Plasmid Transfer by Conjugation in Gram-Negative Bacteria: From the Cellular to the Community Level
Abstract
1. Introduction
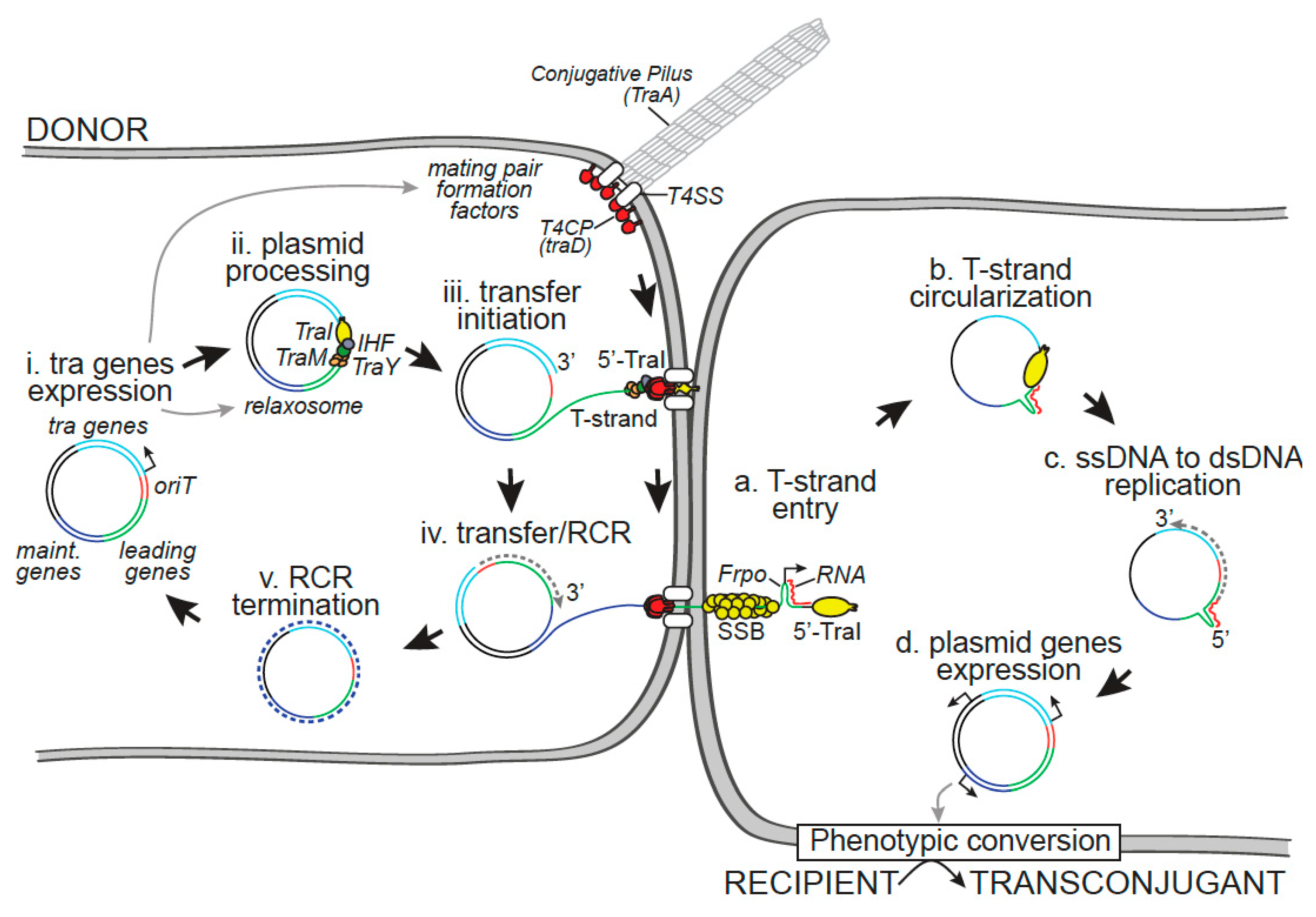
2. Within the Donor Cell
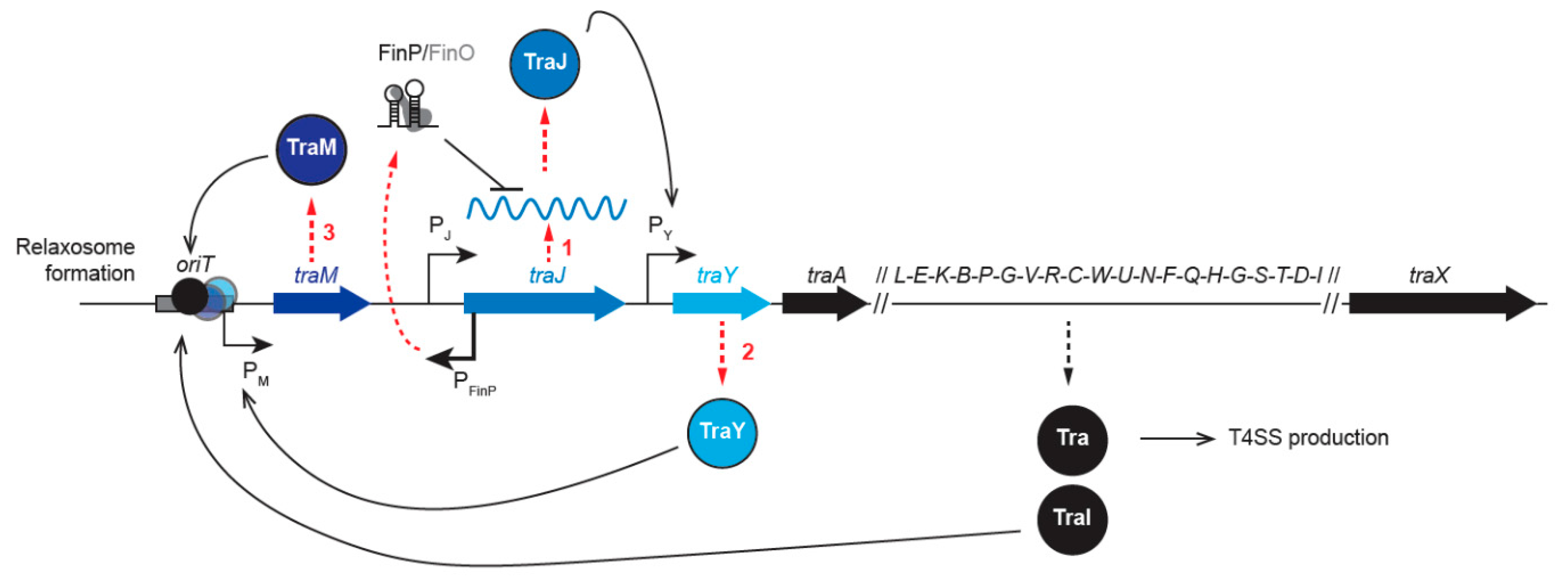
2.1. Transfer Gene Expression
2.1.1. Regulation of tra Gene Expression
2.1.2. Superspreader Mutations
2.2. Conjugative Pilus, Mating Pair Formation, and Stabilization
2.2.1. F-Pilus Structure and Biosynthesis
2.2.2. Pilus Biological Function
2.2.3. Factors Involved in the Specificity of Donor–Recipient Interactions
2.3. Plasmid Processing by the Relaxosome
2.4. Initiation of Rolling Circle Replication in the Donor Cell
2.5. T4CP Connects the Relaxosome to the T4SS
3. Within the Recipient Cell
3.1. Plasmid Circularization by TraI
3.2. Avoiding Host Defense Systems against Foreign DNA
3.3. Role of the Leading Region Genes and Conversion of the ssDNA Plasmid into dsDNA
3.3.1. Early Expression of the Leading Region Genes
3.3.2. PsiB Inhibits the SOS Response
3.3.3. Roles of the Host and Plasmidic SSB Proteins in Plasmid Establishment
3.4. Plasmid Maintenance: Replication and Segregation
3.5. Phenotypic Conversion of the Transconjugant
4. Conjugation in Natural Habitats: The Example of Bacterial Biofilms
4.1. The Biofilm as a Niche Promoting Bacterial Conjugation
4.2. Impact of the Biofilm Structure on Conjugation
4.3. Impact of Conjugative Plasmids on Biofilm Formation
4.4. Influence of Antibiotic Treatment on Conjugation within Biofilms
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lederberg, J.; Tatum, E.L. Gene recombination in Escherichia coli. Nature 1946, 158, 558. [Google Scholar] [CrossRef] [PubMed]
- Grohmann, E.; Muth, G.; Espinosa, M. Conjugative plasmid transfer in gram-positive bacteria. Microbiol. Mol. Biol. Rev. 2003, 67, 277–301. [Google Scholar] [CrossRef] [PubMed]
- Cruz, F.D.L.; Frost, L.S.; Meyer, R.J.; Zechner, E.L. Conjugative DNA metabolism in Gram-negative bacteria. FEMS Microbiol. Rev. 2010, 34, 18–40. [Google Scholar] [CrossRef] [PubMed]
- Davison, J. Genetic exchange between bacteria in the environment. Plasmid 1999, 42, 73–91. [Google Scholar] [CrossRef]
- Lawrence, J.G. Gene transfer, speciation, and the evolution of bacterial genomes. Curr. Opin. Microbiol. 1999, 2, 519–523. [Google Scholar] [CrossRef]
- Ochman, H.; Lawrence, J.G.; Groisman, E.A. Lateral gene transfer and the nature of bacterial innovation. Nature 2000, 405, 299–304. [Google Scholar] [CrossRef]
- Ippen-Ihler, K.; Achtman, M.; Willetts, N. Deletion map of the Escherichia coli K-12 sex factor F: The order of eleven transfer cistrons. J. Bacteriol. 1972, 110, 857–863. [Google Scholar] [CrossRef]
- Penfold, S.S.; Simon, J.; Frost, L.S. Regulation of the expression of the traM gene of the F sex factor of Escherichia coli. Mol. Microbiol. 1996, 20, 549–558. [Google Scholar] [CrossRef] [PubMed]
- Koraimann, G.; Wagner, M.A. Social behavior and decision making in bacterial conjugation. Front. Cell. Infect. Microbiol. 2014, 4. [Google Scholar] [CrossRef]
- Timmis, K.N.; Andrés, I.; Achtman, M. Fertility repression of F-like conjugative plasmids: Physical mapping of the R6--5 finO and finP cistrons and identification of the finO protein. Proc. Natl. Acad. Sci. USA 1978, 75, 5836–5840. [Google Scholar] [CrossRef]
- Willetts, N. The transcriptional control of fertility in F-like plasmids. J. Mol. Biol. 1977, 112, 141–148. [Google Scholar] [CrossRef]
- Frost, L.; Lee, S.; Yanchar, N.; Paranchych, W. finP and fisO mutations in FinP anti-sense RNA suggest a model for FinOP action in the repression of bacterial conjugation by the Flac plasmid JCFL0. Mol. Gen. Genet. 1989, 218, 152–160. [Google Scholar] [CrossRef] [PubMed]
- Koraimann, G.; Teferle, K.; Markolin, G.; Woger, W.; Högenauer, G. The FinOP repressor system of plasmid R1: Analysis of the antisense RNA control of traJ expression and conjugative DNA transfer. Mol. Microbiol. 1996, 21, 811–821. [Google Scholar] [CrossRef] [PubMed]
- van Biesen, T.; Frost, L.S. Differential levels of fertility inhibition among F-like plasmids are related to the cellular concentration of finO mRNA. Mol. Microbiol. 1992, 6, 771–780. [Google Scholar] [CrossRef] [PubMed]
- van Biesen, T.; Frost, L.S. The FinO protein of IncF plasmids binds FinP antisense RNA and its target, traJ mRNA, and promotes duplex formation. Mol. Microbiol. 1994, 14, 427–436. [Google Scholar] [CrossRef]
- Jerome, L.J.; van Biesen, T.; Frost, L.S. Degradation of FinP antisense RNA from F-like plasmids: The RNA-binding protein, FinO, protects FinP from ribonuclease E. J. Mol. Biol. 1999, 285, 1457–1473. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.J.W.; Lu, J.; Glover, J.N.M. Relaxosome function and conjugation regulation in F-like plasmids—A structural biology perspective. Mol. Microbiol. 2012, 85, 602–617. [Google Scholar] [CrossRef]
- Will, W.R.; Frost, L.S. Characterization of the opposing roles of H-NS and TraJ in transcriptional regulation of the F-Plasmid tra Operon. J. Bacteriol. 2006, 188, 507–514. [Google Scholar] [CrossRef]
- Will, W.R.; Lu, J.; Frost, L.S. The role of H-NS in silencing F transfer gene expression during entry into stationary phase. Mol. Microbiol. 2004, 54, 769–782. [Google Scholar] [CrossRef]
- Ali Azam, T.; Iwata, A.; Nishimura, A.; Ueda, S.; Ishihama, A. Growth phase-dependent variation in protein composition of the Escherichia coli nucleoid. J. Bacteriol. 1999, 181, 6361–6370. [Google Scholar] [CrossRef]
- Frost, L.S.; Manchak, J. F-phenocopies: Characterization of expression of the F transfer region in stationary phase. Microbiology 1998, 144 Pt 9, 2579–2587. [Google Scholar] [CrossRef]
- Headd, B.; Bradford, S.A. The conjugation window in an Escherichia coli K-12 strain with an IncFII Plasmid. Appl. Environ. Microbiol. 2020, 86. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Peng, Y.; Wan, S.; Frost, L.S.; Raivio, T.; Glover, J.N.M. Cooperative function of TraJ and ArcA in regulating the F Plasmid tra Operon. J. Bacteriol. 2019, 201. [Google Scholar] [CrossRef] [PubMed]
- Camacho, E.M.; Serna, A.; Madrid, C.; Marqués, S.; Fernández, R.; de la Cruz, F.; Juárez, A.; Casadesús, J. Regulation of finP transcription by DNA Adenine Methylation in the virulence Plasmid of Salmonella enterica. J. Bacteriol. 2005, 187, 5691–5699. [Google Scholar] [CrossRef] [PubMed]
- Zahrl, D.; Wagner, A.; Tscherner, M.; Koraimann, G. GroEL plays a central role in stress-induced negative regulation of bacterial conjugation by promoting proteolytic degradation of the activator protein TraJ. J. Bacteriol. 2007, 189, 5885–5894. [Google Scholar] [CrossRef]
- Dessaux, Y.; Faure, D. Quorum sensing and Quorum quenching in Agrobacterium: A “Go/No Go system”? Genes 2018, 9, 210. [Google Scholar] [CrossRef]
- Lu, Y.; Zeng, J.; Wu, B.; Wang, L.; Cai, R.; Zhang, N.; Li, Y.; Huang, X.; Huang, B.; Chen, C.; et al. Quorum sensing N-acyl homoserine Lactones-SdiA suppresses Escherichia coli-Pseudomonas aeruginosa conjugation through Inhibiting traI expression. Front. Cell. Infect. Microbiol. 2017, 7. [Google Scholar] [CrossRef]
- Cheah, K.C.; Skurray, R. The F plasmid carries an IS3 insertion within finO. J. Gen. Microbiol. 1986, 132, 3269–3275. [Google Scholar] [CrossRef]
- Yoshioka, Y.; Ohtsubo, H.; Ohtsubo, E. Repressor gene finO in plasmids R100 and F: Constitutive transfer of plasmid F is caused by insertion of IS3 into F finO. J. Bacteriol. 1987, 169, 619–623. [Google Scholar] [CrossRef]
- Poidevin, M.; Sato, M.; Altinoglu, I.; Delaplace, M.; Sato, C.; Yamaichi, Y. Mutation in ESBL Plasmid from Escherichia coli O104:H4 leads autoagglutination and enhanced Plasmid dissemination. Front. Microbiol. 2018, 9, 130. [Google Scholar] [CrossRef]
- Yamaichi, Y.; Chao, M.C.; Sasabe, J.; Clark, L.; Davis, B.M.; Yamamoto, N.; Mori, H.; Kurokawa, K.; Waldor, M.K. High-Resolution genetic analysis of the requirements for horizontal transmission of the ESBL plasmid from Escherichia coli O104:H4. Nucleic Acids Res. 2015, 43, 348–360. [Google Scholar] [CrossRef]
- Dmowski, M.; Gołębiewski, M.; Kern-Zdanowicz, I. Characteristics of the conjugative transfer system of the IncM Plasmid pCTX-M3 and identification of Its putative regulators. J. Bacteriol. 2018, 200. [Google Scholar] [CrossRef]
- Kohler, V.; Goessweiner-Mohr, N.; Aufschnaiter, A.; Fercher, C.; Probst, I.; Pavkov-Keller, T.; Hunger, K.; Wolinski, H.; Büttner, S.; Grohmann, E.; et al. TraN: A novel repressor of an Enterococcus conjugative type IV secretion system. Nucleic Acids Res. 2018, 46, 9201–9219. [Google Scholar] [CrossRef] [PubMed]
- Potron, A.; Poirel, L.; Nordmann, P. Derepressed transfer properties leading to the efficient spread of the plasmid encoding carbapenemase OXA-48. Antimicrob. Agents Chemother. 2014, 58, 467–471. [Google Scholar] [CrossRef] [PubMed]
