Hes5.9 Coordinate FGF and Notch Signaling to Modulate Gastrulation via Regulating Cell Fate Specification and Cell Migration in Xenopus tropicalis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Animal Ethics and Embryo Manipulation
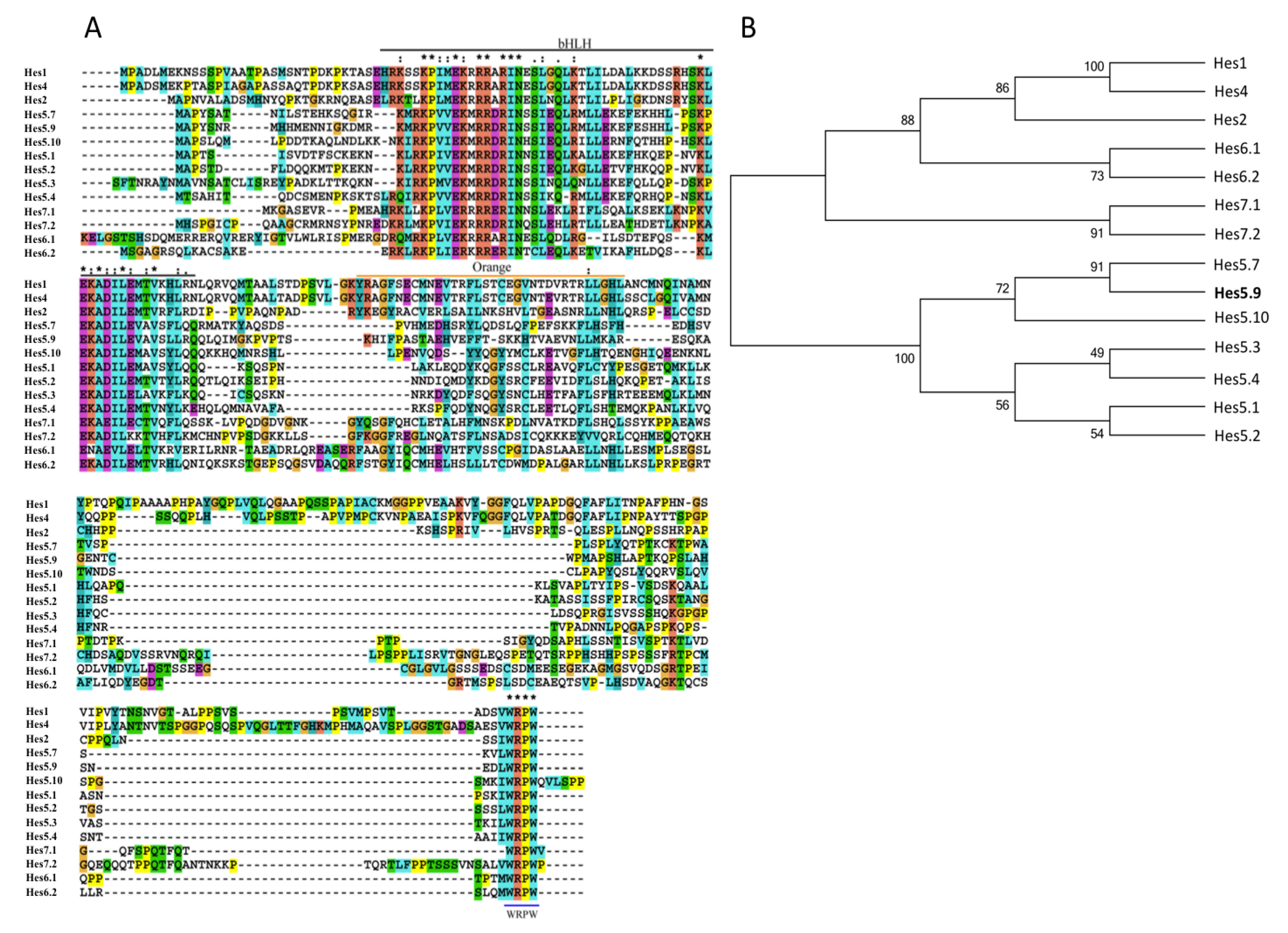
2.2. Multiple Sequence Alignment and Phylogenetic Tree Construction
2.3. RNA Extraction, Reverse Transcription-PCR, and cDNA Cloning
2.4. mRNA Synthesis and Microinjection
2.5. Quantitative Reverse Transcription PCR (RT-qPCR)
2.6. Whole-Mount in situ Hybridization
2.7. Animal Cap and Dorsal Marginal Zone (DMZ) Elongation Assays
2.8. RNA-Sequencing and Data Analyses
3. Results
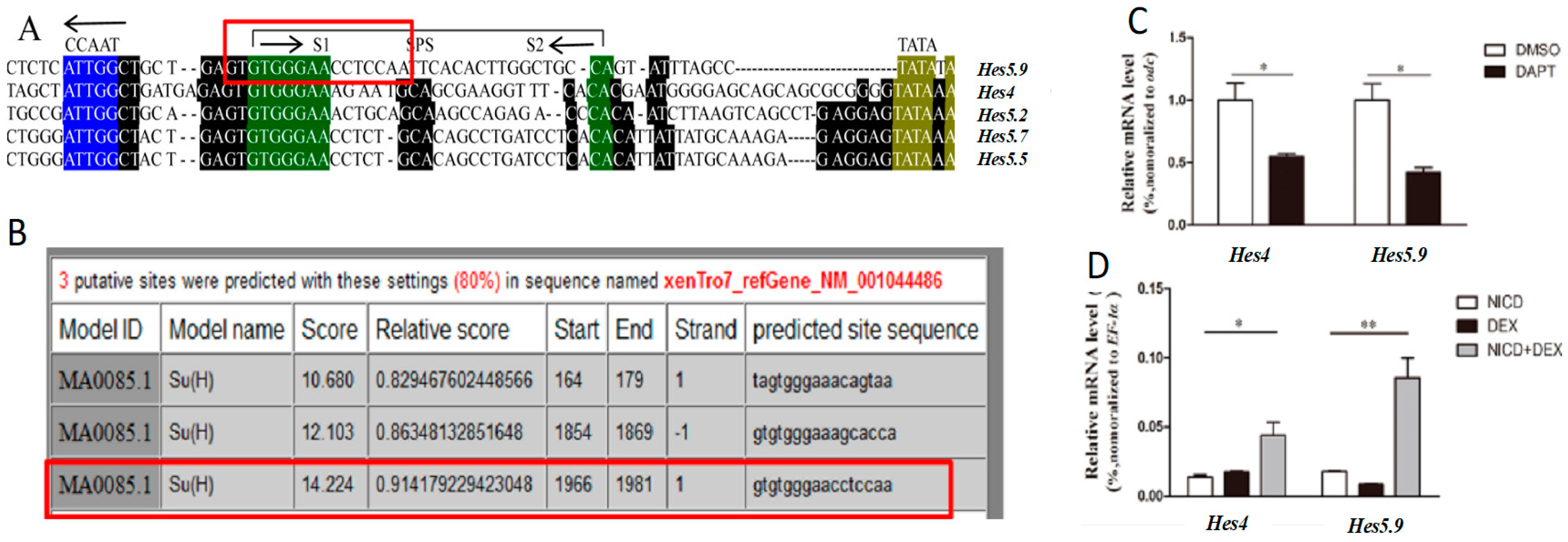
3.1. Hes5.9 is Regulated by the Notch Signaling
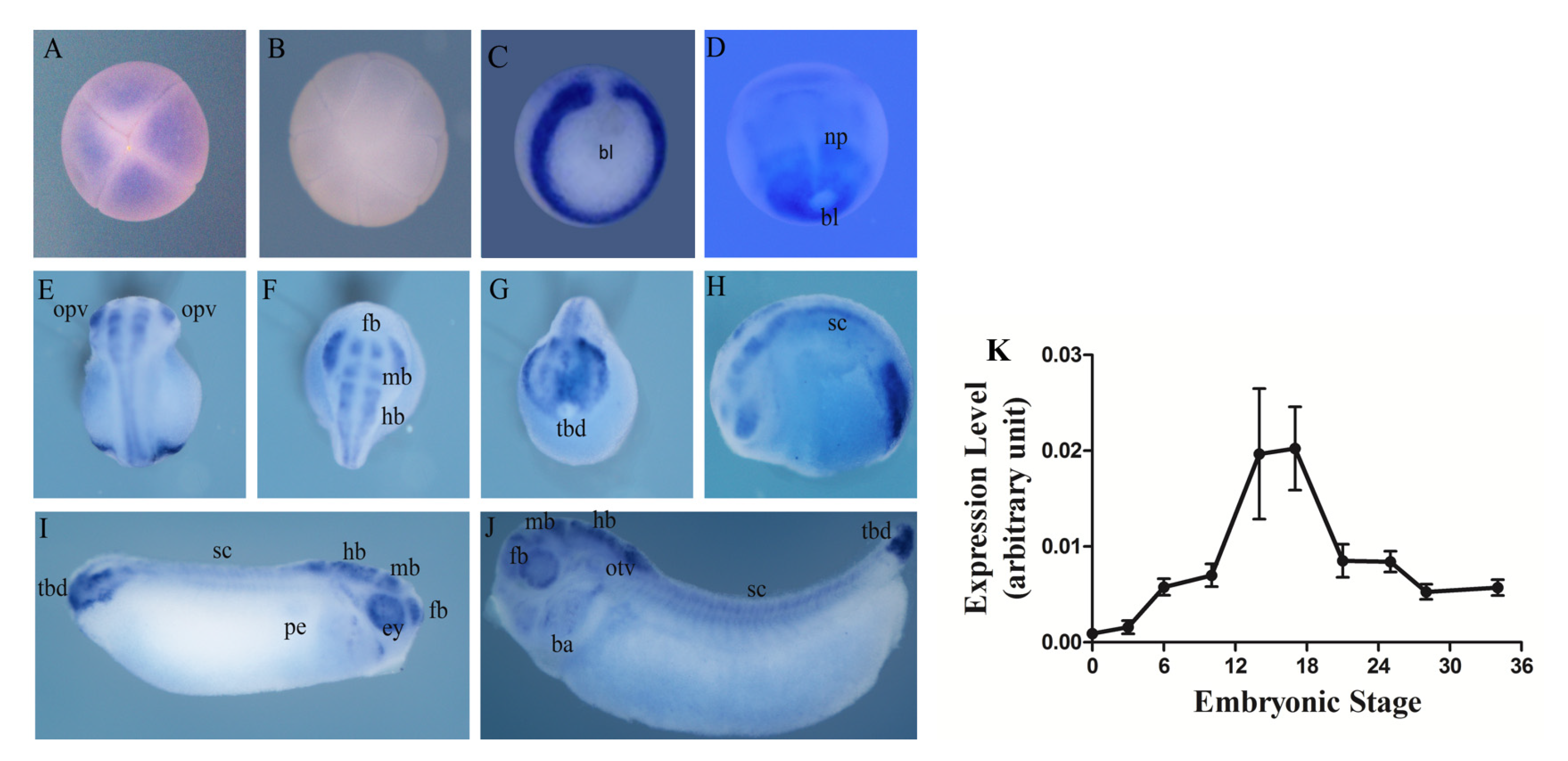
3.2. Spatiotemporal Expression Pattern of Hes5.9 in Embryonic Development
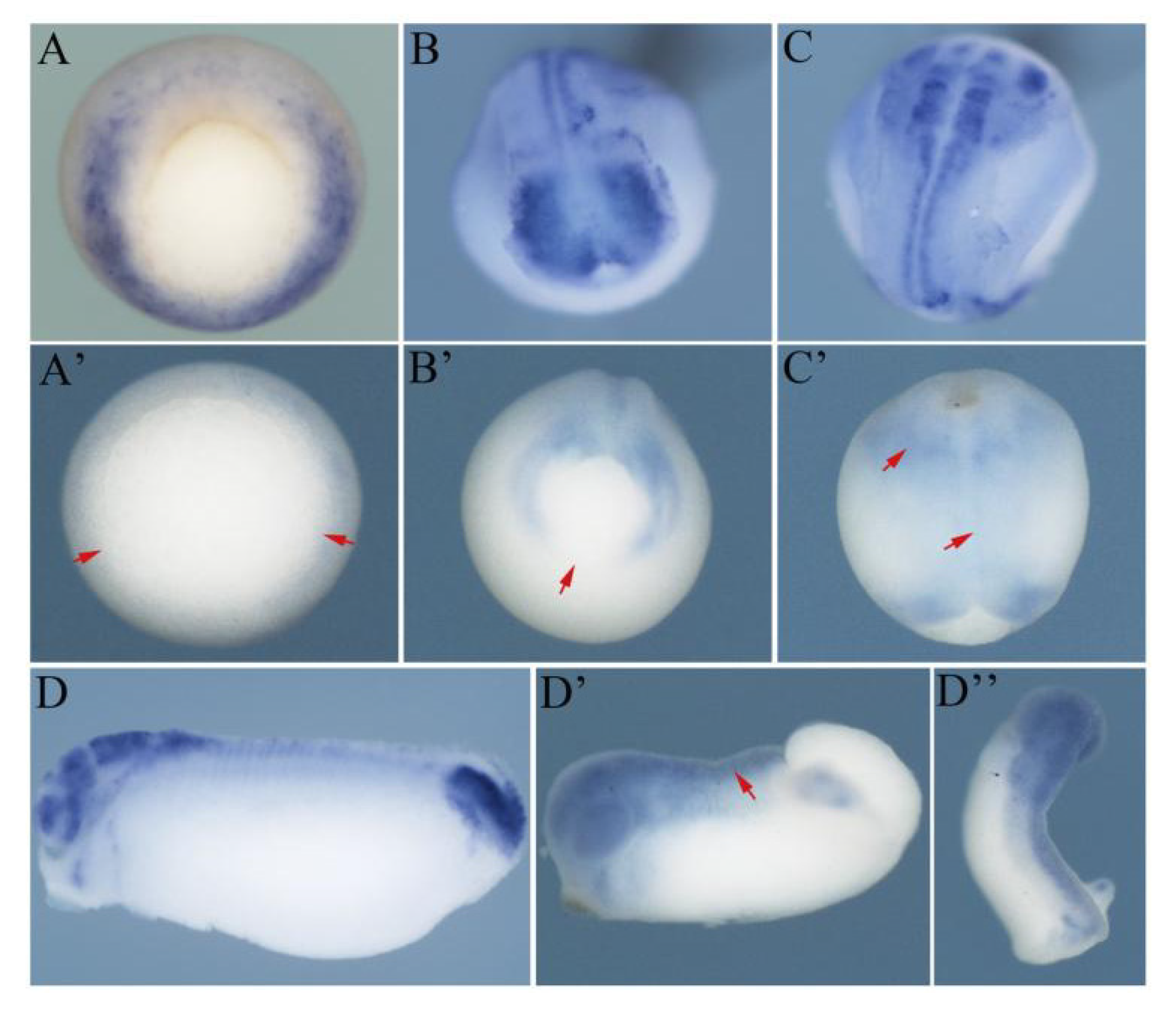
3.3. The FGF Signaling is Required for the Spatiotemporal Expression Pattern of Hes5.9 during Gastrulation
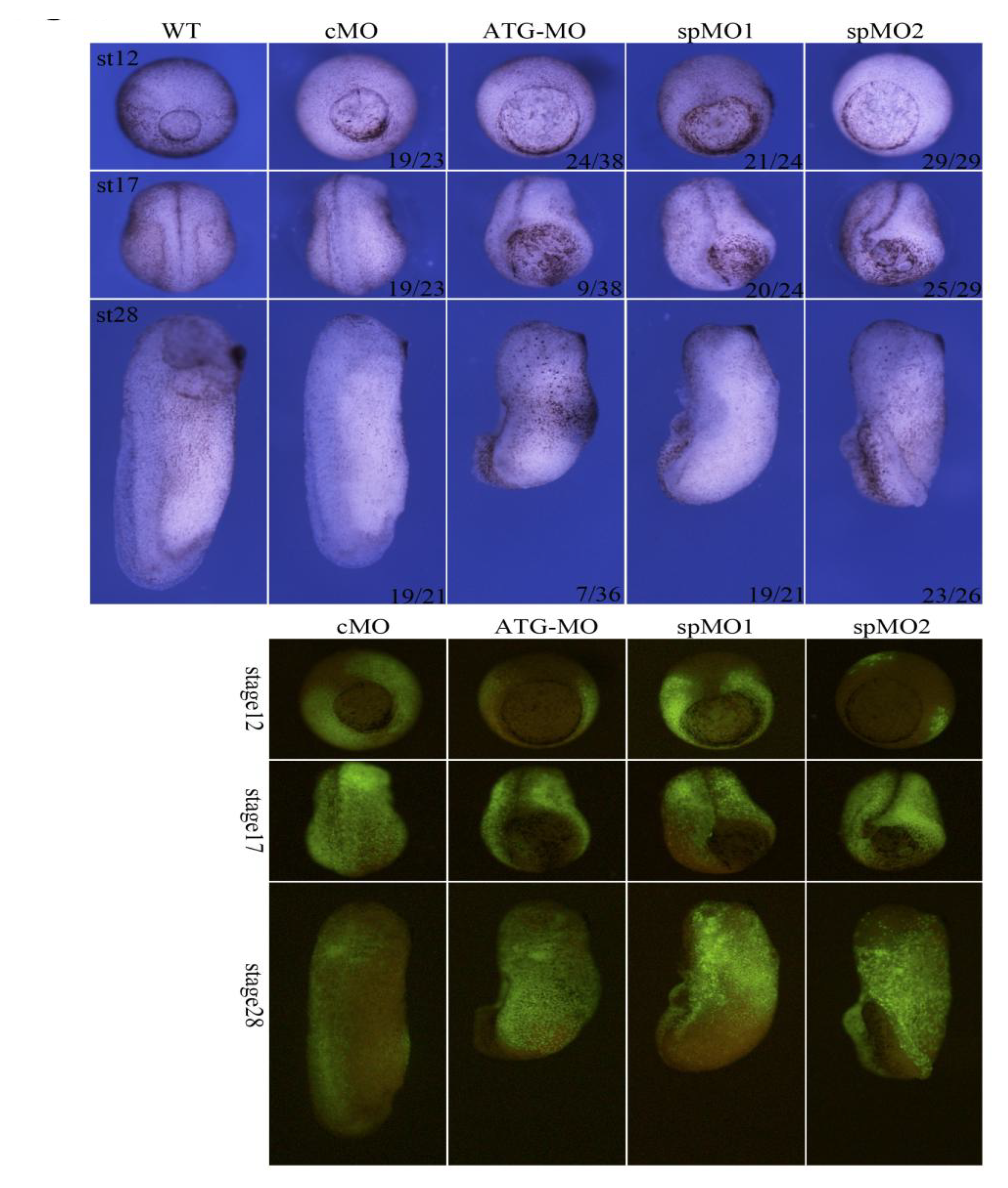
3.4. Knockdown of Hes5.9 Results in Defects on Gastrulation
3.5. Overexpression of Hes5.9 Causes Defects on Gastrulation
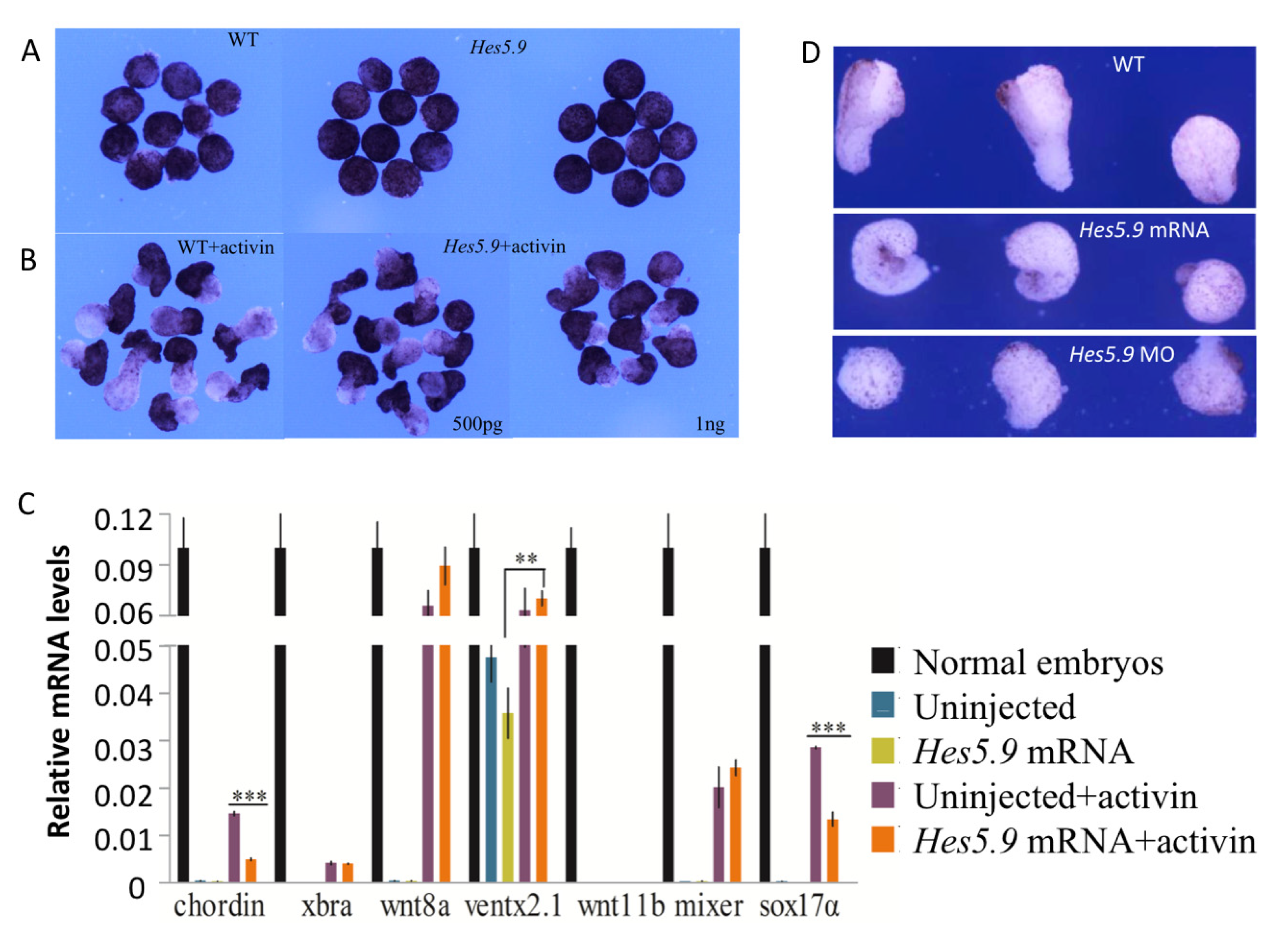
3.6. Hes5.9 Modulates the Elongation of Animal Cap and DMZ Explants
3.7. Transcriptomics Analysis of Hes5.9 Ectopic Expression in Late Gastrula Stage
4. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Kiecker, C.; Bates, T.; Bell, E. Molecular specification of germ layers in vertebrate embryos. Cell. Mol. Life Sci. 2015, 73, 923–947. