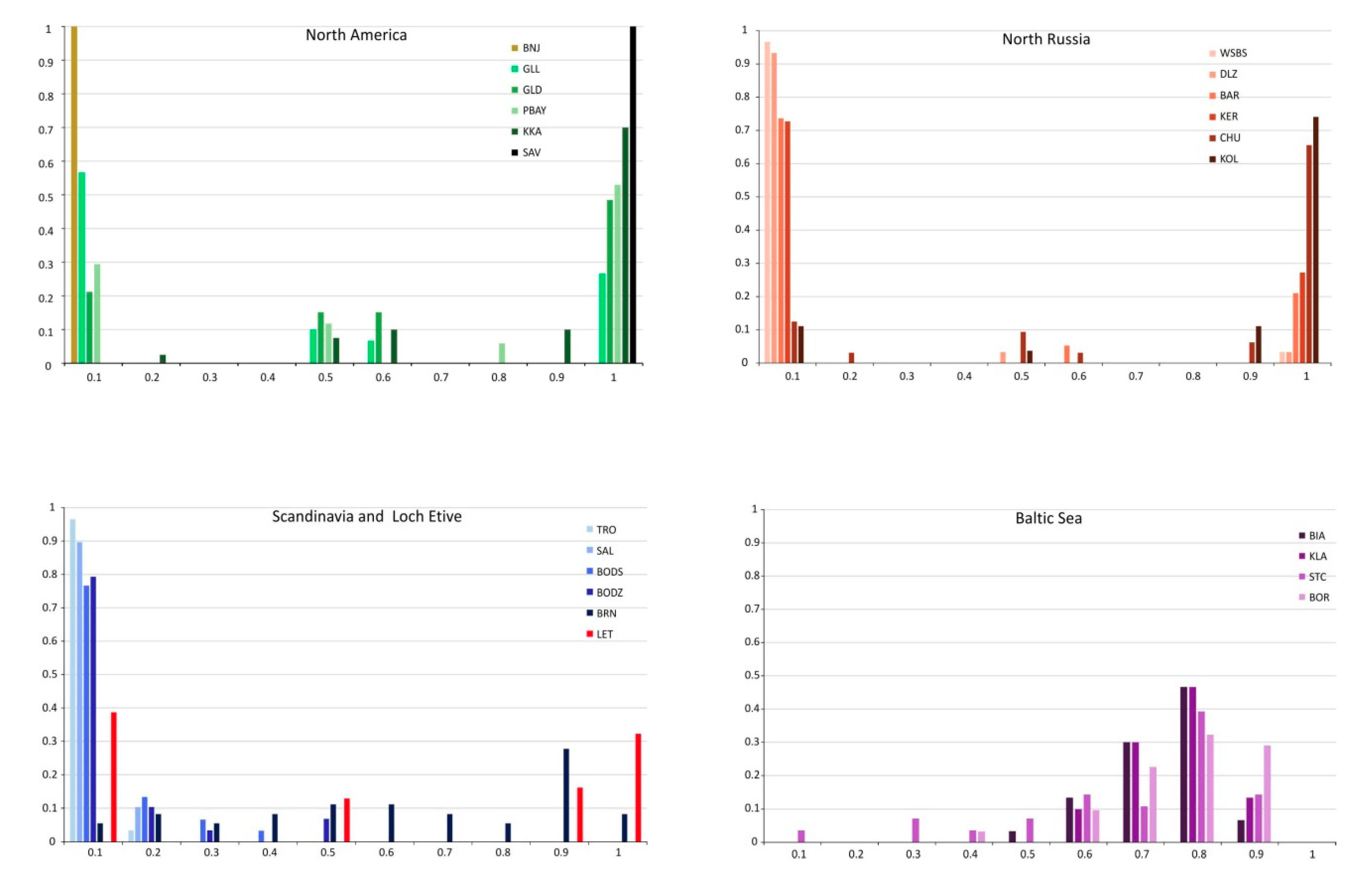
3.4. M. edulis Populations Structure
3.5. M. galloprovincialis Identity and Introgression
3.6. M. trossulus × M. edulis Hybrid Identification
3.7. Relationships between Environment and Allele Frequencies
4. Discussion
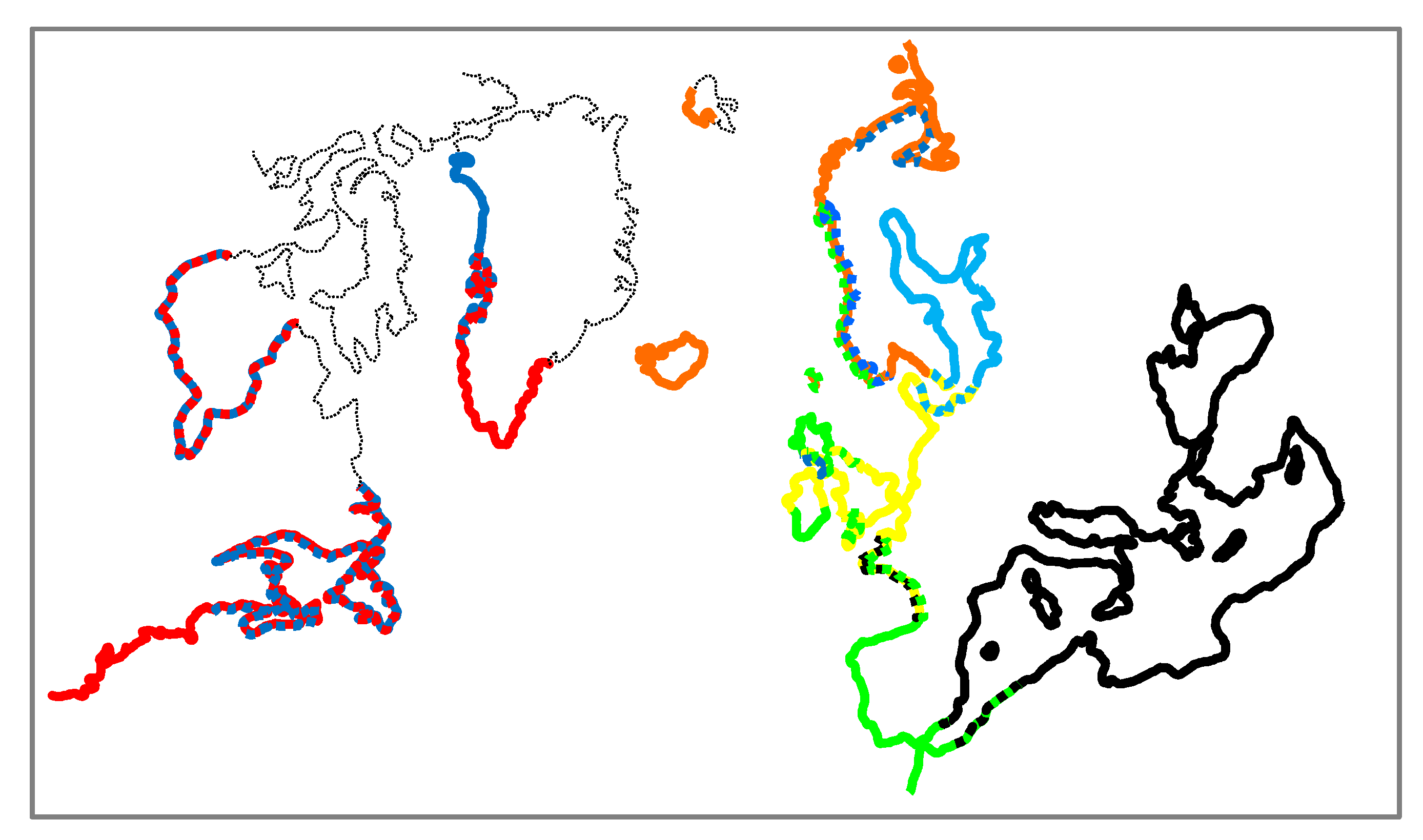
4.1. Distribution of Mytilus Taxa on the Coasts of North Atlantic
4.2. Hybridization and Population Structure
4.3. Relationships between Mytilus Genotypes and Environmental Factors
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Ricklefs, R.E.; Schluter, D. (Eds.) Species diversity: Regional and historical influences. In Species Diversity in Ecological Communities: Historical and Geographical Perspectives; University of Chicago Press: Chicago, IL, USA, 1993; pp. 350–364. [Google Scholar]
- Holyoak, M.; Leibold, M.A.; Moquet, N.; Holt, R.D.; Hoopes, M.F. (Eds.) Metacommunities: A framework for large-scale community ecology. In Metacommunities: Spatial Dynamics and Ecological Communities; University of Chicago Press: Chicago, IL, USA, 2005; pp. 1–34. [Google Scholar]
- Menge, B.A.; Sutherland, J.P. Community Regulation: Variation in Disturbance, Competition, and Predation in Relation to Environmental Stress and Recruitment. Am. Nat. 1987, 130, 730–757. [Google Scholar] [CrossRef]
- Teske, P.R.; Sandoval-Castillo, J.; Golla, T.R.; Emami-Khoyi, A.; Tine, M.; Von Der Heyden, S.; Beheregaray, L.B. Thermal selection as a driver of marine ecological speciation. Proc. R. Soc. B Biol. Sci. 2019, 286, 20182023. [Google Scholar] [CrossRef]
- Vermeij, G.J. Biogeography and Adaptation: Patterns of Marine Life; Harvard University Press: Cambridge, MA, USA, 1978. [Google Scholar]
- Gérard, K.; Bierne, N.; Borsa, P.; Chenuil, A.; Feral, J.-P. Pleistocene separation of mitochondrial lineages of Mytilus spp. mussels from Northern and Southern Hemispheres and strong genetic differentiation among southern populations. Mol. Phylogenetics Evol. 2008, 49, 84–91. [Google Scholar] [CrossRef] [PubMed]
- Álvarez-Varas, R.; Véliz, D.; Vélez-Rubio, G.M.; Fallabrino, A.; Zárate, P.; Heidemeyer, M.; Godoy, D.A.; Benítez, H.A. Identifying genetic lineages through shape: An example in a cosmopolitan marine turtle species using geometric morphometrics. PLoS ONE 2019, 14, e0223587. [Google Scholar] [CrossRef] [PubMed]
- Barahona, S.P.; Vélez-Zuazo, X.; Santa-Maria, M.; Pacheco, A.S. Phylogeography of the rocky intertidal periwinkle Echinolittorina paytensis through a biogeographic transition zone in the Southeastern Pacific. Mar. Ecol. 2019, 40, e12556. [Google Scholar] [CrossRef]
- Moura, C.J.; Collins, A.G.; Santos, R.S.; Lessios, H. Predominant east to west colonizations across major oceanic barriers: Insights into the phylogeographic history of the hydroid superfamily Plumularioidea, suggested by a mitochondrial DNA barcoding marker. Ecol. Evol. 2019, 9, 13001–13016. [Google Scholar] [CrossRef]
- Kawecki, T.J.; Ebert, D. Conceptual issues in local adaptation. Ecol. Lett. 2004, 7, 1225–1241. [Google Scholar] [CrossRef]
- Pelini, S.L.; Keppel, J.A.; Kelley, A.E.; Hellmann, J.J. Adaptation to host plants may prevent rapid insect responses to climate change. Glob. Chang. Biol. 2010, 16, 2923–2929. [Google Scholar] [CrossRef]
- Kijewski, T.; Zbawicka, M.; Strand, J.; Kautsky, H.; Kotta, J.; Rätsep, M.; Wenne, R. Random forest assessment of correlation between environmental factors and genetic differentiation of populations: Case of marine mussels Mytilus. Oceanologia 2019, 61, 131–142. [Google Scholar] [CrossRef]
- Young, E.; Tysklind, N.; Meredith, M.P.; De Bruyn, M.; Belchier, M.; Murphy, E.J.; Carvalho, G.R. Stepping stones to isolation: Impacts of a changing climate on the connectivity of fragmented fish populations. Evol. Appl. 2018, 11, 978–994. [Google Scholar] [CrossRef]
- De Wit, P.; Jonsson, P.R.; Pereyra, R.T.; Panova, M.; André, C.; Johannesson, K. Spatial genetic structure in a crustacean herbivore highlights the need for local considerations in Baltic Sea biodiversity management. Evol. Appl. 2020. Early View. [Google Scholar] [CrossRef]
- Gosling, E. (Ed.) Genetics of Mytilus. In The Mussels Mytilus: Ecology, Physiology, Genetics and Culture; Elsevier: Amsterdam, The Netherlands, 1992; pp. 309–382. [Google Scholar]
- Gosling, E. Bivalve Molluscs: Biology, Ecology and Culture; Fishing News Books; Blackwell Publishing; MPG Books Ltd.: Bodmin, Cornwall, Great Britain, 2003; 443p. [Google Scholar] [CrossRef]
- Kijewski, T.; Śmietanka, B.; Zbawicka, M.; Gosling, E.; Hummel, H.; Wenne, R. Distribution of Mytilus taxa in European coastal areas as inferred from molecular markers. J. Sea Res. 2011, 65, 224–234. [Google Scholar] [CrossRef]
- Mathiesen, S.S.; Thyrring, J.; Hemmer-Hansen, J.; Berge, J.; Sukhotin, A.; Leopold, P.; Bekaert, M.; Sejr, M.K.; Nielsen, E.E. Genetic diversity and connectivity within Mytilus spp. in the subarctic and Arctic. Evol. Appl. 2016, 10, 39–55. [Google Scholar] [CrossRef] [PubMed]
- Blicher, M.; Sejr, M.K.; Høgslund, S. Population structure of Mytilus edulis in the intertidal zone in a sub-Arctic fjord, SW Greenland. Mar. Ecol. Prog. Ser. 2013, 487, 89–100. [Google Scholar] [CrossRef]
- Väinölä, R.; Strelkov, P. Mytilus trossulus in Northern Europe. Mar. Biol. 2011, 158, 817–833. [Google Scholar] [CrossRef]
