Genome-Wide Homozygosity Patterns and Evidence for Selection in a Set of European and Near Eastern Horse Breeds
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling
2.2. SNP Genotyping
2.3. Population Stratification and ROH Analysis
2.4. Gene Ontology Analysis
3. Results
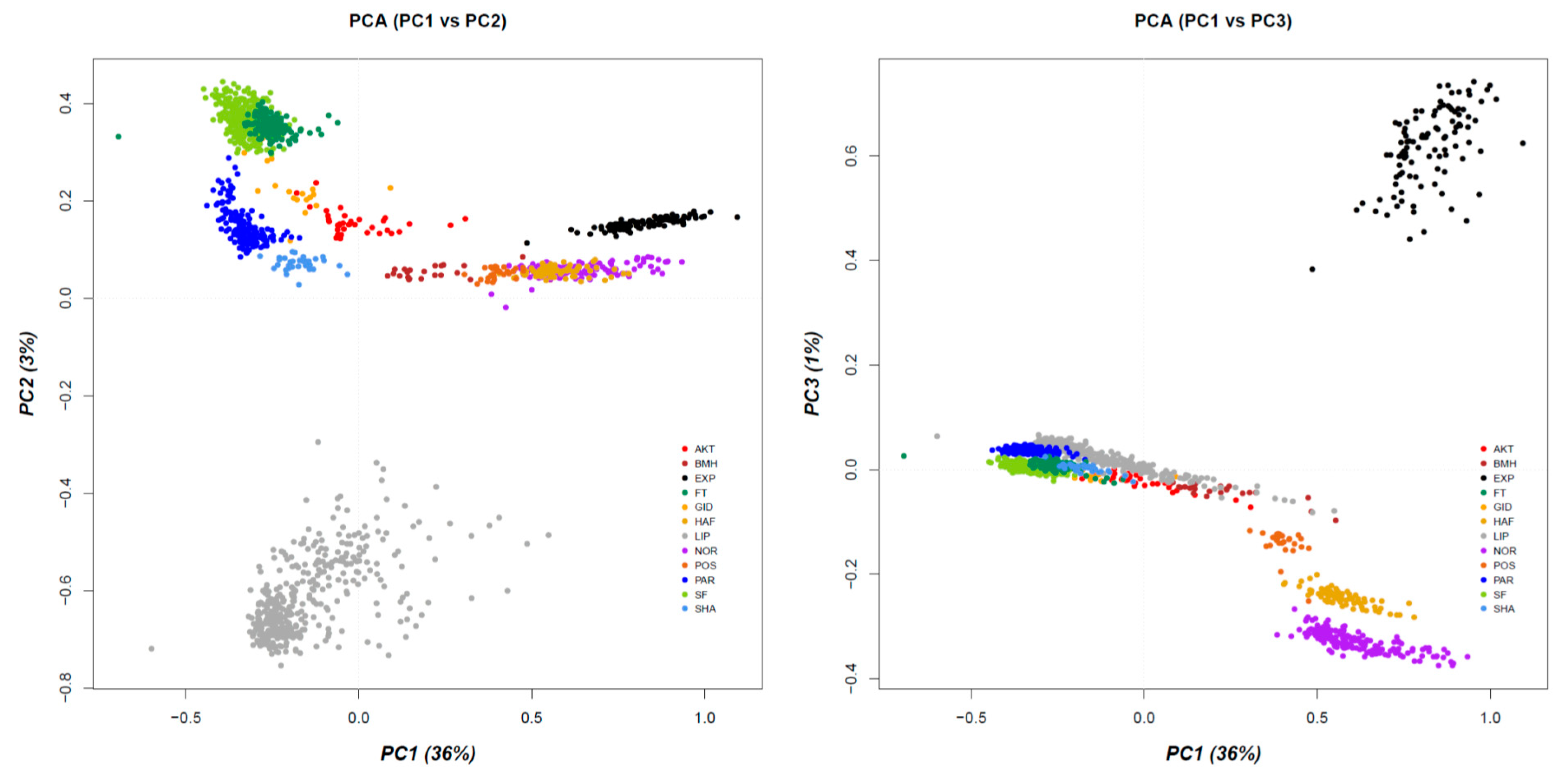
3.1. Population Stratification
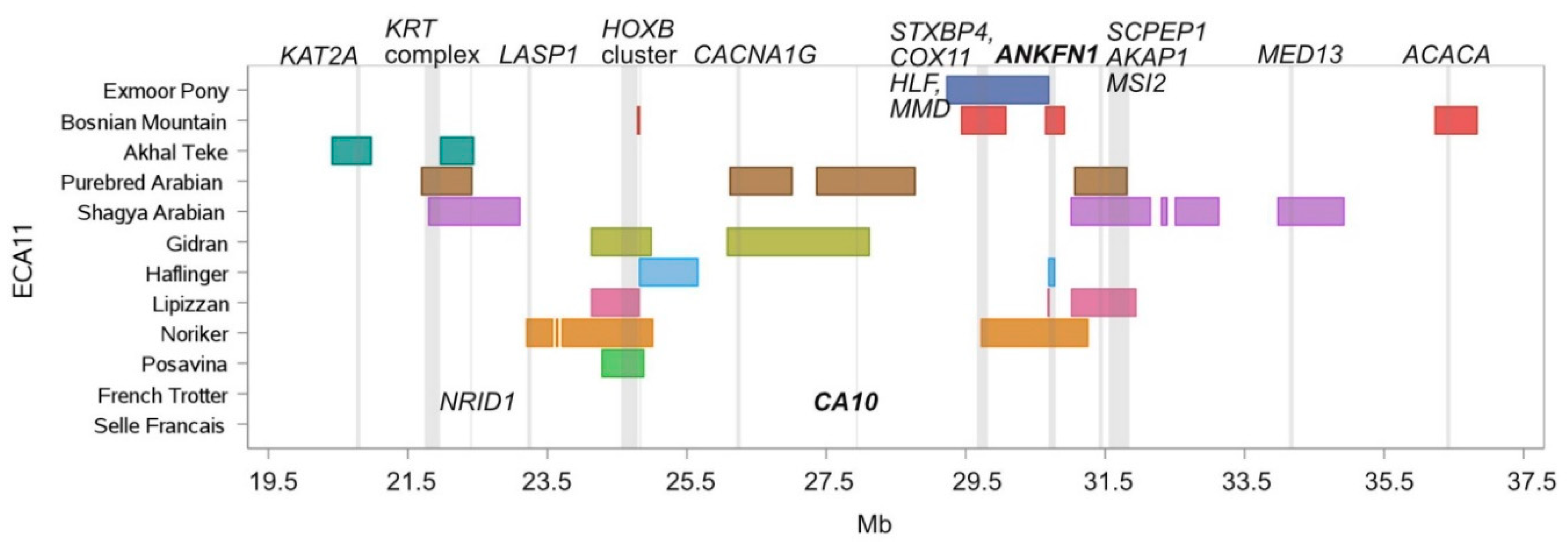
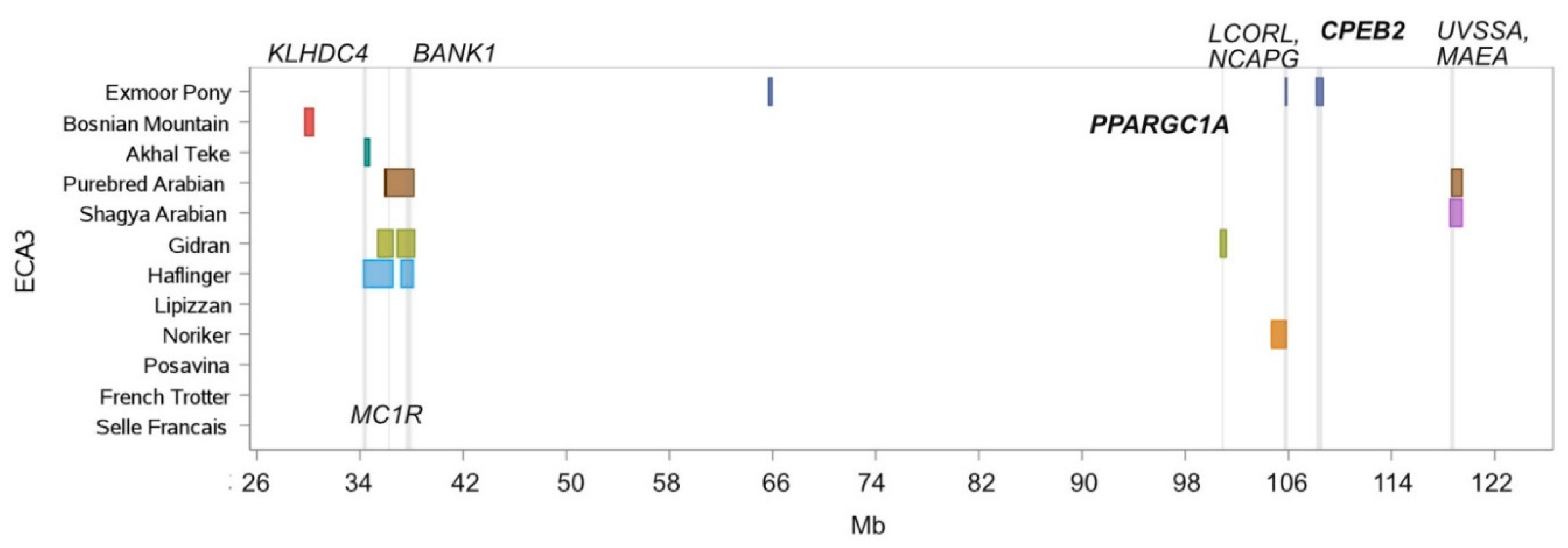
3.2. ROH Island Pattern and Distribution
3.3. Genotype Frequencies of Genes Involved in Coat Color and Body Size
3.4. Gene Ontology and Enrichment Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Data Availability
References
- Gu, J.; Orr, N.; Park, S.D.; Katz, L.M.; Sulimova, G.; MacHugh, D.E.; Hill, E.W. A genome scan for positive selection in thoroughbred horses. PLoS ONE 2009, 4, e5767. [Google Scholar] [CrossRef] [PubMed]
- Petersen, J.L.; Mickelson, J.R.; Cothran, E.G.; Andersson, L.S.; Axelsson, J.; Bailey, E.; da Câmara Machado, A. Genetic diversity in the modern horse illustrated from genome-wide SNP data. PLoS ONE 2013, 8, e54997. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, R.J.; Schubert, M.; Bailey, E.; Bannasch, D.L.; Barrey, E.; Bar-Gal, G.K.; Brem, G.; Brooks, S.A.; Distl, O.; Fries, R. Developing a 670k genotyping array to tag ~2 M SNPs across 24 horse breeds. BMC Genom. 2017, 18, 565–583. [Google Scholar] [CrossRef] [PubMed]
- McCue, M.E.; Bannasch, D.L.; Petersen, J.L.; Gurr, J.; Bailey, E.; Binns, M.M.; Leeb, T. A high density SNP array for the domestic horse and extant Perissodactyla: Utility for association mapping; genetic diversity; and phylogeny studies. PLoS Genet. 2012, 8, e1002451. [Google Scholar] [CrossRef] [PubMed]
- Petersen, J.L.; Mickelson, J.R.; Rendahl, A.K.; Valberg, S.J.; Andersson, L.S.; Axelson, J.; Bailey, E.; Bannesch, D.; Binns, M.M.; Borges, A.S. Genome-wide analysis reveals selection for important traits in domestic horse breeds. PLoS ONE 2013, 9, e10003211. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Ni, P.; Ahmad, H.I.; Gemingguli, M.; Baizilaitibei, A.; Gulibaheti, D.; Fang, Y.; Wang, H.; Asif, A.R.; Xiao, C.; et al. Detecting the population structure and scanning for signatures of selection in horses (Equus caballus) from whole-genome sequencing data. Evol. Bioinform. 2018, 14, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Gurgul, A.; Jasielczuk, I.; Semik-Gurgul, E.; Pawlina-Tyszko, K.; Stefaniuk-Szmukier, M.; Szmatoła, T.; Polak, G.; Tomczyk-Wrona, I.; Bugno-Poniewierska, M. A genome-wide scan for diversifying selection signatures in selected horse breeds. PLoS ONE 2019, 30, e0210751. [Google Scholar] [CrossRef]
- Schubert, M.; Jónsson, H.; Chang, D.; Der Sarkissian, C.; Ermini, L.; Ginolhac, A.; Fumagalli, M. Prehistoric genomes reveal the genetic foundation and cost of horse domestication. Proc. Natl. Acad. Sci. USA 2014, 111, e5661–e5669. [Google Scholar] [CrossRef]
- Librado, P.; Gamba, C.; Gaunitz, C.; Der Sarkissian, C.; Pruvost, M.; Albrechtsen, A.; Serres-Armero, A. Ancient genomic changes associated with domestication of the horse. Science 2017, 356, 442–445. [Google Scholar] [CrossRef]
