AaCOI1, Encoding a CORONATINE INSENSITIVE 1-Like Protein of Artemisia annua L., Is Involved in Development, Defense, and Anthocyanin Synthesis
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Growth Condition
2.2. Bioinformatics Analysis
2.3. Mechanical Wounding and Chemical Treatments
2.4. Complementation Assay of AaCOI1 in Arabidopsis Coi1 Mutants
2.5. Subcellular Localization of AaCOI1 in Nicotiana benthamiana
2.6. Bioassay of Herbivory Resistance
2.7. Analysis of Gene Expression
2.8. Quantification of MeJA-Induced Anthocyanin
3. Results
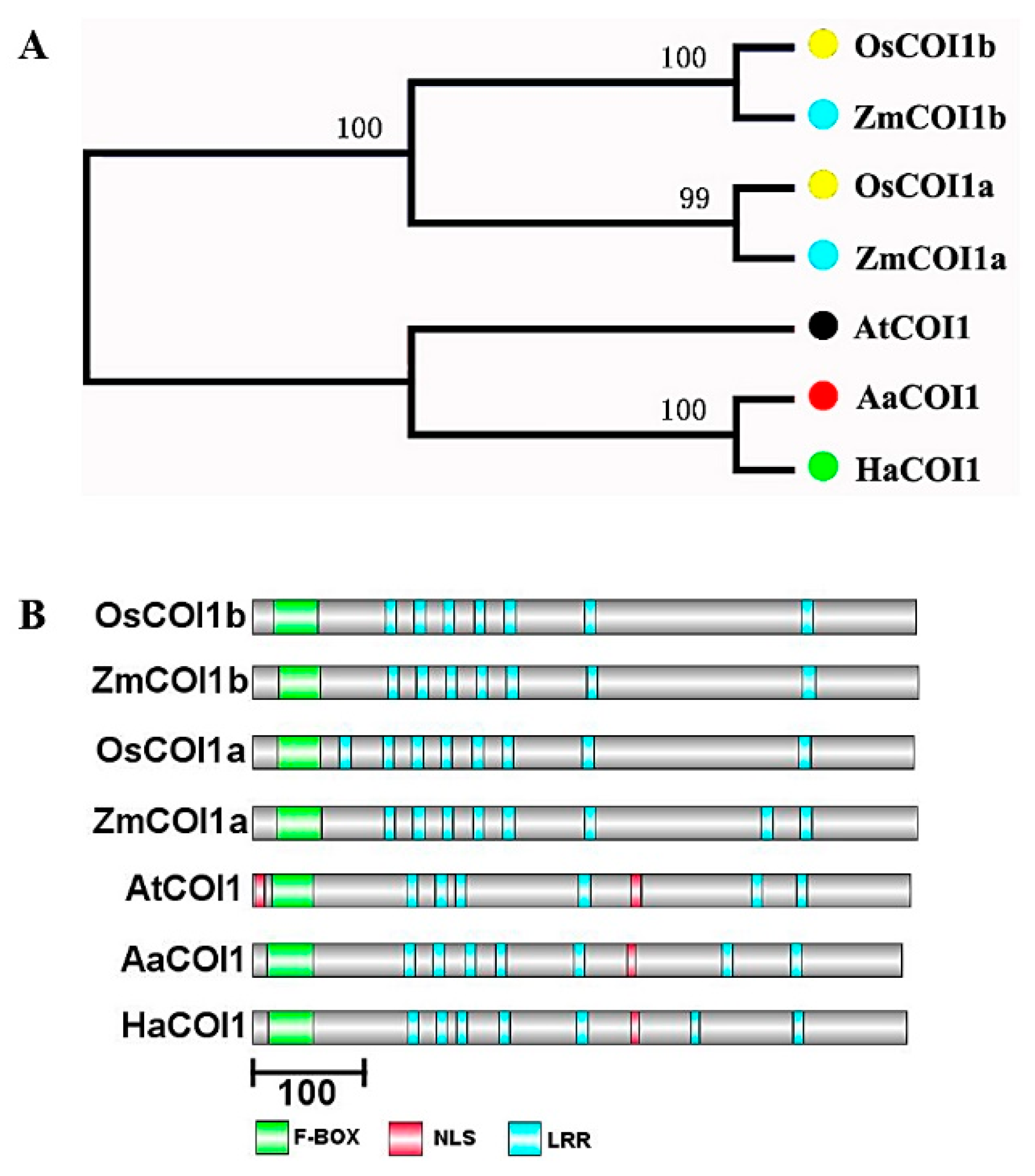
3.1. Identification of AaCOI1 in Artemisia annua
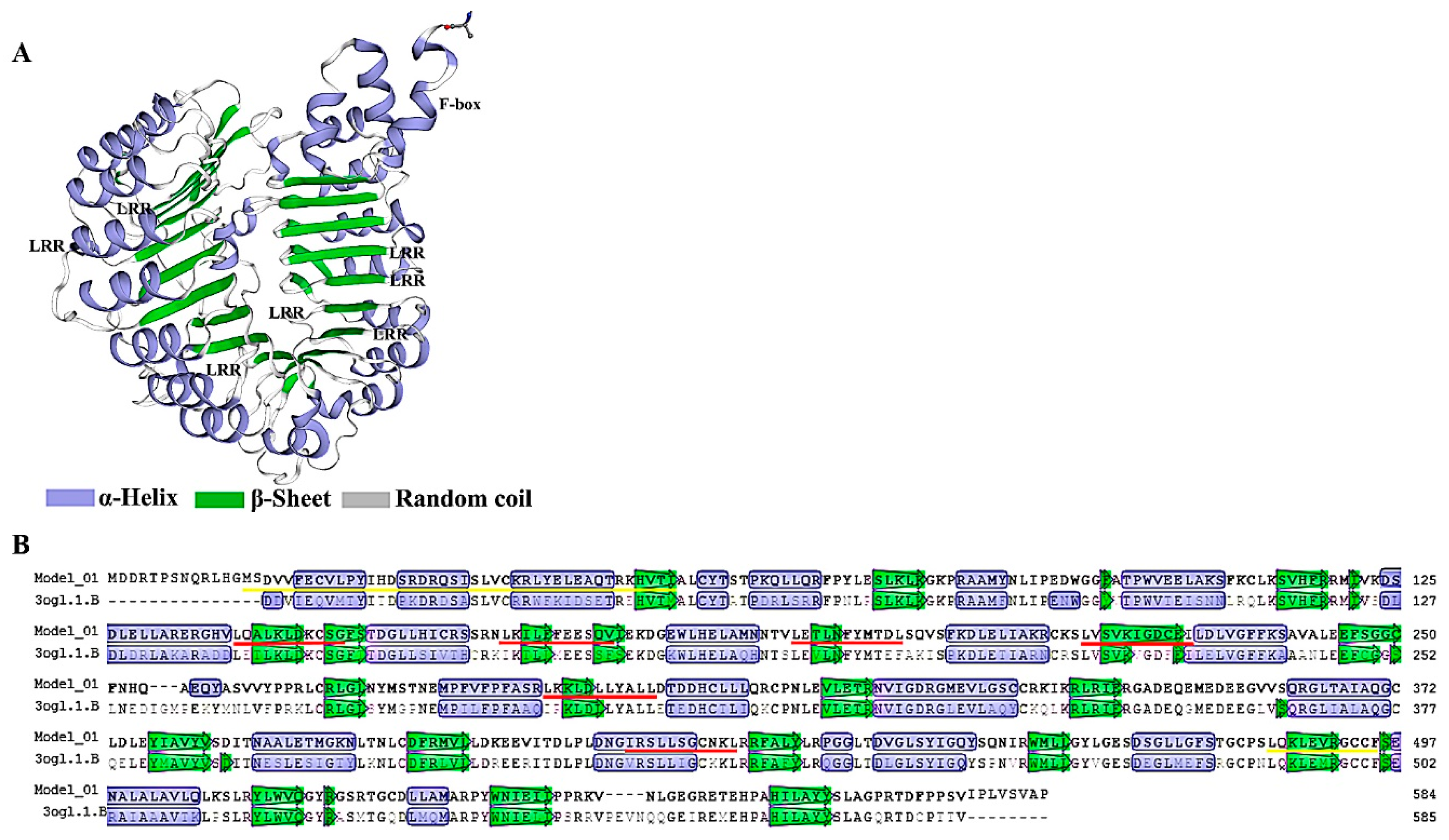
3.2. Sequence Alignment and Structural Analysis
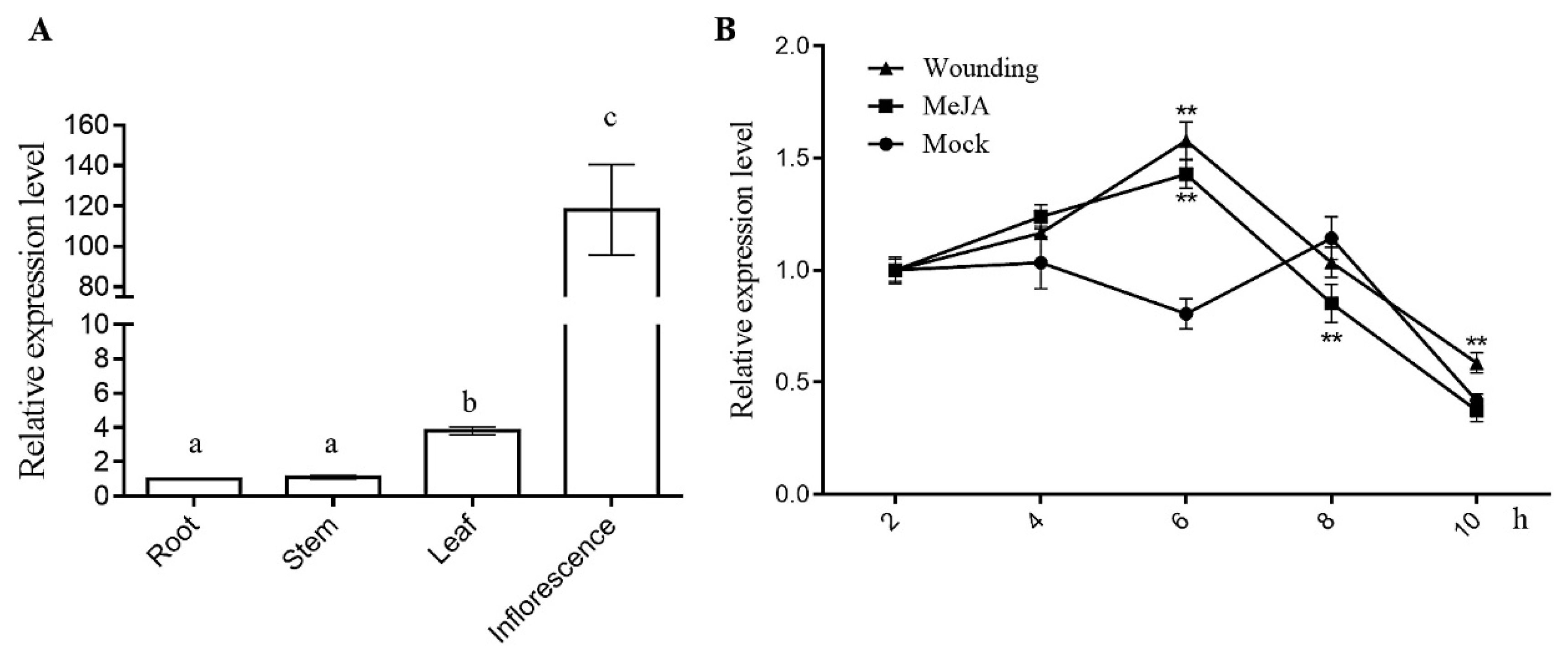
3.3. Expression Patterns of AaCOI1
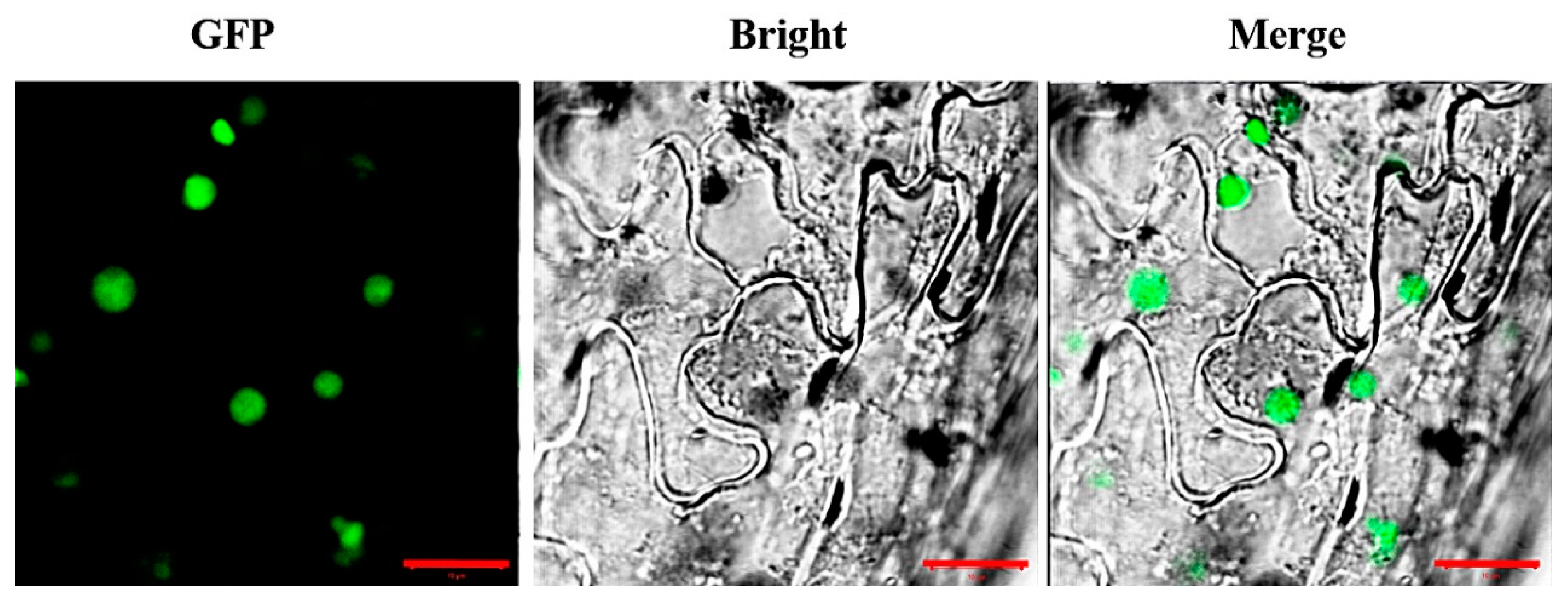
3.4. Subcellular Localization of AaCOI1
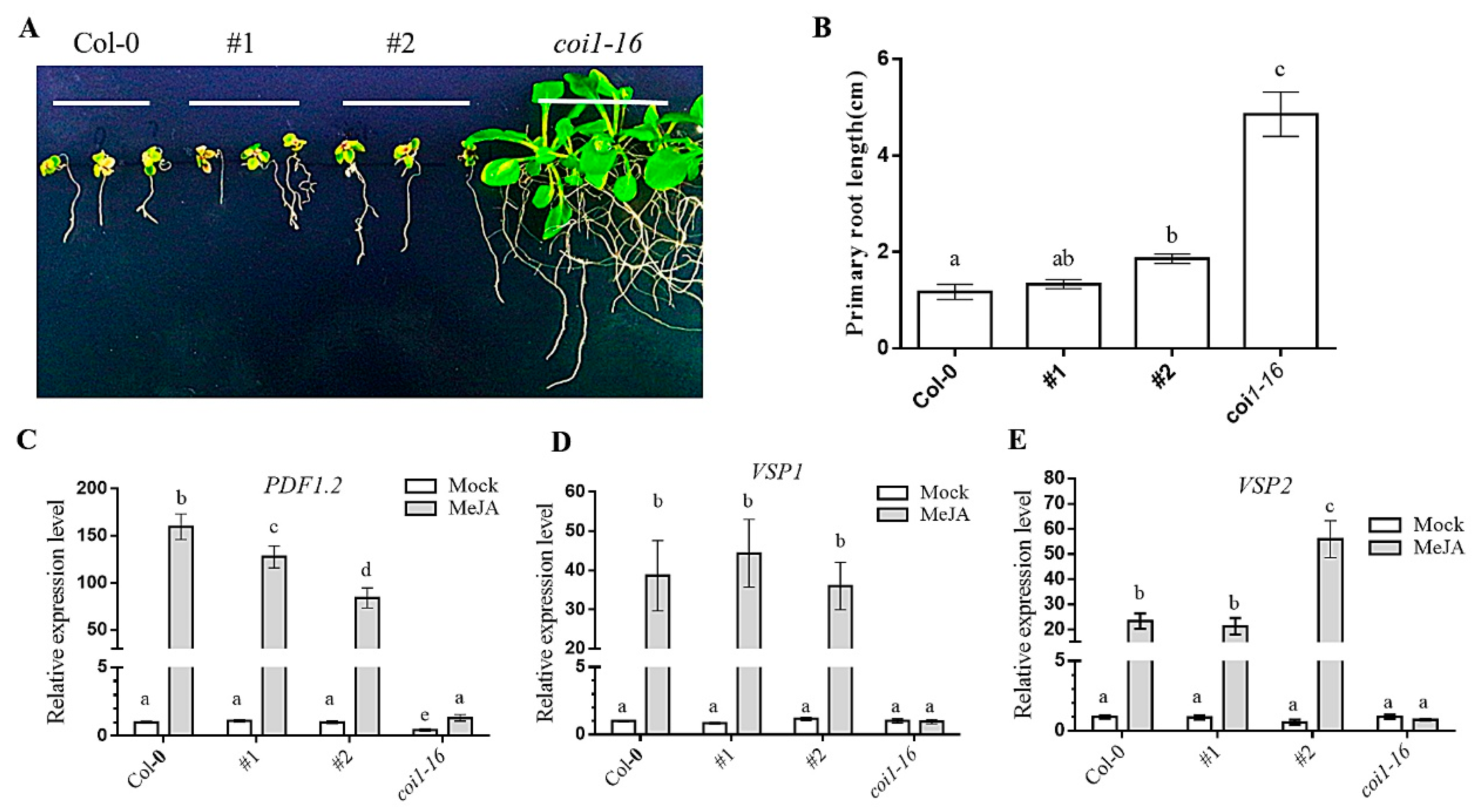
3.5. Ectopic Expression of AaCOI Partially Rescues Developmental Defects of Coi1-16
3.6. Ectopic Expression of AaCOI Restores JA-Sensitivity of Coi1-16
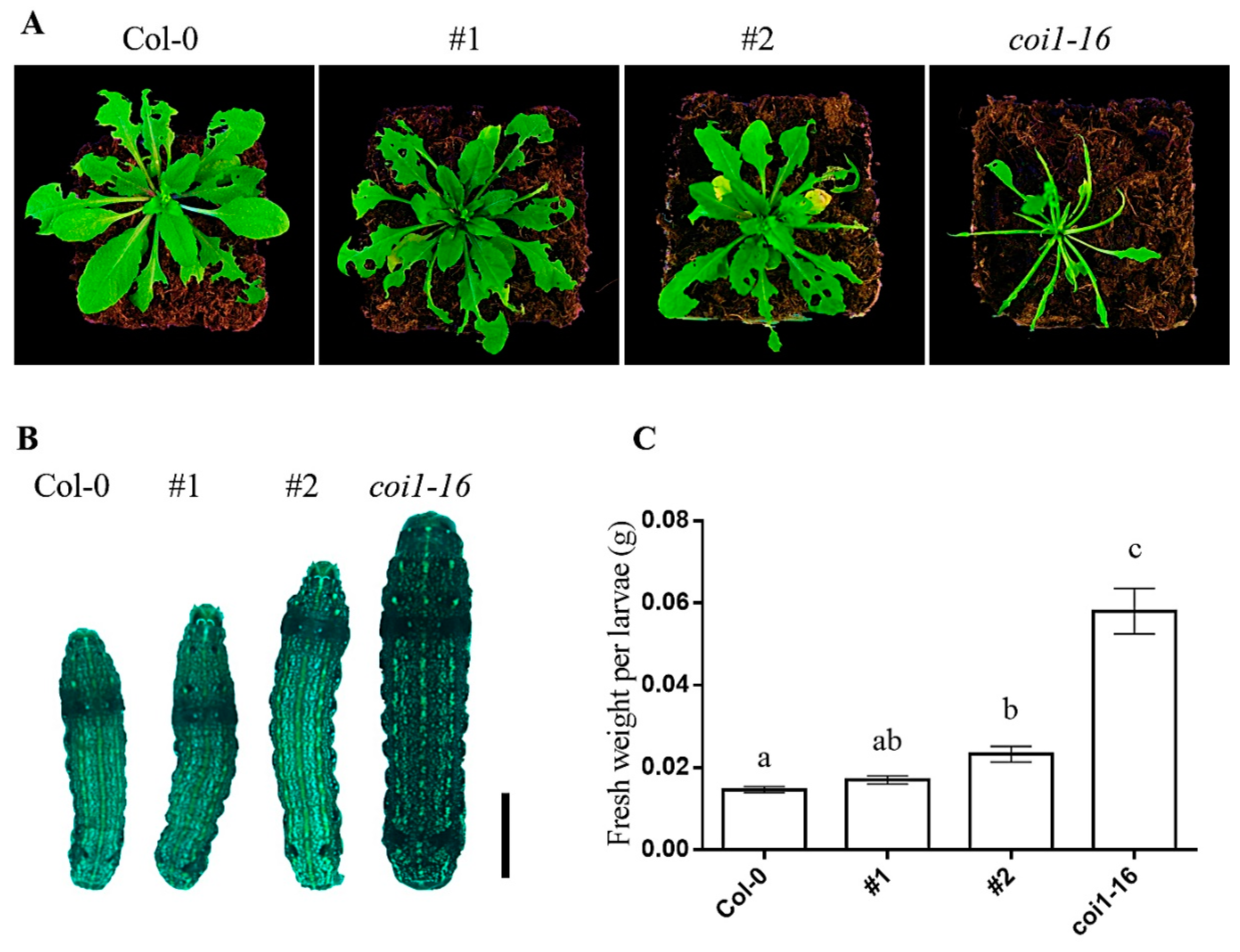
3.7. Ectopic Expression of AaCOI1 Imparts Defense against Herbivory in Coi1-16
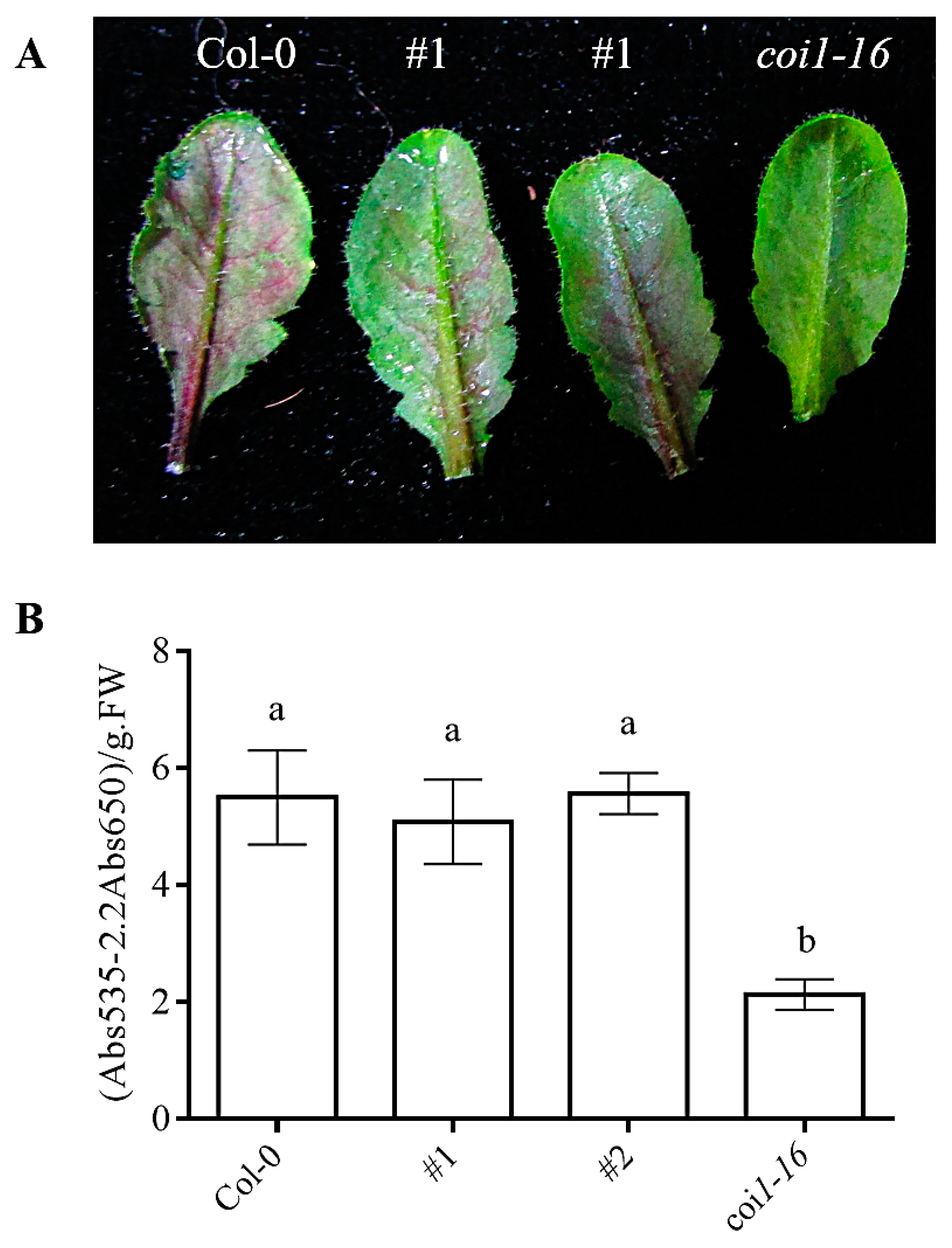
3.8. Ectopic Expression of AaCOI1 Restores the Accumulation of Anthocyanin in Coi1-16
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Wasternack, C.; Kombrink, E. Jasmonates: Structural requirements for lipid-derived signals active in plant stress responses and development. ACS Chem. Biol. 2010, 5, 63–77. [Google Scholar] [CrossRef]
- Fonseca, S.; Chini, A.; Hamberg, M.; Adie, B.; Porzel, A.; Kramell, R.; Miersch, O.; Wasternack, C.; Solano, R. (+)-7-iso-Jasmonoyl-l-isoleucine is the endogenous bioactive jasmonate. Nat. Chem. Biol. 2009, 5, 344–350. [Google Scholar] [CrossRef]
- Wasternack, C.; Song, S. Jasmonates: Biosynthesis, metabolism, and signaling by proteins activating and repressing transcription. J. Exp. Bot. 2017, 68, 1303. [Google Scholar] [CrossRef]
