miR-194 Accelerates Apoptosis of Aβ1–42-Transduced Hippocampal Neurons by Inhibiting Nrn1 and Decreasing PI3K/Akt Signaling Pathway Activity
Abstract
1. Introduction
2. Material and Methods
2.1. Primary Culture of Rat Hippocampal Neurons
2.2. Immunofluorescence Labeling of Hippocampal Neurons
2.3. MTT Assay for Hippocampal Neuron Survival after Induction of Aβ1–42
2.4. Flow Cytometry for Apoptosis
2.5. qRT-PCR
2.6. Luciferase Reporter Gene Assay
2.7. Hippocampal Neuron Transfection
2.8. Detection of Mitochondrial Transmembrane Potential
2.9. Western Blot Analysis
2.10. Statistical Analysis
3. Results
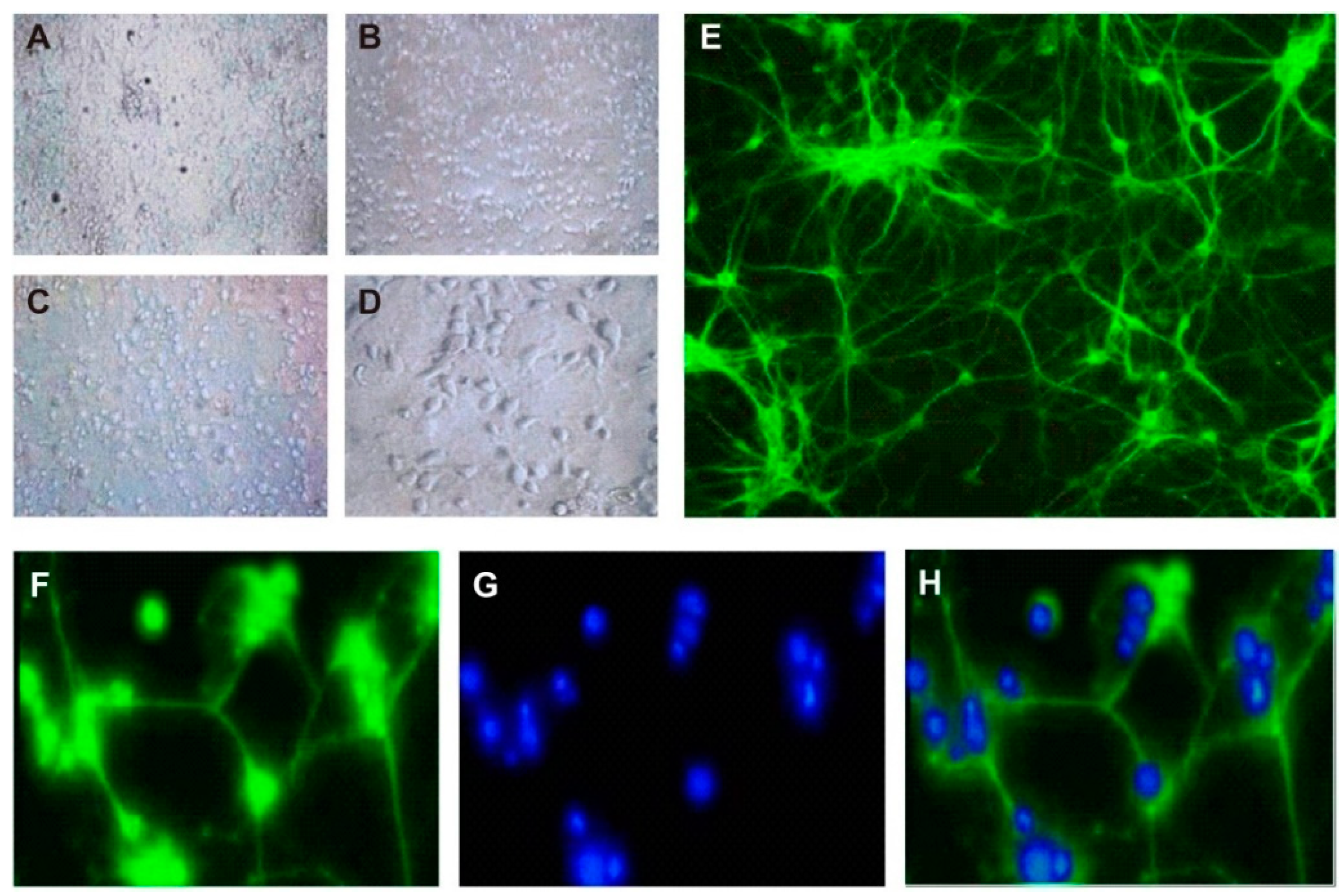
3.1. Culture and Identification of Hippocampal Neurons
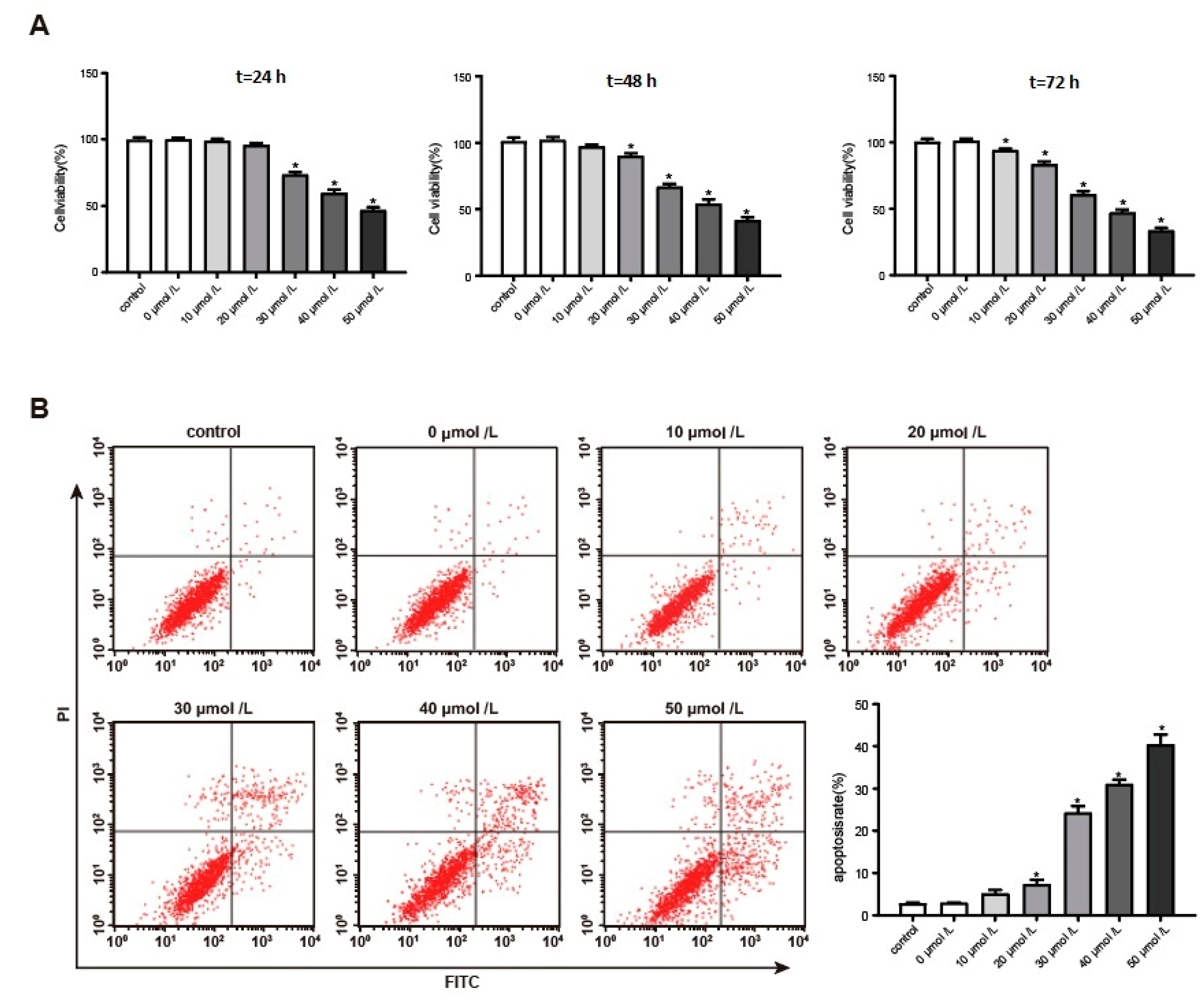
3.2. Aβ1–42 Inhibited Hippocampal Neuron Viability and Promoted Its Apoptosis in a Dose-Dependent Manner
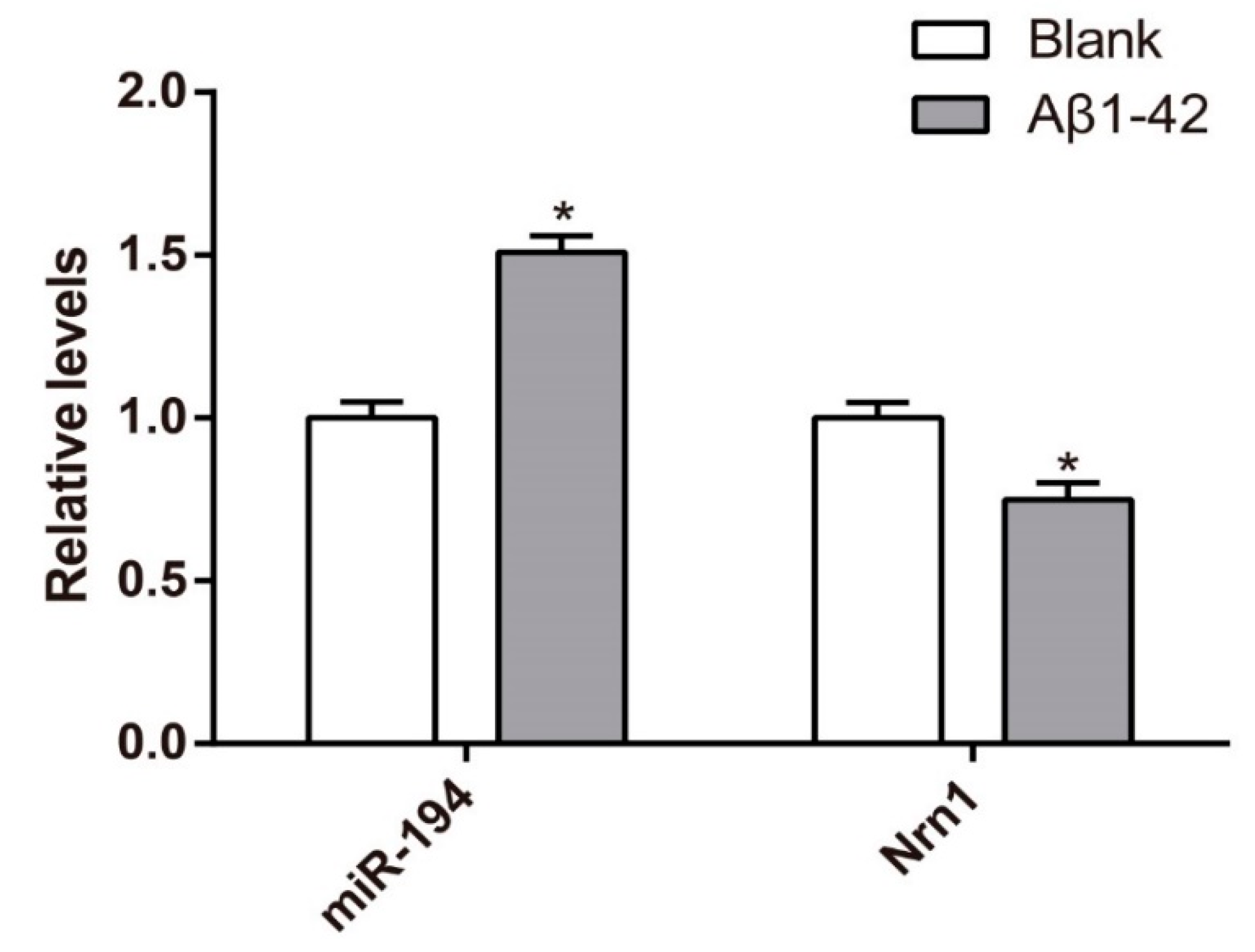
3.3. Upregulation of miR-194 and Downregulation of Nrn1 mRNA in Aβ1–42-Transduced Hippocampal Neurons
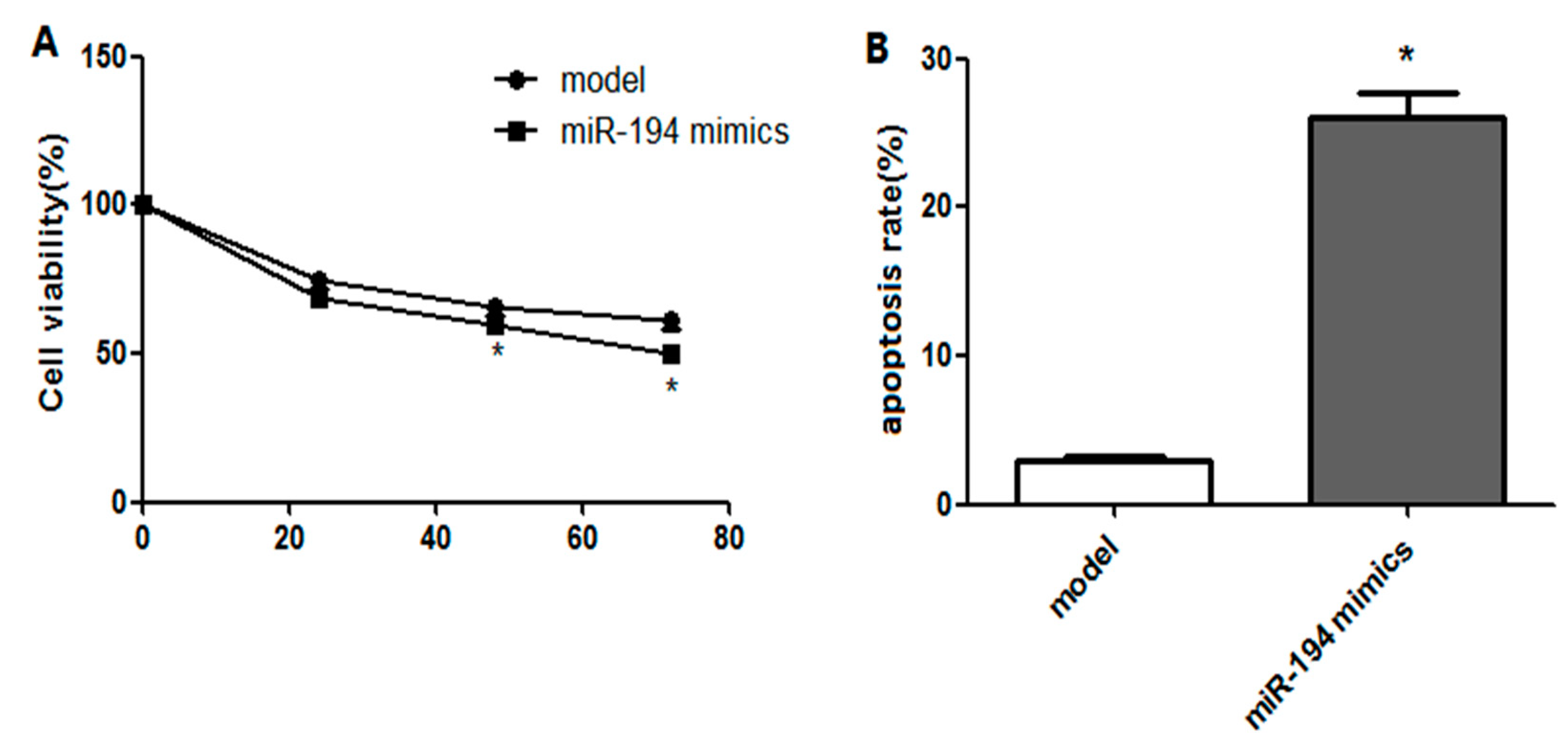
3.4. Upregulated miR-194 Inhibited Hippocampal Neuron Viability and Promoted Its Apoptosis
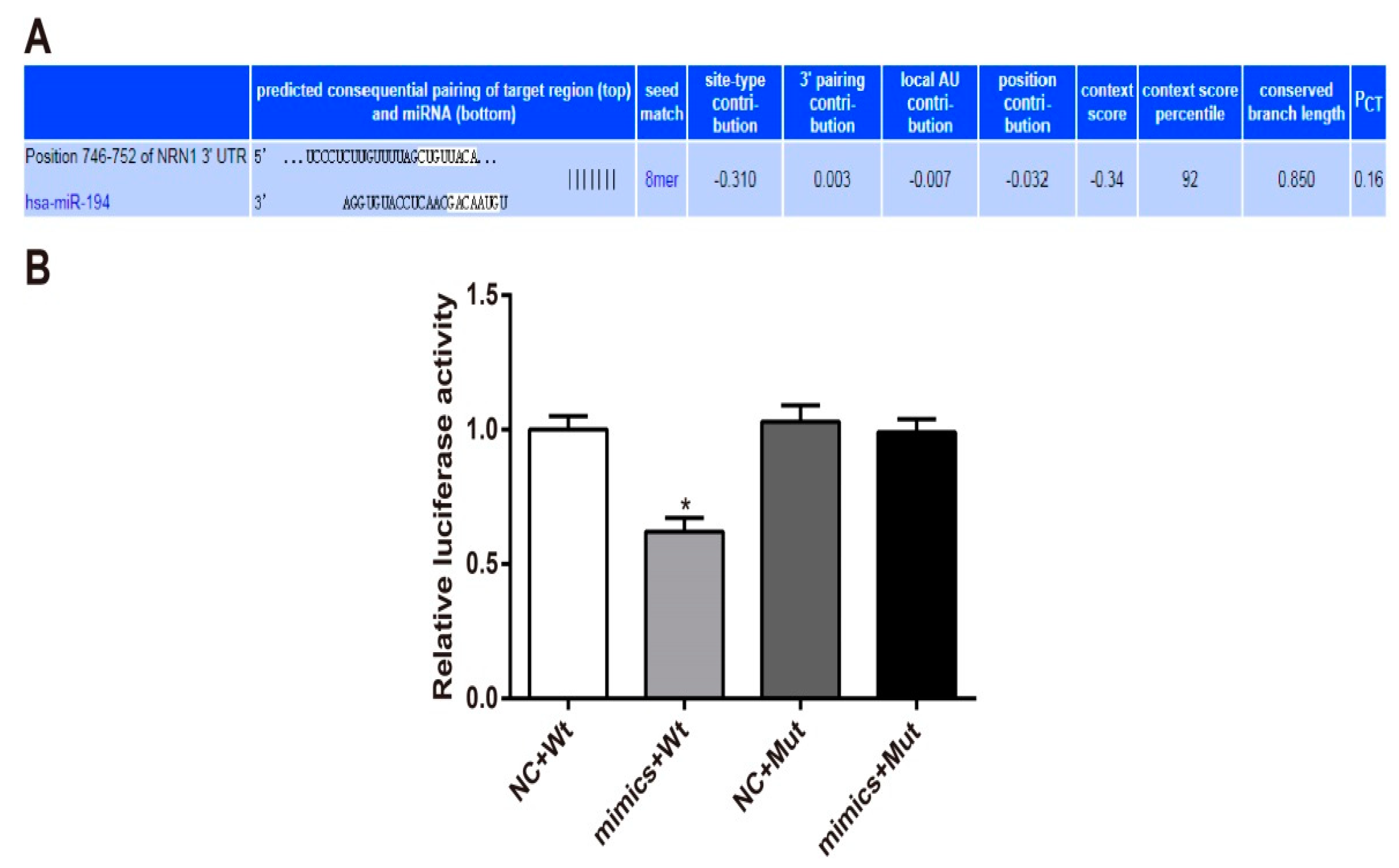
3.5. Nrn1 Is A Target Gene of miR-194
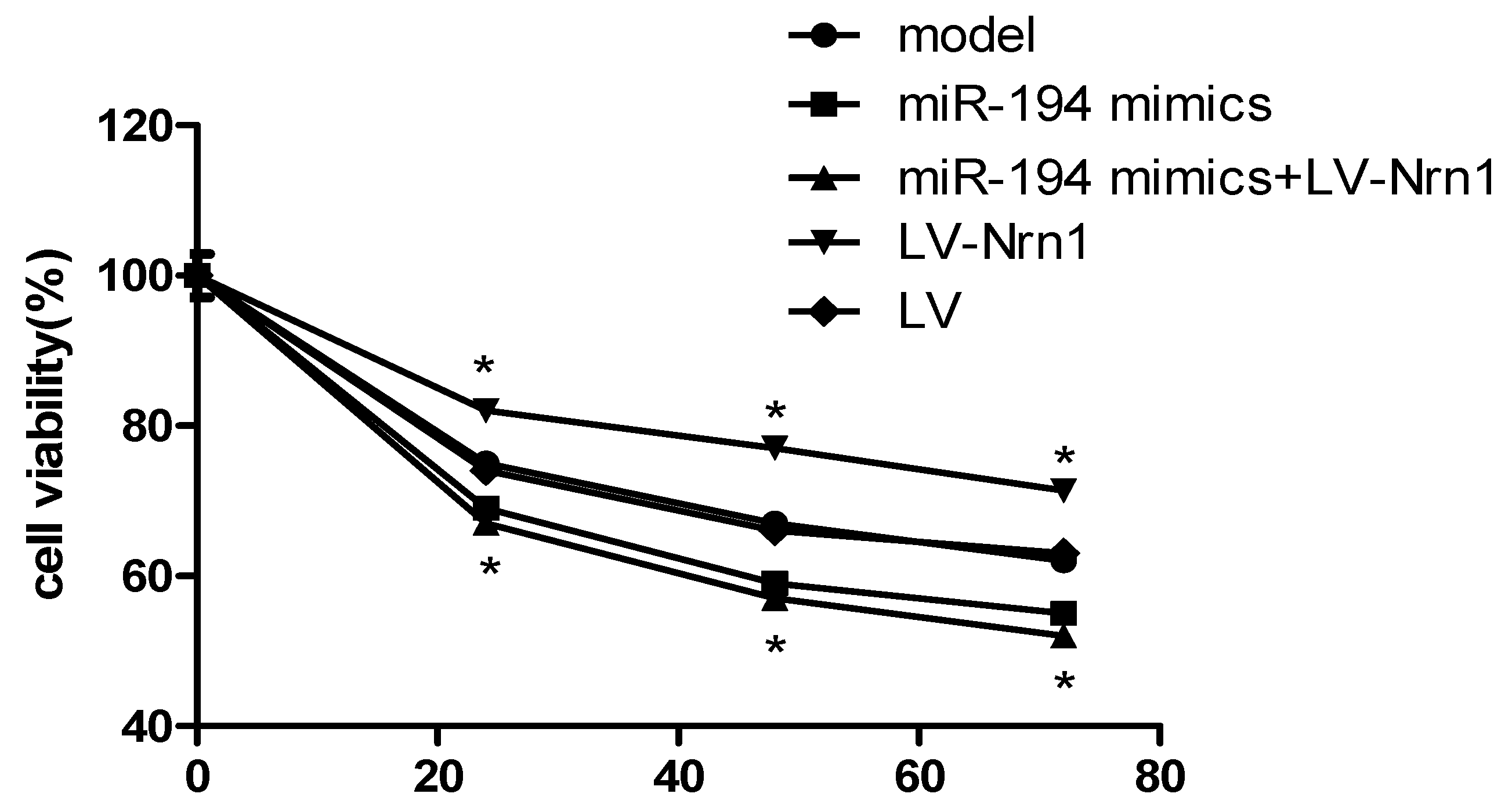
3.6. Nrn1 Repaired the Damage of miR-194 on Aβ1–42-Transduced Hippocampal Neuron Viability
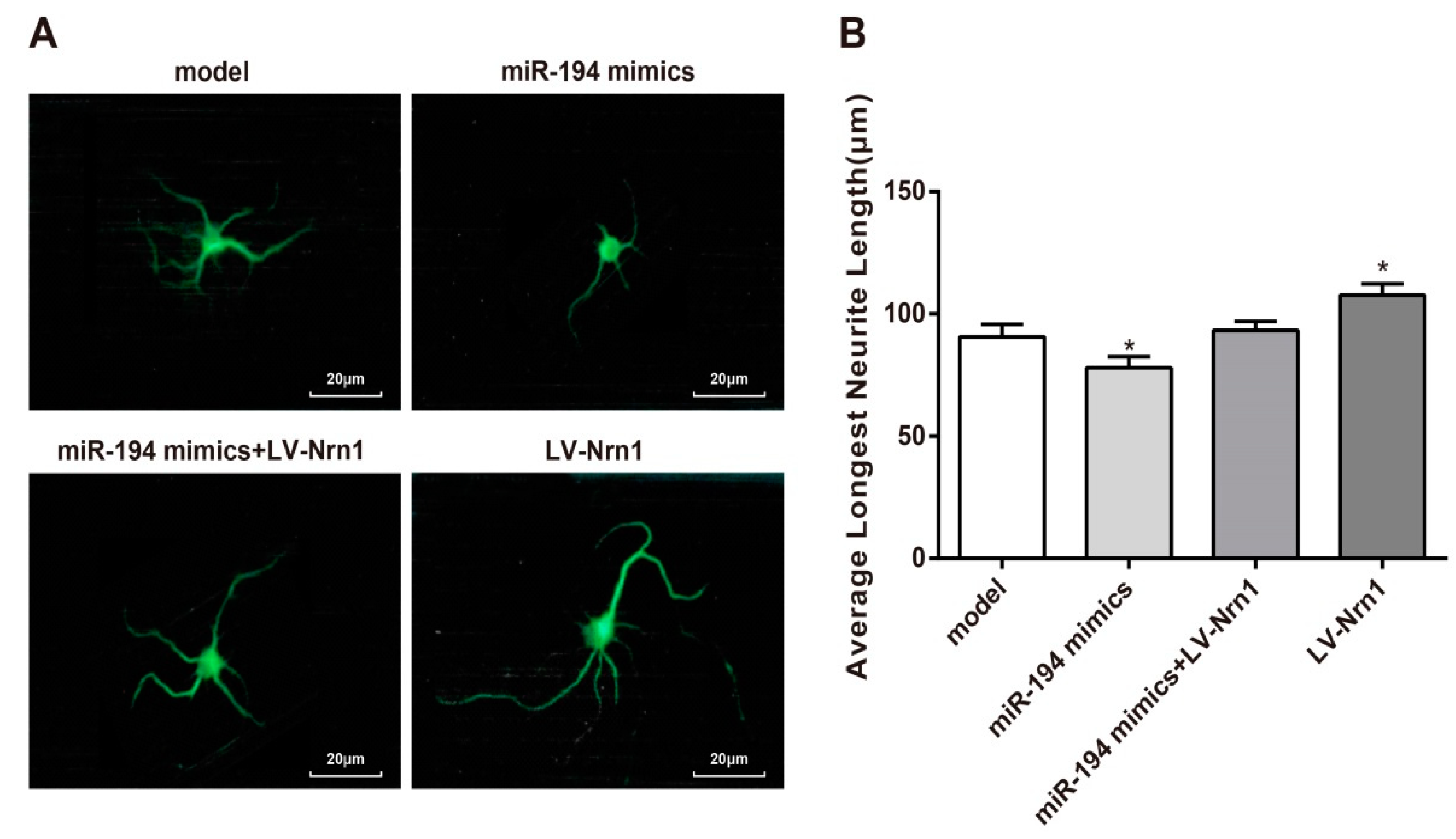
