Diverse Horizontally-Acquired Gene Clusters Confer Sucrose Utilization to Different Lineages of the Marine Pathogen Photobacterium damselae subsp. damselae
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains, Plasmids, and Culture Conditions
2.2. PCR Assays
2.3. Genome Sequencing
2.4. Construction of a scrA Mutant in Pdd DK32
2.5. Growth Assays with Sucrose as a Carbon Source
2.6. Construction of a Transcriptional Fusion of scrA Promoter to a lacZ Reporter Gene, and β-Galactosidase Assays
2.7. Comparative Genomics and Molecular Phylogeny Analyses
2.8. Database Submission
3. Results
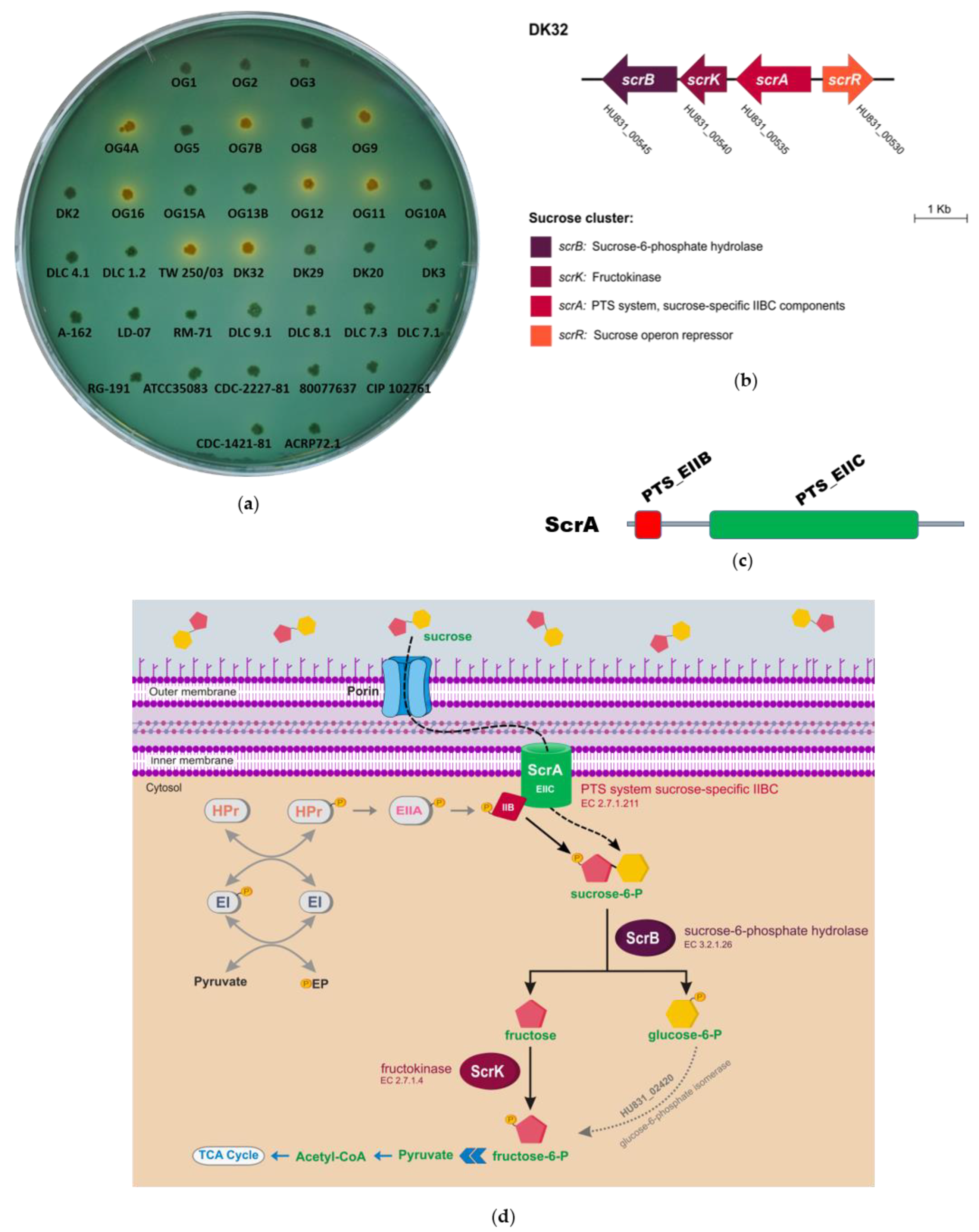
3.1. Identification of Two Different Genetic Variants of a Four Gene Cluster Encoding Functions for Sucrose Uptake and Catabolism in Pdd Strains
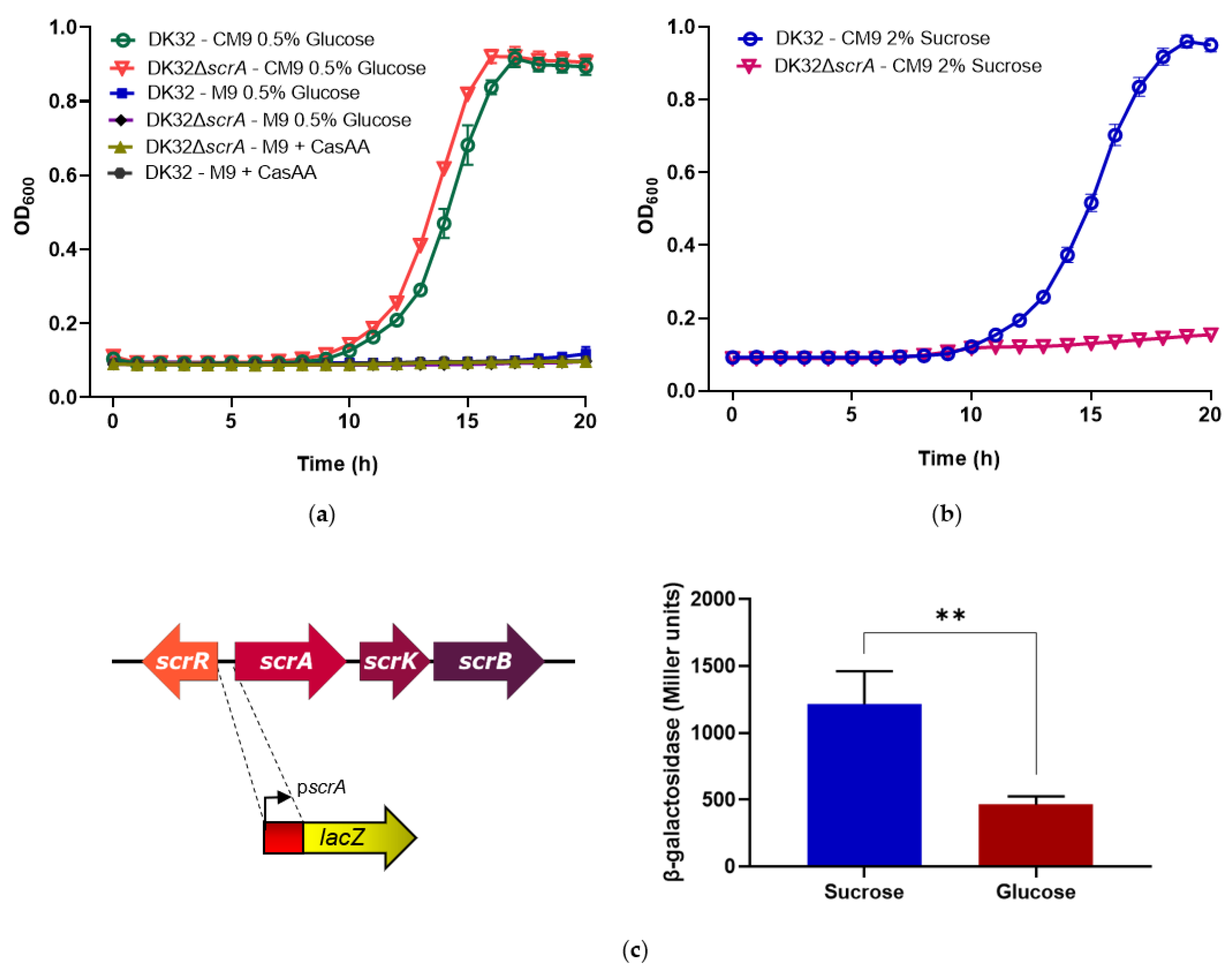
3.2. Deletion of scrA in the scr+ Strain DK32 Causes Green Colonies on TCBS Medium, and Abolishes Growth with Sucrose as Carbon Source
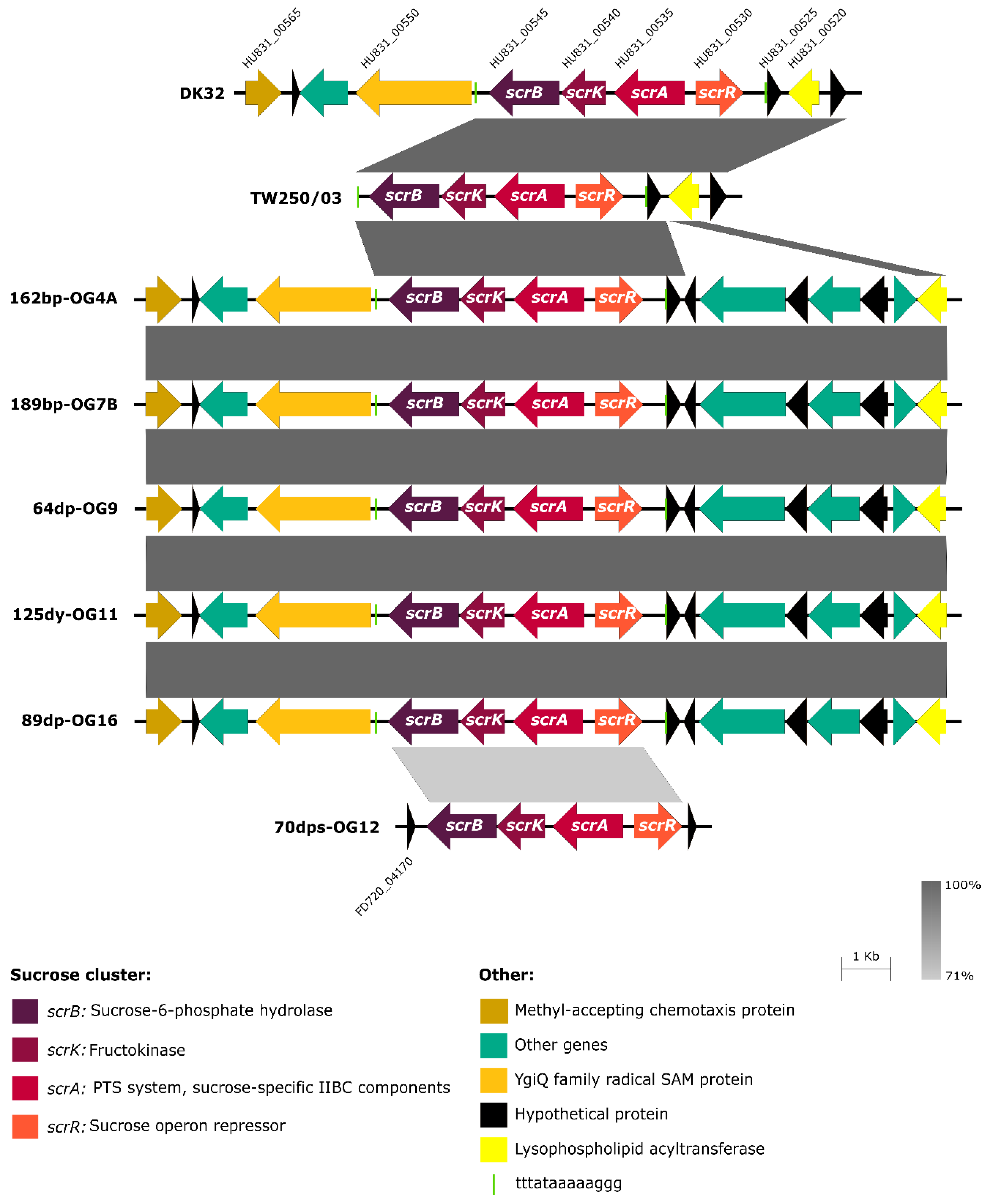
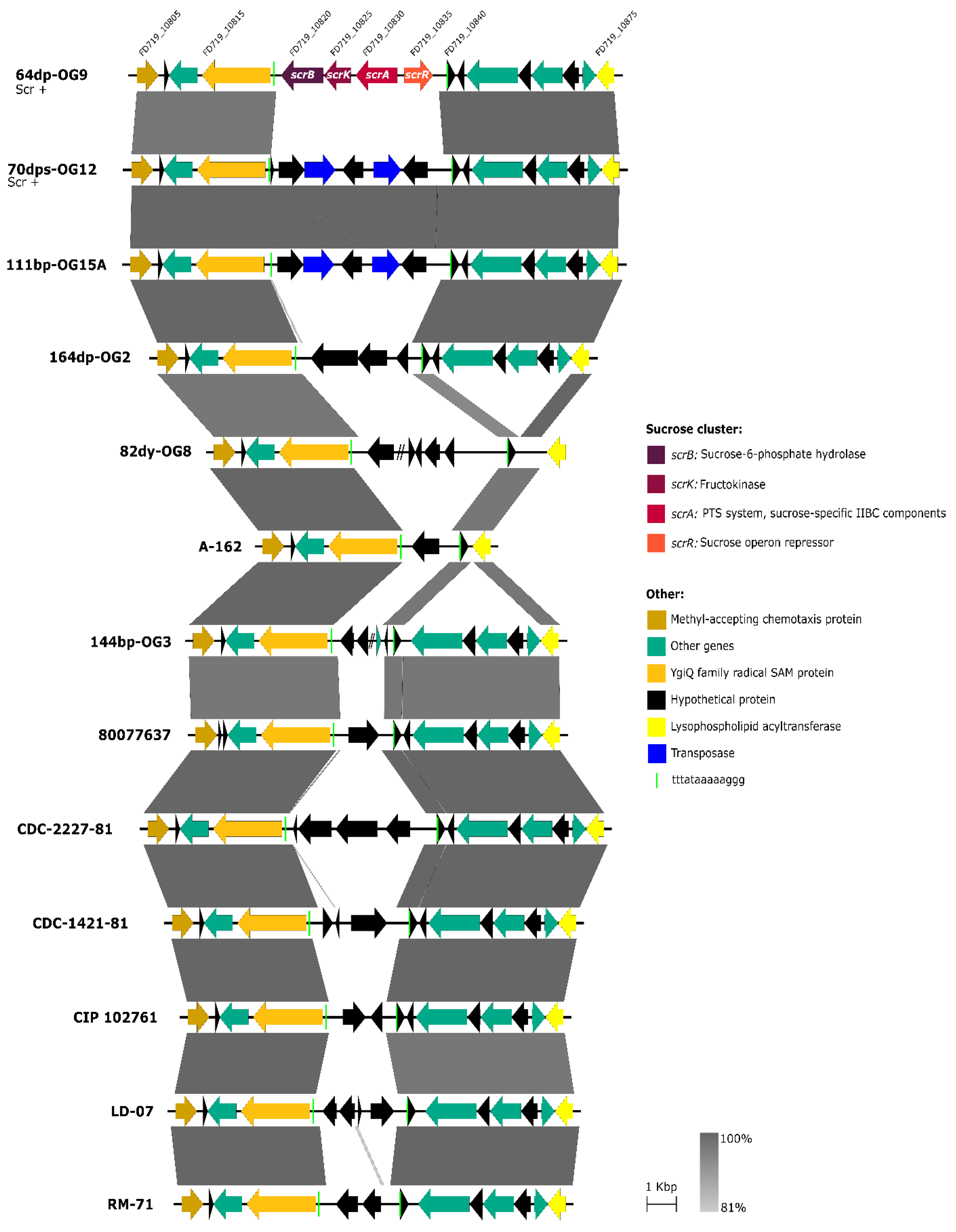
3.3. Scr Clusters Are Inserted into Putative Hot-Spots for DNA Acquisition in Pdd Genomes
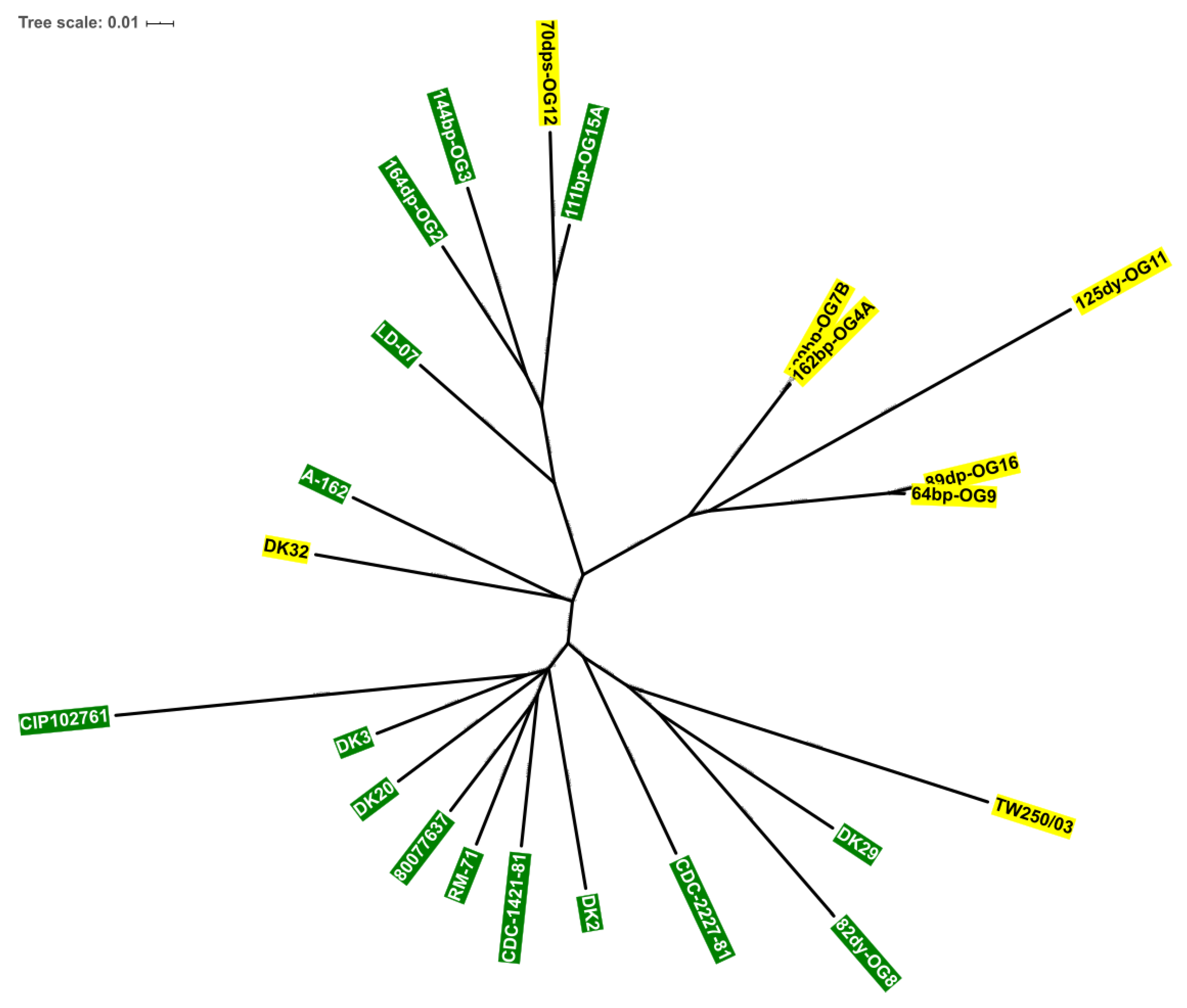
3.4. Scr Clusters Occur in Different Genetic Lineages of Pdd
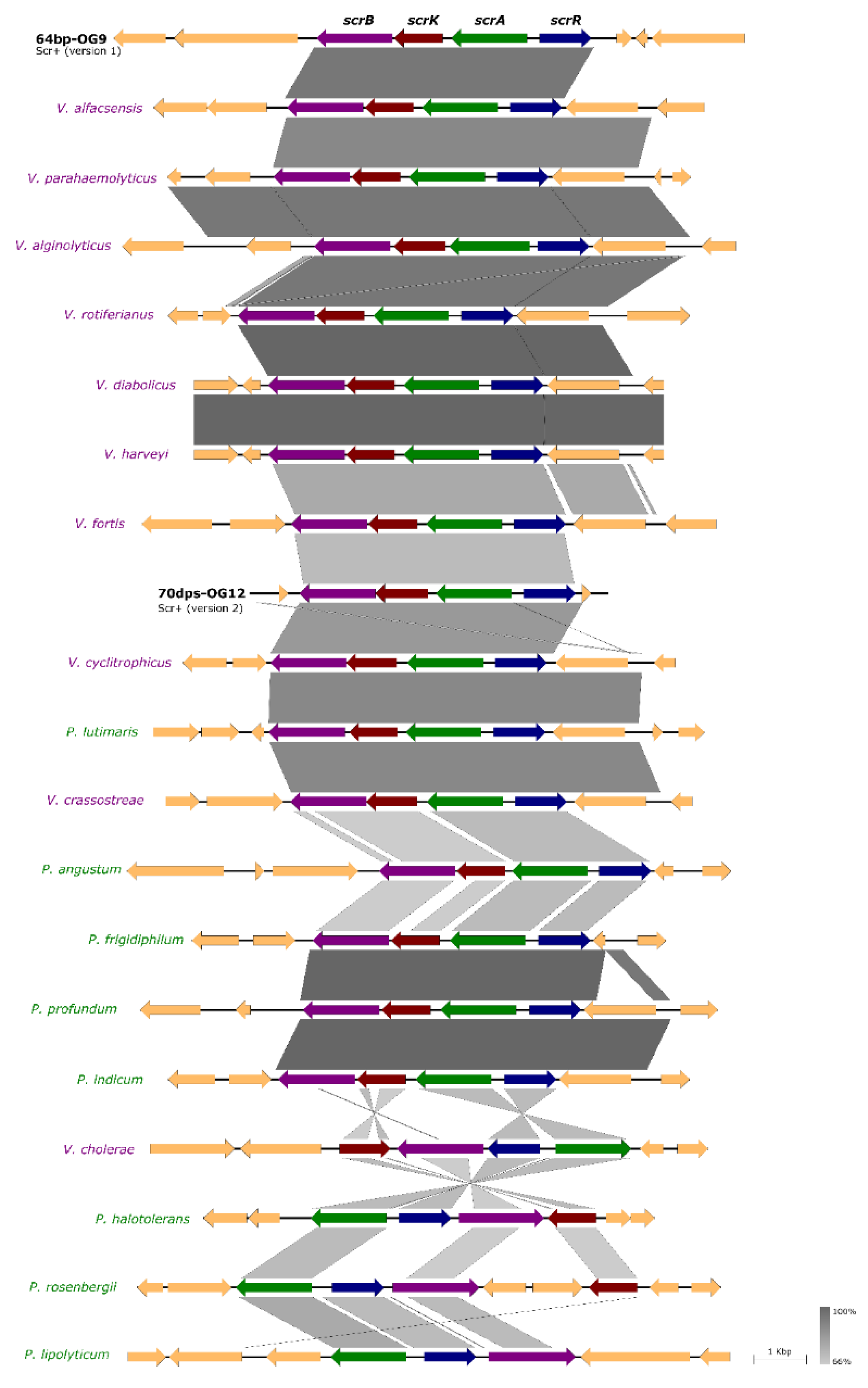
3.5. Incongruences between the Species Tree and the Sucrose Genes Tree Reveal Extensive Horizontal Transfer of scr Genes among Species of Vibrio and Photobacterium
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Thompson, F.L.; Iida, T.; Swings, J. Biodiversity of vibrios. Microbiol. Mol. Biol. Rev. 2004, 68, 403–431. [Google Scholar] [CrossRef]
- Baker-Austin, C.; Oliver, J.D.; Alam, M.; Ali, A.; Waldor, M.K.; Qadri, F.; Martinez-Urtaza, J. Vibrio spp. infections. Nat. Rev. Dis. Prim. 2018, 4, 8. [Google Scholar] [CrossRef] [PubMed]
- Takemura, A.F.; Chien, D.M.; Polz, M.F. Associations and dynamics of Vibrionaceae in the environment, from the genus to the population level. Front. Microbiol. 2014, 5, 38. [Google Scholar] [CrossRef] [PubMed]
- Reen, F.J.; Almagro-Moreno, S.; Ussery, D.; Boyd, E.F. The genomic code: Inferring Vibrionaceae niche specialization. Nat. Rev. Microbiol. 2006, 4, 697–704. [Google Scholar] [CrossRef] [PubMed]
- Thompson, C.C.; Vicente, A.C.P.; Souza, R.C.; Vasconcelos, A.T.R.; Vesth, T.; Alves, N.; Ussery, D.W.; Iida, T.; Thompson, F.L. Genomic taxonomy of vibrios. BMC Evol. Biol. 2009, 9, 258. [Google Scholar] [CrossRef]
- Urbanczyk, H.; Ast, J.C.; Dunlap, P.V. Phylogeny, genomics, and symbiosis of Photobacterium. FEMS Microbiol. Rev. 2011, 35, 324–342. [Google Scholar] [CrossRef]
- Le Roux, F.; Wegner, K.M.; Baker-Austin, C.; Vezzulli, L.; Osorio, C.R.; Amaro, C.; Ritchie, J.M.; Defoirdt, T.; Destoumieux-Garzón, D.; Blokesch, M.; et al. The emergence of Vibrio pathogens in Europe: Ecology, evolution, and pathogenesis (Paris, 11–12 March 2015). Front. Microbiol. 2015, 6, 830. [Google Scholar]
- Hehemann, J.-H.; Arevalo, P.; Datta, M.S.; Yu, X.; Corzett, C.H.; Henschel, A.; Preheim, S.P.; Timberlake, S.; Alm, E.J.; Polz, M.F. Adaptive radiation by waves of gene transfer leads to fine-scale resource partitioning in marine microbes. Nat. Commun. 2016, 7, 1–10. [Google Scholar] [CrossRef]
- Machado, H.; Gram, L. Comparative genomics reveals high genomic diversity in the genus Photobacterium. Front. Microbiol. 2017, 8, 1204. [Google Scholar] [CrossRef]
- Le Roux, F.; Blokesch, M. Eco-evolutionary dynamics linked to horizontal gene transfer in vibrios. Annu. Rev. Microbiol. 2018. [Google Scholar] [CrossRef]