- Wasternack, C.; Strnad, M. Jasmonates are signals in the biosynthesis of secondary metabolites—Pathways, transcription factors and applied aspects—A brief review. New Biotechnol. 2019, 48, 1–11. [Google Scholar] [CrossRef]
- Acosta, I.F.; Farmer, E.E. Jasmonates. Arabidopsis Book 2010, 8, e0129. [Google Scholar] [CrossRef]
- Feys, B.; Benedetti, C.E.; Penfold, C.N.; Turner, J.G. Arabidopsis mutants selected for resistance to the phytotoxin coronatine are male sterile, insensitive to methyl Jasmonate, and resistant to a bacterial pathogen. Plant Cell 1994, 6, 751–759. [Google Scholar] [CrossRef]
- Xie, D.; Feys, B.F.; James, S.; Nieto-Rostro, M.; Turner, J.G. COI1: An Arabidopsis gene required for Jasmonate-regulated defense and fertility. Science 1998, 280, 1091–1094. [Google Scholar] [CrossRef]
- Yan, J.; Zhang, C.; Gu, M.; Bai, Z.; Zhang, W.; Qi, T.; Cheng, Z.; Peng, W.; Luo, H.; Nan, F.; et al. The Arabidopsis CORONATINE INSENSITIVE1 protein is a Jasmonate receptor. Plant Cell 2009, 21, 2220–2236. [Google Scholar] [CrossRef]
- Sheard, L.B.; Tan, X.; Mao, H.; Withers, J.; Ben-Nissan, G.; Hinds, T.R.; Kobayashi, Y.; Hsu, F.; Sharon, M.; Browse, J.; et al. Jasmonate perception by inositol-phosphate-potentiated COI1-JAZ co-receptor. Nature 2010, 468, 400–405. [Google Scholar] [CrossRef]
- Monte, I.; Ishida, S.; Zamarreño, A.M.; Hamberg, M.; Franco-Zorrilla, J.M.; García-Casado, G.; Gouhier-Darimont, C.; Reymond, P.; Takahashi, K.; García-Mina, J.M.; et al. Ligand-Receptor co-evolution shaped the jasmonate pathway in land plants. Nat. Chem. Biol. 2018, 14, 480–488. [Google Scholar] [CrossRef]
- Wang, Z.; Dai, L.; Jiang, Z.; Peng, W.; Zhang, L.; Wang, G.; Xie, D. GmCOI1, a soybean F-box protein gene, shows ability to mediate jasmonate-regulated plant defense and fertility in Arabidopsis. Mol. Plant Microbe Interact. MPMI 2005, 18, 1285. [Google Scholar] [CrossRef]
- Liao, Y.; Wei, J.; Xu, Y.; Zhang, Z. Cloning, expression and characterization of COI1 gene (AsCOI1) from Aquilaria sinensis (Lour.) Gilg. Acta Pharm. Sin. B 2015, 5, 473–481. [Google Scholar] [CrossRef]