- Zbawicka, M.; Drywa, A.; Śmietanka, B.; Wenne, R. Identification and validation of novel SNP markers in European populations of marine Mytilus mussels. Mar. Biol. 2012, 159, 1347–1362. [Google Scholar] [CrossRef]
- Śmietanka, B.; Zbawicka, M.; Sanko, T.; Wenne, R.; Burzyński, A. Molecular population genetics of male and female mitochondrial genomes in subarctic Mytilus trossulus. Mar. Biol. 2013, 160, 1709–1721. [Google Scholar] [CrossRef]
- Śmietanka, B.; Burzyński, A.; Hummel, H.; Wenne, R. Glacial history of the European marine mussels Mytilus, inferred from distribution of mitochondrial DNA lineages. Heredity 2014, 113, 250–258. [Google Scholar] [CrossRef]
- Zbawicka, M.; Sanko, T.; Strand, J.; Wenne, R. New SNP markers reveal largely concordant clinal variation across the hybrid zone between Mytilus spp. in the Baltic Sea. Aquat. Biol. 2014, 21, 25–36. [Google Scholar] [CrossRef]
- Wenne, R.; Bach, L.; Zbawicka, M.; Strand, J.; McDonald, J.H. A first report on coexistence and hybridization of Mytilus trossulus and M. edulis mussels in Greenland. Polar Biol. 2015, 39, 343–355. [Google Scholar] [CrossRef]
- Bach, L.; Zbawicka, M.; Strand, J.; Wenne, R. Mytilus trossulus in NW Greenland is genetically more similar to North Pacific than NW Atlantic populations of the species. Mar. Biodivers. 2018, 49, 1053–1059. [Google Scholar] [CrossRef]
- Rawson, P.D.; Hilbish, T. Evolutionary relationships among the male and female mitochondrial DNA lineages in the Mytilus edulis species complex. Mol. Biol. Evol. 1995, 12, 893–901. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Larraín, M.A.; Zbawicka, M.; Araneda, C.; Gardner, J.P.A.; Wenne, R. Native and invasive taxa on the Pacific coast of South America: Impacts on aquaculture, traceability and biodiversity of blue mussels (Mytilus spp.). Evol. Appl. 2018, 11, 298–311. [Google Scholar] [CrossRef]
- Vermeij, G.J. Anatomy of an invasion: The trans-Arctic interchange. Paleobiology 1991, 17, 281–307. [Google Scholar] [CrossRef]
- Riginos, C.; Henzler, C.M. Erratum to: Patterns of mtDNA diversity in North Atlantic populations of the mussel Mytilus edulis. Mar. Biol. 2009, 156, 2649. [Google Scholar] [CrossRef]
- Riginos, C.; Hickerson, M.J.; Henzler, C.M.; Cunningham, C. Differential patterns of male and female mtDNA exchange across the Atlantic Ocean in the blue mussel, Mytilus edulis. Evolution 2004, 58, 2438–2451. [Google Scholar] [CrossRef] [PubMed]
- Quesada, H.; Wenne, R.; Skibinski, D.O.F. Differential introgression of mitochondrial DNA across species boundaries within the marine mussel genus Mytilus. Proc. R. Soc. B Biol. Sci. 1995, 262, 51–56. [Google Scholar] [CrossRef]
- Paterno, M.; Bat, L.; Ben Souissi, J.; Boscari, E.; Chassanite, A.; Congiu, L.; Guarnieri, G.; Kruschel, C.; Mačić, V.; Marino, I.A.M.; et al. A Genome-Wide Approach to the Phylogeography of the Mussel Mytilus galloprovincialis in the Adriatic and the Black Seas. Front. Mar. Sci. 2019, 6, 566. [Google Scholar] [CrossRef]
- Roux, C.; Fraisse, C.; Castric, V.; Vekemans, X.; Pogson, G.H.; Bierne, N. Can we continue to neglect genomic variation in introgression rates when inferring the history of speciation? A case study in a Mytilus hybrid zone. J. Evol. Biol. 2014, 27, 1662–1675. [Google Scholar] [CrossRef] [PubMed]
- Rawson, P.D.; Harper, F.M. Colonization of the northwest Atlantic by the blue mussel, Mytilus trossulus postdates the last glacial maximum. Mar. Biol. 2009, 156, 1857–1868. [Google Scholar] [CrossRef]
- Riginos, C.; Cunningham, C. INVITED REVIEW: Local adaptation and species segregation in two mussel (Mytilus edulis × Mytilus trossulus) hybrid zones. Mol. Ecol. 2004, 14, 381–400. [Google Scholar] [CrossRef]
- Zbawicka, M.; Trucco, M.I.; Wenne, R. Single nucleotide polymorphisms in native South American Atlantic coast populations of smooth shelled mussels: Hybridization with invasive European Mytilus galloprovincialis. Genet. Sel. Evol. 2018, 50, 5. [Google Scholar] [CrossRef] [PubMed]
- Zbawicka, M.; Gardner, J.P.A.; Wenne, R. Cryptic diversity in smooth-shelled mussels on Southern Ocean islands: Connectivity, hybridisation and a marine invasion. Front. Zool. 2019, 16, 32. [Google Scholar] [CrossRef]
- Toro, J.E.; Ojeda, J.A.; Vergara, A.M.; Castro, G.C.; Alcapan, A.C. Molecular characterization of the Chilean blue mussel (Mytilus chilensis Hupe 1854) demonstrates evidence for the occurrence of Mytilus galloprovincialis in southern Chile. J. Shellfish Res. 2005, 24, 1117–1121. [Google Scholar] [CrossRef]
- Zardi, G.I.; McQuaid, C.D.; Jacinto, R.; Lourenço, C.R.; Serrão, E.Á.; Nicastro, K. Re-assessing the origins of the invasive mussel Mytilus galloprovincialis in southern Africa. Mar. Freshw. Res. 2018, 69, 607–613. [Google Scholar] [CrossRef]
- Lockwood, B.L.; Somero, G.N. Invasive and native blue mussels (genus Mytilus) on the California coast: The role of physiology in a biological invasion. J. Exp. Mar. Biol. Ecol. 2011, 400, 167–174. [Google Scholar] [CrossRef]
- Gardner, J.P.A.; Thompson, R.J. Influence of genotype and geography on shell shape and morphometric trait variation among North Atlantic blue mussel (Mytilus spp.) populations. Biol. J. Linn. Soc. 2009, 96, 875–897. [Google Scholar] [CrossRef]