- Bradley, D.E. Derepressed plasmids of incompatibility group I1 determine two different morphological forms of pilus. Plasmid 1983, 9, 331–334. [Google Scholar] [CrossRef]
- Bradley, D.E. Specification of the conjugative pili and surface mating systems of Pseudomonas plasmids. J. Gen. Microbiol. 1983, 129, 2545–2556. [Google Scholar] [CrossRef]
- Bradley, D.E. Characteristics and function of thick and thin conjugative pili determined by transfer-derepressed plasmids of incompatibility groups I1, I2, I5, B, K and Z. J. Gen. Microbiol. 1984, 130, 1489–1502. [Google Scholar] [CrossRef]
- Bradley, D.E.; Taylor, D.E.; Cohen, D.R. Specification of surface mating systems among conjugative drug resistance plasmids in Escherichia coli K-12. J. Bacteriol. 1980, 143, 1466–1470. [Google Scholar] [CrossRef]
- Costa, T.R.D.; Ilangovan, A.; Ukleja, M.; Redzej, A.; Santini, J.M.; Smith, T.K.; Egelman, E.H.; Waksman, G. Structure of the bacterial sex F Pilus reveals an assembly of a stoichiometric protein-phospholipid complex. Cell 2016, 166, 1436–1444.e10. [Google Scholar] [CrossRef]
- Date, T.; Inuzuka, M.; Tomoeda, M. Purification and characterization of F pili from Escherichia coli. Biochemistry 1977, 16, 5579–5585. [Google Scholar] [CrossRef]
- Folkhard, W.; Leonard, K.R.; Malsey, S.; Marvin, D.A.; Dubochet, J.; Engel, A.; Achtman, M.; Helmuth, R. X-ray diffraction and electron microscope studies on the structure of bacterial F pili. J. Mol. Biol. 1979, 130, 145–160. [Google Scholar] [CrossRef]
- Ippen-Ihler, K.A.; Minkley, E.G. The conjugation system of F, the fertility factor of Escherichia coli. Annu. Rev. Genet. 1986, 20, 593–624. [Google Scholar] [CrossRef] [PubMed]
- Silverman, P.M.; Clarke, M.B. New insights into F-pilus structure, dynamics, and function. Integr. Biol. 2010, 2, 25–31. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.A.; Yu, X.; Silverman, P.M.; Harris, R.L.; Egelman, E.H. The structure of F-pili. J. Mol. Biol. 2009, 385, 22–29. [Google Scholar] [CrossRef]
- Frost, L.S.; Paranchych, W.; Willetts, N.S. DNA sequence of the F traALE region that includes the gene for F pilin. J. Bacteriol. 1984, 160, 395–401. [Google Scholar] [CrossRef]
- Minkley, E.G.; Polen, S.; Brinton, C.C.; Ippen-Ihler, K. Identification of the structural gene for F-pilin. J. Mol. Biol. 1976, 108, 111–121. [Google Scholar] [CrossRef]
- Maneewannakul, K.; Maneewannakul, S.; Ippen-Ihler, K. Synthesis of F pilin. J. Bacteriol. 1993, 175, 1384–1391. [Google Scholar] [CrossRef][Green Version]
- Harris, R.L.; Sholl, K.A.; Conrad, M.N.; Dresser, M.E.; Silverman, P.M. Interaction between the F plasmid TraA (F-pilin) and TraQ proteins. Mol. Microbiol. 1999, 34, 780–791. [Google Scholar] [CrossRef]
- Majdalani, N.; Ippen-Ihler, K. Membrane insertion of the F-pilin subunit is Sec independent but requires leader peptidase B and the proton motive force. J. Bacteriol. 1996, 178, 3742–3747. [Google Scholar] [CrossRef][Green Version]
- Moore, D.; Hamilton, C.M.; Maneewannakul, K.; Mintz, Y.; Frost, L.S.; Ippen-Ihler, K. The Escherichia coli K-12 F plasmid gene traX is required for acetylation of F pilin. J. Bacteriol. 1993, 175, 1375–1383. [Google Scholar] [CrossRef]
- Firth, N.; Berg, T.; Skurray, R.A. Evolution of conjugative plasmids from gram-positive bacteria. Mol. Microbiol. 1999, 31, 1598–1600. [Google Scholar] [PubMed]
- Lawley, T.D.; Klimke, W.A.; Gubbins, M.J.; Frost, L.S. F factor conjugation is a true type IV secretion system. FEMS Microbiol. Lett. 2003, 224, 1–15. [Google Scholar] [CrossRef]
- Manchak, J.; Anthony, K.G.; Frost, L.S. Mutational analysis of F-pilin reveals domains for pilus assembly, phage infection and DNA transfer. Mol. Microbiol. 2002, 43, 195–205. [Google Scholar] [CrossRef] [PubMed]
- Firth, N.; Ippen-Ihler, K.; Skurray, R.A. Gene Transfer: Conjugation Structure and Function of the F Factor and Mechanism of Conjugation; American Society of Microbiology: Washington, DC, USA, 1999. [Google Scholar]
- Koraimann, G. Spread and persistence of virulence and antibiotic resistance genes: A ride on the F Plasmid conjugation module. EcoSal Plus 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Arutyunov, D.; Frost, L.S. F conjugation: Back to the beginning. Plasmid 2013, 70, 18–32. [Google Scholar] [CrossRef]
- Frost, L.S.; Ippen-Ihler, K.; Skurray, R.A. Analysis of the sequence and gene products of the transfer region of the F sex factor. Microbiol. Rev. 1994, 58, 162–210. [Google Scholar] [CrossRef]
- Anthony, K.G.; Kathir, P.; Moore, D.; Ippen-Ihler, K.; Frost, L.S. Analysis of the traLEKBP sequence and the TraP protein from three F-like plasmids: F, R100-1 and ColB2. J. Bacteriol. 1996, 178, 3194–3200. [Google Scholar] [CrossRef][Green Version]
- Anthony, K.G.; Klimke, W.A.; Manchak, J.; Frost, L.S. Comparison of proteins involved in Pilus synthesis and mating pair stabilization from the related Plasmids F and R100-1: Insights into the mechanism of conjugation. J. Bacteriol. 1999, 181, 5149–5159. [Google Scholar] [CrossRef]
- Gopalkrishnan, S.; Ross, W.; Chen, A.Y.; Gourse, R.L. TraR directly regulates transcription initiation by mimicking the combined effects of the global regulators DksA and ppGpp. Proc. Natl. Acad. Sci. USA 2017, 114, E5539–E5548. [Google Scholar] [CrossRef]
- Shala-Lawrence, A.; Bragagnolo, N.; Nowroozi-Dayeni, R.; Kheyson, S.; Audette, G.F. The interaction of TraW and TrbC is required to facilitate conjugation in F-like plasmids. Biochem. Biophys. Res. Commun. 2018, 503, 2386–2392. [Google Scholar] [CrossRef]
- Garcillán-Barcia, M.P.; de la Cruz, F. Why is entry exclusion an essential feature of conjugative plasmids? Plasmid 2008, 60, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Achtman, M. Mating aggregates in Escherichia coli conjugation. J. Bacteriol. 1975, 123, 505–515. [Google Scholar] [CrossRef] [PubMed]
- Lanka, E.; Wilkins, B.M. DNA processing reactions in bacterial conjugation. Annu. Rev. Biochem. 1995, 31. [Google Scholar] [CrossRef]
- Dostál, L.; Shao, S.; Schildbach, J.F. Tracking F plasmid TraI relaxase processing reactions provides insight into F plasmid transfer. Nucleic Acids Res. 2011, 39, 2658–2670. [Google Scholar] [CrossRef] [PubMed]
- Matson, S.W.; Sampson, J.K.; Byrd, D.R.N. F Plasmid conjugative DNA transfer the TraI helicase activity is essential for DNA strand transfer. J. Biol. Chem. 2001, 276, 2372–2379. [Google Scholar] [CrossRef]
- Curtiss, R. Bacterial conjugation. Annu. Rev. Microbiol. 1969, 23, 69–136. [Google Scholar] [CrossRef]
- Marvin, D.A.; Hohn, B. Filamentous bacterial viruses. Bacteriol. Rev. 1969, 33, 172–209. [Google Scholar] [CrossRef]
- Clarke, M.; Maddera, L.; Harris, R.L.; Silverman, P.M. F-pili dynamics by live-cell imaging. Proc. Natl. Acad. Sci. USA 2008, 105, 17978–17981. [Google Scholar] [CrossRef]
- Achtman, M.; Morelli, G.; Schwuchow, S. Cell-Cell interactions in conjugating Escherichia coli: Role of F pili and fate of mating aggregates. J. Bacteriol. 1978, 135, 1053–1061. [Google Scholar] [CrossRef]
- Dürrenberger, M.B.; Villiger, W.; Bächi, T. Conjugational junctions: Morphology of specific contacts in conjugating Escherichia coli bacteria. J. Struct. Biol. 1991, 107, 146–156. [Google Scholar] [CrossRef]
- Brinton, C.C. The structure, function, synthesis and genetic control of bacterial pili and a molecular model for DNA and RNA transport in gram negative bacteria. Trans. N. Y. Acad. Sci. 1965, 27, 1003–1054. [Google Scholar] [CrossRef] [PubMed]
- Harrington, L.C.; Rogerson, A.C. The F pilus of Escherichia coli appears to support stable DNA transfer in the absence of wall-to-wall contact between cells. J. Bacteriol. 1990, 172, 7263–7264. [Google Scholar] [CrossRef] [PubMed]
- Babić, A.; Lindner, A.B.; Vulić, M.; Stewart, E.J.; Radman, M. Direct visualization of horizontal gene transfer. Science 2008, 319, 1533–1536. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.K.; Reeves, P. Colicin tolerance and map location of conjugation-deficient mutants. J. Bacteriol. 1975, 123, 372–373. [Google Scholar] [CrossRef]
- Falkinham, J.O.; Curtiss, R. Isolation and characterization of conjugation-deficient mutants of Escherichia coli K-12. J. Bacteriol. 1976, 126, 1194–1206. [Google Scholar] [CrossRef] [PubMed]
- Havekes, L.M.; Hoekstra, W.P. Characterization of an Escherichia coli K-12 F-Con-mutant. J. Bacteriol. 1976, 126, 593–600. [Google Scholar] [CrossRef]
- Manning, P.A.; Puspurs, A.; Reeves, P. Outer membrane of Escherichia coli K-12: Isolation of mutants with altered protein 3A by using host range mutants of bacteriophage K3. J. Bacteriol. 1976, 127, 1080–1084. [Google Scholar] [CrossRef] [PubMed]
- Monner, D.A.; Jonsson, S.; Boman, H.G. Ampicillin-Resistant mutants of Escherichia coli K-12 with lipopolysaccharide alterations affecting mating ability and susceptibility to sex-specific bacteriophages. J. Bacteriol. 1971, 107, 420–432. [Google Scholar] [CrossRef]