- Regmi, A.; Boyd, E.F. Carbohydrate metabolic systems present on genomic islands are lost and gained in Vibrio parahaemolyticus. BMC Microbiol. 2019, 19, 112. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Enomoto, S.; Sakazaki, R.; Kuwahara, S. A new selective isolation medium for pathogenic vibrios: TCBS agar. Jap. J. Bact. 1963, 18, 387–391. [Google Scholar] [CrossRef]
- Scholle, R.R.; Coyne, V.E.; Maharaj, R.; Robb, F.T.; Woods, D.R. Expression and regulation of a Vibrio alginolyticus sucrose utilization system cloned in Escherichia coli. J. Bacteriol. 1987, 169, 2685–2690. [Google Scholar] [CrossRef]
- Scholle, R.R.; Robb, S.M.; Robb, F.T.; Woods, D.R. Nucleotide sequence and analysis of the Vibrio alginolyticus sucrase gene (scrB). Gene 1989, 80, 49–56. [Google Scholar] [CrossRef]
- Blatch, G.L.; Scholle, R.R.; Woods, D.R. Nucleotide sequence and analysis of the Vibrio alginolyticus sucrose uptake-encoding region. Gene 1990, 95, 17–23. [Google Scholar] [CrossRef]
- Blatch, G.L.; Woods, D.R. Nucleotide sequence and analysis of the Vibrio alginolyticus scr repressor-encoding gene (scrR). Gene 1991, 101, 45–50. [Google Scholar] [CrossRef]
- Houot, L.; Chang, S.; Absalon, C.; Watnick, P.I. Vibrio cholerae phosphoenolpyruvate phosphotransferase system control of carbohydrate transport, biofilm formation, and colonization of the germfree mouse intestine. Infect. Immun. 2010, 78, 1482–1494. [Google Scholar] [CrossRef]
- Wang, D.; Wang, H.; Zhou, Y.; Zhang, Q.; Zhang, F.; Du, P.; Wang, S.; Chen, C.; Kan, B. Genome sequencing reveals unique mutations in characteristic metabolic pathways and the transfer of virulence genes between V. mimicus and V. cholerae. PLoS ONE 2011, 6, e21299. [Google Scholar] [CrossRef]
- Garza, D.R.; Thompson, C.C.; Loureiro, E.C.B.; Dutilh, B.E.; Inada, D.T.; Junior, E.C.S.; Cardoso, J.F.; Nunes, M.R.T.; de Lima, C.P.S.; Silvestre, R.V.D. Genome-wide study of the defective sucrose fermenter strain of Vibrio cholerae from the Latin American cholera epidemic. PLoS ONE 2012, 7, e37283. [Google Scholar] [CrossRef]
- Hayes, C.A.; Dalia, T.N.; Dalia, A.B. Systematic genetic dissection of PTS in Vibrio cholerae uncovers a novel glucose transporter and a limited role for PTS during infection of a mammalian host. Mol. Microbiol. 2017, 104, 568–579. [Google Scholar] [CrossRef]
- Reid, S.J.; Abratt, V.R. Sucrose utilisation in bacteria: Genetic organisation and regulation. Appl. Microbiol. Biotechnol. 2005, 67, 312–321. [Google Scholar] [CrossRef] [PubMed]
- Osorio, C.R.; Vences, A.; Matanza, X.M.; Terceti, M.S. Photobacterium damselae subsp. damselae, a generalist pathogen with unique virulence factors and high genetic diversity. J. Bacteriol. 2018, 15, e00002-18. [Google Scholar] [CrossRef] [PubMed]
- Hundenborn, J.; Thurig, S.; Kommerell, M.; Haag, H.; Nolte, O. Severe wound infection with Photobacterium damselae ssp. damselae and Vibrio harveyi, following a laceration injury in marine environment: A case report and review of the literature. Case Rep. Med. 2013, 2013, 610632. [Google Scholar] [CrossRef] [PubMed]
- Terceti, M.S.; Vences, A.; Matanza, X.M.; Dalsgaard, I.; Pedersen, K.; Osorio, C.R. Molecular epidemiology of Photobacterium damselae subsp. damselae outbreaks in marine rainbow trout farms reveals extensive horizontal gene transfer and high genetic diversity. Front. Microbiol. 2018, 9, 2155. [Google Scholar] [CrossRef] [PubMed]
- Osorio, C.R. Photobacterium damselae: How horizontal gene transfer shaped two different pathogenic lifestyles in a marine bacterium. In Horizontal Gene Transfer: Breaking Borders between Living Kingdoms; Villa, T.G., Viñas, M., Eds.; Springer: Cham, Switzerland, 2019; pp. 175–199. [Google Scholar]