- Zbawicka, M.; Wenne, R.; Burzyński, A. Mitogenomics of recombinant mitochondrial genomes of Baltic Sea Mytilus mussels. Mol. Genet. Genom. 2014, 289, 1275–1287. [Google Scholar] [CrossRef]
- Fraisse, C.; Belkhir, K.; Welch, J.J.; Bierne, N. Local interspecies introgression is the main cause of extreme levels of intraspecific differentiation in mussels. Mol. Ecol. 2015, 25, 269–286. [Google Scholar] [CrossRef]
- Väinölä, R.; Hvilsom, M.M. Genetic divergence and a hybrid zone between Baltic and North Sea Mytilus populations (Mytilidae: Mollusca). Biol. J. Linn. Soc. 1991, 43, 127–148. [Google Scholar] [CrossRef]
- Borsa, P.; Daguin, C.; Caetano, S.R.; Bonhomme, F. Nuclear-DNA evidence that northeastern Atlantic Mytilus trossulus mussels carry M. edulis genes. J. Molluscan Stud. 1999, 65, 504–507. [Google Scholar] [CrossRef]
- Bierne, N.; Borsa, P.; Daguin, C.; Jollivet, D.; Viard, F.; Bonhomme, F.; David, P. Introgression patterns in the mosaic hybrid zone between Mytilus edulis and M. galloprovincialis. Mol. Ecol. 2003, 12, 447–461. [Google Scholar] [CrossRef] [PubMed]
- Beaumont, A.R.; Hawkins, M.P.; Doig, F.L.; Davies, I.M.; Snow, M. Three species of Mytilus and their hybrids identified in a Scottish Loch: Natives, relicts and invaders? J. Exp. Mar. Biol. Ecol. 2008, 367, 100–110. [Google Scholar] [CrossRef]
- Zbawicka, M.; Burzyński, A.; Skibinski, D.; Wenne, R. Scottish Mytilus trossulus mussels retain ancestral mitochondrial DNA: Complete sequences of male and female mtDNA genomes. Gene 2010, 456, 45–53. [Google Scholar] [CrossRef]
- Kaiser, M.J.; Attrill, M.J.; Jennings, S.; Thomas, D.N.; Barnes, D.K.A.; Brierley, A.S.; Hiddink, J.G.; Kaartokallio, H.; Polunin, N.V.C.; Raffaelli, D.G. Marine Ecology: Processes, Systems, and Impacts; Oxford University Press: Oxford, UK, 2011. [Google Scholar]
- Kotta, J.; Oganjan, K.; Lauringson, V.; Pärnoja, M.; Kaasik, A.; Rohtla, L.; Kotta, I.; Orav-Kotta, H. Establishing Functional Relationships between Abiotic Environment, Macrophyte Coverage, Resource Gradients and the Distribution of Mytilus trossulus in a Brackish Non-Tidal Environment. PLoS ONE 2015, 10, e0136949. [Google Scholar] [CrossRef]
- Lucas, J.M.; Vaccaro, E.; Waite, J.H. A molecular, morphometric and mechanical comparison of the structural elements of byssus from Mytilus edulis and Mytilus galloprovincialis. J. Exp. Biol. 2002, 205, 1807–1817. [Google Scholar]
- Braby, C.E.; Somero, G.N. Ecological gradients and relative abundance of native (Mytilus trossulus) and invasive (Mytilus galloprovincialis) blue mussels in the California hybrid zone. Mar. Biol. 2005, 148, 1249–1262. [Google Scholar] [CrossRef]
- Hayhurst, S.; Rawson, P.D. Species-specific variation in larval survival and patterns of distribution for the blue mussels Mytilus edulis and Mytilus trossulus in the Gulf of Maine. J. Molluscan Stud. 2009, 75, 215–222. [Google Scholar] [CrossRef]
- Brooks, S.J.; Farmen, E.; Heier, L.S.; Blanco-Rayón, E.; Izagirre, U. Differences in copper bioaccumulation and biological responses in three Mytilus species. Aquat. Toxicol. 2015, 160, 1–12. [Google Scholar] [CrossRef][Green Version]
- Telesca, L.; Michalek, K.; Sanders, T.; Peck, L.S.; Thyrring, J.; Harper, E.M. Blue mussel shell shape plasticity and natural environments: A quantitative approach. Sci. Rep. 2018, 8, 2865. [Google Scholar] [CrossRef]
- Jones, S.J.; Lima, F.P.; Wethey, D.S. Rising environmental temperatures and biogeography: Poleward range contraction of the blue mussel, Mytilus edulis L., in the western Atlantic. J. Biogeogr. 2010, 37, 2243–2259. [Google Scholar] [CrossRef]
- Thomas, Y.; Bacher, C. Assessing the sensitivity of bivalve populations to global warming using an individual-based modelling approach. Glob. Chang. Biol. 2018, 24, 4581–4597. [Google Scholar] [CrossRef] [PubMed]
- Sorte, C.J.B.; Jones, S.J.; Miller, L.P. Geographic variation in temperature tolerance as an indicator of potential population responses to climate change. J. Exp. Mar. Biol. Ecol. 2011, 400, 209–217. [Google Scholar] [CrossRef]
- Hoarau, G.; Rijnsdorp, A.D.; Van Der Veer, H.W.; Stam, W.T.; Olsen, J.L. Population structure of plaice (Pleuronectes platessa L.) in northern Europe: Microsatellites revealed large-scale spatial and temporal homogeneity. Mol. Ecol. 2002, 11, 1165–1176. [Google Scholar] [CrossRef] [PubMed]
- Śmietanka, B.; Zbawicka, M.; Wołowicz, M.; Wenne, R. Mitochondrial DNA lineages in the European populations of mussels (Mytilus spp.). Mar. Biol. 2004, 146, 79–92. [Google Scholar] [CrossRef]
- Gardner, J.P.A.; Zbawicka, M.; Westfall, K.M.; Wenne, R. Invasive blue mussels threaten regional scale genetic diversity in mainland and remote offshore locations: The need for baseline data and enhanced protection in the Southern Ocean. Glob. Chang. Biol. 2016, 22, 3182–3195. [Google Scholar] [CrossRef] [PubMed]
- Gabriel, S.; Ziaugra, L.; Tabbaa, D. SNP Genotyping Using the Sequenom MassARRAY iPLEX Platform. Curr. Protoc. Hum. Genet. 2009, 60, 2.12.1–2.12.18. [Google Scholar] [CrossRef]
- Excoffier, L.; Lischer, H.E.L. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 2010, 10, 564–567. [Google Scholar] [CrossRef]
- Yekutieli, D.; Benjamini, Y. under dependency. Ann. Stat. 2001, 29, 1165–1188. [Google Scholar] [CrossRef]