- Lewontin, R.C.; Krakauer, J. Distribution of gene frequency as a test of the theory of the selective neutrality of polymorphisms. Genetics 1973, 74, 175–195. [Google Scholar]
- Fariello, M.I.; Boitard, S.; Naya, H.; SanCristobal, M.; Servin, B. Detecting signatures of selection through haplotype differentiation among hierarchically structured populations. Genetics 2013, 193, 929–941. [Google Scholar] [CrossRef] [PubMed]
- Peripolli, E.; Stafuzza, N.B.; Munari, D.P.; Lima, A.L.F.; Irgang, R.; Machado, M.A.; Panetto, J.C.D.C.; Ventura, R.V.; Baldi, F.; da Silva, M.V.G.B. Assessment of runs of homozygosity islands and estimates of genomic inbreeding in Gyr (Bos indicus) dairy cattle. BMC Genom. 2018, 19. [Google Scholar] [CrossRef] [PubMed]
- Peripolli, E.; Metzger, J.; De Lemos, M.V.A.; Stafuzza, N.B.; Kluska, S.; Olivieri, B.F.; Lôbo, R.B. Autozygosity islands and ROH patterns in Nellore lineages: Evidence of selection for functionally important traits. BMC Genom. 2018, 19. [Google Scholar] [CrossRef] [PubMed]
- Bertolini, F.; Cardoso, T.F.; Marras, G.; Nicolazzi, E.L.; Rothschild, M.F.; Amills, M. AdaptMap consortium. Genome-wide patterns of homozygosity provide clues about the population history and adaptation of goats. Genet. Sel. Evol. 2018, 50. [Google Scholar] [CrossRef] [PubMed]
- Mastrangelo, S.; Tolone, M.; Sardina, M.T.; Sottile, G.; Sutera, A.M.; Di Gerlando, R.; Portolano, B. Genome-wide scan for runs of homozygosity identifies potential candidate genes associated with local adaptation in Valle del Belice sheep. Genet. Sel. Evol. 2017, 49. [Google Scholar] [CrossRef] [PubMed]
- Metzger, J.; Karwath, M.; Tonda, R.; Beltran, S.; Águeda, L.; Gut, M.; Gut, L.G.; Distl, O. Runs of homozygostiy reveal signatures of positive selection for reproduction traits in breed and non-breed horses. BMC Genom. 2015, 16. [Google Scholar] [CrossRef] [PubMed]
- Grilz-Seger, G.; Druml, T.; Neuditschko, M.; Mesarič, M.; Cotman, M.; Brem, G. Analysis of ROH patterns in the Noriker horse breed reveal signatures of selection for coat color and body size. Anim. Genet. 2019. [Google Scholar] [CrossRef]
- Grilz-Seger, G.; Druml, T.; Neuditschko, M.; Dobretsberger, M.; Horna, M.; Brem, G. High-resolution population structure and runs of homozygosity reveal the genetic architecture of complex traits in the Lipizzan horse. BMC Genom. 2019, 20, 174. [Google Scholar] [CrossRef]
- Velie, B.; Solé, M.; Jäderkvist Fegraeus, K.; Rosengren, M.K.; Røed, K.H.; Ihler, C.F.; Strand, E.; Lindgren, G. Genomic measures of inbreeding in the Norwegian–Swedish Coldblooded Trotter and their associations with known QTL for reproduction and health traits. Genet. Sel. Evol. 2019, 51, 22. [Google Scholar] [CrossRef]
- Grilz-Seger, G.; Mesarič, M.; Cotman, M.; Neuditschko, M.; Druml, T.; Brem, G. Runs of homozygosity and population history of three horse breeds with small population size. J. Equine Vet. Sci. 2018, 71, 27–34. [Google Scholar] [CrossRef]
- Nolte, W.; Thaller, G.; Khuen, C. Selection signatures in four German warmblood horse breeds: Tracing breeding history in the modern sport horse. PLoS ONE 2019, 14, e0215913. [Google Scholar] [CrossRef] [PubMed]
- Stud-Book Selle Français. Available online: http://en.sellefrancais.fr/la-race-selle-francais-82-rubrique.html (accessed on 17 June 2019).