- Lee, H.Y.; Seo, J.; Cho, J.H.; Jung, H.; Kim, J.; Lee, J.S.; Rhee, S.; Do Choi, Y. Oryza sativa COI homologues restore jasmonate signal transduction in Arabidopsis coi1-1 mutants. PLoS ONE 2013, 8, e52802. [Google Scholar] [CrossRef]
- Lee, S.H.; Sakuraba, Y.; Lee, T.; Kim, K.W.; An, G.; Lee, H.Y.; Paek, N.C. Mutation of Oryza sativa CORONATINE INSENSITIVE 1b (OsCOI1b) delays leaf senescence. J. Integr. Plant Biol. 2015, 57, 562–576. [Google Scholar] [CrossRef]
- Schubert, R.; Grunewald, S.; von Sivers, L.; Hause, B. Effects of Jasmonate on ethylene function during the development of tomato stamens. Plants 2019, 8, 277. [Google Scholar] [CrossRef]
- Bai, J.; Wang, Y.; Wang, P.; Yuan, S.; Gao, J.; Duan, W.; Wang, N.; Zhang, F.; Zhang, W.; Qin, M.; et al. Genome-Wide identification and analysis of the COI gene family in wheat (Triticum aestivum L.). BMC Genom. 2018, 19, 754. [Google Scholar] [CrossRef]
- Tu, Y. The discovery of Artemisinin (qinghaosu) and gifts from Chinese medicine. Nat. Med. 2011, 17, 1217–1220. [Google Scholar] [CrossRef]
- Xiang, M.; Chen, Z.; He, L.; Xiong, G.; Lu, J. Transcription profiling of artemisinin-treated diabetic nephropathy rats using high-throughput sequencing. Life Sci. 2019, 219, 353–363. [Google Scholar] [CrossRef]
- Kumari, K.; Keshari, S.; Sengupta, D.; Sabat, S.C.; Mishra, S.K. Transcriptome analysis of genes associated with breast cancer cell motility in response to Artemisinin treatment. BMC Cancer 2017, 17, 858. [Google Scholar] [CrossRef]
- Wong, Y.K.; Xu, C.; Kalesh, K.A.; He, Y.; Lin, Q.; Wong, W.S.F.; Shen, H.; Wang, J. Artemisinin as an anticancer drug: Recent advances in target profiling and mechanisms of action. Med. Res. Rev. 2017, 37, 1492–1517. [Google Scholar] [CrossRef]
- Zhang, B. Artemisinin-Derived dimers as potential anticancer agents: Current developments, action mechanisms, and structure-activity relationships. Arch. Pharm. 2019, 353. [Google Scholar] [CrossRef] [PubMed]
- Larson, T.R.; Branigan, C.; Harvey, D.; Penfield, T.; Bowles, D.; Graham, I.A. A survey of artemisinic and dihydroartemisinic acid contents in glasshouse and global field-grown populations of the artemisinin-producing plant Artemisia annua L. Ind. Crops Prod. 2013, 45, 1–6. [Google Scholar] [CrossRef]