- Narum, S.R. Beyond Bonferroni: Less conservative analyses for conservation genetics. Conserv. Genet. 2006, 7, 783–787. [Google Scholar] [CrossRef]
- Diz, Á.P.; Skibinski, D.O.F.; Carvajal-Rodriguez, A. Multiple hypothesis testing in proteomics: A strategy for experimental work. Mol. Cell. Proteom. 2010, 10, 10. [Google Scholar] [CrossRef] [PubMed]
- Rousset, F. genepop’007: A complete re-implementation of the genepop software for Windows and Linux. Mol. Ecol. Resour. 2008, 8, 103–106. [Google Scholar] [CrossRef] [PubMed]
- Takezaki, N.; Nei, M.; Tamura, K. POPTREEW: Web version of POPTREE for constructing population trees from allele frequency data and computing some other quantities. Mol. Biol. Evol. 2014, 31, 1622–1624. [Google Scholar] [CrossRef] [PubMed]
- Anderson, E.C.; Thompson, E.A. A model-based method for identifying species hybrids using multilocus genetic data. Genetics 2002, 160, 1217–1229. [Google Scholar]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar]
- Falush, D.; Stephens, M.; Pritchard, J.K. Inference of population structure using multilocus genotype data: Dominant markers and null alleles. Mol. Ecol. Notes 2007, 7, 574–578. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Vähä, J.-P.; Primmer, C.R. Efficiency of model-based Bayesian methods for detecting hybrid individuals under different hybridization scenarios and with different numbers of loci. Mol. Ecol. 2005, 15, 63–72. [Google Scholar] [CrossRef]
- Lecis, R.; Pierpaoli, M.; Birò, Z.S.; Szemethy, L.; Ragni, B.; Vercillo, F.; Randi, E. Bayesian analyses of admixture in wild and domestic cats (Felis silvestris) using linked microsatellite loci. Mol. Ecol. 2005, 15, 119–131. [Google Scholar] [CrossRef]
- Benzécri, J.P. Correspondence analysis handbook. In Statistics: A Series of Textbooks and Monographs; Balakrishnan, N., Schucany, W.R., Garvey, P.R., Eds.; Marcel Dekker: New York, NY, USA, 1992; Volume 125. [Google Scholar]
- Belkhir, K.; Borsa, P.; Chikhi, L.; Raufaste, N.; Bonhomme, F. GENETIX Version 4.04, Logiciel sous Windows™ pour la Genetique des Populations. Laboratoire Genome, Populations, Interactions: CNRS UMR 5000; Université de Montpellier II: Montpellier, France, 2003. [Google Scholar]
- Stackhouse, P.W.; Gupta, S.K. ISLSCP II Cloud and Meteorology Parameters. In ISLSCP Initiative II Collection; Forrest, G., Collatz, G., Meeson, B., Los, S., de Colstoun, E.B., Landis, D., Eds.; Data Set; Oak Ridge National Laboratory Distributed Active Archive Center: Oak Ridge, TN, USA, 2012. [Google Scholar] [CrossRef]
- Fick, S.E.; Hijmans, R.J. WorldClim 2: New 1-km spatial resolution climate surfaces for global land areas. Int. J. Clim. 2017, 37, 4302–4315. [Google Scholar] [CrossRef]
- Armstrong, R.; Knowles, K. ISLSCP II Global Sea Ice Concentration. In ISLSCP Initiative II Collection; Forrest, G., Collatz, G., Meeson, B., Los, S., de Colstoun, E.B., Landis, D., Eds.; Data Set; Oak Ridge National Laboratory Distributed Active Archive Center: Oak Ridge, TN, USA, 2010. [Google Scholar] [CrossRef]
- Shutler, J.; Land, P.; Smyth, T.; Groom, S. Extending the MODIS 1 km ocean colour atmospheric correction to the MODIS 500 m bands and 500 m chlorophyll-a estimation towards coastal and estuarine monitoring. Remote. Sens. Environ. 2007, 107, 521–532. [Google Scholar] [CrossRef]
- Dormann, C.F.; Elith, J.; Bacher, S.; Buchmann, C.; Carl, G.; Carré, G.; Márquez, J.R.G.; Gruber, B.; Lafourcade, B.; Leitão, P.J.; et al. Collinearity: A review of methods to deal with it and a simulation study evaluating their performance. Ecography 2012, 36, 27–46. [Google Scholar] [CrossRef]
- Hastie, T.; Tibshirani, R.; Friedman, J. Overview of Supervised Learning. In The Elements of Statistical Learning; Springer Series in Statistics; Springer: New York, NY, USA, 2009. [Google Scholar]
- Elith, J.; Graham, C.H.; Anderson, R.P.; Dudík, M.; Ferrier, S.; Guisan, A.; Hijmans, R.J.; Huettmann, F.; Leathwick, J.R.; Lehmann, A.; et al. Novel methods improve prediction of species’ distributions from occurrence data. Ecography 2006, 29, 129–151. [Google Scholar] [CrossRef]
- Leathwick, J.R.; Elith, J.; Francis, M.; Hastie, T.; Taylor, P. Variation in demersal fish species richness in the oceans surrounding New Zealand: An analysis using boosted regression trees. Mar. Ecol. Prog. Ser. 2006, 321, 267–281. [Google Scholar] [CrossRef]
- Yang, R.-M.; Zhang, G.-L.; Liu, F.; Lu, Y.-Y.; Yang, F.; Yang, F.; Yang, M.; Zhao, Y.-G.; Li, D.-C. Comparison of boosted regression tree and random forest models for mapping topsoil organic carbon concentration in an alpine ecosystem. Ecol. Indic. 2016, 60, 870–878. [Google Scholar] [CrossRef]
- Elith, J.; Leathwick, J.R.; Hastie, T. A working guide to boosted regression trees. J. Anim. Ecol. 2008, 77, 802–813. [Google Scholar] [CrossRef]
- Kotta, J.; Orav-Kotta, H.; Jänes, H.; Hummel, H.; Arvanitidis, C.; Van Avesaath, P.; Bachelet, G.; Benedetti-Cecchi, L.; Bojanić, N.; Como, S.; et al. Essence of the patterns of cover and richness of intertidal hard bottom communities: A pan-European study. J. Mar. Biol. Assoc. UK. 2016, 97, 525–538. [Google Scholar] [CrossRef]
- Greenwell, B.; Boehmke, B.; Cunningham, J. GBM Developers 2018. gbm: Generalized Boosted Regression Models. R Package Version 2.1.4. Available online: https://CRAN.R-project.org/package=gbm (accessed on 10 December 2019).