- Equipedia Ifce. Available online: https://equipedia.ifce.fr/elevage-et-entretien/race-et-robe/races-dequides-et-stud-book/trotteur-francais.html (accessed on 17 June 2019).
- Lipizzan International Federation. Available online: http://www.lipizzan-online.com/main.asp?kat1=64&kat2=543&kat3=&vid=1 (accessed on 17 June 2019).
- Zentrale Arbeitsgemeinschaft Österreichischer Pferdezüchter. Available online: https://www.pferdezucht-austria.at/main.asp?kat1=2119&kat2=2817&kat3=728 (accessed on 17 June 2019).
- Zentrale Arbeitsgemeinschaft Österreichischer Pferdezüchter. Available online: https://www.pferdezucht-austria.at/main.asp?kat1=2119&kat2=2816&kat3=661 (accessed on 17 June 2019).
- Mesaric, M. Posavina Zucht in Slowenien. In Das Posavina Pferd; Grilz-Seger, G., Druml, T., Eds.; Vehling Verlag: Graz, Austria, 2018; pp. 57–71. [Google Scholar]
- Mihok, S. A Gidran lo Monografiaja; Debrecen, Hungary, 2006. [Google Scholar]
- ISG-Shagya-Araber. Available online: http://isg-shagya-araber.de/index.php?entstehungsgeschichte-der-isg-2 (accessed on 17 June 2019).
- Mesaric, M.; Dolinsek, A.; Dovc, P. Bosnian Mountain Horse; PLANIDO-A.&A. Dolinsek, and the International Association of Bosnian Mountain Horse Breeders: Ljubljana, Slovenia, 2015. [Google Scholar]
- Wade, C.M.; Giulotto, E.; Sigurdsson, S.; Zoli, M.; Gnerre, S.; Imsland, F.; Blöcker, H. Genome sequence, comparative analysis, and population genetics of the domestic horse. Science 2009, 326, 865–867. [Google Scholar] [CrossRef] [PubMed]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.; Daly, M.J.; et al. PLINK: A tool set for whole genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef] [PubMed]
- SAS Institute. SAS Version 9.1; SAS Institute: Cary, NC, USA, 2009. [Google Scholar]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID Bioinformatics Resources. Nat. Prot. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Makvandi-Nejad, S.; Hoffman, G.E.; Allen, J.J.; Chu, E.; Gu, E.; Chandler, A.M.; Loredo, A.I.; Bellone, R.R.; Mezey, J.G.; Brooks, S.A.; et al. Four loci explain 83% of size variation in the horse. PLoS ONE 2012, 7, e39929. [Google Scholar] [CrossRef] [PubMed]
- Pearson, J.C.; Lemons, D.; McGinnis, W. Modulating Hox gene functions during animal body patterning. Nat. Rev. Genet. 2005, 6, 893–904. [Google Scholar] [CrossRef] [PubMed]
- Andersson, L.S.; Larhammar, M.; Memic, F.; Wootz, H.; Schwochow, D.; Rubin, C.J. Mutations in DMRT3 affect locomotion in horses and spinal circuit function in mice. Nature 2012, 488, 642–646. [Google Scholar] [CrossRef] [PubMed]
- Rieder, S.; Taourit, S.; Mariat, D.; Langlois, B.; Guerin, G. Mutation in the agouti (ASIP); the extension (MC1R); and the brown (TYRP1) loci and their association to coat color phenotypes in horses (Equus caballus). Mamm. Gen. 2001, 12, 450–455. [Google Scholar] [CrossRef] [PubMed]
- Higa, M.; Zhang, X.; Tanaka, K.; Saijo, M. Stabilization of Ultraviolet (UV)-stimulated Scaffold Protein A by interaction with Ubiquitin-specific Peptidase 7 is essential for Transcription-coupled Nucleotide Excision Repair. J. Biol. Chem. 2016, 291, 13771–13779. [Google Scholar] [CrossRef]
- Donnelly, M.P.; Paschou, P.; Grigorenko, E.; Gurwitz, D.; Barta, C.; Lu, R.B.; Li, H. A global view of the OCA2-HERC2 region and pigmentation. Hum. Genet. 2012, 131, 683–696. [Google Scholar] [CrossRef]
- Signer-Hasler, H.; Flury, C.; Haase, B.; Burger, D.; Simianer, H.; Leeb, T.; Rieder, S. A genome-wide association study reveals loci influencing height and other conformation traits in horses. PLoS ONE 2012, 7, e37282. [Google Scholar] [CrossRef] [PubMed]
- Cunningham, P. The Genetics of Thoroughbred Horses. Sci. Am. 1991, 264, 92–99. [Google Scholar] [CrossRef]
- Bower, M.A.; McGivney, B.A.; Campana, M.G.; Gu, J.; Andersson, L.S.; Barrett, E.; Bradley, D.G. The genetic origin and history of speed in the Thoroughbred racehorse. Nature 2012, 3, 643. [Google Scholar] [CrossRef]