- Vences, A.; Abushattal, S.; Matanza, X.M.; Dubert, J.; Uzun, E.; Ogut, H.; Osorio, C.R. Highly transferable pAQU-related plasmids encoding multidrug resistance are widespread in the human and fish pathogen Photobacterium damselae subsp. damselae in aquaculture areas in the Black Sea. Microb. Ecol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Farmer, J.J., III; Hickman-Brenner, F.W.; Kelly, M.T. Vibrio. In Manual of Clinical Microbiology, 4th ed.; Lennette, E.H., Balows, A., Hausler, W.J., Shadomy, H.J., Eds.; American Society for Microbiology: Washington, DC, USA, 1985; pp. 282–301. [Google Scholar]
- Botella, S.; Pujalte, M.; Macián, M.; Ferrús, M.; Hernández, J.; Garay, E. Amplified fragment length polymorphism (AFLP) and biochemical typing of Photobacterium damselae subsp. damselae. J. Appl. Microbiol. 2002, 93, 681–688. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.H.; Sun, J.J.; Liu, L.; Zhao, H.H.; Wang, H.F.; Liang, L.Q.; Liu, L.B.; Li, G.F. Characterization of two phenotypes of Photobacterium damselae subsp. damselae isolated from diseased juvenile Trachinotus ovatus reared in cage mariculture. J. World Aquac. Soc. 2009, 40, 281–289. [Google Scholar] [CrossRef]
- Terceti, M.S.; Ogut, H.; Osorio, C.R. Photobacterium damselae subsp. damselae, an emerging fish pathogen in the Black Sea: Evidence of a multiclonal origin. Appl. Environ. Microbiol. 2016, 82, 3736–3745. [Google Scholar] [CrossRef]
- Tao, Z.; Shen, C.; Zhou, S.-M.; Yang, N.; Wang, G.-L.; Wang, Y.-J.; Xu, S.-L. An outbreak of Photobacterium damselae subsp. damselae infection in cultured silver pomfret Pampus argenteus in Eastern China. Aquaculture 2018, 492, 201–205. [Google Scholar] [CrossRef]
- Love, M.; Teebken-Fisher, D.; Hose, J.E.; Farmer, J.J., 3rd; Hickman, F.W.; Fanning, G.R. Vibrio damsela, a marine bacterium, causes skin ulcers on the damselfish Chromis punctipinnis. Science 1981, 214, 1139–1140. [Google Scholar] [CrossRef]
- Fouz, B.; Larsen, J.L.; Nielsen, B.; Barja, J.L.; Toranzo, A.E. Characterization of Vibrio damsela strains isolated from turbot Scophthalmus maximus in Spain. Dis. Aquat. Organ. 1992, 12, 155–166. [Google Scholar] [CrossRef]
- Vences, A.; Rivas, A.J.; Lemos, M.L.; Husmann, M.; Osorio, C.R. Chromosome-encoded hemolysin, phospholipase, and collagenase in plasmidless isolates of Photobacterium damselae subsp. damselae contribute to virulence for fish. Appl. Environ. Microbiol. 2017, 83, e00401-17. [Google Scholar] [CrossRef] [PubMed]
- Vera, P. First isolation of Vibrio damsela from sea bream (Sparus aurata). Bull. Eur. Assoc. Fish Pathol. 1991, 11, 112. [Google Scholar]
- Pedersen, K.; Dalsgaard, I.; Larsen, J.L. Vibrio damsela associated with diseased fish in Denmark. Appl. Environ. Microbiol. 1997, 63, 3711–3715. [Google Scholar] [CrossRef]
- Pedersen, K.; Skall, H.F.; Lassen-Nielsen, A.M.; Bjerrum, L.; Olesen, N.J. Photobacterium damselae subsp. damselae, an emerging pathogen in Danish rainbow trout, Oncorhynchus mykiss (Walbaum), mariculture. J. Fish Dis. 2009, 32, 465–472. [Google Scholar] [CrossRef]
- Kreger, A.S. Cytolytic activity and virulence of Vibrio damsela. Infect. Immun. 1984, 44, 326–331. [Google Scholar] [CrossRef]
- Grimes, D.J.; Colwell, R.R.; Stemmler, J.; Hada, H.; Maneval, D.; Hetrick, F.M.; May, E.B.; Jones, R.T.; Stoskopf, M. Vibrio species as agents of elasmobranch disease. Helgoländer Meeresunters 1984, 37, 309–315. [Google Scholar] [CrossRef][Green Version]
- Matanza, X.M.; Osorio, C.R. Exposure of the opportunistic marine pathogen Photobacterium damselae subsp. damselae to human body temperature is a stressful condition that shapes the transcriptome, viability, cell morphology, and virulence. Front. Microbiol. 2020, 11, 1771. [Google Scholar] [CrossRef]