- Clarke, K.R.; Gorley, R.N. PRIMER v7: User Manual/Tutorial; PRIMER-E: Plymouth, UK, 2015. [Google Scholar]
- McDonald, J.H.; Seed, R.; Koehn, R.K. Allozymes and morphometric characters of three species of Mytilus in the Northern and Southern Hemispheres. Mar. Biol. 1991, 111, 323–333. [Google Scholar] [CrossRef]
- Bates, J.A.; Innes, D. Genetic variation among populations of Mytilus spp. in eastern Newfoundland. Mar. Biol. 1995, 124, 417–424. [Google Scholar] [CrossRef]
- Dewaard, J.R.; Ratnasingham, S.; Zakharov, E.V.; Borisenko, A.V.; Steinke, D.; Telfer, A.C.; Perez, K.H.J.; Sones, J.E.; Young, M.R.; Levesque-Beaudin, V.; et al. A reference library for Canadian invertebrates with 1.5 million barcodes, voucher specimens, and DNA samples. Sci. Data 2019, 6, 308–312. [Google Scholar] [CrossRef]
- Katolikova, M.; Khaitov, V.; Väinölä, R.; Gantsevich, M.M.; Strelkov, P. Genetic, Ecological and Morphological Distinctness of the Blue Mussels Mytilus trossulus Gould and M. edulis L. in the White Sea. PLoS ONE 2016, 11, e0152963. [Google Scholar] [CrossRef] [PubMed]
- Brooks, S.J.; Farmen, E. The Distribution of the Mussel Mytilus Species Along the Norwegian Coast. J. Shellfish. Res. 2013, 32, 265–270. [Google Scholar] [CrossRef][Green Version]
- Dias, P.J.; Piertney, S.; Snow, M.; Davies, I.M. Survey and management of mussel Mytilus species in Scotland. Hydrobiologia 2011, 670, 127–140. [Google Scholar] [CrossRef]
- Śmietanka, B.; Burzyński, A. Disruption of doubly uniparental inheritance of mitochondrial DNA associated with hybridization area of European Mytilus edulis and Mytilus trossulus in Norway. Mar. Biol. 2017, 164, 209. [Google Scholar] [CrossRef][Green Version]
- Simon, A.; Arbiol, C.; Nielsen, E.E.; Couteau, J.; Sussarellu, R.; Burgeot, T.; Bernard, I.; Coolen, J.W.; Lamy, J.-B.; Robert, S.; et al. Replicated anthropogenic hybridisations reveal parallel patterns of admixture in marine mussels. Evol. Appl. 2019, 13, 575–599. [Google Scholar] [CrossRef]
- Varvio, S.-L.; Koehn, R.K. Evolutionary genetics of the Mytilus edulis complex in the North Atlantic region. Mar. Biol. 1988, 98, 51–60. [Google Scholar] [CrossRef]
- Wenne, R.; Skibinski, D.O.F. Mitochondrial DNA heteroplasmy in European populations of the mussel Mytilus trossulus. Mar. Biol. 1995, 122, 619–624. [Google Scholar] [CrossRef]
- Rawson, P.D.; Hilbish, T. Asymetric introgression of mitochondrial DNA among European populations of blue mussels (Mytilus spp.). Evolution 1998, 52, 100–108. [Google Scholar] [CrossRef]
- Śmietanka, B.; Burzyński, A.; Wenne, R. Molecular population genetics of male and female mitochondrial genomes in European mussels Mytilus. Mar. Biol. 2009, 156, 913–925. [Google Scholar] [CrossRef]
- Skibinski, D.O.F.; Beardmore, J.A.; Cross, T.F. Aspects of the population genetics of Mytilus (Mytilidae; Mollusca) in the British Isles. Biol. J. Linn. Soc. Lond. 1983, 19, 137–183. [Google Scholar] [CrossRef]
- Araujo, M.B.; Peterson, A.T. Uses and misuses of bioclimatic envelope modeling. Ecology 2012, 93, 1527–1539. [Google Scholar] [CrossRef] [PubMed]
- Jiggins, C.D.; Mallet, J. Bimodal hybrid zones and speciation. Trends Ecol. Evol. 2000, 15, 250–255. [Google Scholar] [CrossRef]
- Leopold, P.; Renaud, P.E.; Ambrose, W.G.; Berge, J. High Arctic Mytilus spp.: Occurrence, distribution and history of dispersal. Polar Biol. 2018, 42, 237–244. [Google Scholar] [CrossRef]
- Hansen, J.; Hanken, N.; Nielsen, J.K.; Nielsen, J.K.; Thomsen, E. Late Pleistocene and Holocene distribution of Mytilus edulis in the Barents Sea region and its palaeoclimatic implications. J. Biogeogr. 2011, 38, 1197–1212. [Google Scholar] [CrossRef]
- Briski, E.; Ghabooli, S.; Bailey, S.; MacIsaac, H.J. Invasion risk posed by macroinvertebrates transported in ships’ ballast tanks. Biol. Invasions 2012, 14, 1843–1850. [Google Scholar] [CrossRef]
- Wenne, R. Single nucleotide polymorphism markers with applications in aquaculture and assessment of its impact on natural populations. Aquat. Living Resour. 2017, 31, 2. [Google Scholar] [CrossRef]
- Ridgway, G.; Nævdal, G. Genotypes of Mytilus from waters of different salinity around Bergen, Norway. Helgol. Mar. Res. 2004, 58, 104–109. [Google Scholar] [CrossRef]
- Qiu, J.; Tremblay, R.; Bourget, E. Ontogenetic changes in hyposaline tolerance in the mussels Mytilus edulis and M. trossulus: Implications for distribution. Mar. Ecol. Prog. Ser. 2002, 228, 143–152. [Google Scholar] [CrossRef]
- Moreau, V.; Tremblay, R.; Bourget, E. Distribution of Mytilus edulis and M. trossulus on the Gaspe Coast in relation to spatial scale. J. Shellfish Res. 2005, 24, 545–555. [Google Scholar] [CrossRef]
- Tedengren, M.; Kautsky, N. Comparative study of the physiology and its probable effect on size in Blue Mussels (Mytilus edulis L.) from the North Sea and the Northern Baltic Proper. Ophelia 1986, 25, 147–155. [Google Scholar] [CrossRef]
- Riisgård, H.U.; Egede, P.; Saavedra, I.B.; Riisgå Rd, H.U. Feeding Behaviour of the Mussel, Mytilus edulis: New Observations, with a Minireview of Current Knowledge. J. Mar. Biol. 2011, 2011, 312459. [Google Scholar] [CrossRef]
- Bennike, O.; Wagner, B. Holocene range of Mytilus edulis in central East Greenland. Polar Rec. 2012, 49, 291–296. [Google Scholar] [CrossRef]
- Toro, J.E.; Thompson, R.J.; Innes, D. Reproductive isolation and reproductive output in two sympatric mussel species (Mytilus edulis, M. trossulus) and their hybrids from Newfoundland. Mar. Biol. 2002, 141, 897–909. [Google Scholar] [CrossRef]
- Toro, J.; Thompson, R.J.; Innes, D. Fertilization success and early survival in pure and hybrid larvae of Mytilus edulis (Linnaeus, 1758) and M. trossulus (Gould, 1850) from laboratory crosses. Aquac. Res. 2006, 37, 1703–1708. [Google Scholar] [CrossRef]