- Ma, D.M.; Wang, Z.; Wang, L.; Alejos-Gonzales, F.; Sun, M.A.; Xie, D.Y. A Genome-Wide scenario of terpene pathways in self-pollinated Artemisia annua. Mol. Plant 2015, 8, 1580–1598. [Google Scholar] [CrossRef] [PubMed]
- Shen, Q.; Zhang, L.; Liao, Z.; Wang, S.; Yan, T.; Shi, P.; Liu, M.; Fu, X.; Pan, Q.; Wang, Y.; et al. The genome of Artemisia annua provides insight into the evolution of Asteraceae family and Artemisinin biosynthesis. Mol. Plant 2018, 11, 776–788. [Google Scholar] [CrossRef] [PubMed]
- Xie, D.; Ma, D.; Judd, R.; Jones, A.L. Artemisinin biosynthesis in Artemisia annua and metabolic engineering: Questions, challenges, and perspectives. Phytochem. Rev. 2016, 15, 1093–1114. [Google Scholar] [CrossRef]
- Pu, G.; Ma, D.; Chen, J.; Ma, L.; Wang, H.; Li, G.; Ye, H.; Liu, B. Salicylic acid activates Artemisinin biosynthesis in Artemisia annua L. Plant Cell Rep. 2009, 28, 1127–1135. [Google Scholar] [CrossRef] [PubMed]
- Maes, L.; Van Nieuwerburgh, F.C.; Zhang, Y.; Reed, D.W.; Pollier, J.; Vande, C.S.; Inze, D.; Covello, P.S.; Deforce, D.L.; Goossens, A. Dissection of the phytohormonal regulation of trichome formation and biosynthesis of the antimalarial compound Artemisinin in Artemisia annua plants. New Phytol. 2011, 189, 176–189. [Google Scholar] [CrossRef]
- Banyai, W.; Mii, M.; Supaibulwatana, K. Enhancement of artemisinin content and biomass in Artemisia annua by exogenous GA3 treatment. Plant Growth Regul. 2011, 63, 45–54. [Google Scholar] [CrossRef]
- Guo, X.; Yang, X.; Yang, R.; Zeng, Q. Salicylic acid and methyl jasmonate but not Rose Bengal enhance Artemisinin production through invoking burst of endogenous singlet oxygen. Plant Sci. 2010, 178, 390–397. [Google Scholar] [CrossRef]
- Dangash, A.; Pandya, N.; Bharillya, A.; Jhala, A.; Jain, D. Impact of exogenous elicitors on Artemisinin production and trichome density in Artemisia annua L. under subtropical conditions. Not. Sci. Biol. 2014, 6. [Google Scholar] [CrossRef]