- Binns, M.M.; Boehler, D.A.; Bailey, E.; Lear, T.L.; Cardwell, J.M.; Lambert, D.H. Inbreeding in the Thoroughbred horse. Anim. Genet. 2012, 43, 340–342. [Google Scholar] [CrossRef] [PubMed]
- Hill, E.W.; Gu, J.; McGivney, B.A.; MacHugh, D.E. Targets of selection in the Thoroughbred genome contain exercise-relevant gene SNPs associated with elite racecourse performance. Anim. Genet. 2010, 41, 56–63. [Google Scholar] [CrossRef]
- Tomlinson, D.J.; Mulling, C.H.; Fakler, T.M. Formation of keratins in the bovine claw: Roles of hormones, minerals, and vitamins in functional claw integrity. J. Dairy Sci. 2004, 87, 797–809. [Google Scholar] [CrossRef]
- Thomer, A.; Gottschalk, M.; Christmann, A.; Naccache, F.; Jung, K.; Hewicker-Trautwein, M.; Distl, O.; Metzger, J. An epistatic effect of KRT25 on SP6 is involved in curly coat in horses. Sci. Rep. 2018. [Google Scholar] [CrossRef]
- Morgenthaler, C.; Diribarne, M.; Capitan, A.; Legendre, R.; Saintilan, R.; Gilles, M.; Esquerré, D.; Juras, J.; Khanshour, A.; Schibler, L.; et al. A missense variant in the coil1A domain of the keratin 25 gene is associated with the dominant curly hair coat trait (Crd) in horse. Genet. Sel. Evol. 2017, 49. [Google Scholar] [CrossRef]
- Ludwig, A.; Pruvost, M.; Reissmann, M.; Benecke, N.; Brockmann, G.A.; Castaños, P.; Slatkin, M. Coat color variation at the beginning of horse domestication. Science 2009, 324. [Google Scholar] [CrossRef]
- Wutke, S.; Benecke, N.; Sandoval-Castellanos, E.; Döhle, H.J.; Friederich, S.; Gonzalez, J.; Morales-Muniz, A. Spotted phenotypes in horses lost attractiveness in the Middle Ages. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef]
- Marklund, L.; Moller, M.J.; Sandberg, K.; Andersson, L. A missense mutation in the gene for melanocyte-stimulating hormone receptor (MC1R) is associated with the chestnut coat color in horses. Mamm. Gen. 1996, 7, 895–899. [Google Scholar] [CrossRef]
- Zechner, P.; Sölkner, J.; Bodo, I.; Druml, T.; Baumung, R.; Achmann, R.; Marti, E.; Habe, F.; Brem, G. Analysis of diversity and population structure in the Lipizzan horse breed based on pedigree information. Livest. Prod. Sci. 2002, 77, 137–146. [Google Scholar] [CrossRef]
- Takagishi, T.; Hara, T.; Fukada, T. Recent advances in the role of SLC39A/ZIP zinc transporters in vivo. Int. J. Mol. Sci. 2017, 18, 2708. [Google Scholar] [CrossRef] [PubMed]
- Kozyrev, S.V.; Abelson, A.K.; Wojcik, J.; Zaghlool, A.; Linga, R. Functional variants in the B-cell gene BANK1 are associated with systemic lupus erythematosus. Nat. Genet. 2008, 40, 211–216. [Google Scholar] [CrossRef] [PubMed]
- Wullaert, A.; Bonnet, M.C.; Pasparakis, M. NF-κB in the regulation of epithelial homeostasis and inflammation. Cell Res. 2011, 21, 146–158. [Google Scholar] [CrossRef]
- Bellone, R.R.; Liu, J.; Petersen, J.L.; Mack, M.; Singer-Berk, M.; Drögemüller, C.; Malvick, J.; Wallner, B.; Brem, G.; Penedo, M.C.; et al. A missense mutation in damage-specific DNA binding protein 2 is a genetic risk factor for limbal squamous cell carcinoma in horses. Int. J. Cancer 2017, 141, 342–353. [Google Scholar] [CrossRef]
- Pielberg, R.G.; Golovko, A.; Sundström, E.; Curik, I.; Lennartsson, J. A cis-acting regulatory mutation causes premature hair graying and susceptibility to melanoma in the horse. Nat. Genet. 2008, 40, 1004–1009. [Google Scholar] [CrossRef]
- Pielberg, R.G.; Mikko, S.; Sandberg, K.; Andersson, L. Comparative linkage mapping of the Gray coat colour gene in horses. Anim. Genet. 2005, 36, 390–395. [Google Scholar] [CrossRef]
- Fernández, A.; Silió, L.; Rodríguez, C.; Óvilo, C. Characterization of OCA2 cDNA in different porcine breeds and analysis of its potential effect on skin pigmentation in a red Iberian strain. Anim. Genet. 2006, 37, 166–170. [Google Scholar] [CrossRef]
- Bellone, R.; Lear, T.; Adelson, D.L.; Bailey, E. Comparative mapping of oculocutaneous albinism type II (OCA2); transient receptor potential cation channel; subfamily M member 1 (TRPM1) and two equine microsatellites; ASB08 and 1CA43; among four equid species by fluorescence in situ hybridization. Cytogenet. Genome Res. 2006, 114, 93. [Google Scholar] [CrossRef]