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Revinski, D.R.; Paganelli, A.R.; Carrasco, A.E.; López, S.L. Delta-Notch signaling is involved in the segregation of the three germ layers in Xenopus laevis. Dev. Biol. 2010, 339, 477–492. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Solnica-Krezel, L.; Sepich, D.S. Gastrulation: Making and Shaping Germ Layers. Annu. Rev. Cell Dev. Biol. 2012, 28, 687–717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takeuchi, M.; Nakabayashi, J.; Sakaguchi, T.; Yamamoto, T.S.; Takahashi, H.; Takeda, H.; Ueno, N. The prickle-Related Gene in Vertebrates Is Essential for Gastrulation Cell Movements. Curr. Biol. 2003, 13, 674–679. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goto, T.; Davidson, L.; Asashima, M.; Keller, R. Planar Cell Polarity Genes Regulate Polarized Extracellular Matrix Deposition during Frog Gastrulation. Curr. Biol. 2005, 15, 787–793. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tada, M.; Heisenberg, C.-P. Convergent extension: Using collective cell migration and cell intercalation to shape embryos. Development 2012, 139, 3897–3904. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dorey, K.; Amaya, E. FGF signalling: Diverse roles during early vertebrate embryogenesis. Development 2010, 137, 3731–3742. [Google Scholar] [CrossRef] [Green Version]
- Böttcher, R.T.; Niehrs, C. Fibroblast Growth Factor Signaling during Early Vertebrate Development. Endocr. Rev. 2004, 26, 63–77. [Google Scholar] [CrossRef]
- Itoh, N. The Fgf Families in Humans, Mice, and Zebrafish: Their Evolutional Processes and Roles in Development, Metabolism, and Disease. Biol. Pharm. Bull. 2007, 30, 1819–1825. [Google Scholar] [CrossRef] [Green Version]