- Ikram, N.K.B.K.; Simonsen, H.T. A Review of Biotechnological Artemisinin Production in Plants. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Ma, C.; Li, Z.; Ma, L.; Wang, H.; Ye, H.; Xu, G.; Liu, B. Effects of exogenous methyl Jasmonate on Artemisinin biosynthesis and secondary metabolites in Artemisia annua L. Ind. Crops Prod. 2010, 31, 214–218. [Google Scholar] [CrossRef]
- Xiang, L.; Zhu, S.; Zhao, T.; Zhang, M.; Liu, W.; Chen, M.; Lan, X.; Liao, Z. Enhancement of Artemisinin content and relative expression of genes of artemisinin biosynthesis in Artemisia annua by exogenous MeJA treatment. Plant Growth Regul. 2015, 75, 435–441. [Google Scholar] [CrossRef]
- Hao, X.; Zhong, Y.; Fu, X.; Lv, Z.; Shen, Q.; Yan, T.; Shi, P.; Ma, Y.; Chen, M.; Lv, X.; et al. Transcriptome analysis of genes associated with the Artemisinin biosynthesis by Jasmonic acid treatment under the light in Artemisia annua. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Adie, B.A.T.; Pérez-Pérez, J.; Pérez-Pérez, M.M.; Godoy, M.; Sánchez-Serrano, J.; Schmelz, E.A.; Solano, R. ABA is an essential signal for plant resistance to pathogens affecting JA biosynthesis and the activation of defenses in Arabidopsis. Plant Cell 2007, 19, 1665–1681. [Google Scholar] [CrossRef] [PubMed]
- Ellis, C.; Karafyllidis, I.; Turner, J.G. Constitutive activation of jasmonate signaling in an Arabidopsis mutant correlates with enhanced resistance to Erysiphe cichoracearum, Pseudomonas syringae, and Myzus persicae. Mol. Plant Microbe Interact. MPMI 2002, 15, 1025. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Rombauts, S.; Dehais, P.; Van Montagu, M.; Rouze, P. PlantCARE, a plant cis-acting regulatory element database. Nucleic Acids Res. 1999, 27, 295–296. [Google Scholar] [CrossRef]
- Ishimaru, Y.; Hayashi, K.; Suzuki, T.; Fukaki, H.; Prusinska, J.; Meester, C.; Quareshy, M.; Egoshi, S.; Matsuura, H.; Takahashi, K.; et al. Jasmonic acid inhibits auxin-induced lateral rooting independently of the CORONATINE INSENSITIVE1 receptor. Plant Physiol. 2018, 177, 1704–1716. [Google Scholar] [CrossRef]
- Xu, F.; Copeland, C.; Li, X. Protein immunoprecipitation using Nicotiana benthamiana transient expression system. Bio Protoc. 2015, 5. [Google Scholar] [CrossRef]