- Skurray, R.A.; Hancock, R.E.; Reeves, P. Con--mutants: Class of mutants in Escherichia coli K-12 lacking a major cell wall protein and defective in conjugation and adsorption of a bacteriophage. J. Bacteriol. 1974, 119, 726–735. [Google Scholar] [CrossRef]
- Manoil, C.; Rosenbusch, J.P. Conjugation-deficient mutants of Escherichia coli distinguish classes of functions of the outer membrane OmpA protein. Mol. Gen. Genet. 1982, 187, 148–156. [Google Scholar] [CrossRef]
- Morona, R.; Klose, M.; Henning, U. Escherichia coli K-12 outer membrane protein (OmpA) as a bacteriophage receptor: Analysis of mutant genes expressing altered proteins. J. Bacteriol. 1984, 159, 570–578. [Google Scholar] [CrossRef] [PubMed]
- Achtman, M.; Schwuchow, S.; Helmuth, R.; Morelli, G.; Manning, P.A. Cell-Cell interactions in conjugating Escherichia coli: Con− mutants and stabilization of mating aggregates. Mol. Gen. Genet. MGG 1978, 164, 171–183. [Google Scholar] [CrossRef]
- Anthony, K.G.; Sherburne, C.; Sherburne, R.; Frost, L.S. The role of the pilus in recipient cell recognition during bacterial conjugation mediated by F-like plasmids. Mol. Microbiol. 1994, 13, 939–953. [Google Scholar] [CrossRef] [PubMed]
- Klimke, W.A.; Frost, L.S. Genetic analysis of the role of the transfer gene, traN, of the F and R100-1 plasmids in mating pair stabilization during conjugation. J. Bacteriol. 1998, 180, 4036–4043. [Google Scholar] [CrossRef]
- Klimke, W.A.; Rypien, C.D.; Klinger, B.; Kennedy, R.A.; Rodriguez-Maillard, J.M.; Frost, L.S. The mating pair stabilization protein, TraN, of the F plasmid is an outer-membrane protein with two regions that are important for its function in conjugation. Microbiology 2005, 151, 3527–3540. [Google Scholar] [CrossRef] [PubMed]
- Maneewannakul, S.; Kathir, P.; Ippen-Ihler, K. Characterization of the F plasmid mating aggregation gene traN and of a new F transfer region locus trbE. J. Mol. Biol. 1992, 225, 299–311. [Google Scholar] [CrossRef]
- Willetts, N.; Achtman, M. Genetic analysis of transfer by the Escherichia coli sex factor F, using P1 transductional complementation. J. Bacteriol. 1972, 110, 843–851. [Google Scholar] [CrossRef]
- Audette, G.F.; Manchak, J.; Beatty, P.; Klimke, W.A.; Frost, L.S. Entry exclusion in F-like plasmids requires intact TraG in the donor that recognizes its cognate TraS in the recipient. Microbiology (Read. Engl.) 2007, 153, 442–451. [Google Scholar] [CrossRef]
- Marrero, J.; Waldor, M.K. Determinants of entry exclusion within Eex and TraG are cytoplasmic. J. Bacteriol. 2007, 189, 6469–6473. [Google Scholar] [CrossRef]
- Low, W.W.; Wong, J.; Pena, A.; Seddon, C.; Costa, T.; Beis, K.; Frankel, G. OmpK36 and TraN facilitate conjugal transfer of the Klebsiella pneumoniae carbapenem resistance plasmid pKpQIL. Microbiology 2020. [Google Scholar] [CrossRef]
- Havekes, L.; Tommassen, J.; Hoekstra, W.; Lugtenberg, B. Isolation and characterization of Escherichia coli K-12 F- mutants defective in conjugation with an I-type donor. J. Bacteriol. 1977, 129, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Ishiwa, A.; Komano, T. PilV Adhesins of Plasmid R64 Thin Pili specifically bind to the lipopolysaccharides of recipient cells. J. Mol. Biol. 2004, 343, 615–625. [Google Scholar] [CrossRef] [PubMed]
- Frost, L.S.; Simon, J. Studies on the pili of the promiscuous plasmid RP4. In Molecular Mechanisms of Bacterial Virulence; Kado, C.I., Crosa, J.H., Eds.; Developments in Plant Pathology; Springer: Dordrecht, The Netherlands, 1994; Volume 3, pp. 47–65. ISBN 978-94-010-4322-9. [Google Scholar]
- Moriguchi, K.; Zoolkefli, F.I.R.M.; Abe, M.; Kiyokawa, K.; Yamamoto, S.; Suzuki, K. Targeting antibiotic resistance genes is a better approach to block acquisition of antibiotic resistance than blocking conjugal transfer by recipient cells: A genome-wide screening in Escherichia coli. Front. Microbiol. 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Mendoza, D.; de la Cruz, F. Escherichia coli genes affecting recipient ability in plasmid conjugation: Are there any? BMC Genom. 2009, 10, 71. [Google Scholar] [CrossRef]
- Dodsworth, J.A.; Li, L.; Wei, S.; Hedlund, B.P.; Leigh, J.A.; de Figueiredo, P. Interdomain conjugal transfer of DNA from bacteria to Archaea. Appl. Environ. Microbiol. 2010, 76, 5644–5647. [Google Scholar] [CrossRef]
- Heinemann, J.A.; Sprague, G.F. Bacterial conjugative plasmids mobilize DNA transfer between bacteria and yeast. Nature 1989, 340, 205–209. [Google Scholar] [CrossRef]
- Waters, V.L. Conjugation between bacterial and mammalian cells. Nat. Genet. 2001, 29, 375–376. [Google Scholar] [CrossRef]
- Llosa, M.; Gomis-Rüth, F.X.; Coll, M.; de la Cruz Fd, F. Bacterial conjugation: A two-step mechanism for DNA transport. Mol. Microbiol. 2002, 45, 1–8. [Google Scholar] [CrossRef]
- Clewell, D.B.; Helinski, D.E. Existence of the colicinogenic factor-sex factor ColI-b-P9 as a supercoiled circular DNA-protein relaxation complex. Biochem. Biophys. Res. Commun. 1970, 41, 150–156. [Google Scholar] [CrossRef]
- Cohen, A.; Fisher, W.D.; Curtiss, R.; Adler, H.I. DNA isolated from Escherichia coli minicells mated with F+ cells. Proc. Natl. Acad. Sci. USA 1968, 61, 61–68. [Google Scholar] [CrossRef]
- Willetts, N.; Skurray, R. The conjugation system of F-like plasmids. Annu. Rev. Genet. 1980, 14, 41–76. [Google Scholar] [CrossRef] [PubMed]
- Dostál, L.; Schildbach, J.F. Single-Stranded DNA binding by F TraI relaxase and helicase domains is coordinately regulated. J. Bacteriol. 2010, 192, 3620–3628. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Everett, R.; Willetts, N. Characterisation of an In Vivo system for nicking at the origin of conjugal DNA transfer of the sex factor F. J. Mol. Biol. 1980, 136, 129–150. [Google Scholar] [CrossRef]
- Matson, S.W.; Morton, B.S. Escherichia coli DNA helicase I catalyzes a site- and strand-specific nicking reaction at the F plasmid oriT. J. Biol. Chem. 1991, 266, 16232–16237. [Google Scholar]
- Matson, S.W.; Ragonese, H. The F-plasmid TraI protein contains three functional domains required for conjugative DNA strand transfer. J. Bacteriol. 2005, 187, 697–706. [Google Scholar] [CrossRef]
- Reygers, U.; Wessel, R.; Müller, H.; Hoffmann-Berling, H. Endonuclease activity of Escherichia coli DNA helicase I directed against the transfer origin of the F factor. EMBO J. 1991, 10, 2689–2694. [Google Scholar] [CrossRef]
- Traxler, B.A.; Minkley, E.G. Evidence that DNA helicase I and oriT site-specific nicking are both functions of the F TraI protein. J. Mol. Biol. 1988, 204, 205–209. [Google Scholar] [CrossRef]
- Howard, M.T.; Nelson, W.C.; Matson, S.W. Stepwise assembly of a relaxosome at the F plasmid origin of transfer. J. Biol. Chem. 1995, 270, 28381–28386. [Google Scholar]
- Nelson, W.C.; Morton, B.S.; Lahue, E.E.; Matson, S.W. Characterization of the Escherichia coli F factor traY gene product and its binding sites. J. Bacteriol. 1993, 175, 2221–2228. [Google Scholar] [CrossRef][Green Version]
- Schildbach, J.F.; Robinson, C.R.; Sauer, R.T. Biophysical characterization of the TraY protein of Escherichia coli F factor. J. Biol. Chem. 1998, 273, 1329–1333. [Google Scholar] [CrossRef]
- Luo, Y.; Gao, Q.; Deonier, R.C. Mutational and physical analysis of F plasmid traY protein binding to oriT. Mol. Microbiol. 1994, 11, 459–469. [Google Scholar] [CrossRef] [PubMed]
- Nelson, W.C.; Howard, M.T.; Sherman, J.A.; Matson, S.W. The traY gene product and integration host factor stimulate Escherichia coli DNA helicase I-catalyzed nicking at the F plasmid oriT. J. Biol. Chem. 1995, 270, 28374–28380. [Google Scholar] [PubMed]
- Rice, P.A.; Yang, S.; Mizuuchi, K.; Nash, H.A. Crystal structure of an IHF-DNA complex: A protein-induced DNA U-turn. Cell 1996, 87, 1295–1306. [Google Scholar] [CrossRef]
- Tsai, M.M.; Fu, Y.H.; Deonier, R.C. Intrinsic bends and integration host factor binding at F plasmid oriT. J. Bacteriol. 1990, 172, 4603–4609. [Google Scholar] [CrossRef]
- Williams, S.L.; Schildbach, J.F. TraY and integration host factor oriT binding sites and F conjugal transfer: Sequence variations, but not altered spacing, are tolerated. J. Bacteriol. 2007, 189, 3813–3823. [Google Scholar] [CrossRef]
- Di Laurenzio, L.; Frost, L.S.; Paranchych, W. The TraM protein of the conjugative plasmid F binds to the origin of transfer of the F and ColE1 plasmids. Mol. Microbiol. 1992, 6, 2951–2959. [Google Scholar] [CrossRef]
- Fu, Y.H.; Tsai, M.M.; Luo, Y.N.; Deonier, R.C. Deletion analysis of the F plasmid oriT locus. J. Bacteriol. 1991, 173, 1012–1020. [Google Scholar] [CrossRef]
- Kupelwieser, G.; Schwab, M.; Högenauer, G.; Koraimann, G.; Zechner, E.L. Transfer protein TraM stimulates TraI-catalyzed cleavage of the transfer origin of plasmid R1 In Vivo. J. Mol. Biol. 1998, 275, 81–94. [Google Scholar] [CrossRef]