- Sivak, J.M.; Petersen, L.F.; Amaya, E. FGF Signal Interpretation Is Directed by Sprouty and Spred Proteins during Mesoderm Formation. Dev. Cell 2005, 8, 689–701. [Google Scholar] [CrossRef]
- Shi, S.; Stanley, P. Evolutionary Origins of Notch Signaling in Early Development. Cell Cycle 2006, 5, 274–278. [Google Scholar] [CrossRef] [PubMed]
- Hori, K.; Sen, A.; Artavanis-Tsakonas, S. Notch signaling at a glance. J. Cell Sci. 2013, 126, 2135–2140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ohtsuka, T.; Sakamoto, M.; Guillemot, F.; Kageyama, R. Roles of the Basic Helix-Loop-Helix GenesHes1andHes5in Expansion of Neural Stem Cells of the Developing Brain. J. Biol. Chem. 2001, 276, 30467–30474. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hong, C.-S.; Saint-Jeannet, J.-P. The b-HLH transcription factor Hes3 participates in neural plate border formation by interfering with Wnt/β-catenin signaling. Dev. Biol. 2018, 442, 162–172. [Google Scholar] [CrossRef] [PubMed]
- Glavic, A.; Silva, F.; Aybar, M.J.; Bastidas, F.; Mayor, R. Interplay between Notch signaling and the homeoprotein Xiro1 is required for neural crest induction in Xenopus embryos. Development (Camb. Engl.) 2003, 131, 347–359. [Google Scholar] [CrossRef] [Green Version]
- Coffman, C.; Harris, W.; Kintner, C. Xotch, the Xenopus homolog of Drosophila notch. Science 1990, 249, 1438–1441. [Google Scholar] [CrossRef]
- Ito, M.; Katada, T.; Miyatani, S.; Kinoshita, T. XSu(H)2 is an essential factor for gene expression and morphogenesis of the Xenopus gastrula embryo. Int. J. Dev. Biol. 2007, 51, 27–36. [Google Scholar] [CrossRef]