- Xu, F.; Kapos, P.; Cheng, Y.T.; Li, M.; Zhang, Y.; Li, X. NLR-Associating transcription factor bHLH84 and its paralogs function redundantly in plant immunity. PLoS Pathog. 2014, 10, e1004312. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Wang, W.; Gao, J.; Yin, K.; Wang, R.; Wang, C.; Petersen, M.; Mundy, J.; Qiu, J. MYB75 phosphorylation by MPK4 Is required for light-induced Anthocyanin accumulation in Arabidopsis. Plant Cell 2016, 28, 2866–2883. [Google Scholar] [CrossRef] [PubMed]
- Winter, D.; Vinegar, B.; Nahal, H.; Ammar, R.; Wilson, G.V.; Provart, N.J. An “Electronic fluorescent pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2007, 2, e718. [Google Scholar] [CrossRef] [PubMed]
- Gou, J.; Felippes, F.F.; Liu, C.; Weigel, D.; Wang, J. Negative regulation of Anthocyanin biosynthesis in Arabidopsis by a miR156-targeted SPL transcription factor. Plant Cell 2011, 23, 1512–1522. [Google Scholar] [CrossRef]
- Gan, Y.; Li, H.; Xie, Y.; Wu, W.; Li, M.; Wang, X.; Huang, J. THF1 mutations lead to increased basal and wound-induced levels of oxylipins that stimulate anthocyanin biosynthesis via COI1 signaling inArabidopsis. J. Integr. Plant Biol. 2014, 56, 916–927. [Google Scholar] [CrossRef]
- Devoto, A.; Ellis, C.; Magusin, A.; Chang, H.; Chilcott, C.; Zhu, T.; Turner, J.G. Expression profiling reveals COI1 to be a key regulator of genesinvolved in wound-and methyl Jasmonate-induced secondary metabolism, defence, and hormone interactions. Plant Mol. Biol. 2005, 58, 497–513. [Google Scholar] [CrossRef]
- Ai, Y.; Zhu, Z. Melatonin antagonizes Jasmonate-triggered Anthocyanin biosynthesis in Arabidopsis thaliana. J. Agric. Food Chem. 2018, 66, 5392–5400. [Google Scholar] [CrossRef]
- Ma, Y.N.; Xu, D.B.; Li, L.; Zhang, F.; Fu, X.Q.; Shen, Q.; Lyu, X.Y.; Wu, Z.K.; Pan, Q.F.; Shi, P.; et al. Jasmonate promotes Artemisinin biosynthesis by activating the TCP14-ORA complex in Artemisia annua. Sci. Adv. 2018, 4, s9357. [Google Scholar] [CrossRef]