- Kowalski, E.J.A.; Bellone, R. Investigation of HERC2 and OCA2 SNP for iris color variation in Puerto Rican Paso Fino horses. J. Equine Vet. Sci. 2011, 31, 319. [Google Scholar] [CrossRef]
- Mack, M.; Kowalski, E.; Grahn, R.; Bras, D.; Cecilia, M.; Penedo, T.; Bellone, R. Two variants in SLC24A5 are associated with “Tiger-Eye” iris pigmentation in Puerto Rican Paso Fino horses. G3 2017, 7, 2799–2806. [Google Scholar] [CrossRef] [PubMed]
- Jäderkvist Fegraeus, K.; Velie, B.; Axelsson, J.; Ang, R.; Hamilton, N.A.; Andersson, L.; Meadows, J.R.S.; Lindgren, G. A potential regulatory region near the EDN3 gene may control both harness racing performance and coat color variation in horses. Physiol. Rep. 2018, 6, e13700. [Google Scholar] [CrossRef] [PubMed]
- Brooks, S.; Bailey, E. Exon skipping in the KIT gene causes a Sabino spotting pattern in Horses. Mamm. Gen. 2005, 16, 893–902. [Google Scholar] [CrossRef] [PubMed]
- Haase, B.; Rieder, S.; Tozaki, T.; Hasegawa, T.; Penedo, M.; Hasegawa, T.; Penedo, M.C.T.; Jude, R.; Leeb, T. Five novel KIT mutations in horses with white coat colour phenotypes. Anim. Gen. 2011, 42, 337–339. [Google Scholar] [CrossRef] [PubMed]
- Hauswirth, R.; Jude, R.; Haase, B.; Bellone, R.R.; Archer, S.; Holl, H.; Leeb, T. Novel variants in the KIT and PAX3 genes in horses with white-spotted coat colour phenotypes. Anim. Gen. 2013, 4, 763–765. [Google Scholar] [CrossRef] [PubMed]
- Dominy, J.E.; Puigserver, P. Mitochondrial biogenesis through activation of nuclear signaling proteins cold spring harb. Perspect. Biol. 2013, 5, a015008. [Google Scholar]
- Pilegaard, H.; Saltin, B.; Neufer, P.D. Exercise induces transient transcriptional activation of the PGC-1α gene in human skeletal muscle. J. Physiol. 2003, 546, 851–858. [Google Scholar] [CrossRef]
- Eivers, S.S.; McGivney, B.A.; Gu, J.; MacHugh, D.E.; Katz, L.M.; Hill, E.M. PGC-1α encoded by the PPARGC1A gene regulates oxidative energy metabolism in equine skeletal muscle during exercise. Anim. Genet. 2012, 43, 153–162. [Google Scholar] [CrossRef]
- Ricard, A. Does heterozygosity at the DMRT3 gene make French trotters better racers? Gen. Sel. Evol. 2015, 47, 10. [Google Scholar] [CrossRef]
- Liu, F.; Arias-Vásquez, A.; Sleegers, K.; Aulchenko, Y.S.; Kayser, M.; Sanchez-Juan, P.; Heutink, P. A genome wide screen for late-onset Alzheimer disease in a genetically isolated Dutch population. Am. J. Hum. Genet. 2007, 81, 17–31. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Wang, K.S.; Aragam, N. NTM and NR3C2 polymorphisms influencing intelligence: Family-based association studies. Prog. Neuro Psychopharmacol. Biol. Psychiatry 2011, 35, 154–160. [Google Scholar] [CrossRef] [PubMed]
- Avila, F.; Mickelson, J.M.; Schaefer, R.J.; McCue, M. Genome-wide signatures of selection reveal genes associated with performance in American Quarter Horse subpopulations. Front. Genet. 2018, 9, 249. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.P.; Shimizu, E.; Dube, G.R.; Rampon, C.; Kerchner, G.A.; Min, Z.; Liu, G.; Tsien, J.Z. Genetic enhancement of learning and memory in mice. Nature 1999, 401, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Velie, B.D.; Fegraeus, K.J.; Solé, M.; Rosengren, M.K.; Røed, K.H.; Ihler, C.F.; Strand, E.; Lindgren, G. A genome-wide association study for harness racing success in the Norwegian-Swedish coldblooded trotter reveals genes for learning and energy metabolism. BMC Genet. 2018, 29, 80. [Google Scholar] [CrossRef] [PubMed]
- Hendrickson, S.L. A genome wide study of genetic adaptation to high altitude in feral Andean Horses of the páramo. BMC Evol. Biol. 2013, 13. [Google Scholar] [CrossRef] [PubMed]
- Eskelinen, E.L.; Saftig, P. Autophagy: A lysosomal degradation pathway with a central role in health and disease. Biochimica et Biophysica Acta (BBA). Mol. Cell Res. 2009, 17937, 664–673. [Google Scholar]
- Xu, T.; Nicolson, S.; Denton, D.; Kumar, S. Distinct requirements of Autophagy-related genes in programmed cell death. Cell Death Differ. 2015, 22, 1792–1802. [Google Scholar] [CrossRef] [PubMed]