- Ma, Q.; Kintner, C.; Anderson, D.J. Identification of neurogenin, a Vertebrate Neuronal Determination Gene. Cell 1996, 87, 43–52. [Google Scholar] [CrossRef] [Green Version]
- López, S.L.; Paganelli, A.R.; Siri, M.V.R.; Ocaña, O.H.; Franco, P.; Carrasco, A.E. Notch activates sonic hedgehog and both are involved in the specification of dorsal midline cell-fates in Xenopus. Development 2003, 130, 2225–2238. [Google Scholar] [CrossRef] [Green Version]
- Shook, D.R.; Keller, R. Mechanisms, mechanics and function of epithelial–mesenchymal transitions in early development. Mech. Dev. 2003, 120, 1351–1383. [Google Scholar] [CrossRef]
- Favarolo, M.B.; López, S.L. Notch signaling in the division of germ layers in bilaterian embryos. Mech. Dev. 2018, 154, 122–144. [Google Scholar] [CrossRef] [PubMed]
- Modrell, M.S.; Tidswell, O.R.; Baker, C.V.H. Notch and Fgf signaling during electrosensory versus mechanosensory lateral line organ development in a non-teleost ray-finned fish. Dev. Biol. 2017, 431, 48–58. [Google Scholar] [CrossRef] [PubMed]
- Lloyd-Lewis, B.; Mourikis, P.; Fre, S. Notch signalling: Sensor and instructor of the microenvironment to coordinate cell fate and organ morphogenesis. Curr. Opin. Cell Biol. 2019, 61, 16–23. [Google Scholar] [CrossRef] [PubMed]
- Kawamura, A.; Koshida, S.; Hijikata, H.; Sakaguchi, T.; Kondoh, H.; Takada, S. Zebrafish Hairy/Enhancer of split protein links FGF signaling to cyclic gene expression in the periodic segmentation of somites. Genes Dev. 2005, 19, 1156–1161. [Google Scholar] [CrossRef] [Green Version]