- Herrero, M.; de Lorenzo, V.; Timmis, K.N. Transposon vectors containing non-antibiotic resistance selection markers for cloning and stable chromosomal insertion of foreign genes in gram-negative bacteria. J. Bacteriol. 1990, 172, 6557–6567. [Google Scholar] [CrossRef]
- Parales, R.E.; Harwood, C.S. Construction and use of a new broad-host-range lacZ transcriptional fusion vector, pHRP309, for Gram− bacteria. Gene 1993, 133, 23–30. [Google Scholar] [CrossRef]
- Wang, R.F.; Kushner, S.R. Construction of versatile low-copy-number vectors for cloning, sequencing and gene expression in Escherichia coli. Gene 1991, 100, 195–199. [Google Scholar] [CrossRef]
- Mouriño, S.; Osorio, C.R.; Lemos, M.L. Characterization of heme uptake cluster genes in the fish pathogen Vibrio anguillarum. J. Bacteriol. 2004, 186, 6159–6167. [Google Scholar] [CrossRef] [PubMed]
- Nurk, S.; Bankevich, A.; Antipov, D.; Gurevich, A.A.; Korobeynikov, A.; Lapidus, A.; Prjibelski, A.D.; Pyshkin, A.; Sirotkin, A.; Sirotkin, Y.; et al. Assembling single-cell genomes and mini-metagenomes from chimeric MDA products. J. Comput. Biol. 2013, 20, 714–737. [Google Scholar] [CrossRef]
- Tatusova, T.; DiCuccio, M.; Badretdin, A.; Chetvernin, V.; Nawrocki, E.P.; Zaslavsky, L.; Lomsadze, A.; Pruitt, K.D.; Borodovsky, M.; Ostell, J. NCBI Prokaryotic Genome Annotation Pipeline. Nucleic Acids Res. 2016, 44, 6614–6624. [Google Scholar] [CrossRef]
- Miller, J.H. Experiments in Molecular Genetics; Cold Spring Harbor Laboratory: Cold Spring Harbor, NY, USA, 1972. [Google Scholar]
- Miller, J.H. A Short Course in Bacterial Genetics; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1992. [Google Scholar]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST Server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef]
- El-Gebali, S.; Mistry, J.; Bateman, A.; Eddy, S.R.; Luciani, A.; Potter, S.C.; Qureshi, M.; Richardson, L.J.; Salazar, G.A.; Smart, A.; et al. The Pfam protein families database in 2019. Nucleic Acids Res. 2019, 47, D427–D432. [Google Scholar] [CrossRef]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A genome comparison visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef]
- Darling, A.C.; Mau, B.; Blattner, F.R.; Perna, N.T. Mauve: Multiple alignment of conserved genomic sequence with rearrangements. Genome Res. 2004, 14, 1394–1403. [Google Scholar] [CrossRef]
- Darling, A.E.; Mau, B.; Perna, N.T. progressiveMauve: Multiple genome alignment with gene gain, loss and rearrangement. PLoS ONE 2010, 5, e11147. [Google Scholar] [CrossRef]
- Pérez-Cataluña, A.; Lucena, T.; Tarazona, E.; Arahal, D.R.; Macián, M.C.; Pujalte, M.J. An MLSA approach for the taxonomic update of the Splendidus clade, a lineage containing several fish and shellfish pathogenic Vibrio spp. Syst. Appl. Microbiol. 2016, 39, 361–369. [Google Scholar] [CrossRef]
- Pascual, J.; Macián, M.C.; Arahal, D.R.; Garay, E.; Pujalte, M.J. Multilocus sequence analysis of the central clade of the genus Vibrio by using the 16S rRNA, recA, pyrH, rpoD, gyrB, rctB and toxR genes. Int. J. Syst. Evol. Microbiol. 2010, 60, 154–165. [Google Scholar] [CrossRef]
- Maddison, W.P.; Maddison, D.R. Mesquite: A Modular System for Evolutionary Analysis. Version 3.61. Available online: http://www.mesquiteproject.org (accessed on 29 February 2020).
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar]
- Tamura, K.; Nei, M.; Kumar, S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc. Natl. Acad. Sci. USA 2004, 101, 11030–11035. [Google Scholar] [CrossRef] [PubMed]