3.7. Nrn1 Reversed the Inhibitory Effect of miR-194 on Neurite Growth of Aβ1–42-Transduced Hippocampal Neurons
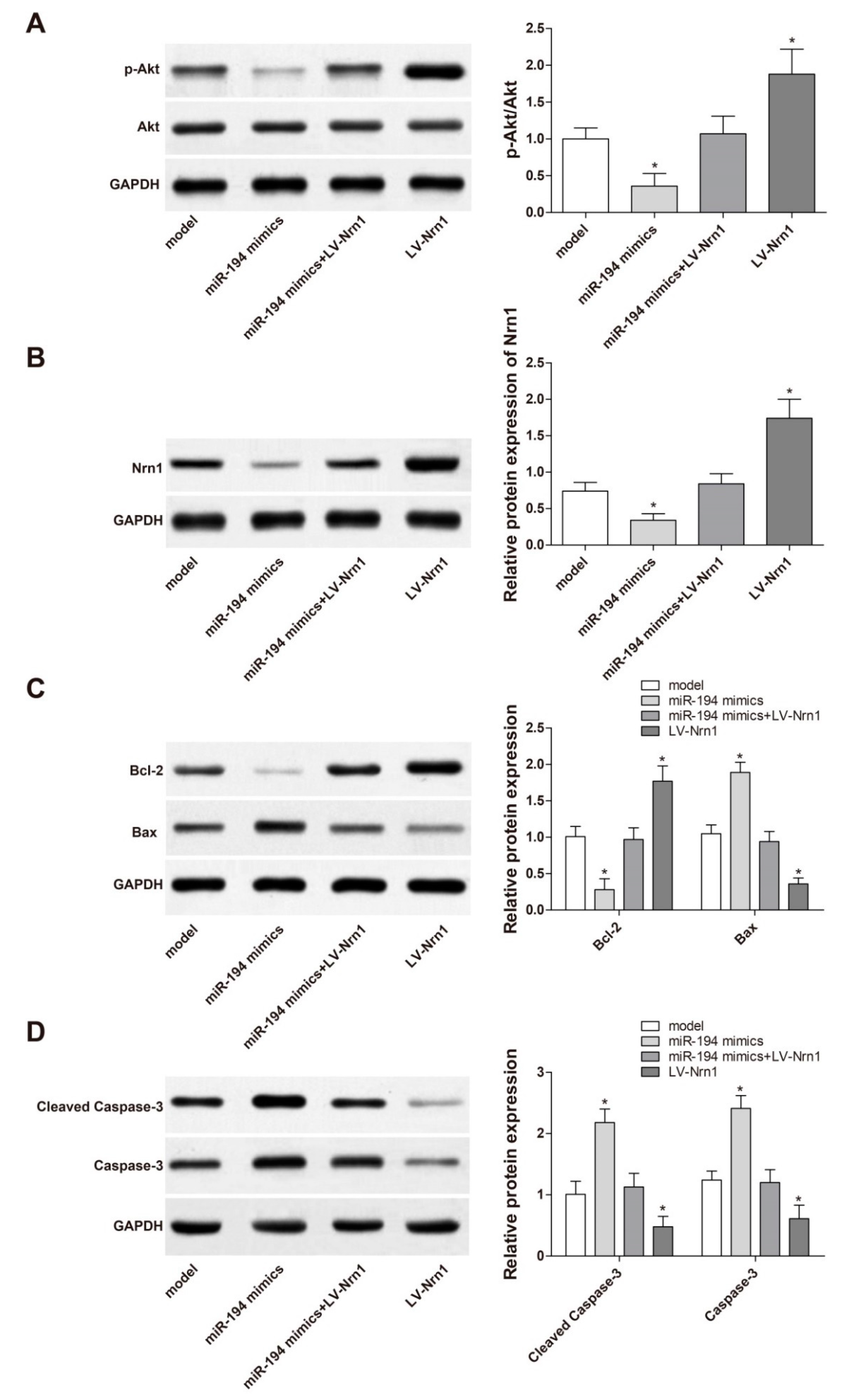
3.8. miR-194 Regulated Apoptosis-Related Protein Expression in Aβ1–42-Transduced Hippocampal Neurons by Inhibiting the Activity of PI3K/Akt Signaling Pathway
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Xu, J.; Begley, P.; Church, S.J.; Patassini, S.; Hollywood, K.A.; Jüllig, M.; Curtis, M.A.; Waldvogel, H.J.; Faull, R.L.M.; Unwin, R.D. Graded perturbations of metabolism in multiple regions of human brain in Alzheimer’s disease: Snapshot of a pervasive metabolic disorder. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2016, 1862, 1084–1092. [Google Scholar] [CrossRef] [PubMed]
- Förstl, H.; Sattel, H.; Bahro, M. Alzheimer’s disease: Clinical features. Int. Rev. Psychiatry 2009, 5, 327–349. [Google Scholar] [CrossRef]
- Lupton, M.K.; Strike, L.; Hansell, N.K.; Wei, W.; Mather, K.A.; Armstrong, N.J.; Thalamuthu, A.; Mcmahon, K.L.; Zubicaray, G.I.D.; Assareh, A.A. The effect of increased genetic risk for Alzheimer’s disease on hippocampal and amygdala volume. Neurobiol. Aging 2016, 40, 68–77. [Google Scholar] [PubMed]
- Kim, H.J.; Cha, J.; Lee, J.M.; Shin, J.S.; Jung, N.Y.; Kim, Y.J.; Choe, Y.S.; Lee, K.H.; Kim, S.T.; Kim, J.S. Distinctive Resting State Network Disruptions Among Alzheimer’s Disease, Subcortical Vascular Dementia, and Mixed Dementia Patients. Alzheimers Dement. J. Alzheimers Assoc. 2016, 11, P412. [Google Scholar] [CrossRef]