- Doetzlhofer, A.; Basch, M.L.; Ohyama, T.; Gessler, M.; Groves, A.K.; Segil, N. Hey2 Regulation by FGF Provides a Notch-Independent Mechanism for Maintaining Pillar Cell Fate in the Organ of Corti. Dev. Cell 2009, 16, 58–69. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Voelkel, J.E.; Harvey, J.A.; Adams, J.S.; Lassiter, R.N.; Stark, M.R. FGF and Notch signaling in sensory neuron formation: A multifactorial approach to understanding signaling pathway hierarchy. Mech. Dev. 2014, 134, 55–66. [Google Scholar] [CrossRef]
- Davis, R.L.; Turner, D.L.; Evans, L.M.; Kirschner, M.W. Molecular Targets of Vertebrate Segmentation: Two mechanisms control segmental expression of xenopus hairy2 during somite formation. Dev. Cell 2001, 1, 553–565. [Google Scholar] [CrossRef] [Green Version]
- Lamar, E.; Kintner, C. The Notch targets Esr1 and Esr10 are differentially regulated in Xenopus neural precursors. Development 2005, 132, 3619–3630. [Google Scholar] [CrossRef] [Green Version]
- Davis, R.L.; Turner, D.L. Vertebrate hairy and Enhancer of split related proteins: Transcriptional repressors regulating cellular differentiation and embryonic patterning. Oncogene 2001, 20, 8342–8357. [Google Scholar] [CrossRef]
- Hopwood, N.; Pluck, A.; Gurdon, J. Myod expression in the forming somites is an early response to mesoderm induction in xenopus embryos. EMBO J. 1989, 8, 3409. [Google Scholar] [CrossRef]
- Groves, J.A.; Hammond, C.L.; Hughes, S.M. Fgf8 drives myogenic progression of a novel lateral fast muscle fibre population in zebrafish. Development. 2005, 132, 4211–4222. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stylianou, P.; Skourides, P.A. Imaging morphogenesis, in Xenopus with Quantum Dot nanocrystals. Mech. Dev. 2009, 126, 828–841. [Google Scholar] [CrossRef] [PubMed]