- Popovic, I.; Riginos, C. Comparative genomics reveals divergent thermal selection in warm- and cold-tolerant marine mussels. Mol. Ecol. 2020, 29, 519–535. [Google Scholar] [CrossRef] [PubMed]
- Zippay, M.L.; Helmuth, B. Effects of temperature change on mussel, Mytilus. Integr. Zool. 2012, 7, 312–327. [Google Scholar] [CrossRef] [PubMed]
- Widdows, J. Physiological adaptation of Mytilus edulis to cyclic temperatures. J. Comp. Physiol. B 1976, 105, 115–128. [Google Scholar] [CrossRef]
- Thyrring, J.; Rysgaard, S.; Blicher, M.E.; Sejr, M.K. Metabolic cold adaptation and aerobic performance of blue mussels (Mytilus edulis) along a temperature gradient into the High Arctic region. Mar. Biol. 2014, 162, 235–243. [Google Scholar] [CrossRef]
- Tomanek, L.; Zuzow, M.J. The proteomic response of the mussel congeners Mytilus galloprovincialis and M. trossulus to acute heat stress: Implications for thermal tolerance limits and metabolic costs of thermal stress. J. Exp. Biol. 2010, 213, 3559–3574. [Google Scholar] [CrossRef]
- Malachowicz, M.; Wenne, R. Mantle transcriptome sequencing of Mytilus spp. and identification of putative biomineralization genes. PeerJ 2019, 6, e6245. [Google Scholar] [CrossRef]
- Martino, P.A.; Carlon, D.B.; Kingston, S.E. Blue Mussel (Genus Mytilus) Transcriptome Response to Simulated Climate Change in the Gulf of Maine. J. Shellfish. Res. 2019, 38, 587–602. [Google Scholar] [CrossRef]
- Mlouka, R.; Cachot, J.; Sforzini, S.; Oliveri, C.; Boukadida, K.; Clerandeau, C.; Pacchioni, B.; Millino, C.; Viarengo, A.; Banni, M. Molecular mechanisms underlying the effects of temperature increase on Mytilus sp. and their hybrids at early larval stages. Sci. Total. Environ. 2020, 708, 135200. [Google Scholar] [CrossRef] [PubMed]
- Lamb, H. 1977. Climatic History and the Future; Princeton University Press: Princeton, NJ, USA, 1985; Volume 2. [Google Scholar]
- Hellmann, J.J.; Byers, J.E.; Bierwagen, B.G.; Dukes, J.S. Five Potential Consequences of Climate Change for Invasive Species. Conserv. Biol. 2008, 22, 534–543. [Google Scholar] [CrossRef]
- Saarman, N.P.; Pogson, G.H. Introgression between invasive and native blue mussels (genus Mytilus) in the central California hybrid zone. Mol. Ecol. 2015, 24, 4723–4738. [Google Scholar] [CrossRef] [PubMed]
- Oyarzún, P.; Toro, J.; Navarro, J.M. Comparison of the physiological energetics between Mytilus chilensis, Mytilus galloprovincialis and their hybrids, under laboratory conditions. Aquac. Res. 2012, 44, 1805–1814. [Google Scholar] [CrossRef]
| Sample Name | Localisation | Country | Water Area | No. of Individuals | PO | FIS | Loci with HWE Departure | HO | HE | MAF | Average Gene Diversity over Loci | Average No. of Pairwise Differences within Population | Coordinates | Sample Collection | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BNJ | Belmar, New Jersey | USA | Atlantic | 32 | 46.30 | 0.111 | 2 | 0.233 | 0.275 | 0.086 | 0.122 | 5.23 | 40°11′13.56″ N | 74°0′36.36″ W | 2012 |
| IRD * 1, 2, 3, 4 | Indian River Inlet, Delaware, | USA | Atlantic | 30 | 44.44 | 0.108 | 0 | 0.239 | 0.271 | 0.081 | 0.116 | 4.96 | 36°52′6.19″ N | 75°58′2.16″ W | 2012 |
| KPV | Kiptopeke State Park, Virginia, | USA | Atlantic | 31 | 44.44 | 0.110 | 1 | 0.229 | 0.259 | 0.074 | 0.106 | 4.67 | 37° 9′51.12″ N | 75°59′29.40″ W | 2012 |
| MID | Mispillion Inlet, Delaware, | USA | Atlantic | 30 | 44.44 | 0.222 | 1 | 0.215 | 0.282 | 0.084 | 0.123 | 5.16 | 38°56′42.00″ N | 75°18′38.88″ W | 2012 |
| PNY | Point Lookout, New York | USA | Atlantic | 33 | 48.15 | 0.059 | 2 | 0.215 | 0.243 | 0.079 | 0.106 | 4.73 | 40°35′34.80″ N | 73°34′30.72″ W | 2012 |
| SNY | Stony Brook, New York | USA | Atlantic | 30 | 48.15 | 0.058 | 0 | 0.236 | 0.254 | 0.081 | 0.110 | 4.95 | 40°55′15.96″ N | 73°9′0.36″ W | 2012 |
| NWF | North coast of New Foundland | Canada | Atlantic | 24 | 51.85 | 0.109 | 1 | 0.198 | 0.232 | 0.075 | 0.102 | 4.57 | 49°30′5.32″ N | 55°41′44.21″ W | 2012 |
| PEI | Prince Edward Island | Canada | Atlantic | 31 | 46.30 | −0.008 | 1 | 0.226 | 0.250 | 0.078 | 0.093 | 4.35 | 46°26′11.10″ N | 62°40′24.06″ W | 2012 |
| SNS | Ship Harbour, Nova Scotia | Canada | Atlantic | 23 | 51.85 | 0.046 | 1 | 0.241 | 0.246 | 0.087 | 0.111 | 5.05 | 44°48′5.13″ N | 62°50′13.55″ W | 2012 |
| KKA | Halifax, Nova Scotia | Canada | Atlantic | 40 | 85.19 | 0.120 | 4 | 0.218 | 0.261 | 0.158 | 0.198 | 10.44 | 44°30′33.79″ N | 63°29′24.91″ W | 1996 |
| PBAY | Placentia Bay, New Foundland | Canada | Atlantic | 17 | 81.48 | 0.535 | 18 | 0.172 | 0.388 | 0.248 | 0.312 | 15.29 | 47° 2′40.05″ N | 54°11′34.72″ W | 2012 |
| GLD 1 | North-west Greenland, Maarmorilik, 17 | Denmark | Atlantic | 33 | 79.63 | 0.328 | 10 | 0.233 | 0.373 | 0.229 | 0.266 | 13.45 | 71°8′42.96″ N | 51°16′31.99″ W | 2012 |
| GLL 1 | North-west Greenland, Maarmorilik, L | Denmark | Atlantic | 30 | 77.78 | 0.493 | 21 | 0.193 | 0.382 | 0.214 | 0.291 | 14.15 | 70°59′42.42″ N | 52°16′41.37″ W | 2012 |
| SAV * 2 | North Greenland, Savissivik | Denmark | Atlantic | 27 | 40.38 | 0.144 | 1 | 0.243 | 0.284 | 0.082 | 0.089 | 4.59 | 76°1′5.26″ N | 65°7′4.18″ W | 2015 |
| NUU | South-west Greenland, Nuuk | Denmark | Atlantic | 25 | 37.04 | 0.003 | 1 | 0.296 | 0.336 | 0.081 | 0.113 | 4.90 | 64°10′24.36″ N | 51°29′25.86″ W | 2015 |
| ISLB | Reykjavik | Iceland | Atlantic | 29 | 42.59 | 0.083 | 2 | 0.282 | 0.322 | 0.087 | 0.127 | 5.74 | 64°8′59.03″ N | 21°53′16.61″ W | 1986 |
| ISLR | Reykjavik | Iceland | Atlantic | 30 | 46.30 | −0.043 | 1 | 0.283 | 0.290 | 0.086 | 0.123 | 5.57 | 64°5′44.18″ N | 21°56′48.83″ W | 2004 |
| SPI | Spitsbergen, Smerenburg | Norway | Atlantic | 4 | 37.04 | NA | NA | NA | NA | NA | NA | NA | 79°38′46.94″ N | 11°14′10.38″ E | 2014 |