- Koyama, A.; Hashimoto, M.; Tanaka, H.; Fujise, N.; Matsushita, M.; Miyagawa, Y.; Hatada, Y.; Fukuhara, R.; Hasegawa, N.; Todani, S. Malnutrition in Alzheimer’s Disease, Dementia with Lewy Bodies, and Frontotemporal Lobar Degeneration: Comparison Using Serum Albumin, Total Protein, and Hemoglobin Level. PLoS ONE 2016, 11, e0157053. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Lee, J.E. miR-155 is involved in Alzheimer’s disease by regulating T lymphocyte function. Fronti. Aging Neurosci. 2015, 7, 61. [Google Scholar] [CrossRef]
- Asai, M.; Kawakubo, T.; Mori, R.; Iwata, N. Elucidating Pathogenic Mechanisms of Early-onset Alzheimer’s Disease in Down Syndrome Patients. Yakugaku Zasshi J. Pharm. Soc. Jpn. 2017, 137, 801. [Google Scholar] [CrossRef] [PubMed]
- Ruizriquelme, A.; Lau, H.; Stuart, E.; Goczi, A.N.; Wang, Z.; Schmittulms, G.; Watts, J.C. Prion-like propagation of β-amyloid aggregates in the absence of APP overexpression. Acta Neuropathol. Commun. 2018, 6, 26. [Google Scholar] [CrossRef] [PubMed]
- Kelly, B.L.; Ferreira, A. beta-Amyloid-induced dynamin 1 degradation is mediated by N-methyl-d-aspartate receptors in hippocampal neurons. J. Biol. Chem. 2016, 281, 28079. [Google Scholar] [CrossRef]
- Ma, B.; Meng, X.; Wang, J.; Sun, J.; Ren, X.; Qin, M.; Sun, G.; Sun, X. Notoginsenoside R1 attenuates amyloid-β-induced damage in neurons by inhibiting reactive oxygen species and modulating MAPK activation. Int. Immunopharmacol. 2014, 22, 151–159. [Google Scholar] [CrossRef] [PubMed]