- Deutscher, J.; Aké, F.M.D.; Derkaoui, M.; Zébré, A.C.; Cao, T.N.; Bouraoui, H.; Kentache, T.; Mokhtari, A.; Milohanic, E.; Joyet, P. The bacterial phosphoenolpyruvate: Carbohydrate phosphotransferase system: Regulation by protein phosphorylation and phosphorylation-dependent protein-protein interactions. Microbiol. Mol. Biol. Rev. 2014, 78, 231–256. [Google Scholar] [CrossRef]
- McCoy, J.G.; Ren, Z.; Stanevich, V.; Lee, J.; Mitra, S.; Levin, E.J.; Poget, S.; Quick, M.; Im, W.; Zhou, M. The structure of a sugar transporter of the glucose EIIC superfamily provides insight into the elevator mechanism of membrane transport. Structure 2016, 24, 956–964. [Google Scholar] [CrossRef]
- Jahrels, K.; Lengeler, J.W. Molecular analysis of two ScrR repressors and of a ScrR–FruR hybrid repressor for sucrose and D-fructose specific regulons from enteric bacteria. Mol. Microbiol. 1993, 9, 195–209. [Google Scholar] [CrossRef]
- Bogs, J.; Geider, K. Molecular analysis of sucrose metabolism of Erwinia amylovora and influence on bacterial virulence. J. Bacteriol. 2000, 182, 5351–5358. [Google Scholar] [CrossRef]
- Lambert, B.; Dassanayake, M.; Oh, D.-H.; Garrett, S.B.; Lee, S.-Y.; Pettis, G.S. A novel phase variant of the cholera pathogen shows stress-adaptive cryptic transcriptomic signatures. BMC Genom. 2016, 17, 914. [Google Scholar] [CrossRef][Green Version]
- Nácher-Vázquez, M.; Iturria, I.; Zarour, K.; Mohedano, M.L.; Aznar, R.; Pardo, M.Á.; López, P. Dextran production by Lactobacillus sakei MN1 coincides with reduced autoagglutination, biofilm formation and epithelial cell adhesion. Carbohydr. Polym. 2017, 168, 22–31. [Google Scholar] [CrossRef]
- Durica-Mitic, S.; Goepel, Y.; Görke, B. Carbohydrate utilization in bacteria: Making the most out of sugars with the help of small regulatory RNAs. Microbiol. Spectr. 2018, 6. [Google Scholar] [CrossRef]
- Ochman, H.; Lawrence, J.G.; Groisman, E.A. Lateral gene transfer and the nature of bacterial innovation. Nature 2000, 405, 299–304. [Google Scholar] [CrossRef]
- Hazen, T.H.; Pan, L.; Gu, J.-D.; Sobecky, P.A. The contribution of mobile genetic elements to the evolution and ecology of Vibrios. FEMS Microbiol. Ecol. 2010, 74, 485–499. [Google Scholar] [CrossRef] [PubMed]
- Adin, D.M.; Visick, K.L.; Stabb, E.V. Identification of a cellobiose utilization gene cluster with cryptic beta-galactosidase activity in Vibrio fischeri. Appl. Environ. Microbiol. 2008, 74, 4059–4069. [Google Scholar] [CrossRef]
- Treviño-Quintanilla, L.G.; Escalante, A.; Caro, A.D.; Martínez, A.; González, R.; Puente, J.L.; Bolívar, F.; Gosset, G. The phosphotransferase system-dependent sucrose utilization regulon in enteropathogenic Escherichia coli strains is located in a variable chromosomal region containing iap sequences. J. Mol. Microbiol. Biotechnol. 2007, 13, 117–125. [Google Scholar] [CrossRef]
- Alaeddinoglu, N.G.; Charles, H.P. Transfer of a gene for sucrose utilization into Escherichia coli k12, and consequent failure of expression of genes for D-serine utilization. Microbiology 1979, 110, 47–59. [Google Scholar] [CrossRef]
- Bockmann, J.; Heuel, H.; Lengeler, J.W. Characterization of a chromosomally encoded, non-PTS metabolic pathway for sucrose utilization in Escherichia coli EC3132. Mol. Gen. Genet. MGG 1992, 235, 22–32. [Google Scholar] [CrossRef]
- Wohlhieter, J.A.; Lazere, J.R.; Snellings, N.J.; Johnson, E.M.; Synenki, R.M.; Baron, L.S. Characterization of transmissible genetic elements from sucrose-fermenting Salmonella strains. J. Bacteriol. 1975, 122, 401–406. [Google Scholar] [CrossRef]