| BRN | Bergen | Norway | Atlantic | 36 | 88.89 | 0.219 | 7 | 0.281 | 0.379 | 0.262 | 0.325 | 15.71 | 60°23′25.65″ N | 5°12′27.08″ E | 2012 |
| BODS | Bodø, Rundholmen | Norway | Atlantic | 30 | 85.19 | −0.004 | 0 | 0.194 | 0.201 | 0.106 | 0.156 | 7.54 | 67°17′0.61″ N | 14°21′48.02″ E | 2013 |
| BODZ | Bodø, Rønvikleira | Norway | Atlantic | 29 | 85.19 | 0.067 | 0 | 0.197 | 0.223 | 0.120 | 0.185 | 8.52 | 67°17′45.25″ N | 14°23′49.53″ E | 2013 |
| TRO | Tromsø | Norway | Atlantic | 29 | 64.81 | 0.068 | 1 | 0.198 | 0.213 | 0.089 | 0.123 | 5.92 | 69°35′27.68″ N | 18°53′20.62″ E | 2006 |
| BAR | Barents Sea | Russia | Barents Sea | 19 | 79.63 | 0.562 | 18 | 0.157 | 0.348 | 0.185 | 0.268 | 13.02 | 69°20′18″ N | 34°01′28″ E | 2004 |
| KOL | Kola Bay, Abram Mys | Russia | Barents Sea | 27 | 85.19 | 0.444 | 21 | 0.151 | 0.290 | 0.165 | 0.228 | 11.64 | 68°58′56.47″ N | 33°1′36.08″ E | 2014 |
| DLZ | Dalnie Zelentsy, Yarnyshnaya | Russia | Barents Sea | 30 | 83.33 | 0.309 | 2 | 0.135 | 0.201 | 0.108 | 0.162 | 7.35 | 69°5′16.56″ N | 36°3′3.42″ E | 2014 |
| WSBS | White Sea Biological Station | Russia | White Sea | 30 | 83.33 | 0.278 | 2 | 0.142 | 0.186 | 0.093 | 0.146 | 6.85 | 66°33′5.62″ N | 33°6′50.58″ E | 2014 |
| CHU | Chupa Inlet, Kandalaksha Bay | Russia | White Sea | 32 | 81.48 | 0.465 | 19 | 0.168 | 0.325 | 0.186 | 0.234 | 12.11 | 66°16′12.31″ N | 33°4′12.93″ E | 2014 |
| ONE | Chupa | Russia | White Sea | 28 | 53.70 | 0.210 | 4 | 0.202 | 0.247 | 0.089 | 0.128 | 5.59 | 66°15′51.67″ N | 33°2′54.21″ E | 1997 |
| KER | Keret, Kandalaksha Bay | Russia | White Sea | 33 | 83.33 | 0.647 | 26 | 0.135 | 0.357 | 0.207 | 0.297 | 14.36 | 66°17′22.66″ N | 33°40′6.28″ E | 2014 |
| BIA | Białogóra | Poland | Baltic Sea | 30 | 85.19 | 0.098 | 6 | 0.273 | 0.339 | 0.215 | 0.261 | 12.92 | 54°49′55.99″ N | 17°57′9.02″ E | 2014 |
| BOR | Bornholm | Denmark | Baltic Sea | 30 | 85.19 | 0.075 | 4 | 0.297 | 0.339 | 0.216 | 0.272 | 13.82 | 55°4′25.89″ N | 14°43′56.56″ E | 2013 |
| KLA | Klaipeda | Lithuania | Baltic Sea | 30 | 81.48 | 0.094 | 4 | 0.290 | 0.348 | 0.209 | 0.262 | 13.33 | 55°49′4″ N | 20°30′2″ E | 2013 |
| STC | Stevns Klint | Denmark | Baltic Sea | 28 | 87.04 | 0.120 | 5 | 0.305 | 0.374 | 0.241 | 0.306 | 15.03 | 55°16′50.25″ N | 12°26′49.85″ E | 2014 |
| CIS | Cullivoe intertidal Shetland | Great Britain | Atlantic | 33 | 46.30 | −0.004 | 1 | 0.318 | 0.339 | 0.101 | 0.117 | 6.22 | 60°40′0.37″ N | 0°56′40.85″ W | 2012 |
| SCO | Malage, Scotland | Great Britain | Atlantic | 29 | 53.70 | 0.044 | 1 | 0.262 | 0.290 | 0.099 | 0.149 | 6.74 | 57°4′24.00″ N | 5°47′24.00″ W | 2014 |
| LET | Loch Etive, Scotland | Great Britain | Atlantic | 31 | 85.19 | 0.527 | 27 | 0.182 | 0.403 | 0.287 | 0.324 | 15.96 | 56°27′21.35″ N | 5°18′26.62″ W | 2008 |
| OBA | Oban, Scotland | Great Britain | Atlantic | 29 | 48.15 | 0.127 | 2 | 0.209 | 0.270 | 0.091 | 0.112 | 5.28 | 56°24′49.40″ N | 5°28′23.00″ W | 2014 |
| IONA | Iona, Inner Hebrides, Scotland | Great Britain | Atlantic | 29 | 53.70 | 0.128 | 3 | 0.274 | 0.318 | 0.111 | 0.147 | 7.30 | 56°19′52.72″ N | 6°23′29.93″ W | 2014 |
| KRR | Kerrera, Inner Hebrides, Scotland | Great Britain | Atlantic | 30 | 46.30 | 0.128 | 2 | 0.257 | 0.312 | 0.095 | 0.127 | 5.95 | 56°22′42.56″ N | 5°33′17.14″ W | 2014 |
| STA | Staffa, Inner Hebrides, Scotland | Great Britain | Atlantic | 30 | 48.15 | 0.090 | 0 | 0.310 | 0.347 | 0.111 | 0.144 | 7.06 | 56°26′9.98″ N | 6°20′15.43″ W | 2014 |
| GBLO 5 | Lowestoft | Great Britain | Atlantic | 11 | 35.19 | 0.012 | 0 | 0.297 | 0.330 | 0.078 | 0.114 | 4.60 | 52°20′44.07″ N | 1°45′27.63″ E | 2000 |
| LGF 3, 4 | Lough Foyle | Ireland | Atlantic | 28 | 48.15 | 0.062 | 1 | 0.232 | 0.266 | 0.088 | 0.096 | 5.14 | 55°5′35.50″ N | 7°4′48.92″ W | 2006 |
| SAL | Saltö | Sweden | Atlantic | 29 | 64.81 | 0.084 | 1 | 0.194 | 0.217 | 0.088 | 0.132 | 6.02 | 58°52′45.38″ N | 11° 7′13.18″ E | 2014 |
| NLOO 5 | Oosterschelde | Netherlands | Atlantic | 17 | 42.59 | −0.033 | 0 | 0.261 | 0.271 | 0.075 | 0.113 | 4.62 | 51°50′7.10″ N | 3°49′18.21″ E | 2000 |
| MSM | Mont Saint-Michel | France | Atlantic | 4 | 22.22 | NA | NA | NA | NA | NA | NA | NA | 48°39′0.06″ N | 1°31′40.26″ W | 2013 |
| LOI * 1 | Loire | France | Atlantic | 30 | 50.00 | 0.097 | 0 | 0.229 | 0.251 | 0.088 | 0.108 | 5.08 | 47°14′43.83″ N | 2°13′48.88″ W | 2004 |
| BID * 1 | Bidasoa | Spain | Atlantic | 30 | 50.00 | 0.033 | 1 | 0.331 | 0.345 | 0.123 | 0.159 | 7.75 | 43°21′38.71″ N | 1°51′11.15″ W | 2004 |
| VIG | Vigo | Spain | Atlantic | 30 | 53.70 | 0.081 | 2 | 0.285 | 0.318 | 0.125 | 0.161 | 7.65 | 42°13′54.12″ N | 8°45′7.22″ W | 2004 |
| CAS | Cascais | Portugal | Atlantic | 30 | 50.00 | 0.070 | 0 | 0.282 | 0.311 | 0.108 | 0.150 | 7.20 | 38°34′14.89″ N | 9°19′8.95″ W | 2013 |
| CAM 3, 4 | Camarinal | Spain | Atlantic | 29 | 46.30 | 0.035 | 0 | 0.329 | 0.345 | 0.118 | 0.147 | 7.10 | 36° 3′30.09″ N | 5°46′8.88″ W | 2004 |
| IMC | Torre Grande port | Italy | Mediterranean Sea | 20 | 53.70 | 0.055 | 1 | 0.259 | 0.284 | 0.102 | 0.151 | 7.40 | 39°47′59.88″ N | 8°31′9.72″ E | 2004 |
| NEA | Gulf of Naples | Italy | Mediterranean Sea | 30 | 51.85 | 0.038 | 1 | 0.280 | 0.308 | 0.108 | 0.154 | 7.55 | 40°46′44.64″ N | 14°5′28.20″ E | 2014 |
| AZO * | Azov sea | Russia | Azow Sea | 30 | 48.15 | 0.004 | 1 | 0.277 | 0.279 | 0.083 | 0.132 | 6.56 | 45°43′51.71″ N | 35°5′0.26″ E | 1997 |
| Variable | Unit | Temporal Resolution | Spatial Resolution | Data Range |
|---|---|---|---|---|
| Sea ice concentration | % coverage | monthly | 0.25° | 1986–1995 |
| Cloud amount | percent total cloud amount | monthly | 2° | 1986–1995 |
| Wind speed | m s−1 | monthly | 1° | 1983–1993 |
| Solar radiation | kJ m−2 day−1 | monthly | 0.04167° | 1970–2000 |
| Precipitation | mm | monthly | 0.04167° | 1970–2000 |