- Zhao, F.; Han, J.; Chen, X.; Wang, J.; Wang, X.; Sun, J.; Chen, Z. miR-223 enhances the sensitivity of non-small cell lung cancer cells to erlotinib by targeting the insulin-like growth factor-1 receptor. Int. J. Mol. Med. 2016, 38, 183–191. [Google Scholar] [CrossRef]
- Jia, L.H.; Liu, Y.N. Downregulated serum miR-223 servers as biomarker in Alzheimer’s disease. Cell Biochem. Funct. 2016, 34, 233–237. [Google Scholar] [CrossRef]
- Moncini, S.; Lunghi, M.; Valmadre, A.; Grasso, M.; Del, V.V.; Riva, P.; Denti, M.A.; Venturin, M. The miR-15/107 Family of microRNA Genes Regulates CDK5R1/p35 with Implications for Alzheimer’s Disease Pathogenesis. Mol. Neurobiol. 2017, 54, 4329–4354. [Google Scholar] [CrossRef] [PubMed]
- An, F.; Gong, G.; Wang, Y.; Bian, M.; Yu, L.; Wei, C. MiR-124 acts as a target for Alzheimer’s disease by regulating BACE1. Oncotarget 2017, 8, 114065–114071. [Google Scholar] [CrossRef] [PubMed]
- Pierre, L.; Koen, B.; Rekin’S, J.; Evgenia, S.; Sala, F.C.; Shahar, B.; Roy, R.; Sierksma, A.S.R.; Amantha, T.; David, G. Alteration of the microRNA network during the progression of Alzheimer’s disease. Embo Mol. Med. 2013, 5, 1613–1634. [Google Scholar]