- Mihajlovic, S.; Lang, S.; Sut, M.V.; Strohmaier, H.; Gruber, C.J.; Koraimann, G.; Cabezón, E.; Moncalián, G.; de la Cruz, F.; Zechner, E.L. Plasmid r1 conjugative DNA processing is regulated at the coupling protein interface. J. Bacteriol. 2009, 191, 6877–6887. [Google Scholar] [CrossRef] [PubMed]
- Ragonese, H.; Haisch, D.; Villareal, E.; Choi, J.-H.; Matson, S.W. The F plasmid-encoded TraM protein stimulates relaxosome-mediated cleavage at oriT through an interaction with TraI. Mol. Microbiol. 2007, 63, 1173–1184. [Google Scholar] [CrossRef]
- Wawrzyniak, P.; Płucienniczak, G.; Bartosik, D. The different faces of rolling-circle replication and its multifunctional initiator proteins. Front. Microbiol. 2017, 8, 2353. [Google Scholar] [CrossRef] [PubMed]
- Byrd, D.R.; Matson, S.W. Nicking by transesterification: The reaction catalysed by a relaxase. Mol. Microbiol. 1997, 25, 1011–1022. [Google Scholar] [CrossRef] [PubMed]
- Guiney, D.G.; Helinski, D.R. Relaxation complexes of poasmid DNA and protein. III. Association of protein with the 5′ terminus of the broken DNA strand in the relaxed complex of plasmid ColE1. J. Biol. Chem. 1975, 250, 8796–8803. [Google Scholar]
- Matson, S.W.; Nelson, W.C.; Morton, B.S. Characterization of the reaction product of the oriT nicking reaction catalyzed by Escherichia coli DNA helicase I. J. Bacteriol. 1993, 175, 2599–2606. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pansegrau, W.; Ziegelin, G.; Lanka, E. Covalent association of the traI gene product of plasmid RP4 with the 5′-terminal nucleotide at the relaxation nick site. J. Biol. Chem. 1990, 265, 10637–10644. [Google Scholar] [PubMed]
- Khan, S.A. Rolling-Circle replication of bacterial plasmids. Microbiol. Mol. Biol. Rev. 1997, 61, 442–455. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.A. Plasmid rolling-circle replication: Highlights of two decades of research. Plasmid 2005, 53, 126–136. [Google Scholar] [CrossRef]
- Ruiz-Masó, J.A.; MachóN, C.; Bordanaba-Ruiseco, L.; Espinosa, M.; Coll, M.; Del Solar, G. Plasmid rolling-circle replication. Microbiol. Spectr. 2015, 3. [Google Scholar] [CrossRef]
- Waters, V.L.; Guiney, D.G. Processes at the nick region link conjugation, T-DNA transfer and rolling circle replication. Mol. Microbiol. 1993, 9, 1123–1130. [Google Scholar] [CrossRef]
- Lorenzo-Díaz, F.; Fernández-López, C.; Garcillán-Barcia, M.P.; Espinosa, M. Bringing them together: Plasmid pMV158 rolling circle replication and conjugation under an evolutionary perspective. Plasmid 2014, 74, 15–31. [Google Scholar] [CrossRef]
- Draper, O.; César, C.E.; Machón, C.; de la Cruz, F.; Llosa, M. Site-Specific recombinase and integrase activities of a conjugative relaxase in recipient cells. Proc. Natl. Acad. Sci. USA 2005, 102, 16385–16390. [Google Scholar] [CrossRef] [PubMed]
- Trokter, M.; Waksman, G. Translocation through the conjugative type IV secretion system requires unfolding of its protein substrate. J. Bacteriol. 2018, 200. [Google Scholar] [CrossRef] [PubMed]
- Ilangovan, A.; Kay, C.W.M.; Roier, S.; El Mkami, H.; Salvadori, E.; Zechner, E.L.; Zanetti, G.; Waksman, G. Cryo-EM structure of a relaxase reveals the molecular basis of DNA unwinding during bacterial conjugation. Cell 2017, 169, 708–721.e12. [Google Scholar] [CrossRef] [PubMed]
- Cabezón, E.; Sastre, J.I.; de la Cruz, F. Genetic evidence of a coupling role for the TraG protein family in bacterial conjugation. Mol. Gen. Genet. 1997, 254, 400–406. [Google Scholar] [CrossRef] [PubMed]
- Cascales, E.; Christie, P.J. Definition of a bacterial type IV secretion pathway for a DNA substrate. Science 2004, 304, 1170–1173. [Google Scholar] [CrossRef] [PubMed]
- Christie, P.J. Type IV secretion: The Agrobacterium VirB/D4 and related conjugation systems. Biochim. Biophys. Acta 2004, 1694, 219–234. [Google Scholar] [CrossRef]
- Christie, P.J. The mosaic type IV secretion systems. EcoSal Plus 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Christie, P.J.; Whitaker, N.; González-Rivera, C. BBA review revised mechanism and structure of the bacterial type IV secretion systems. Biochim. Biophys. Acta 2014, 1843, 1578. [Google Scholar] [CrossRef]
- Beranek, A.; Zettl, M.; Lorenzoni, K.; Schauer, A.; Manhart, M.; Koraimann, G. Thirty-Eight C-terminal amino acids of the coupling protein TraD of the F-like conjugative resistance plasmid R1 are required and sufficient to confer binding to the substrate selector protein TraM. J. Bacteriol. 2004, 186, 6999–7006. [Google Scholar] [CrossRef][Green Version]
- Disqué-Kochem, C.; Dreiseikelmann, B. The cytoplasmic DNA-binding protein TraM binds to the inner membrane protein TraD In Vitro. J. Bacteriol. 1997, 179, 6133–6137. [Google Scholar] [CrossRef][Green Version]
- Lu, J.; Frost, L.S. Mutations in the C-terminal region of TraM provide evidence for In Vivo TraM-TraD interactions during F-plasmid conjugation. J. Bacteriol. 2005, 187, 4767–4773. [Google Scholar] [CrossRef][Green Version]
- Lu, J.; Zhao, W.; Frost, L.S. Mutational analysis of TraM correlates oligomerization and DNA binding with autoregulation and conjugative DNA transfer. J. Biol. Chem. 2004, 279, 55324–55333. [Google Scholar] [CrossRef]
- Lu, J.; Wong, J.J.W.; Edwards, R.A.; Manchak, J.; Frost, L.S.; Glover, J.N.M. Structural basis of specific TraD-TraM recognition during F plasmid-mediated bacterial conjugation. Mol. Microbiol. 2008, 70, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Schröder, G.; Krause, S.; Zechner, E.L.; Traxler, B.; Yeo, H.-J.; Lurz, R.; Waksman, G.; Lanka, E. TraG-like proteins of DNA transfer systems and of the Helicobacter pylori type IV secretion system: Inner membrane gate for exported substrates? J. Bacteriol. 2002, 184, 2767–2779. [Google Scholar] [CrossRef] [PubMed]
- Llosa, M.; Zunzunegui, S.; de la Cruz, F. Conjugative coupling proteins interact with cognate and heterologous VirB10-like proteins while exhibiting specificity for cognate relaxosomes. Proc. Natl. Acad. Sci. USA 2003, 100, 10465–10470. [Google Scholar] [CrossRef]
- Haft, R.J.F.; Palacios, G.; Nguyen, T.; Mally, M.; Gachelet, E.G.; Zechner, E.L.; Traxler, B. General mutagenesis of F plasmid TraI reveals its role in conjugative regulation. J. Bacteriol. 2006, 188, 6346–6353. [Google Scholar] [CrossRef][Green Version]
- Gilmour, M.W.; Gunton, J.E.; Lawley, T.D.; Taylor, D.E. Interaction between the IncHI1 plasmid R27 coupling protein and type IV secretion system: TraG associates with the coiled-coil mating pair formation protein TrhB. Mol. Microbiol. 2003, 49, 105–116. [Google Scholar] [CrossRef]
- Egelman, E.H. Structural biology. Pumping DNA. Nature 2001, 409, 573–575. [Google Scholar] [CrossRef]
- Gomis-Rüth, F.X.; Moncalián, G.; Pérez-Luque, R.; González, A.; Cabezón, E.; de la Cruz, F.; Coll, M. The bacterial conjugation protein TrwB resembles ring helicases and F1-ATPase. Nature 2001, 409, 637–641. [Google Scholar] [CrossRef] [PubMed]
- Moncalián, G.; Cabezón, E.; Alkorta, I.; Valle, M.; Moro, F.; Valpuesta, J.M.; Goñi, F.M.; de La Cruz, F. Characterization of ATP and DNA binding activities of TrwB, the coupling protein essential in plasmid R388 conjugation. J. Biol. Chem. 1999, 274, 36117–36124. [Google Scholar] [CrossRef][Green Version]
- Schröder, G.; Lanka, E. TraG-like proteins of type IV secretion systems: Functional dissection of the multiple activities of TraG (RP4) and TrwB (R388). J. Bacteriol. 2003, 185, 4371–4381. [Google Scholar] [CrossRef] [PubMed]
- Haft, R.J.F.; Gachelet, E.G.; Nguyen, T.; Toussaint, L.; Chivian, D.; Traxler, B. In Vivo oligomerization of the F conjugative coupling protein TraD. J. Bacteriol. 2007, 189, 6626–6634. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Tato, I.; Zunzunegui, S.; de la Cruz, F.; Cabezon, E. TrwB, the coupling protein involved in DNA transport during bacterial conjugation, is a DNA-dependent ATPase. Proc. Natl. Acad. Sci. USA 2005, 102, 8156–8161. [Google Scholar] [CrossRef] [PubMed]
- Christie, P.J.; Atmakuri, K.; Krishnamoorthy, V.; Jakubowski, S.; Cascales, E. Biogenesis, architecture, and function of bacterial type IV secretion systems. Annu. Rev. Microbiol. 2005, 59, 451–485. [Google Scholar] [CrossRef] [PubMed]
- Garcillán-Barcia, M.P.; Jurado, P.; González-Pérez, B.; Moncalián, G.; Fernández, L.A.; de la Cruz, F. Conjugative transfer can be inhibited by blocking relaxase activity within recipient cells with intrabodies. Mol. Microbiol. 2007, 63, 404–416. [Google Scholar] [CrossRef] [PubMed]
- Chandler, M.; de la Cruz, F.; Dyda, F.; Hickman, A.B.; Moncalian, G.; Ton-Hoang, B. Breaking and joining single-stranded DNA: The HUH endonuclease superfamily. Nat. Rev. Microbiol. 2013, 11, 525–538. [Google Scholar] [CrossRef]
- Becker, E.C.; Meyer, R. Origin and fate of the 3′ ends of single-stranded DNA generated by conjugal transfer of plasmid R1162. J. Bacteriol. 2012, 194, 5368–5376. [Google Scholar] [CrossRef][Green Version]