- Bodemann, B.O.; Orvedahl, A.; Cheng, T.; Ram, R.R.; Ou, Y.; Formstecher, E.; Maiti, M.; Hazelett, C.C.; Wauson, E.M.; Balakireva, M.; et al. RalB and the exocyst mediate the cellular starvation response by direct activation of autophagosome assembly. Cell 2011, 144, 253–267. [Google Scholar] [CrossRef]
- Mrschtik, M.; Ryan, K.M. Lysosomal proteins in cell death and autophagy. FEBS J. 2015, 282, 1858–1870. [Google Scholar] [CrossRef]
- Stolz, A.; Ernst, A.; Dikic, I. Cargo recognition and trafficking in selective autophagy. Nat. Cell Biol. 2014, 16, 495–501. [Google Scholar] [CrossRef] [PubMed]
- Meiri, S.; Dayan, T.; Simberloff, D. The generality of the island rule re-examined. J. Biogeogr. 2006, 33, 1571–1577. [Google Scholar] [CrossRef]
- Norton, E.M.; Avila, F.; Schultz, N.E.; Mickelson, J.R.; Geor, R.J.; McCue, M.E. Evaluation of an HMGA2 variant for pleiotropic effects on height and metabolic traits in ponies. J. Vet. Intern. Med. 2019. [Google Scholar] [CrossRef] [PubMed]
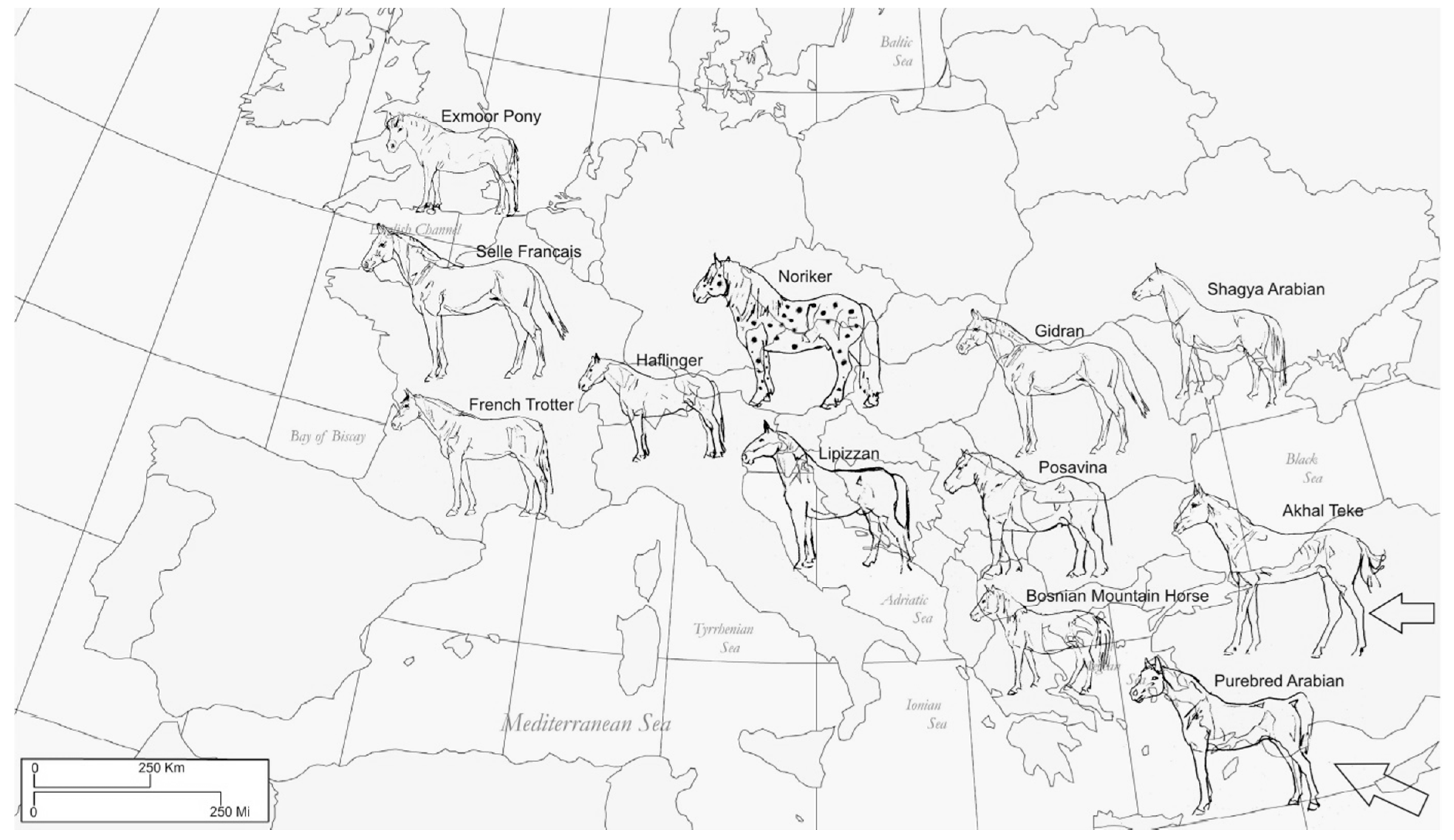

| Breed/Classification | Gene Pool | Region of Origin/Sample Origin | Population Notes and Census |
|---|---|---|---|
| Exmoor Pony small sized, multipurpose working horse | Native English breed | British Isles/UK | Closed stud book since 1921, ca. 2.000 animals [2] |
| Selle Francais Riding horse | French Trotter, Thoroughbred, Arabian | France/FRA | Stud book founded 1958; ca. 12.700 breeding animals [22] |
| French Trotter harness racing (trot) | Anglo Norman, Standardbred | France/FRA | Stud book closed since 1937, allows Standardbred; 15.500 breeding animals [23] |
| Lipizzan riding, driving | Old-Spanish and Oriental breeds | Austro—Hungarian/Lipizzan Stud farms AT, SLK, HUN, HR | Founded 1580, closed studbook since 1880; ca. 2.000 breeding animals in European state stud farms [24] |
| Noriker heavy working draught horse | Native Austrian breed | Austria/AT | Very old breed, closed stud book since 1880; ca. 4.000 breeding animals [25] |
| Haflinger light draught horse, multipurpose | Derived from local and Galizian mares, and influenced by Arabian, Gidran, Noriker | Austria/AT | Founded 1898, closed studbook since 1928; ca. 6.000 breeding animals in Austria [26] |
| Posavina small draught horse, heavy working | Native Croatian breed, influenced by Ardenner, Bosnian Mountain Horse | Croatia/SLO | Closed studbook since 1994; ca. 600 breeding animals in Slovenia [27] |
| Gidran multipurpose riding horse | Arabian, English Thoroughbred, Old-Spanish | Stud farms Mezöhegyes, Radautz (Austro - Hungarian empire)/HUN | Closed studbook since 1860, introgression of some Arabian and Thoroughbred horses; ca. 250 breeding animals in Hungary [28] |
| Shagya Arabian endurance and multipurpose riding horse | Original Arabians from Syria, influence by Russian founder mares | Stud farms Babolna, Mezöhegyes, Radautz (Austro - Hungarian empire)/SLK | Closed studbook since 1830, Purebred Arabians allowed; ca. 2.000 breeding animals [29] |
| Bosnian Mountain Horse small multipurpose working horse | Native breed of the Balkan Peninsula, influenced by Arabian | Bosnia and Herzegovina/BIH, SLO | Purebred, very old breed; 114 registered animals in the International Association of Bosnian Mountain Horse Breeders [30] |
| Purebred Arabian endurance and multipurpose riding horse | Original Arabian | Egypt, Near East/FRA | Purebred, very old breed; worldwide |
| Akhal Teke Endurance horse | Native Middle East | Turkmenistan/RUS | Purebred, stud book closed 1941; ca. 3.500 breeding animals [2] |
| Breed | SROH Mb | Sum ROH Islands Mb | n ROH Islands | n Genes in ROH Islands | n Private ROH Islands |
|---|---|---|---|---|---|
| Exmoor Pony | 506.1 | 24.5 | 33 | 171 | 28 on ECA1,2,3,4,6,16,18,19,22,23,28,30 |
| French Trotter | 352.8 | 20.5 | 22 | 156 | 14 on ECA2,4,6,7,8,9,15,23 |
| Selle Francais | 297.1 | 2.1 | 5 | 8 | 1 on ECA1 |
| Lipizzan | 297.0 | 1.9 | 5 | 23 | 1 on ECA14 |