| Water temperature | °C | monthly | 0.25° | 2005–2017 |
| Salinity | psu | monthly | 0.25° | 2005–2017 |
| Swell height | m | monthly | 2° | 1970–2015 |
| Tide height | cm | monthly | 0.0625° | modeled data |
| Chlorophyll-a concentration | mg m−3 | monthly | 0.04167° | 1997–2017 |
| Concentration of nitrates | µmol L−2 | monthly | 1° | 1955–2012 |
| Concentration of phosphates | µmol L−3 | monthly | 1° | 1955–2012 |
| Structure K = 2 | NewHybrids | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample | Location | HI | VHI | M. edulis | M. trossulus | Hybrids | M. edulis | M. trossulus | F1 hybrids | F2 hybrids | tr_BAX | edu_BAX | |
| North America | % | % | % | % | % | % | % | % | % | ||||
| BNJ | USA | Atlantic | 0.0254 | 0.0256 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| KPV | USA | Atlantic | 0.0182 | 0.0210 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| MID | USA | Atlantic | 0.0125 | 0.0194 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| PNY | USA | Atlantic | 0.0114 | 0.0171 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| SNY | USA | Atlantic | 0.0168 | 0.0229 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| NWF | Canada | Atlantic | 0.0336 | 0.0281 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| PEI | Canada | Atlantic | 0.0319 | 0.0320 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| SNS | Canada | Atlantic | 0.0287 | 0.0282 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| KKA | Canada | Atlantic | 0.8469 | 0.1996 | 2.5 | 80 | 17.5 | 0 | 77.5 | 17.5 | 0 | 2.5 | 2.5 |
| PBAY | Canada | Atlantic | 0.6132 | 0.4315 | 29.41 | 52.94 | 17.65 | 29.41 | 52.94 | 11.76 | 0 | 5.88 | 0 |
| GLD | Greenland | Atlantic | 0.6399 | 0.3781 | 21.21 | 48.48 | 30.30 | 21.21 | 48.48 | 24.24 | 0 | 6.06 | 0 |
| GLL | Greenland | Atlantic | 0.3728 | 0.4122 | 56.67 | 26.67 | 16.67 | 56.67 | 26.67 | 16.67 | 0 | 0 | 0 |
| NUU | Greenland | Atlantic | 0.0444 | 0.0306 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| North Russia | |||||||||||||
| ONE | Russia | White Sea | 0.0202 | 0.0309 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| WSBS | Russia | White Sea | 0.0531 | 0.1811 | 96.67 | 3.33 | 0 | 96.67 | 3.33 | 0 | 0 | 0 | 0 |
| KER | Russia | White Sea | 0.2804 | 0.4247 | 72.73 | 27.27 | 0 | 72.73 | 27.27 | 0 | 0 | 0 | 0 |
| CHU | Russia | White Sea | 0.7558 | 0.3486 | 15.63 | 71.88 | 12.50 | 15.63 | 71.88 | 9.38 | 0 | 3.13 | 0 |
| DLZ | Russia | Barents Sea | 0.0634 | 0.1958 | 93.33 | 3.33 | 3.33 | 93.33 | 3.33 | 3.33 | 0 | 0 | 0 |
| BAR | Russia | Barents Sea | 0.2503 | 0.4056 | 73.68 | 21.05 | 5.26 | 73.68 | 21.05 | 5.26 | 0 | 0 | 0 |
| KOL | Russia | Barents Sea | 0.8235 | 0.3100 | 11.11 | 85.19 | 3.70 | 11.11 | 77.78 | 3.70 | 0 | 7.41 | 0 |
| Scandinavia and Loch Etive | |||||||||||||
| SAL | Sweden | Atlantic | 0.0405 | 0.0465 | 100 | 0 | 0 | 96.55 | 0 | 0 | 0 | 0 | 3.45 |
| TRO | Norway | Atlantic | 0.0304 | 0.0295 | 100 | 0 | 0 | 96.55 | 0 | 0 | 0 | 0 | 3.45 |
| BODS | Norway | Atlantic | 0.0604 | 0.0921 | 96.67 | 0 | 3.33 | 80.00 | 0 | 0 | 3.33 | 0 | 16.67 |
| BODZ | Norway | Atlantic | 0.0752 | 0.1291 | 86.21 | 0 | 13.79 | 82.76 | 0 | 3.45 | 3.45 | 0 | 10.34 |
| BRN | Norway | Atlantic | 0.5867 | 0.2780 | 13.89 | 36.11 | 50.00 | 5.56 | 30.56 | 13.89 | 8.33 | 25.00 | 16.67 |
| LET | Great Britain | Atlantic | 0.5111 | 0.4374 | 38.71 | 48.39 | 12.90 | 38.71 | 41.94 | 12.90 | 0 | 6.45 | 0 |
| Baltic Sea | |||||||||||||
| KLA | Lithuania | Baltic Sea | 0.7255 | 0.0763 | 0 | 60 | 40 | 0 | 50 | 0 | 10 | 40 | 0 |
| BIA | Poland | Baltic Sea | 0.7010 | 0.0893 | 0 | 50 | 50 | 0 | 36,67 | 0 | 10 | 53.33 | 0 |
| BOR | Denmark | Baltic Sea | 0.7259 | 0.1152 | 0 | 63.33 | 36.67 | 0 | 43.33 | 0 | 6.67 | 50 | 0 |
| STC | Denmark | Baltic Sea | 0.6321 | 0.2004 | 3.57 | 46.43 | 50 | 3.57 | 32.14 | 0 | 14.29 | 39.29 | 10.71 |
| reference M. edulis | 0 | ||||||||||||
| LOI | France | Atlantic | 0.0073 | 0.0158 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| IRD | USA | Atlantic | 0.0199 | 0.0255 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 |
| reference M. trossulus | |||||||||||||
| SAV | Greenland | Atlantic | 0.9629 | 0.0313 | 0 | 100 | 0 | 0 | 100 | 0 | 0 | 0 | 0 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wenne, R.; Zbawicka, M.; Bach, L.; Strelkov, P.; Gantsevich, M.; Kukliński, P.; Kijewski, T.; McDonald, J.H.; Sundsaasen, K.K.; Árnyasi, M.; et al. Trans-Atlantic Distribution and Introgression as Inferred from Single Nucleotide Polymorphism: Mussels Mytilus and Environmental Factors. Genes 2020, 11, 530. https://doi.org/10.3390/genes11050530
Wenne R, Zbawicka M, Bach L, Strelkov P, Gantsevich M, Kukliński P, Kijewski T, McDonald JH, Sundsaasen KK, Árnyasi M, et al. Trans-Atlantic Distribution and Introgression as Inferred from Single Nucleotide Polymorphism: Mussels Mytilus and Environmental Factors. Genes. 2020; 11(5):530. https://doi.org/10.3390/genes11050530
Chicago/Turabian StyleWenne, Roman, Małgorzata Zbawicka, Lis Bach, Petr Strelkov, Mikhail Gantsevich, Piotr Kukliński, Tomasz Kijewski, John H. McDonald, Kristil Kindem Sundsaasen, Mariann Árnyasi, and et al. 2020. "Trans-Atlantic Distribution and Introgression as Inferred from Single Nucleotide Polymorphism: Mussels Mytilus and Environmental Factors" Genes 11, no. 5: 530. https://doi.org/10.3390/genes11050530
APA StyleWenne, R., Zbawicka, M., Bach, L., Strelkov, P., Gantsevich, M., Kukliński, P., Kijewski, T., McDonald, J. H., Sundsaasen, K. K., Árnyasi, M., Lien, S., Kaasik, A., Herkül, K., & Kotta, J. (2020). Trans-Atlantic Distribution and Introgression as Inferred from Single Nucleotide Polymorphism: Mussels Mytilus and Environmental Factors. Genes, 11(5), 530. https://doi.org/10.3390/genes11050530