- Sørensen, S.S.; Nygaard, A.B.; Christensen, T. miRNA expression profiles in cerebrospinal fluid and blood of patients with Alzheimer’s disease and other types of dementia—An exploratory study. Transl. Neurodegener. 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Geng, D.; Kang, L.; Su, Y.; Jia, J.; Ma, J.; Li, S.; Du, J.; Cui, H. Protective effects of EphB2 on Abeta1-42 oligomer-induced neurotoxicity and synaptic NMDA receptor signaling in hippocampal neurons. Neurochem. Int. 2013, 63, 283–290. [Google Scholar] [CrossRef]
- Luikart, B.W.; Zhang, W.; Wayman, G.A.; Kwon, C.H.; Westbrook, G.L.; Parada, L.F. Neurotrophin-Dependent Dendritic Filopodial Motility: A Convergence on PI3K Signaling. J. Neurosci. 2008, 28, 7006–7012. [Google Scholar] [CrossRef]
- Zhang, M.; Liu, Y.; Liu, M.; Liu, B.; Li, N.; Dong, X.; Hong, Z.; Chai, Y. UHPLC-QTOF/MS-based metabolomics investigation for the protective mechanism of Danshen in Alzheimer’s disease cell model induced by Abeta1-42. Metabolomics 2019, 15, 13. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Lei, B.; Wu, H.; Zhang, X.; Zheng, N. Tumor suppressive role of miR-194-5p in glioblastoma multiforme. Mol. Med. Rep. 2017, 16, 9317–9322. [Google Scholar] [CrossRef]
- Li, P.; Yang, Y.; Liu, H.; Yang, A.K.; Di, J.M.; Tan, G.M.; Wang, H.F.; Qiu, J.G.; Zhang, W.J.; Jiang, Q.W. MiR-194 functions as a tumor suppressor in laryngeal squamous cell carcinoma by targeting Wee1. J. Hematol. Oncol. 2016, 10, 32. [Google Scholar] [CrossRef] [PubMed]