- Wallingford, J.B.; Fraser, S.E.; Harland, R.M. Convergent extension: The molecular control of polarized cell movement during embryonic development. Dev. Cell 2002, 2, 695–706. [Google Scholar] [CrossRef]
- Wallingford, J.B. Planar Cell Polarity and the Developmental Control of Cell Behavior in Vertebrate Embryos. Annu. Rev. Cell Dev. Biol. 2012, 28, 627–653. [Google Scholar] [CrossRef] [PubMed]
- Harima, Y.; Imayoshi, I.; Shimojo, H.; Kobayashi, T.; Kageyama, R. The roles and mechanism of ultradian oscillatory expression of the mouse hes genes. Semin. Cell Dev. Biol. 2014, 34, 85–90. [Google Scholar] [CrossRef]
- Takada, H.; Hattori, D.; Kitayama, A.; Ueno, N.; Taira, M. Identification of target genes for the Xenopus Hes-related protein XHR1, a prepattern factor specifying the midbrain–hindbrain boundary. Dev. Biol. 2005, 283, 253–267. [Google Scholar] [CrossRef] [Green Version]
- Watanabe, M.; Yasuoka, Y.; Mawaribuchi, S.; Kuretani, A.; Ito, M.; Kondo, M.; Ochi, H.; Ogino, H.; Fukui, A.; Taira, M.; et al. Conservatism and variability of gene expression profiles among homeologous transcription factors in Xenopus laevis. Dev. Biol. 2017, 426, 301–324. [Google Scholar] [CrossRef]
- Fletcher, R.B.; Harland, R.M. The role of FGF signaling in the establishment and maintenance of mesodermal gene expression inXenopus. Dev. Dyn. 2008, 237, 1243–1254. [Google Scholar] [CrossRef] [Green Version]
- Wahi, K.; Bochter, M.S.; Cole, S.E. The many roles of Notch signaling during vertebrate somitogenesis. Semin. Cell Dev. Biol. 2016, 49, 68–75. [Google Scholar] [CrossRef]