- Olofsson, L.; Engström, A.; Lundgren, A.; Brodelius, P.E. Relative expression of genes of terpene metabolism in different tissues of Artemisia annua L. BMC Plant Biol. 2011, 11, 45. [Google Scholar] [CrossRef] [PubMed]
- Kosugi, S.; Hasebe, M.; Tomita, M.; Yanagawa, H. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl. Acad. Sci. USA 2009, 106, 10171–10176. [Google Scholar] [CrossRef] [PubMed]
- Bej, A.; Sahoo, B.R.; Swain, B.; Basu, M.; Jayasankar, P.; Samanta, M. LRRsearch: An asynchronous server-based application for the prediction of leucine-rich repeat motifs and an integrative database of NOD-like receptors. Comput. Biol. Med. 2014, 53, 164–170. [Google Scholar] [CrossRef] [PubMed]
- Utsugi, S.; Sakamoto, W.; Murata, M.; Motoyoshi, F. Arabidopsis thaliana vegetative storage protein (VSP) genes: Gene organization and tissue-specific expression. Plant Mol. Biol. 1998, 38, 565–576. [Google Scholar] [CrossRef] [PubMed]
- Hiraoka, Y.; Kawamata, K.; Haraguchi, T.; Chikashige, Y. Codon usage bias is correlated with gene expression levels in the fission yeast Schizosaccharomyces pombe. Genes Cells 2009, 14, 499–509. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Lu, X.; Lv, Z.; Zhang, L.; Zhu, M.; Jiang, W.; Wang, G.; Sun, X.; Tang, K. Overexpression of the Artemisia orthologue of ABA receptor, AaPYL9, enhances ABA sensitivity and improves Artemisinin content in Artemisia annua L. PLoS ONE 2013, 8, e56697. [Google Scholar] [CrossRef]
- Winkel-Shirley, B. Biosynthesis of flavonoids and effects of stress. Curr. Opin. Plant Biol. 2002, 5, 218–223. [Google Scholar] [CrossRef]
- Gould, K.S. Nature’s swiss army knife: The diverse protective roles of Anthocyanins in leaves. J. Biomed. Biotechnol. 2004, 2004, 314–320. [Google Scholar] [CrossRef]
- Matsumoto, H.; Nakamura, Y.; Hirayama, M.; Yoshiki, Y.; Okubo, K. Antioxidant activity of black currant Anthocyanin aglycons and their glycosides measured by chemiluminescence in a neutral pH region and in human plasma. J. Agric. Food Chem. 2002, 50, 5034–5037. [Google Scholar] [CrossRef]