| Noriker | 215.5 | 5.0 | 5 | 50 | 1 on ECA11 |
| Posavina | 192.7 | 2.2 | 5 | 16 | 1 on ECA28 |
| Haflinger | 279.7 | 4.9 | 5 | 60 | - |
| Gidran | 321.9 | 19.2 | 30 | 143 | 18 on ECA2,3,4,6,9,14,17,22,25,27,28 |
| Purebred Arabian | 368.1 | 15.5 | 19 | 141 | 5 on ECA2,4,5,14,22 |
| Shagya Arabian | 355.1 | 22.4 | 32 | 242 | 23 on ECA1,2,6,8,9,11,15,16,18,19, 23,25 |
| Akhal Teke | 246.1 | 4.8 | 9 | 90 | 4 on ECA1,11,19 |
| Bosnian Mountain Horse | 296.3 | 4.2 | 10 | 35 | 4 on ECA3,6,11,23 |
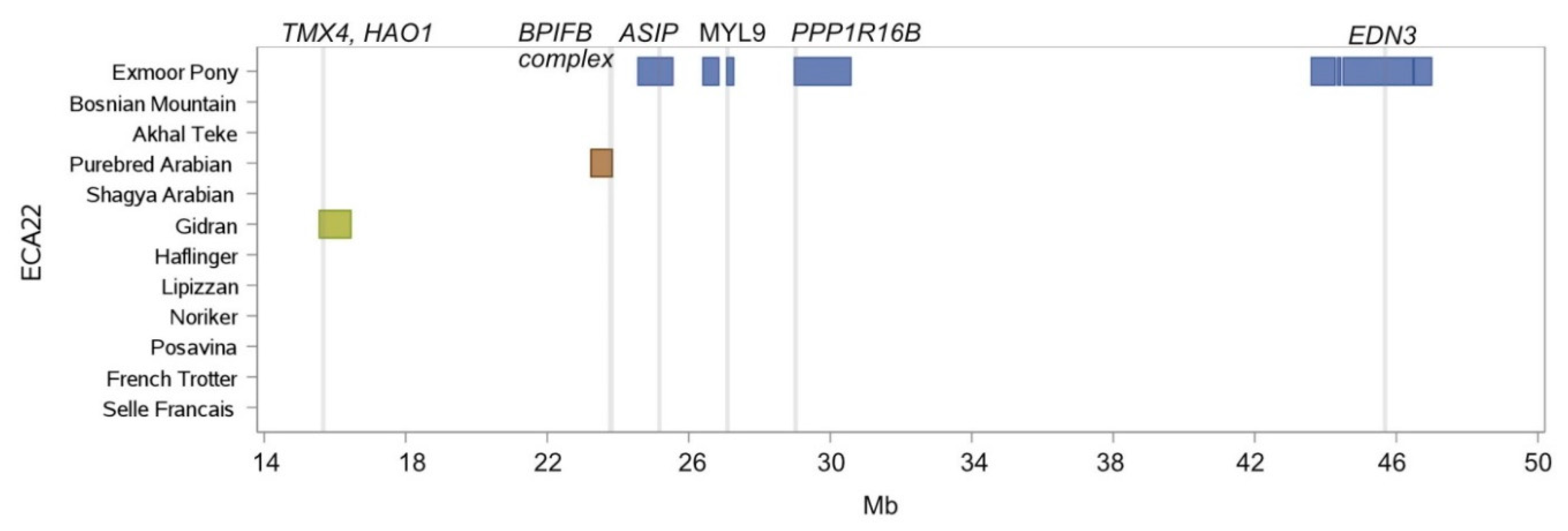
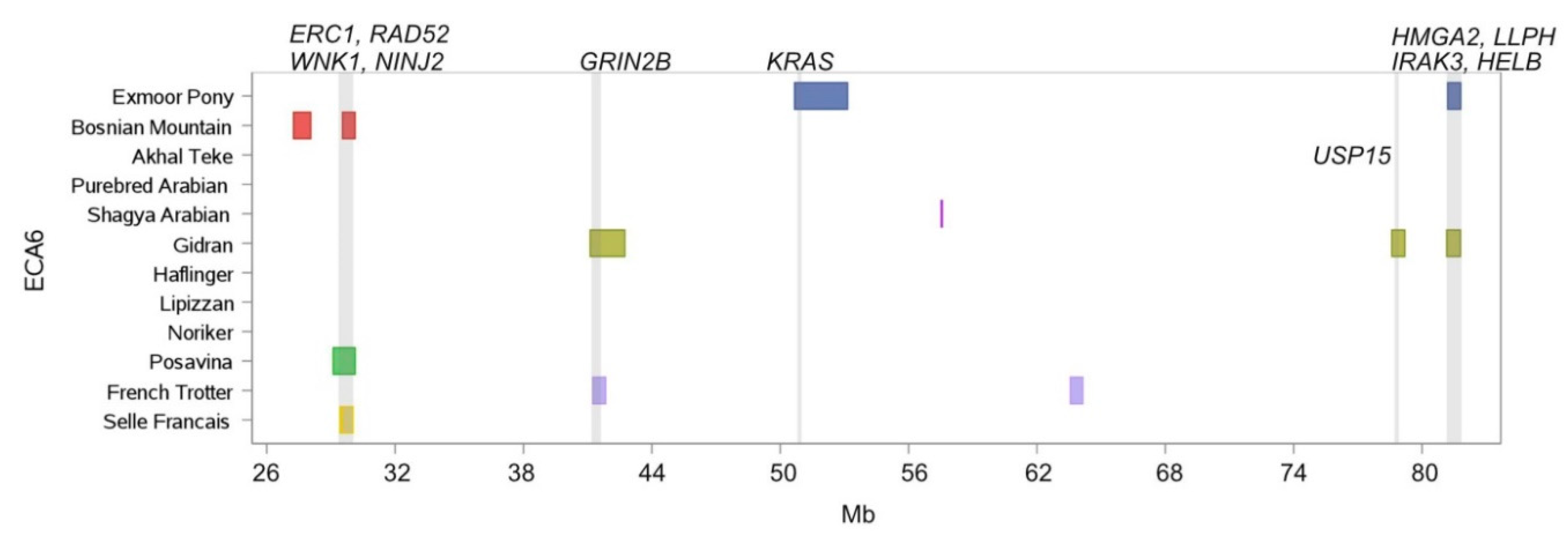
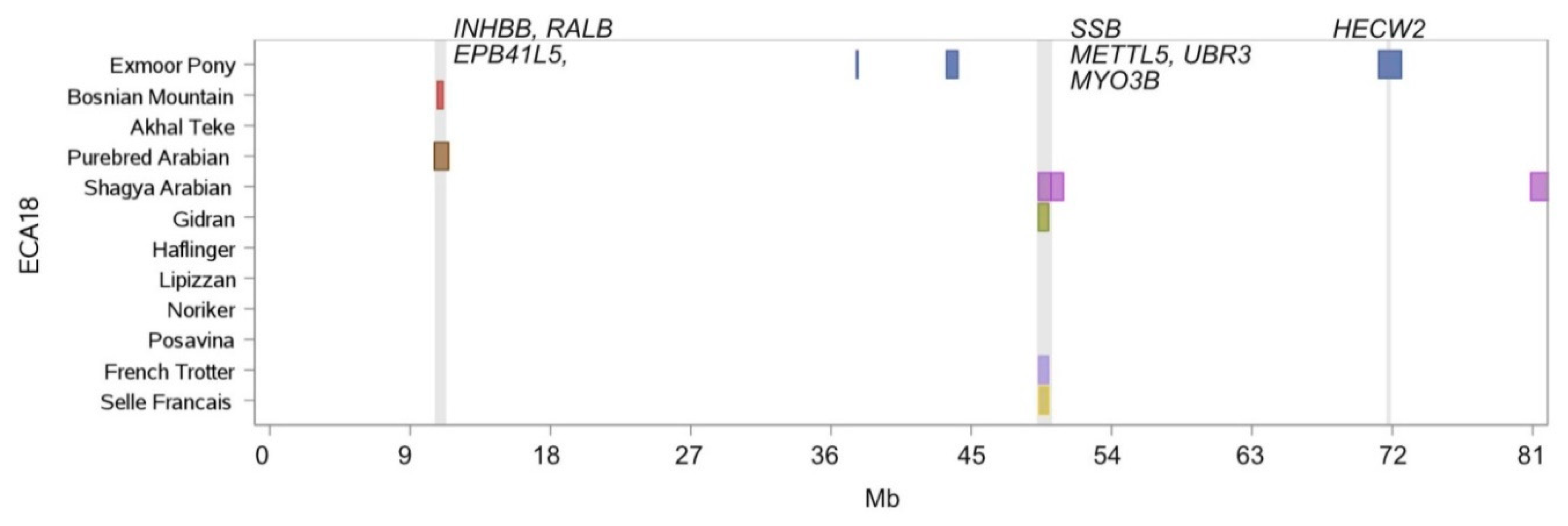
| Chr. | Begin | End | Breed Shared ROH Islands | Annotated Genes within ROH Islands |
|---|---|---|---|---|
| 1 | 148,443.474 | 148,516.796 | Akhal Teke, Bosnian Mountain Horse | EIF2AK4 |
| 3 | 34,386.385 | 34,732.351 | Akhal Teke, Haflinger | KLHDC4, SLC7A5, CA5A, BANP |
| 3 | 36,047.262 | 36,548.306 | Gidran, Haflinger, Purebred Arabian | CDK10, SPATA2L, ZNF276, VPS9D1, FANCA, SPIRE2, TCF25, DBNDD1, GAS8, MC1R, DEF8 |
| 3 | 37,179.496 | 38,127.255 | Gidran, Haflinger, Purebred Arabian | MANBA, NFKB1, SLC39A8, BANK1 |
| 3 | 105,767.304 | 105,832.553 | Exmoor Pony, Noriker | NCAPG, DCAF16 |
| 3 | 118,669.793 | 118,769.903 | Lipizzan, Shagya Arabian, Purebred Arabian | UVSSA, MAEA |
| 3 | 118,809.979 | 118,893.880 | Lipizzan, Shagya Arabian, Purebred Arabian | CTBP1 |
| 3 | 118,659.800 | 119,479.623 | Shagya Arabian, Purebred Arabian | UVSSA, MAEA, CTBP1, SPON2, FGFRL1, IDUA, SLC26A1, DGKQ, TMEM175, GAK, CPLX1, PCGF3, SLC49A3, MYL5, ATP5ME, PDE6B, PIGG |
| 4 | 15,163.634 | 15,817.917 | Akhal Teke, French Trotter | NUDCD3, NPC1L1, DDX56, TMED4, OGDH, ZMIZ2, H2AFV, MYO1G, CCM2, TBRG4, RAMO3 |
| 4 | 50,883.038 | 50,904.590 | Gidran, Shagya Arabian | HDAC9 |
| 4 | 52,692.165 | 52,840.002 | Gidran, French Trotter | ABCB5 |
| 6 | 29,424.361 | 30,040.425 | Bosnian Mountain Horse, Posavina, Selle Francais | ERC1, RAD52, WNK1, NINJ2 |
| 6 | 41,240.508 | 41,843.164 | Gidran, French Trotter | GRIN2B |
| 6 | 81,156.975 | 81,795.964 | Exmoor Pony, Gidran | HMGA2, LLPH, IRAK3 |
| 7 | 39,587.420 | 41,127.737 | French Trotter, Purebred Arabian, Shagya Arabian | NTM, OPCML |
| 7 | 48,157.328 | 48,930.905 | French Trotter, Haflinger | ACP5, ELOF1, CNN1, ECSIT, ZNF653, PRKCSH, RGL3, EPOR, SWSAP1 |
| 7 | 50,167.155 | 50,932.454 | French Trotter, Purebred Arabian | ZNF699 |
| 9 | 31,329.939 | 32,034.084 | Purebred Arabian, Shagya Arabian | PCMTD1, PXDNL |
| 9 | 44,411.007 | 44,562.960 | Purebred Arabian, Shagya Arabian, Exmoor Pony, (Haflinger) | KCNS2 |
| 9 | 75,054.137 | 75,434.658 | Noriker, Posavina | ZFAT |
| 11 | 21,801.064 | 21,938.218 | Purebred Arabian, Shagya Arabian | KRT28, KRT27, KRT26, KRT25, KRT24 |
| 11 | 21,966.387 | 22,416.682 | Akhal-Teke, Purebred Arabian, Shagya Arabian | SMARCE1, CCR7, TNS4, GFBP4, TOP2A, RARA, CDC6, WIPF2, RAPGEFL1, CASC3, MSL1, NR1D1, THRA |
| 11 | 24,285.730 | 24,816.254 | Noriker, Posavina, Gidran, Lipizzan | HOXB1, HOXB2, HOXB3, HOXB5, HOXB6, HOXB7, HOXB8, HOXB13, TTLL6 |
| 11 | 24,793.573 | 24,822.752 | Noriker, Posavina, Bosnian Mountain Horse, Haflinger, Gidran | CALCOCO2 |
| 11 | 26,117.473 | 27,008.802 | Gidran, Purebred Arabian | ACSF2, CHAD, RSAD1, MYCABPAP, EPN3, SPATA20, CACNA1G, ABCC3, ANKRD40, LUC7L3, ANKRD40CL, WFIKKN2, TOB1, SPAG9, NME1-NME2, MBTD1, UTP18 |
| 11 | 27,363.048 | 28,118.531 | Gidran, Purebred Arabian | CA10 |
| 11 | 29,747.581 | 30,078.455 | Bosnian Mountain Horse, Exmoor Pony, Noriker, | STXBP4, COX11, HLF, MMD |
| 11 | 30,690.860 | 30,774.494 | Bosnian Mountain Horse, Haflinger, Lipizzan, Noriker | ANKFN1 |
| 11 | 31,062.702 | 31,250.934 | Lipizzan, Noriker, Purebred Arabian, Shagya Arabian | C11H17orf67 |
| 11 | 31,062.702 | 31,810.316 | Purebred Arabian, Shagya Arabian, Lipizzan | C11H17orf67, DGKE, COIL, SCPEP1, AKAP1, MSI2 |
| 15 | 79,489.871 | 79,964.006 | French Trotter, Shagya Arabian | - |
| 18 | 10,733.279 | 11,116.742 | Bosnian Mountain Horse, Purebred Arabian | INHBB, RALB, EPB41L5 |