- Bao, J.; Zou, J.H.; Li, C.Y.; Zheng, G.Q. miR-194 inhibits gastric cancer cell proliferation and tumorigenesis by targeting KDM5B. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 4487. [Google Scholar] [PubMed]
- Nedivi, E.; Fieldust, S.; Theill, L.E.; Hevron, D. A set of genes expressed in response to light in the adult cerebral cortex and regulated during development. Proc. Natl. Acad. Sci. USA 1996, 93, 2048–2053. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Zhou, J. Neuritin, a neurotrophic factor in nervous system physiology. Curr. Med. Chem. 2014, 21, 1212. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Li, X.; Shan, L.; Zhu, J.; Chen, R.; Li, Y.; Yuan, W.; Yang, L.; Huang, J. Corrigendum: Recombinant hNeuritin Promotes Structural and Functional Recovery of Sciatic Nerve Injury in Rats. Front. Neurosci. 2016, 10, 589. [Google Scholar] [CrossRef] [PubMed]
- Sharma, T.P.; Liu, Y.; Wordinger, R.J.; Pang, I.H.; Clark, A.F. Neuritin 1 promotes retinal ganglion cell survival and axonal regeneration following optic nerve crush. Cell Death Dis. 2015, 6, e1661. [Google Scholar] [CrossRef] [PubMed]
- Karamoysoyli, E.; Burnand, R.C.; Tomlinson, D.R.; Gardiner, N.J. Neuritin mediates nerve growth factor-induced axonal regeneration and is deficient in experimental diabetic neuropathy. Diabetes 2008, 57, 181. [Google Scholar] [CrossRef]