- Forsberg, K.J.; Malik, H.S. Microbial genomics: The expanding universe of bacterial defense systems. Curr. Biol. 2018, 28, R361–R364. [Google Scholar] [CrossRef]
- Casjens, S. Prophages and bacterial genomics: What have we learned so far? Mol. Microbiol. 2003, 49, 277–300. [Google Scholar] [CrossRef]
- Narra, H.P.; Ochman, H. Of what use is sex to bacteria? Curr. Biol. 2006, 16, R705–R710. [Google Scholar] [CrossRef]
- Johnston, C.D.; Cotton, S.L.; Rittling, S.R.; Starr, J.R.; Borisy, G.G.; Dewhirst, F.E.; Lemon, K.P. Systematic evasion of the restriction-modification barrier in bacteria. Proc. Natl. Acad. Sci. USA 2019, 116, 11454–11459. [Google Scholar] [CrossRef] [PubMed]
- Bickle, T.A. Restricting restriction. Mol. Microbiol. 2004, 51, 3–5. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Reuter, M.; Sanghvi, B.; Roberts, G.A.; Cooper, L.P.; Tilling, M.; Blakely, G.W.; Dryden, D.T.F. ArdA proteins from different mobile genetic elements can bind to the EcoKI Type I DNA methyltransferase of E. coli K12. Biochim. Biophys. Acta 2014, 1844, 505–511. [Google Scholar] [CrossRef] [PubMed]
- McMahon, S.A.; Roberts, G.A.; Johnson, K.A.; Cooper, L.P.; Liu, H.; White, J.H.; Carter, L.G.; Sanghvi, B.; Oke, M.; Walkinshaw, M.D.; et al. Extensive DNA mimicry by the ArdA anti-restriction protein and its role in the spread of antibiotic resistance. Nucleic Acids Res. 2009, 37, 4887–4897. [Google Scholar] [CrossRef]
- Wilkins, B.M. Plasmid promiscuity: Meeting the challenge of DNA immigration control. Environ. Microbiol. 2002, 4, 495–500. [Google Scholar] [CrossRef]
- Belogurov, A.A.; Delver, E.P.; Agafonova, O.V.; Belogurova, N.G.; Lee, L.Y.; Kado, C.I. Antirestriction protein Ard (Type C) encoded by IncW plasmid pSa has a high similarity to the “protein transport” domain of TraC1 primase of promiscuous plasmid RP4. J. Mol. Biol. 2000, 296, 969–977. [Google Scholar] [CrossRef]
- Roy, D.; Huguet, K.T.; Grenier, F.; Burrus, V. IncC conjugative plasmids and SXT/R391 elements repair double-strand breaks caused by CRISPR-Cas during conjugation. Nucleic Acids Res. 2020, 8815–8827. [Google Scholar] [CrossRef]
- Wilkins, B.M.; Chilley, P.M.; Thomas, A.T.; Pocklington, M.J. Distribution of restriction enzyme recognition sequences on broad host range plasmid RP4: Molecular and evolutionary implications. J. Mol. Biol. 1996, 258, 447–456. [Google Scholar] [CrossRef]
- Grissa, I.; Vergnaud, G.; Pourcel, C. The CRISPRdb database and tools to display CRISPRs and to generate dictionaries of spacers and repeats. BMC Bioinform. 2007, 8, 172. [Google Scholar] [CrossRef]
- Makarova, K.S.; Wolf, Y.I.; Alkhnbashi, O.S.; Costa, F.; Shah, S.A.; Saunders, S.J.; Barrangou, R.; Brouns, S.J.J.; Charpentier, E.; Haft, D.H.; et al. An updated evolutionary classification of CRISPR-Cas systems. Nat. Rev. Microbiol. 2015, 13, 722–736. [Google Scholar] [CrossRef]
- Garneau, J.E.; Dupuis, M.-È.; Villion, M.; Romero, D.A.; Barrangou, R.; Boyaval, P.; Fremaux, C.; Horvath, P.; Magadán, A.H.; Moineau, S. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 2010, 468, 67–71. [Google Scholar] [CrossRef] [PubMed]
- Bondy-Denomy, J.; Pawluk, A.; Maxwell, K.L.; Davidson, A.R. Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature 2013, 493, 429–432. [Google Scholar] [CrossRef] [PubMed]
- Bondy-Denomy, J.; Garcia, B.; Strum, S.; Du, M.; Rollins, M.F.; Hidalgo-Reyes, Y.; Wiedenheft, B.; Maxwell, K.L.; Davidson, A.R. Multiple mechanisms for CRISPR-Cas inhibition by anti-CRISPR proteins. Nature 2015, 526, 136–139. [Google Scholar] [CrossRef] [PubMed]
- Mahendra, C.; Christie, K.A.; Osuna, B.A.; Pinilla-Redondo, R.; Kleinstiver, B.P.; Bondy-Denomy, J. Broad-Spectrum anti-CRISPR proteins facilitate horizontal gene transfer. Nat. Microbiol. 2020, 5, 620–629. [Google Scholar] [CrossRef] [PubMed]
- Cram, D.; Ray, A.; O’Gorman, L.; Skurray, R. Transcriptional analysis of the leading region in F plasmid DNA transfer. Plasmid 1984, 11, 221–233. [Google Scholar] [CrossRef]
- Jones, A.L.; Barth, P.T.; Wilkins, B.M. Zygotic induction of plasmid ssb and psiB genes following conjugative transfer of Incl1 plasmid Collb-P9. Mol. Microbiol. 1992, 6, 605–613. [Google Scholar] [CrossRef] [PubMed]
- Althorpe, N.J.; Chilley, P.M.; Thomas, A.T.; Brammar, W.J.; Wilkins, B.M. Transient transcriptional activation of the Incl1 plasmid anti-restriction gene (ardA) and SOS inhibition gene (psiB) early in conjugating recipient bacteria. Mol. Microbiol. 1999, 31, 133–142. [Google Scholar] [CrossRef]
- Masai, H.; Arai, K. Frpo: A novel single-stranded DNA promoter for transcription and for primer RNA synthesis of DNA replication. Cell 1997, 89, 897–907. [Google Scholar] [CrossRef]
- Honda, Y.; Sakai, H.; Komano, T. Two single-strand DNA initiation signals located in the oriV region of plasmid RSF1010. Gene 1988, 68, 221–228. [Google Scholar] [CrossRef]
- Masai, H.; Arai, K. Mechanisms of primer RNA synthesis and D-loop/R-loop-dependent DNA replication in Escherichia coli. Biochimie 1996, 78, 1109–1117. [Google Scholar] [CrossRef]
- Nomura, N.; Low, R.L.; Ray, D.S. Selective cloning of Co1E1 DNA initiation sequences using the cloning vector M13 delta E101. Gene 1982, 18, 239–246. [Google Scholar] [CrossRef]
- Nomura, N.; Masai, H.; Inuzuka, M.; Miyazaki, C.; Ohtsubo, E.; Itoh, T.; Sasamoto, S.; Matsui, M.; Ishizaki, R.; Arai, K. Identification of eleven single-strand initiation sequences (ssi) for priming of DNA replication in the F, R6K, R100 and ColE2 plasmids. Gene 1991, 108, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Díaz, A.; Lacks, S.A.; López, P. Multiple roles for DNA polymerase I in establishment and replication of the promiscuous plasmid pLS1. Mol. Microbiol. 1994, 14, 773–783. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, B.M.; Hollom, S.E. Conjugational synthesis of F lac+ and Col I DNA in the presence of rifampicin and in Escherichia coli K12 mutants defective in DNA synthesis. Mol. Gen. Genet. 1974, 134, 143–156. [Google Scholar] [CrossRef]
- Baharoglu, Z.; Mazel, D. SOS, the formidable strategy of bacteria against aggressions. FEMS Microbiol. Rev. 2014, 38, 1126–1145. [Google Scholar] [CrossRef]
- Baharoglu, Z.; Bikard, D.; Mazel, D. Conjugative DNA transfer induces the bacterial SOS response and promotes antibiotic resistance development through Integron activation. PLoS Genet. 2010, 6, e1001165. [Google Scholar] [CrossRef]
- Maslowska, K.H.; Makiela-Dzbenska, K.; Fijalkowska, I.J. The SOS system: A complex and tightly regulated response to DNA damage. Environ. Mol. Mutagen. 2019, 60, 368–384. [Google Scholar] [CrossRef]
- Golub, E.; Bailone, A.; Devoret, R. A gene encoding an SOS inhibitor is present in different conjugative plasmids. J. Bacteriol. 1988, 170, 4392–4394. [Google Scholar] [CrossRef]
- Bailone, A.; Bäckman, A.; Sommer, S.; Célérier, J.; Bagdasarian, M.M.; Bagdasarian, M.; Devoret, R. PsiB polypeptide prevents activation of RecA protein in Escherichia coli. Mol. Gen. Genet. 1988, 214, 389–395. [Google Scholar] [CrossRef]
- Petrova, V.; Chitteni-Pattu, S.; Drees, J.C.; Inman, R.B.; Cox, M.M. An SOS inhibitor that binds to free RecA protein: The PsiB protein. Mol. Cell 2009, 36, 121–130. [Google Scholar] [CrossRef]
- Meyer, R.R.; Laine, P.S. The single-stranded DNA-binding protein of Escherichia coli. Microbiol. Rev. 1990, 54, 342–380. [Google Scholar] [CrossRef] [PubMed]
- Shereda, R.D.; Kozlov, A.G.; Lohman, T.M.; Cox, M.M.; Keck, J.L. SSB as an organizer/mobilizer of genome maintenance complexes. Crit. Rev. Biochem. Mol. Biol. 2008, 43, 289–318. [Google Scholar] [CrossRef]
- Sigal, N.; Delius, H.; Kornberg, T.; Gefter, M.L.; Alberts, B. A DNA-unwinding protein isolated from Escherichia coli: Its interaction with DNA and with DNA polymerases. Proc. Natl. Acad. Sci. USA 1972, 69, 3537–3541. [Google Scholar] [CrossRef] [PubMed]
- Nolivos, S.; Cayron, J.; Dedieu, A.; Page, A.; Delolme, F.; Lesterlin, C. Role of AcrAB-TolC multidrug efflux pump in drug-resistance acquisition by plasmid transfer. Science 2019, 364, 778–782. [Google Scholar] [CrossRef]
- Golub, E.I.; Low, K.B. Conjugative plasmids of enteric bacteria from many different incompatibility groups have similar genes for single-stranded DNA-binding proteins. J. Bacteriol. 1985, 162, 235–241. [Google Scholar] [CrossRef] [PubMed]
- Kolodkin, A.L.; Capage, M.A.; Golub, E.I.; Low, K.B. F sex factor of Escherichia coli K-12 codes for a single-stranded DNA binding protein. Proc. Natl. Acad. Sci. USA 1983, 80, 4422–4426. [Google Scholar] [CrossRef]
- Golub, E.I.; Low, K.B. Derepression of single-stranded DNA-binding protein genes on plasmids derepressed for conjugation, and complementation of an E. coli ssb-mutation by these genes. Mol. Gen. Genet. 1986, 204, 410–416. [Google Scholar] [CrossRef]