| 18 | 49,323.427 | 49,940.945 | French Trotter, Gidran, Selle Francais, Shagya Arabian | SSB, METTL5, UBR3, MYO3B |
| 19 | 50,554.541 | 50,890.075 | Gidran, Purebred Arabian | CBLB, ALCAM |
| 23 | 54,975.635 | 54,984.719 | French Trotter, Gidran | FGD3 |
| 25 | 6,409.342 | 6,812.785 | Lipizzan, Gidran | NR4A3, STX17, ERP44, INVS |
| GO Biological process | Breeds sharing term | p-value | Genes | BH p-value |
| GO:0009952~anterior/posterior pattern specification | Lipizzan/Posavina/Gidran | 0.000 | HOXB3, HOXB1, HOXB2, HOXB7, HOXB8, HOXB5, HOXB6 | < = 0.001 |
| Noriker | 0.000 | HOXB3, HOXB1, PCGF2, HOXB2, HOXB7, HOXB8, HOXB5, HOXB6 | ||
| GO:0021570~rhombomere 4 development | Lipizzan/Posavina/Noriker/Gidran | 0.003 | HOXB1, HOXB2 | 0.06–0.99 |
| GO:0021612~facial nerve structural organization | Lipizzan/Posavina/Noriker/Gidran | 0.011 | HOXB1, HOXB2 | 0.18–0.98 |
| GO:0048704~embryonic skeletal system morphogenesis | Lipizzan/Posavina/Gidran | 0.000 | HOXB3, HOXB1, HOXB2, HOXB7, HOXB8, HOXB5, HOXB6 | <0.001 |
| Noriker | 0.000 | HOXB3, HOXB1, PCGF2, HOXB2, HOXB7, HOXB8, HOXB5, HOXB6 | ||
| GO:0071222~cellular response to lipopolysaccharide | Akhal Teke | 0.034 | IL6, NR1D1, RARA | 0.99 |
| Purebred Arabian | 0.011 | NR1D1, NFKB1, RARA, SPON2 | ||
| GO Cellular component | Breeds sharing term | p-value | Genes | BH p-value |
| GO:0005654~nucleoplasm | Akhal Teke | 0.010 | COASY, CDC6, HSD17B1, BBX, ACLY, BANP, CNP, STAT3, SMARCE1, ZMIZ2, DNAJC7, ATP6V0A1, TOP2A | 0.36–0.57 |
| Noriker | 0.010 | CWC25, MRPL10, HOXB7, PSMB3, SNF8, PNPO, HOXB13, KPNB1, PIP4K2B | ||
| GO:0005882~intermediate filament | Shagya Arabian | 0.002 | KRT26, KRT25, KRT28, KRT27, KRT24 | <0.001–0.02 |
| Purebred Arabian | 0.000 | KRT26, KRT25, KRT28, KRT27, KRT12, KRT20, KRT23, KRT24 | ||
| GO Molecular function | Breeds sharing term | p-value | Genes | BH p-value |
| GO:0003700~transcription factor activity, sequence-specific DNA binding | Gidran | 0.033 | HOXB2, HOXB7, HOXB8, HOXB6, NFKB1, CBFA2T3, TCF25, FOXP2 | 0.11–0.99 |
| Lipizzan/Posavina | 0.007 | HOXB2, HOXB7, HOXB8, HOXB6 | ||
| Noriker | 0.012 | HOXB2, HOXB7, HOXB8, HOXB6, ZFAT | ||
| GO:0005198~structural molecule activity | Shagya Arabian | 0.032 | KRT26, KRT25, KRT28, KRT27, EPB41, KRT24 | 0.02–0.99 |
| Purebred Arabian | 0.000 | KRT26, KRT25, KRT28, KRT27, KRT12, KRT20, KRT23, KRT24 | ||
| GO:0015299~solute:proton antiporter activity | Purebred Arabian/Gidran | 0.044 | SLC9B1, SLC9B2 | 0.99 |
| GO:0043565~sequence-specific DNA binding | Gidran | 0.019 | HOXB1, HOXB2, HOXB7, HOXB6, HOXB13, PPARGC1A, FOXP2 | 0.007–0.92 |
| Lipizzan/Posavina/Noriker | 0.000 | HOXB1, HOXB2, HOXB7, HOXB6, HOXB13 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grilz-Seger, G.; Neuditschko, M.; Ricard, A.; Velie, B.; Lindgren, G.; Mesarič, M.; Cotman, M.; Horna, M.; Dobretsberger, M.; Brem, G.; et al. Genome-Wide Homozygosity Patterns and Evidence for Selection in a Set of European and Near Eastern Horse Breeds. Genes 2019, 10, 491. https://doi.org/10.3390/genes10070491
Grilz-Seger G, Neuditschko M, Ricard A, Velie B, Lindgren G, Mesarič M, Cotman M, Horna M, Dobretsberger M, Brem G, et al. Genome-Wide Homozygosity Patterns and Evidence for Selection in a Set of European and Near Eastern Horse Breeds. Genes. 2019; 10(7):491. https://doi.org/10.3390/genes10070491
Chicago/Turabian StyleGrilz-Seger, Gertrud, Markus Neuditschko, Anne Ricard, Brandon Velie, Gabriella Lindgren, Matjaz Mesarič, Marko Cotman, Michaela Horna, Max Dobretsberger, Gottfried Brem, and et al. 2019. "Genome-Wide Homozygosity Patterns and Evidence for Selection in a Set of European and Near Eastern Horse Breeds" Genes 10, no. 7: 491. https://doi.org/10.3390/genes10070491
APA StyleGrilz-Seger, G., Neuditschko, M., Ricard, A., Velie, B., Lindgren, G., Mesarič, M., Cotman, M., Horna, M., Dobretsberger, M., Brem, G., & Druml, T. (2019). Genome-Wide Homozygosity Patterns and Evidence for Selection in a Set of European and Near Eastern Horse Breeds. Genes, 10(7), 491. https://doi.org/10.3390/genes10070491