- Schmid, K.; Ebner, R.; Altenbuchner, J.; Schmitt, R.; Lengeler, J.W. Plasmid-mediated sucrose metabolism in Escherichia coli K12: Mapping of the scr genes of pUR400. Mol. Microbiol. 1988, 2, 1–8. [Google Scholar] [CrossRef]
- Hardesty, C.; Ferran, C.; DiRienzo, J.M. Plasmid-mediated sucrose metabolism in Escherichia coli: Characterization of scrY, the structural gene for a phosphoenolpyruvate-dependent sucrose phosphotransferase system outer membrane porin. J. Bacteriol. 1991, 173, 449–456. [Google Scholar] [CrossRef]
- Maruo, B.; Hattori, T.; Takahashi, H. Excretion of ribitol and sucrose by green algae into the culture medium. Agric. Biol. Chem. 1965, 29, 1084–1089. [Google Scholar] [CrossRef]
- Lunn, J.E. Evolution of sucrose synthesis. Plant Physiol. 2002, 128, 1490–1500. [Google Scholar] [CrossRef] [PubMed]
- Hagemann, M. Molecular biology of cyanobacterial salt acclimation. FEMS Microbiol. Rev. 2011, 35, 87–123. [Google Scholar] [CrossRef]
- Bremauntz, M.; Torres-Bustillos, L.G.; Cañizares-Villanueva, R.; Duran-Paramo, E.; Fernández-Linares, L. Trehalose and sucrose osmolytes accumulated by algae as potential raw material for bioethanol. Nat. Resour. 2011, 2, 173. [Google Scholar] [CrossRef]
- Du, W.; Liang, F.; Duan, Y.; Tan, X.; Lu, X. Exploring the photosynthetic production capacity of sucrose by cyanobacteria. Metab. Eng. 2013, 19, 17–25. [Google Scholar] [CrossRef]
- Kolman, M.A.; Nishi, C.N.; Perez-Cenci, M.; Salerno, G.L. Sucrose in cyanobacteria: From a salt-response molecule to play a key role in nitrogen fixation. Life 2015, 5, 102–126. [Google Scholar] [CrossRef]
- Kolman, M.A.; Salerno, G.L. Sucrose in bloom-forming cyanobacteria: Loss and gain of genes involved in its biosynthesis. Environ. Microbiol. 2016, 18, 439–449. [Google Scholar] [CrossRef]
- Kirsch, F.; Klähn, S.; Hagemann, M. Salt-regulated accumulation of the compatible solutes sucrose and glucosylglycerol in cyanobacteria and its biotechnological potential. Front. Microbiol. 2019, 10, 2139. [Google Scholar] [CrossRef]
- Carrasco-Reinado, R.; Escobar, A.; Carrera, C.; Guarnizo, P.; Vallejo, R.A.; Fernández-Acero, F.J. Valorization of microalgae biomass as a potential source of high-value sugars and polyalcohols. LWT-Food Sci. Technol. 2019, 114, 108385. [Google Scholar] [CrossRef]
- Sogin, E.M.; Puskas, E.; Dubilier, N.; Liebeke, M. Marine metabolomics: A method for nontargeted measurement of metabolites in seawater by gas chromatography–mass spectrometry. mSystems 2019, 4. [Google Scholar] [CrossRef]
- Martínez-Crego, B.; Vizzini, S.; Califano, G.; Massa-Gallucci, A.; Andolina, C.; Gambi, M.C.; Santos, R. Resistance of seagrass habitats to ocean acidification via altered interactions in a tri-trophic chain. Sci. Rep. 2020, 10, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.; Bassler, B.L.; Roseman, S. Chemotaxis of the marine bacterium Vibrio furnissii to sugars. A potential mechanism for initiating the chitin catabolic cascade. J. Biol. Chem. 1993, 268, 9405–9409. [Google Scholar] [PubMed]


| Strain | Isolation Source | Strain Reference | Scr Phenotype a | Presence of Sucrose Genes b | Genbank Acc. No. c | Genome Reference |
|---|---|---|---|---|---|---|
| 154dp-OG1 | European sea bass, Turkey | [30] | − | − | NA | NA |
| 164dp-OG2 | European sea bass, Turkey | [30] | − | − | VAUU00000000 | [26] |
| 144bp-OG3 | European sea bass, Turkey | [30] | − | − | VAND00000000 | [26] |
| 162bp-OG4A | European sea bass, Turkey | [30] | + | + | JABWTP000000000 | This study |
| 158dp-OG5 | European sea bass, Turkey | [30] | − | − | NA | NA |