- Jing, P.; Bomser, J.A.; Schwartz, S.J.; He, J.; Magnuson, B.A.; Giusti, M.M. Structure-Function relationships of Anthocyanins from various Anthocyanin-rich extracts on the inhibition of colon cancer cell growth. J. Agric. Food Chem. 2008, 56, 9391–9398. [Google Scholar] [CrossRef]
- Hwang, J.; Kim, E.; Lee, S.; Kim, Y.; Moon, S.; Jeon, B.; Sung, S.; Kim, E.; Park, P. Antioxidant Activity and protective effect of Anthocyanin oligomers on H2O2-triggered G2/M arrest in retinal cells. J. Agric. Food Chem. 2012, 60, 4282–4288. [Google Scholar] [CrossRef] [PubMed]
- Taverniti, V.; Fracassetti, D.; Del Bo, C.; Lanti, C.; Minuzzo, M.; Klimis-Zacas, D.; Riso, P.; Guglielmetti, S. Immunomodulatory effect of a wild blueberry Anthocyanin-rich extract in human Caco-2 intestinal cells. J. Agric. Food Chem. 2014, 62, 8346–8351. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Liu, L.; Liu, Q.; Shen, Q.; Wang, C.; Yang, P.; Zhu, C.; Wang, Q. ZmMYC2 exhibits diverse functions and enhances JA signaling in transgenic Arabidopsis. Plant Cell Rep. 2019, 39, 1–16. [Google Scholar] [CrossRef] [PubMed]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, R.; Wang, J.; Xiao, M.; Gao, X.; Chen, J.; Dai, Y. AaCOI1, Encoding a CORONATINE INSENSITIVE 1-Like Protein of Artemisia annua L., Is Involved in Development, Defense, and Anthocyanin Synthesis. Genes 2020, 11, 221. https://doi.org/10.3390/genes11020221
Liu R, Wang J, Xiao M, Gao X, Chen J, Dai Y. AaCOI1, Encoding a CORONATINE INSENSITIVE 1-Like Protein of Artemisia annua L., Is Involved in Development, Defense, and Anthocyanin Synthesis. Genes. 2020; 11(2):221. https://doi.org/10.3390/genes11020221
Chicago/Turabian StyleLiu, Rong, Jinbiao Wang, Mu Xiao, Xiewang Gao, Jin Chen, and Yanjiao Dai. 2020. "AaCOI1, Encoding a CORONATINE INSENSITIVE 1-Like Protein of Artemisia annua L., Is Involved in Development, Defense, and Anthocyanin Synthesis" Genes 11, no. 2: 221. https://doi.org/10.3390/genes11020221
APA StyleLiu, R., Wang, J., Xiao, M., Gao, X., Chen, J., & Dai, Y. (2020). AaCOI1, Encoding a CORONATINE INSENSITIVE 1-Like Protein of Artemisia annua L., Is Involved in Development, Defense, and Anthocyanin Synthesis. Genes, 11(2), 221. https://doi.org/10.3390/genes11020221