- Zhang, H.; He, X.; Wang, Y.; Sun, X.; Zhu, L.; Lei, C.; Yin, J.; Li, X.; Hou, F.; He, W. Neuritin attenuates early brain injury in rats after experimental subarachnoid hemorrhage. Int. J. Neurosci. 2017, 127, 1087–1095. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Zhang, H.; Xu, J.; Zhao, D. Neuritin Provides Neuroprotection Against Experimental Traumatic Brain Injury in Rats. Int. J. Neurosci. 2018, 26, 811–820. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhi, X.; Shi, S.; Tao, R.; Chen, P.; Sun, S.; Bian, L.; Xu, Z.; Ma, L. SPOCK1 is up-regulated and promotes tumor growth via the PI3K/AKT signaling pathway in colorectal cancer. Biochem. Biophys. Res. Commun. 2017, 482, 870–876. [Google Scholar] [CrossRef]
- Xie, Q.; Wen, H.; Zhang, Q.; Zhou, W.; Lin, X.; Xie, D.; Liu, Y. Inhibiting PI3K-AKt signaling pathway is involved in antitumor effects of ginsenoside Rg3 in lung cancer cell. Biomed. Pharmacother. 2017, 85, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Kang, T.; Qi, B.; Kong, L.; Jiao, Y.; Cao, Y.; Zhang, J.; Yang, J. Neuroprotective effects of ginseng protein on PI3K/Akt signaling pathway in the hippocampus of d-galactose/AlCl3 inducing rats model of Alzheimer’s disease. J. Ethnopharmacol. 2016, 179, 162–169. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Wang, Y.; Li, H.; Xiong, R.; Zhao, Z.; Chu, X.; Li, Q.; Sun, S.; Chen, S. Neuroprotective effects of salidroside through PI3K/Akt pathway activation in Alzheimer’s disease models. Drug Des. Dev. Ther. 2016, 10, 1335–1343. [Google Scholar]