| 189bp-OG7B | European sea bass, Turkey | [30] | + | + | JABXOP000000000 | This study |
| 82dy-OG8 | European sea bass, Turkey | [30] | − | − | JABXOQ000000000 | This study |
| 64bp-OG9 | European sea bass, Turkey | [30] | + | + | VANE00000000 | [26] |
| 156dp-OG10A | European sea bass, Turkey | [30] | − | − | NA | NA |
| 125dy-OG11 | European sea bass, Turkey | [30] | + | + | JACFTX000000000 | This study |
| 70dps-OG12 | European sea bass, Turkey | [30] | + | + | VANF00000000 | [26] |
| 164dpbuy-OG13B | European sea bass, Turkey | [30] | − | − | NA | NA |
| 111bp-OG15A | European sea bass, Turkey | [30] | − | − | VANG00000000 | [26] |
| 89dp-OG16 | European sea bass, Turkey | [30] | + | + | VANH00000000 | [26] |
| CIP 102761 | Damselfish, United Estates | [32] | − | − | ADBS00000000 | Unpublished |
| TW250/03 | Gilthead seabream | Laboratory collection | + | + | JABXOR000000000 | This study |
| RM-71 | Turbot, Spain | [33] | − | − | LYBT00000000 | [34] |
| A-162 | Eel, Belgium | NA | − | − | LZFN00000000 | [34] |
| LD-07 | Gilthead seabream, Spain | [35] | − | − | LYBU00000000 | [34] |
| DK2 | Rainbow trout, Denmark | [24,36] | − | − | PVXF00000000 | [24] |
| DK3 | Rainbow trout, Denmark | [24,36] | − | − | PVXG00000000 | [24] |
| DK20 | Rainbow trout, Denmark | [24,37] | − | − | PVXH00000000 | [24] |
| DK29 | Rainbow trout, Denmark | [24,37] | − | − | PVXI00000000 | [24] |
| DK32 | Rainbow trout, Denmark | [24,37] | + | + | JABWTO000000000 | This study |
| CDC-1421-81 | Fish, Senegal | [38] | − | − | JABXYE000000000 | This study |
| RG-191 | Turbot, Spain | [33] | − | − | NA | NA |
| ATCC35083 | Brown shark, United States | [39] | − | − | NA | NA |
| CDC-2227-81 | Human, United States | [38] | − | − | VZUQ00000000 | [40] |
| 80077637 | Human, Australia | [23] | − | − | WAEO00000000 | [40] |
| ACRP-72.1 | Turbot, Portugal | Laboratory collection | − | − | NA | NA |
| DLC 1.2 | Gilthead seabream, Spain | Laboratory collection | − | − | NA | NA |
| DLC 4.1 | Gilthead seabream, Spain | Laboratory collection | − | − | NA | NA |
| DLC 7.1 | Gilthead seabream, Spain | Laboratory collection | − | − | NA | NA |
| DLC 7.3 | Gilthead seabream, Spain | Laboratory collection | − | − | NA | NA |
| DLC 8.1 | Gilthead seabream, Spain | Laboratory collection | − | − | NA | NA |
| DLC 9.1 | Gilthead seabream, Spain | Laboratory collection | − | − | NA | NA |
| Strain or Plasmid | Description a | Reference/Source |
|---|---|---|
| Strains | ||
| P. damselae subsp. damselae | ||
| DK32 | Isolated from rainbow trout; ferments sucrose (Scr+) | [24,37] |
| SSS165 | DK32 ΔscrA. Does not ferment sucrose (Scr−) | This study |
| SSS250 | DK32 with plasmid pSSS250 | This study |
| E. coli | ||
| DH5α | Cloning strain | Laboratory stock |
| S17-1-λpir | RP4-2(Km::Tn7, Tc::Mu-1) pro-82 λpir recA1 endA1 thiE1 hsdR17 creC510 | [41] |
| Plasmids | ||
| pHRP309 | lacZ reporter plasmid, mob Gmr | [42] |
| pSSS250 | pHRP309 with a transcriptional fusion of scrA promoter to lacZ | This study |
| pWKS30 | Low-copy-number cloning vector; Apr | [43] |
| pNidKan | Suicide vector derived from pCVD442; Kmr | [44] |
| Name of Primer Pair | Oligonucleotide Sequence 1 | Amplicon Size (bp) |
|---|---|---|
| scrA-1-2 | F: 5’-GCTCTAGAGCCATTCGCACAACACTTTG-3’ R: 5’-GCGGATCCGTTCGCTAGATCAGTCAATC-3’ | 2106 |
| scrA-3-4 | F: 5’-GCGGATCCTCAAGGTGCTGCCGCTTTAG-3’ R: 5’-GCGAATTCAGGACCTTTATGCTGCCACG-3’ | 2122 |
| scrA-mutant-test | F: 5’-GGCTCAGGCATAGTAAACCA-3’ R: 5’-CCGCGATAAATGGGTAACGT-3’ | 1024 |
| scrA-promoter | F: 5’-GCTCTAGAACATCATGCAGACTCGCCAT-3’ R: 5’-GCGGATCCCTCTTTAGCTACTGCCGGAT-3’ | 281 |
| scrK | F: 5’-TTACGCGACTCACCTCGACA-3’ R: 5’-ATCGGTCGCGCAGAACAAAC-3’ | 373 |