- Chung, H.A.; Hyodo-Miura, J.; Kitayama, A.; Terasaka, C.; Nagamune, T.; Ueno, N. Screening of FGF target genes in Xenopusby microarray: Temporal dissection of the signalling pathway using a chemical inhibitor. Genes Cells Devoted Mol. Cell. Mech. 2004, 9, 749–761. [Google Scholar] [CrossRef]
- Dubrulle, J.; Pourquié, O. Fgf8 mRNA decay establishes a gradient that couples axial elongation to patterning in the vertebrate embryo. Nature 2004, 427, 419–422. [Google Scholar] [CrossRef] [PubMed]
- Shi, W.; Peyrot, S.M.; Munro, E.; Levine, M. FGF3 in the floor plate directs notochord convergent extension in the Ciona tadpole. Development 2008, 136, 23–28. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chung, H.A.; Hyodo-Miura, J.; Nagamune, T.; Ueno, N. FGF signal regulates gastrulation cell movements and morphology through its target NRH. Dev. Biol. 2005, 282, 95–110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bae, Y.-K.; Trisnadi, N.; Kadam, S.; Stathopoulos, A. The role of FGF signaling in guiding coordinate movement of cell groups: Guidance cue and cell adhesion regulator? Cell Adhes. Migr. 2012, 6, 397–403. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Perris, R.; Perissinotto, D. Role of the extracellular matrix during neural crest cell migration. Mech. Dev. 2000, 95, 3–21. [Google Scholar] [CrossRef]
- Vega-Lopez, G.; Bonano, M.; Tríbulo, C.; Fernández, J.P.; Agüero, T.H.; Aybar, M.J. Functional analysis ofHairygenes inXenopusneural crest initial specification and cell migration. Dev. Dyn. 2015, 244, 988–1013. [Google Scholar] [CrossRef]