- Howland, C.J.; Rees, C.E.; Barth, P.T.; Wilkins, B.M. The ssb gene of plasmid ColIb-P9. J. Bacteriol. 1989, 171, 2466–2473. [Google Scholar] [CrossRef]
- Porter, R.D.; Black, S. The single-stranded-DNA-binding protein encoded by the Escherichia coli F factor can complement a deletion of the chromosomal ssb gene. J. Bacteriol. 1991, 173, 2720–2723. [Google Scholar] [CrossRef]
- Ruvolo, P.P.; Keating, K.M.; Williams, K.R.; Chase, J.W. Single-Stranded DNA binding proteins (SSBs) from prokaryotic transmissible plasmids. Proteins 1991, 9, 120–134. [Google Scholar] [CrossRef]
- Jovanovic, O.S.; Ayres, E.K.; Figurski, D.H. The replication initiator operon of promiscuous plasmid RK2 encodes a gene that complements an Escherichia coli mutant defective in single-stranded DNA-binding protein. J. Bacteriol. 1992, 174, 4842–4846. [Google Scholar] [CrossRef] [PubMed]
- del Solar, G.; Giraldo, R.; Ruiz-Echevarría, M.J.; Espinosa, M.; Díaz-Orejas, R. Replication and control of circular bacterial plasmids. Microbiol. Mol. Biol. Rev. 1998, 62, 434–464. [Google Scholar] [CrossRef]
- Pinto, U.M.; Pappas, K.M.; Winans, S.C. The ABCs of plasmid replication and segregation. Nat. Rev. Microbiol. 2012, 10, 755–765. [Google Scholar] [CrossRef] [PubMed]
- Rakowski, S.A.; Filutowicz, M. Plasmid R6K replication control. Plasmid 2013, 69, 231–242. [Google Scholar] [CrossRef] [PubMed]
- Nordström, K. Plasmid R1--replication and its control. Plasmid 2006, 55, 1–26. [Google Scholar] [CrossRef]
- Guiney, D.G. Host range of conjugation and replication functions of the Escherichia coli sex plasmid Flac. J. Mol. Biol. 1982, 162, 699–703. [Google Scholar] [CrossRef]
- Zhong, Z.; Helinski, D.; Toukdarian, A. Plasmid host-range: Restrictions to F replication in Pseudomonas. Plasmid 2005, 54, 48–56. [Google Scholar] [CrossRef]
- Adamczyk, M.; Jagura-Burdzy, G. Spread and survival of promiscuous IncP-1 plasmids. Acta Biochim. Pol. 2003, 50, 425–453. [Google Scholar] [CrossRef]
- Baxter, J.C.; Funnell, B.E. Plasmid partition mechanisms. Microbiol. Spectr. 2014, 2. [Google Scholar] [CrossRef]
- Ebersbach, G.; Gerdes, K. Plasmid segregation mechanisms. Annu. Rev. Genet. 2005, 39, 453–479. [Google Scholar] [CrossRef]
- Gerdes, K.; Howard, M.; Szardenings, F. Pushing and pulling in prokaryotic DNA segregation. Cell 2010, 141, 927–942. [Google Scholar] [CrossRef] [PubMed]
- Møller-Jensen, J.; Jensen, R.B.; Gerdes, K. Plasmid and chromosome segregation in prokaryotes. Trends Microbiol. 2000, 8, 313–320. [Google Scholar] [CrossRef]
- Schumacher, M.A. Bacterial plasmid partition machinery: A minimalist approach to survival. Curr. Opin. Struct. Biol. 2012, 22, 72–79. [Google Scholar] [CrossRef]
- Deonier, R.C.; Davidson, N. The sequence organization of the integrated F plasmid in two Hfr strains of Escherichia coli. J. Mol. Biol. 1976, 107, 207–222. [Google Scholar] [CrossRef]
- Deonier, R.C.; Mirels, L. Excision of F plasmid sequences by recombination at directly repeated insertion sequence 2 elements: Involvement of recA. Proc. Natl. Acad. Sci. USA 1977, 74, 3965–3969. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.M.; Grossman, A.D. Integrative and Conjugative Elements (ICEs): What they do and how they work. Annu. Rev. Genet. 2015, 49, 577–601. [Google Scholar] [CrossRef]
- Shao, Q.; Hawkins, A.; Zeng, L. Phage DNA dynamics in cells with different fates. Biophys. J. 2015, 108, 2048–2060. [Google Scholar] [CrossRef] [PubMed]
- Gago-Córdoba, C.; Val-Calvo, J.; Miguel-Arribas, A.; Serrano, E.; Singh, P.K.; Abia, D.; Wu, L.J.; Meijer, W.J.J. Surface exclusion revisited: Function related to differential expression of the surface exclusion system of Bacillus subtilis Plasmid pLS20. Front. Microbiol. 2019, 10, 1502. [Google Scholar] [CrossRef] [PubMed]
- Galli, D.M.; Chen, J. Entry exclusion activity on conjugative plasmid pVT745. Plasmid 2006, 55, 158–163. [Google Scholar] [CrossRef]
- Gunton, J.E.; Ussher, J.E.R.; Rooker, M.M.; Wetsch, N.M.; Alonso, G.; Taylor, D.E. Entry exclusion in the IncHI1 plasmid R27 is mediated by EexA and EexB. Plasmid 2008, 59, 86–101. [Google Scholar] [CrossRef]
- Haase, J.; Kalkum, M.; Lanka, E. TrbK, a small cytoplasmic membrane lipoprotein, functions in entry exclusion of the IncP α plasmid RP4. J. Bacteriol. 1996, 178, 6720–6729. [Google Scholar] [CrossRef] [PubMed]
- Humbert, M.; Huguet, K.T.; Coulombe, F.; Burrus, V. Entry exclusion of conjugative Plasmids of the IncA, IncC, and related untyped incompatibility groups. J. Bacteriol. 2019, 201. [Google Scholar] [CrossRef]
- Kraushaar, B.; Appel, B.; Lanka, E.; Strauch, E. Entry exclusion and oriT of a conjugative system encoded by the cryptic plasmid p29930 of Yersinia enterocolitica. Plasmid 2010, 64, 79–84. [Google Scholar] [CrossRef] [PubMed]
- Pohlman, R.F.; Genetti, H.D.; Winans, S.C. Entry exclusion of the IncN plasmid pKM101 is mediated by a single hydrophilic protein containing a lipid attachment motif. Plasmid 1994, 31, 158–165. [Google Scholar] [CrossRef]
- Possoz, C.; Gagnat, J.; Sezonov, G.; Guérineau, M.; Pernodet, J.-L. Conjugal immunity of Streptomyces strains carrying the integrative element pSAM2 is due to the pif gene (pSAM2 immunity factor). Mol. Microbiol. 2003, 47, 1385–1393. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.M.; Nielsen, K.M. Mechanisms of, and barriers to, horizontal gene transfer between bacteria. Nat. Rev. Microbiol. 2005, 3, 711–721. [Google Scholar] [CrossRef] [PubMed]
- Achtman, M.; Kennedy, N.; Skurray, R. Cell-Cell interactions in conjugating Escherichia coli: Role of traT protein in surface exclusion. Proc. Natl. Acad. Sci. USA 1977, 74, 5104–5108. [Google Scholar] [CrossRef]
- Manning, P.A.; Beutin, L.; Achtman, M. Outer membrane of Escherichia coli: Properties of the F sex factor traT protein which is involved in surface exclusion. J. Bacteriol. 1980, 142, 285–294. [Google Scholar] [CrossRef]
- Minkley, E.G.; Ippen-Ihler, K. Identification of a membrane protein associated with expression of the surface exclusion region of the F transfer operon. J. Bacteriol. 1977, 129, 1613–1622. [Google Scholar] [CrossRef]
- Riede, I.; Eschbach, M.L. Evidence that TraT interacts with OmpA of Escherichia coli. FEBS Lett. 1986, 205, 241–245. [Google Scholar] [CrossRef]
- Jalajakumari, M.B.; Guidolin, A.; Buhk, H.J.; Manning, P.A.; Ham, L.M.; Hodgson, A.L.; Cheah, K.C.; Skurray, R.A. Surface exclusion genes traS and traT of the F sex factor of Escherichia coli K-12. Determination of the nucleotide sequence and promoter and terminator activities. J. Mol. Biol. 1987, 198, 1–11. [Google Scholar] [CrossRef]
- San Millan, A.; MacLean, R.C. Fitness costs of Plasmids: A limit to Plasmid transmission. Microbiol. Spectr. 2017, 5. [Google Scholar] [CrossRef]
- Barlow, M. What antimicrobial resistance has taught us about horizontal gene transfer. Methods Mol. Biol. 2009, 532, 397–411. [Google Scholar] [CrossRef] [PubMed]
- Hall-Stoodley, L.; Costerton, J.W.; Stoodley, P. Bacterial biofilms: From the natural environment to infectious diseases. Nat. Rev. Microbiol. 2004, 2, 95–108. [Google Scholar] [CrossRef] [PubMed]
- Bale, M.J.; Fry, J.C.; Day, M.J. Plasmid transfer between strains of Pseudomonas aeruginosa on membrane filters attached to river stones. J. Gen. Microbiol. 1987, 133, 3099–3107. [Google Scholar] [CrossRef]
- Lilley, A.K.; Bailey, M.J. Impact of Plasmid pQBR103 acquisition and carriage on the phytosphere fitness of Pseudomonas fluorescens SBW25: Burden and benefit. Appl. Environ. Microbiol. 1997, 63, 1584–1587. [Google Scholar] [CrossRef]
- Munck, C.; Sheth, R.U.; Freedberg, D.E.; Wang, H.H. Recording mobile DNA in the gut microbiota using an Escherichia coli CRISPR-Cas spacer acquisition platform. Nat. Commun. 2020, 11, 95. [Google Scholar] [CrossRef]
- Ronda, C.; Chen, S.P.; Cabral, V.; Yaung, S.J.; Wang, H.H. Metagenomic engineering of the mammalian gut microbiome in situ. Nat. Methods 2019, 16, 167–170. [Google Scholar] [CrossRef]
- Ehlers, L.J.; Bouwer, E.J. RP4 plasmid transfer among species of pseudomonas in a biofilm reactor. Water Sci. Technol. 1999, 39, 163–171. [Google Scholar] [CrossRef]
- Yang, D.; Wang, J.; Qiu, Z.; Jin, M.; Shen, Z.; Chen, Z.; Wang, X.; Zhang, B.; Li, J.-W. Horizontal transfer of antibiotic resistance genes in a membrane bioreactor. J. Biotechnol. 2013, 167, 441–447. [Google Scholar] [CrossRef]
- Hausner, M.; Wuertz, S. High rates of conjugation in bacterial biofilms as determined by quantitative in situ analysis. Appl. Environ. Microbiol. 1999, 65, 3710–3713. [Google Scholar] [CrossRef] [PubMed]
- Christensen, B.B.; Sternberg, C.; Molin, S. Bacterial plasmid conjugation on semi-solid surfaces monitored with the green fluorescent protein (GFP) from Aequorea victoria as a marker. Gene 1996, 173, 59–65. [Google Scholar] [CrossRef]
- Haagensen, J.A.J.; Hansen, S.K.; Johansen, T.; Molin, S. In situ detection of horizontal transfer of mobile genetic elements. FEMS Microbiol. Ecol. 2002, 42, 261–268. [Google Scholar] [CrossRef] [PubMed]