| scrB | F: 5’-GACCAAGACTACGATTCACA-3’ R: 5’-ACACTCCCACATGTACCCAA-3’ | 372 |
| Species | Accession Number |
|---|---|
| Vibrio alfacsensis | CP032093.1 |
| V. parahaemolyticus | QPIY01000005 |
| V. alginolyticus | AAPS01000004 |
| Vibrio rotiferianus | NZ_KV861318 |
| Vibrio diabolicus | CP014133 |
| Vibrio harveyi | CP014038 |
| Vibrio fortis | NZ_JFFR01000009 |
| Vibrio cyclitrophicus | VUKB01000001 |
| Photobacterium lutimaris | NZ_SNZO01000003 |
| Vibrio crassostreae | NZ_AJZB02000137 |
| Photobacterium angustum | NZ_PYOK01000006 |
| Photobacterium frigidiphilum | NZ_PYMJ01000001 |
| Photobacterium profundum | NZ_PYOD01000001 |
| Photobacterium indicum | NZ_PYOC01000002 |
| V. cholerae | NZ_VTLI01000001 |
| Photobacterium halotolerans | NZ_AULG01000013 |
| Photobacterium rosenbergii | NZ_PYMB01000001 |
| Photobacterium lipolyticum | NZ_PYMC01000002 |
| Attribute | 162bp-OG4A (Scr+) | 189bp-OG7B (Scr+) | 82dy-OG8 (Scr−) | 125dy-OG11 (Scr+) | DK32 (Scr+) | TW250/03 (Scr+) | CDC-1421-81 (Scr−) |
|---|---|---|---|---|---|---|---|
| Accession no. | JABWTP000000000 | JABXOP000000000 | JABXOQ000000000 | JACFTX000000000 | JABWTO000000000 | JABXOR000000000 | JABXYE000000000 |
| Genome size (bp) | 4,306,101 bp | 4,302,857 bp | 4,627,325 bp | 4,450,948 bp | 4,248,331 bp | 4,695,503 bp | 4,432,211 bp |
| Contigs | 133 | 127 | 138 | 147 | 106 | 1649 | 81 |
| % GC | 40.70% | 40.70% | 40.70% | 40.70% | 40.60% | 39.80% | 40.40% |
| Genes (total) | 3809 | 3807 | 4096 | 4089 | 3750 | 5089 | 3916 |
| CDSs | 3756 | 3758 | 4038 | 3879 | 3699 | 5041 | 3860 |
| Strain | 162bp-OG4A | 189bp-OG7B | 64bp-OG9 | 125dy-OG11 | 70dps-OG12 | 89dp-OG16 | DK32 | TW250/03 |
|---|---|---|---|---|---|---|---|---|
| Accession no. | JABWTP000000000 | JABXOP000000000 | VANE00000000 | JACFTX000000000 | VANF00000000 | VANH00000000 | JABWTO000000000 | JABXOR000000000 |
| Sucrose operon genes (locus_tag) | ||||||||
| scrA | HU985_14400 | HVV26_09235 | FD719_10830 | H3N34_00965 | FD720_04185 | FD722_11585 | HU831_00535 | HWA77_17195 |
| scrB | HU985_14390 | HVV26_09225 | FD719_10820 | H3N34_00955 | FD720_04175 | FD722_11575 | HU831_00545 | HWA77_17185 |
| scrK | HU985_14395 | HVV26_09230 | FD719_10825 | H3N34_00960 | FD720_04180 | FD722_11580 | HU831_00540 | HWA77_17190 |
| scrR | HU985_14405 | HVV26_09240 | FD719_10835 | H3N34_00970 | FD720_04190 | FD722_11590 | HU831_00530 | HWA77_17200 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abushattal, S.; Vences, A.; Barca, A.V.; Osorio, C.R. Diverse Horizontally-Acquired Gene Clusters Confer Sucrose Utilization to Different Lineages of the Marine Pathogen Photobacterium damselae subsp. damselae. Genes 2020, 11, 1244. https://doi.org/10.3390/genes11111244
Abushattal S, Vences A, Barca AV, Osorio CR. Diverse Horizontally-Acquired Gene Clusters Confer Sucrose Utilization to Different Lineages of the Marine Pathogen Photobacterium damselae subsp. damselae. Genes. 2020; 11(11):1244. https://doi.org/10.3390/genes11111244
Chicago/Turabian StyleAbushattal, Saqr, Ana Vences, Alba V. Barca, and Carlos R. Osorio. 2020. "Diverse Horizontally-Acquired Gene Clusters Confer Sucrose Utilization to Different Lineages of the Marine Pathogen Photobacterium damselae subsp. damselae" Genes 11, no. 11: 1244. https://doi.org/10.3390/genes11111244
APA StyleAbushattal, S., Vences, A., Barca, A. V., & Osorio, C. R. (2020). Diverse Horizontally-Acquired Gene Clusters Confer Sucrose Utilization to Different Lineages of the Marine Pathogen Photobacterium damselae subsp. damselae. Genes, 11(11), 1244. https://doi.org/10.3390/genes11111244