- Colabianchi, A.M.C.; Revinski, D.R.; Encinas, P.; Baez, M.V.; Monti, R.J.; Abinal, M.R.; Kodjabachian, L.; Franchini, L.F.; López, S.L. Notch1 is asymmetrically distributed from the beginning of embryogenesis and controls the ventral center. Development 2018, 145, dev159368. [Google Scholar] [CrossRef] [Green Version]
- Yokota, C.; Kofron, M.; Zuck, M.; Houston, D.W.; Isaacs, H.; Asashima, M.; Wylie, C.; Heasman, J. A novel role for a nodal-related protein; Xnr3 regulates convergent extension movements via the FGF receptor. Development 2003, 130, 2199–2212. [Google Scholar] [CrossRef] [Green Version]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, X.; Zhang, L.; Yang, S.; Zhang, Y.; Wu, M.; Chen, P. Hes5.9 Coordinate FGF and Notch Signaling to Modulate Gastrulation via Regulating Cell Fate Specification and Cell Migration in Xenopus tropicalis. Genes 2020, 11, 1363. https://doi.org/10.3390/genes11111363
Huang X, Zhang L, Yang S, Zhang Y, Wu M, Chen P. Hes5.9 Coordinate FGF and Notch Signaling to Modulate Gastrulation via Regulating Cell Fate Specification and Cell Migration in Xenopus tropicalis. Genes. 2020; 11(11):1363. https://doi.org/10.3390/genes11111363
Chicago/Turabian StyleHuang, Xiao, Liyue Zhang, Shanshan Yang, Yongpu Zhang, Mingjiang Wu, and Peichao Chen. 2020. "Hes5.9 Coordinate FGF and Notch Signaling to Modulate Gastrulation via Regulating Cell Fate Specification and Cell Migration in Xenopus tropicalis" Genes 11, no. 11: 1363. https://doi.org/10.3390/genes11111363
APA StyleHuang, X., Zhang, L., Yang, S., Zhang, Y., Wu, M., & Chen, P. (2020). Hes5.9 Coordinate FGF and Notch Signaling to Modulate Gastrulation via Regulating Cell Fate Specification and Cell Migration in Xenopus tropicalis. Genes, 11(11), 1363. https://doi.org/10.3390/genes11111363