- Reisner, A.; Wolinski, H.; Zechner, E.L. In situ monitoring of IncF plasmid transfer on semi-solid agar surfaces reveals a limited invasion of plasmids in recipient colonies. Plasmid 2012, 67, 155–161. [Google Scholar] [CrossRef]
- Lilley, A.K.; Bailey, M.J. The transfer dynamics of Pseudomonas sp. plasmid pQBR11 in biofilms. FEMS Microbiol. Ecol. 2002, 42, 243–250. [Google Scholar] [CrossRef] [PubMed]
- Aspray, T.J.; Hansen, S.K.; Burns, R.G. A soil-based microbial biofilm exposed to 2,4-D: Bacterial community development and establishment of conjugative plasmid pJP4. FEMS Microbiol. Ecol. 2005, 54, 317–327. [Google Scholar] [CrossRef]
- Christensen, B.B.; Sternberg, C.; Andersen, J.B.; Eberl, L.; Moller, S.; Givskov, M.; Molin, S. Establishment of new genetic traits in a microbial biofilm community. Appl. Environ. Microbiol. 1998, 64, 2247–2255. [Google Scholar] [CrossRef]
- Nancharaiah, Y.V.; Wattiau, P.; Wuertz, S.; Bathe, S.; Mohan, S.V.; Wilderer, P.A.; Hausner, M. Dual labeling of Pseudomonas putida with fluorescent proteins for in situ monitoring of conjugal transfer of the TOL plasmid. Appl. Environ. Microbiol. 2003, 69, 4846–4852. [Google Scholar] [CrossRef]
- Abe, K.; Nomura, N.; Suzuki, S. Biofilms: Hot spots of horizontal gene transfer (HGT) in aquatic environments, with a focus on a new HGT mechanism. FEMS Microbiol. Ecol. 2020, 96. [Google Scholar] [CrossRef]
- Madsen, J.S.; Burmølle, M.; Hansen, L.H.; Sørensen, S.J. The interconnection between biofilm formation and horizontal gene transfer. FEMS Immunol. Med. Microbiol. 2012, 65, 183–195. [Google Scholar] [CrossRef]
- Molin, S.; Tolker-Nielsen, T. Gene transfer occurs with enhanced efficiency in biofilms and induces enhanced stabilisation of the biofilm structure. Curr. Opin. Biotechnol. 2003, 14, 255–261. [Google Scholar] [CrossRef]
- Nesse, L.L.; Simm, R. Biofilm: A hotspot for emerging bacterial genotypes. Adv. Appl. Microbiol. 2018, 103, 223–246. [Google Scholar] [CrossRef] [PubMed]
- Stalder, T.; Top, E. Plasmid transfer in biofilms: A perspective on limitations and opportunities. NPJ Biofilms Microbiomes 2016, 2. [Google Scholar] [CrossRef] [PubMed]
- Fuchs, F.M.; Holland, G.; Moeller, R.; Laue, M. Directed freeze-fracturing of Bacillus subtilis biofilms for conventional scanning electron microscopy. J. Microbiol. Methods 2018, 152, 165–172. [Google Scholar] [CrossRef] [PubMed]
- Serra, D.O.; Richter, A.M.; Klauck, G.; Mika, F.; Hengge, R. Microanatomy at cellular resolution and spatial order of physiological differentiation in a bacterial biofilm. mBio 2013, 4, e00103–00113. [Google Scholar] [CrossRef]
- Seoane, J.; Yankelevich, T.; Dechesne, A.; Merkey, B.; Sternberg, C.; Smets, B.F. An individual-based approach to explain plasmid invasion in bacterial populations. FEMS Microbiol. Ecol. 2011, 75, 17–27. [Google Scholar] [CrossRef]
- Licht, T.R.; Christensen, B.B.; Krogfelt, K.A.; Molin, S. Plasmid transfer in the animal intestine and other dynamic bacterial populations: The role of community structure and environment. Microbiology 1999, 145 Pt 9, 2615–2622. [Google Scholar] [CrossRef]
- Serra, D.O.; Hengge, R. Stress responses go three dimensional—The spatial order of physiological differentiation in bacterial macrocolony biofilms. Environ. Microbiol. 2014, 16, 1455–1471. [Google Scholar] [CrossRef]
- Stewart, P.S.; Franklin, M.J. Physiological heterogeneity in biofilms. Nat. Rev. Microbiol. 2008, 6, 199–210. [Google Scholar] [CrossRef]
- Ghigo, J.M. Natural conjugative plasmids induce bacterial biofilm development. Nature 2001, 412, 442–445. [Google Scholar] [CrossRef]
- Lim, J.Y.; La, H.J.; Sheng, H.; Forney, L.J.; Hovde, C.J. Influence of plasmid pO157 on Escherichia coli O157:H7 Sakai biofilm formation. Appl. Environ. Microbiol. 2010, 76, 963–966. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Que, F.; Liao, L.; Zhou, M.; You, L.; Zhao, Q.; Li, Y.; Niu, H.; Wu, S.; Huang, R. Study on the promotion of bacterial biofilm formation by a Salmonella conjugative plasmid and the underlying mechanism. PLoS ONE 2014, 9, e109808. [Google Scholar] [CrossRef] [PubMed]
- Reisner, A.; Höller, B.M.; Molin, S.; Zechner, E.L. Synergistic effects in mixed Escherichia coli biofilms: Conjugative plasmid transfer drives biofilm expansion. J. Bacteriol. 2006, 188, 3582–3588. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Zhou, X.; Zou, W.; Wang, Y.; Lei, C.; Xiang, R.; Zhou, L.; Liu, B.; Zhang, A.; Wang, H. Co-Occurrence of biofilm formation and quinolone resistance in Salmonella enterica serotype typhimurium carrying an IncHI2-type oqxAB-positive plasmid. Microb. Pathog. 2018, 123, 68–73. [Google Scholar] [CrossRef]
- Beloin, C.; Roux, A.; Ghigo, J.M. Escherichia coli biofilms. Curr. Top. Microbiol. Immunol. 2008, 322, 249–289. [Google Scholar] [CrossRef]
- Craig, L.; Forest, K.T.; Maier, B. Type IV pili: Dynamics, biophysics and functional consequences. Nat. Rev. Microbiol. 2019, 17, 429–440. [Google Scholar] [CrossRef]
- Burmølle, M.; Bahl, M.I.; Jensen, L.B.; Sørensen, S.J.; Hansen, L.H. Type 3 fimbriae, encoded by the conjugative plasmid pOLA52, enhance biofilm formation and transfer frequencies in Enterobacteriaceae strains. Microbiology 2008, 154, 187–195. [Google Scholar] [CrossRef]
- Dudley, E.G.; Abe, C.; Ghigo, J.-M.; Latour-Lambert, P.; Hormazabal, J.C.; Nataro, J.P. An IncI1 plasmid contributes to the adherence of the atypical enteroaggregative Escherichia coli strain C1096 to cultured cells and abiotic surfaces. Infect. Immun. 2006, 74, 2102–2114. [Google Scholar] [CrossRef]
- Bhatty, M.; Cruz, M.R.; Frank, K.L.; Gomez, J.A.L.; Andrade, F.; Garsin, D.A.; Dunny, G.M.; Kaplan, H.B.; Christie, P.J. Enterococcus faecalis pCF10-encoded surface proteins PrgA, PrgB (aggregation substance) and PrgC contribute to plasmid transfer, biofilm formation and virulence. Mol. Microbiol. 2015, 95, 660–677. [Google Scholar] [CrossRef]
- Coburn, P.S.; Baghdayan, A.S.; Craig, N.; Burroughs, A.; Tendolkar, P.; Miller, K.; Najar, F.Z.; Roe, B.A.; Shankar, N. A novel conjugative plasmid from Enterococcus faecalis E99 enhances resistance to ultraviolet radiation. Plasmid 2010, 64, 18–25. [Google Scholar] [CrossRef]
- Tendolkar, P.M.; Baghdayan, A.S.; Shankar, N. Putative surface proteins encoded within a novel transferable locus confer a high-biofilm phenotype to Enterococcus faecalis. J. Bacteriol. 2006, 188, 2063–2072. [Google Scholar] [CrossRef] [PubMed]
- Reisner, A.; Haagensen, J.A.J.; Schembri, M.A.; Zechner, E.L.; Molin, S. Development and maturation of Escherichia coli K-12 biofilms. Mol. Microbiol. 2003, 48, 933–946. [Google Scholar] [CrossRef] [PubMed]
- May, T.; Okabe, S. Escherichia coli harboring a natural IncF conjugative F plasmid develops complex mature biofilms by stimulating synthesis of colanic acid and Curli. J. Bacteriol. 2008, 190, 7479–7490. [Google Scholar] [CrossRef] [PubMed]
- Barrios, A.F.G.; Zuo, R.; Ren, D.; Wood, T.K. Hha, YbaJ, and OmpA regulate Escherichia coli K12 biofilm formation and conjugation plasmids abolish motility. Biotechnol. Bioeng. 2006, 93, 188–200. [Google Scholar] [CrossRef]
- Yang, X.; Ma, Q.; Wood, T.K. The R1 conjugative plasmid increases Escherichia coli biofilm formation through an envelope stress response. Appl. Environ. Microbiol. 2008, 74, 2690–2699. [Google Scholar] [CrossRef]
- Okshevsky, M.; Meyer, R.L. The role of extracellular DNA in the establishment, maintenance and perpetuation of bacterial biofilms. Crit. Rev. Microbiol. 2015, 41, 341–352. [Google Scholar] [CrossRef]
- D’Alvise, P.W.; Sjøholm, O.R.; Yankelevich, T.; Jin, Y.; Wuertz, S.; Smets, B.F. TOL plasmid carriage enhances biofilm formation and increases extracellular DNA content in Pseudomonas putida KT2440. FEMS Microbiol. Lett. 2010, 312, 84–92. [Google Scholar] [CrossRef]
- Røder, H.L.; Hansen, L.H.; Sørensen, S.J.; Burmølle, M. The impact of the conjugative IncP-1 plasmid pKJK5 on multispecies biofilm formation is dependent on the plasmid host. FEMS Microbiol. Lett. 2013, 344, 186–192. [Google Scholar] [CrossRef]
- Hoffman, L.R.; D’Argenio, D.A.; MacCoss, M.J.; Zhang, Z.; Jones, R.A.; Miller, S.I. Aminoglycoside antibiotics induce bacterial biofilm formation. Nature 2005, 436, 1171–1175. [Google Scholar] [CrossRef]
- Linares, J.F.; Gustafsson, I.; Baquero, F.; Martinez, J.L. Antibiotics as intermicrobial signaling agents instead of weapons. Proc. Natl. Acad. Sci. USA 2006, 103, 19484–19489. [Google Scholar] [CrossRef]
- Salcedo, D.E.; Lee, J.H.; Ha, U.H.; Kim, S.P. The effects of antibiotics on the biofilm formation and antibiotic resistance gene transfer. Desalin. Water Treat. 2015, 54, 3582–3588. [Google Scholar] [CrossRef]
- Penesyan, A.; Nagy, S.S.; Kjelleberg, S.; Gillings, M.R.; Paulsen, I.T. Rapid microevolution of biofilm cells in response to antibiotics. NPJ Biofilms Microbiomes 2019, 5, 34. [Google Scholar] [CrossRef]