- Chong, Y.H.; Shin, Y.J.; Lee, E.O.; Kayed, R.; Glabe, C.G.; Tenner, A.J. ERK1/2 activation mediates Abeta oligomer-induced neurotoxicity via caspase-3 activation and tau cleavage in rat organotypic hippocampal slice cultures. J. Biol. Chem. 2006, 281, 20315–20325. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, T.; Cheng, Y.; Han, H.; Liu, J.; Tian, B.; Liu, X. miR-194 Accelerates Apoptosis of Aβ1–42-Transduced Hippocampal Neurons by Inhibiting Nrn1 and Decreasing PI3K/Akt Signaling Pathway Activity. Genes 2019, 10, 313. https://doi.org/10.3390/genes10040313
Wang T, Cheng Y, Han H, Liu J, Tian B, Liu X. miR-194 Accelerates Apoptosis of Aβ1–42-Transduced Hippocampal Neurons by Inhibiting Nrn1 and Decreasing PI3K/Akt Signaling Pathway Activity. Genes. 2019; 10(4):313. https://doi.org/10.3390/genes10040313
Chicago/Turabian StyleWang, Tingting, Yaling Cheng, Haibin Han, Jie Liu, Bo Tian, and Xiaocui Liu. 2019. "miR-194 Accelerates Apoptosis of Aβ1–42-Transduced Hippocampal Neurons by Inhibiting Nrn1 and Decreasing PI3K/Akt Signaling Pathway Activity" Genes 10, no. 4: 313. https://doi.org/10.3390/genes10040313
APA StyleWang, T., Cheng, Y., Han, H., Liu, J., Tian, B., & Liu, X. (2019). miR-194 Accelerates Apoptosis of Aβ1–42-Transduced Hippocampal Neurons by Inhibiting Nrn1 and Decreasing PI3K/Akt Signaling Pathway Activity. Genes, 10(4), 313. https://doi.org/10.3390/genes10040313