- Bagge, N.; Schuster, M.; Hentzer, M.; Ciofu, O.; Givskov, M.; Greenberg, E.P.; Høiby, N. Pseudomonas aeruginosa biofilms exposed to imipenem exhibit changes in global gene expression and β-lactamase and alginate production. Antimicrob. Agents Chemother. 2004, 48, 1175–1187. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Pascual, F.; Hartmann, R.; Lempp, M.; Vidakovic, L.; Song, B.; Jeckel, H.; Thormann, K.M.; Yildiz, F.H.; Dunkel, J.; Link, H.; et al. Breakdown of Vibrio cholerae biofilm architecture induced by antibiotics disrupts community barrier function. Nat. Microbiol. 2019, 4, 2136–2145. [Google Scholar] [CrossRef]
- Al-Masaudi, S.B.; Day, M.J.; Russell, A.D. Effect of some antibiotics and biocides on plasmid transfer in Staphylococcus aureus. J. Appl. Bacteriol. 1991, 71, 239–243. [Google Scholar] [CrossRef]
- Barr, V.; Barr, K.; Millar, M.R.; Lacey, R.W. β-Lactam antibiotics increase the frequency of plasmid transfer in Staphylococcus aureus. J. Antimicrob. Chemother. 1986, 17, 409–413. [Google Scholar] [CrossRef] [PubMed]
- Feld, L.; Schjørring, S.; Hammer, K.; Licht, T.R.; Danielsen, M.; Krogfelt, K.; Wilcks, A. Selective pressure affects transfer and establishment of a Lactobacillus plantarum resistance plasmid in the gastrointestinal environment. J. Antimicrob. Chemother. 2008, 61, 845–852. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Bogaj, K.; Bortolaia, V.; Olsen, J.E.; Thomsen, L.E. Antibiotic-Induced, increased conjugative transfer is common to diverse naturally occurring ESBL Plasmids in Escherichia coli. Front. Microbiol. 2019, 10, 2119. [Google Scholar] [CrossRef]
- Lu, Y.; Zeng, J.; Wang, L.; Lan, K.; Shunmei, E.; Wang, L.; Xiao, Q.; Luo, Q.; Huang, X.; Huang, B.; et al. Antibiotics promote Escherichia coli-Pseudomonas aeruginosa conjugation through inhibiting quorum sensing. Antimicrob. Agents Chemother. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.; Bryers, J.D. Non-Invasive determination of conjugative transfer of plasmids bearing antibiotic-resistance genes in biofilm-bound bacteria: Effects of substrate loading and antibiotic selection. Appl. Microbiol. Biotechnol. 2013, 97, 317–328. [Google Scholar] [CrossRef] [PubMed]
- Ohlsen, K.; Ternes, T.; Werner, G.; Wallner, U.; Löffler, D.; Ziebuhr, W.; Witte, W.; Hacker, J. Impact of antibiotics on conjugational resistance gene transfer in Staphylococcus aureus in sewage. Environ. Microbiol. 2003, 5, 711–716. [Google Scholar] [CrossRef]
- Xia, Z.-J.; Wang, J.; Hu, W.; Liu, H.; Gao, X.-Z.; Wu, Z.-H.; Zhang, P.-Y.; Li, Y.-Z. Improving conjugation efficacy of Sorangium cellulosum by the addition of dual selection antibiotics. J. Ind. Microbiol. Biotechnol. 2008, 35, 1157–1163. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.-Y.; Xu, P.-P.; Xia, Z.-J.; Wang, J.; Xiong, J.; Li, Y.-Z. Combined treatment with the antibiotics kanamycin and streptomycin promotes the conjugation of Escherichia coli. FEMS Microbiol. Lett. 2013, 348, 149–156. [Google Scholar] [CrossRef]
- Møller, T.S.B.; Liu, G.; Boysen, A.; Thomsen, L.E.; Lüthje, F.L.; Mortensen, S.; Møller-Jensen, J.; Olsen, J.E. Treatment with Cefotaxime affects expression of conjugation associated proteins and conjugation transfer frequency of an IncI1 Plasmid in Escherichia coli. Front. Microbiol. 2017, 8, 2365. [Google Scholar] [CrossRef]
- Shun-Mei, E.; Zeng, J.-M.; Yuan, H.; Lu, Y.; Cai, R.-X.; Chen, C. Sub-Inhibitory concentrations of fluoroquinolones increase conjugation frequency. Microb. Pathog. 2018, 114, 57–62. [Google Scholar] [CrossRef] [PubMed]
- Cantas, L.; Midtlyng, P.J.; Sørum, H. Impact of antibiotic treatments on the expression of the R plasmid tra genes and on the host innate immune activity during pRAS1 bearing Aeromonas hydrophila infection in zebrafish (Danio rerio). BMC Microbiol. 2012, 12, 37. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Yun, Z.; Ha, U.-H.; Lee, S.; Park, H.; Kwon, E.E.; Cho, Y.; Choung, S.; Oh, J.; Medriano, C.A.; et al. Transfer of antibiotic resistance plasmids in pure and activated sludge cultures in the presence of environmentally representative micro-contaminant concentrations. Sci. Total Environ. 2014, 468–469, 813–820. [Google Scholar] [CrossRef] [PubMed]
- Lopatkin, A.J.; Huang, S.; Smith, R.P.; Srimani, J.K.; Sysoeva, T.A.; Bewick, S.; Karig, D.K.; You, L. Antibiotics as a selective driver for conjugation dynamics. Nat. Microbiol. 2016, 1, 16044. [Google Scholar] [CrossRef]
- Li, B.; Qiu, Y.; Zhang, J.; Huang, X.; Shi, H.; Yin, H. Real-Time study of rapid spread of antibiotic resistance plasmid in biofilm using microfluidics. Environ. Sci. Technol. 2018, 52, 11132–11141. [Google Scholar] [CrossRef]
- Qiu, Y.; Zhang, J.; Li, B.; Wen, X.; Liang, P.; Huang, X. A novel microfluidic system enables visualization and analysis of antibiotic resistance gene transfer to activated sludge bacteria in biofilm. Sci. Total Environ. 2018, 642, 582–590. [Google Scholar] [CrossRef]
| Protein | Proposed Function | Description | Localization | Homolog | Reference |
|---|---|---|---|---|---|
| TraM | Relaxosome | oriT binding, TraI stimulation, | C | [17,52,55,56] | |
| Interaction with TraD | |||||
| TraJ | Regulation | Transcription factor (anti-silencer/activator of PY) | C | [17,55,57] | |
| TraY | Relaxosome Regulation | oriT binding, Transcription factor (activator of PM) | C | [17,52,55,57] | |
| TraA | Pilin | Major subunit of the pilus | IM | VirB2 (pTi) | [52,55,56] |
| TrbC (RP4) | |||||
| TraL | Pilus assembly | Pilus assembly | OM | VirB3 (pTi) | [52,53,55,57,58] |
| TrbD (RP4) | |||||
| TraE | Pilus assembly | Pilus assembly | IM/P | VirB5 (pTi) | [52,53,55] |
| TraK | Pilus assembly | Cell envelope-spanning channel | IM/P | VirB9 (pTi) | [52,53,55,56] |
| TraB | Pilus extension | Cell envelope-spanning channel | IM | VirB10 (pTi) | [52,53,55,56,58] |
| TrbI (RP4) | |||||
| TraP | Pilus extension | Extended pilus stabilization | IM | [55,57] | |
| TraG | Pilus assembly | Pilus tip assembly | IM | VirB6/VirB8 (pTi) | [52,53,55,57,59] |
| Mating pair stabilization | Stabilization via C-terminal Interaction with TraN, | ||||
| Exclusion | Interaction with TraS | ||||
| TraV | Pilus extension | Lipoprotein | OM/P | VirB7 (pTi) | [52,53,55,56] |
| TraR | Regulation | Transcription regulator by binding to RNA polymerase | C | [58,60] | |
| TraC | Pilus assembly | NTPase | IM | VirB4 (pTi) | [52,53,55,56] |
| TrbE (RP4) | |||||
| TraW | Pilus extension | Pilus synthesis | P | [52,53,56,57,61] | |
| TraU | DNA transfer | DNA transfer | P | [52,55,56] | |
| TraN | Mating pair stabilization | Stabilization of OmpA and Lps binding | OM | [52,55,56] | |
| Exclusion system | Interaction with TraG | ||||
| TraF | Pilus extension | Disulfide bonds for T4SS assembly | P | [52,53,55,56] | |
| TraQ | Pilin maturation | Chaperone-like | IM | [55,56,57] | |
| TraH | Pilus extension | Interaction with TraF and TraU | P | [52,55] | |
| TraG | Pilus assembly | Pilus tip assembly | IM | VirB6/VirB8 (pTi) | [52,55,57,59] |
| Mating pair stabilization | Stabilization via C-terminal Interaction with TraN, | ||||
| Exclusion | Interaction with TraS | ||||
| TraS | Entry Exclusion (Eex) | Interaction with TraG | IM | [55,56] | |
| TraT | Surface exclusion (Sfx) | Disaggregation of mating pair after DNA transfer, Interferes with TraN–OmpA interaction | OM | [55,56,62,63] | |
| TraD | T4CP | Coupling protein/DNA dependent ATPase Interaction with TraM | IM | VirD4 (pTi) | [55,56,57,64] |
| TraG (RP4) | |||||
| TrwB (R388) | |||||
| TraI | Relaxosome | Relaxase, transesterase and helicase | C | VirD2 (pTi) | [55,57,65,66] |
| TrwC (R388) | |||||
| TraX | Pilin maturation | N-terminal acetylase | IM | TrbP (RP4) | [55,56,57](1)(3)(8) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Virolle, C.; Goldlust, K.; Djermoun, S.; Bigot, S.; Lesterlin, C. Plasmid Transfer by Conjugation in Gram-Negative Bacteria: From the Cellular to the Community Level. Genes 2020, 11, 1239. https://doi.org/10.3390/genes11111239
Virolle C, Goldlust K, Djermoun S, Bigot S, Lesterlin C. Plasmid Transfer by Conjugation in Gram-Negative Bacteria: From the Cellular to the Community Level. Genes. 2020; 11(11):1239. https://doi.org/10.3390/genes11111239
Chicago/Turabian StyleVirolle, Chloé, Kelly Goldlust, Sarah Djermoun, Sarah Bigot, and Christian Lesterlin. 2020. "Plasmid Transfer by Conjugation in Gram-Negative Bacteria: From the Cellular to the Community Level" Genes 11, no. 11: 1239. https://doi.org/10.3390/genes11111239
APA StyleVirolle, C., Goldlust, K., Djermoun, S., Bigot, S., & Lesterlin, C. (2020). Plasmid Transfer by Conjugation in Gram-Negative Bacteria: From the Cellular to the Community Level. Genes, 11(11), 1239. https://doi.org/10.3390/genes11111239






