DNA Barcode Reveals the Bycatch of Endangered Batoids Species in the Southwest Atlantic: Implications for Sustainable Fisheries Management and Conservation Efforts
Abstract
1. Introduction
2. Materials and Methods
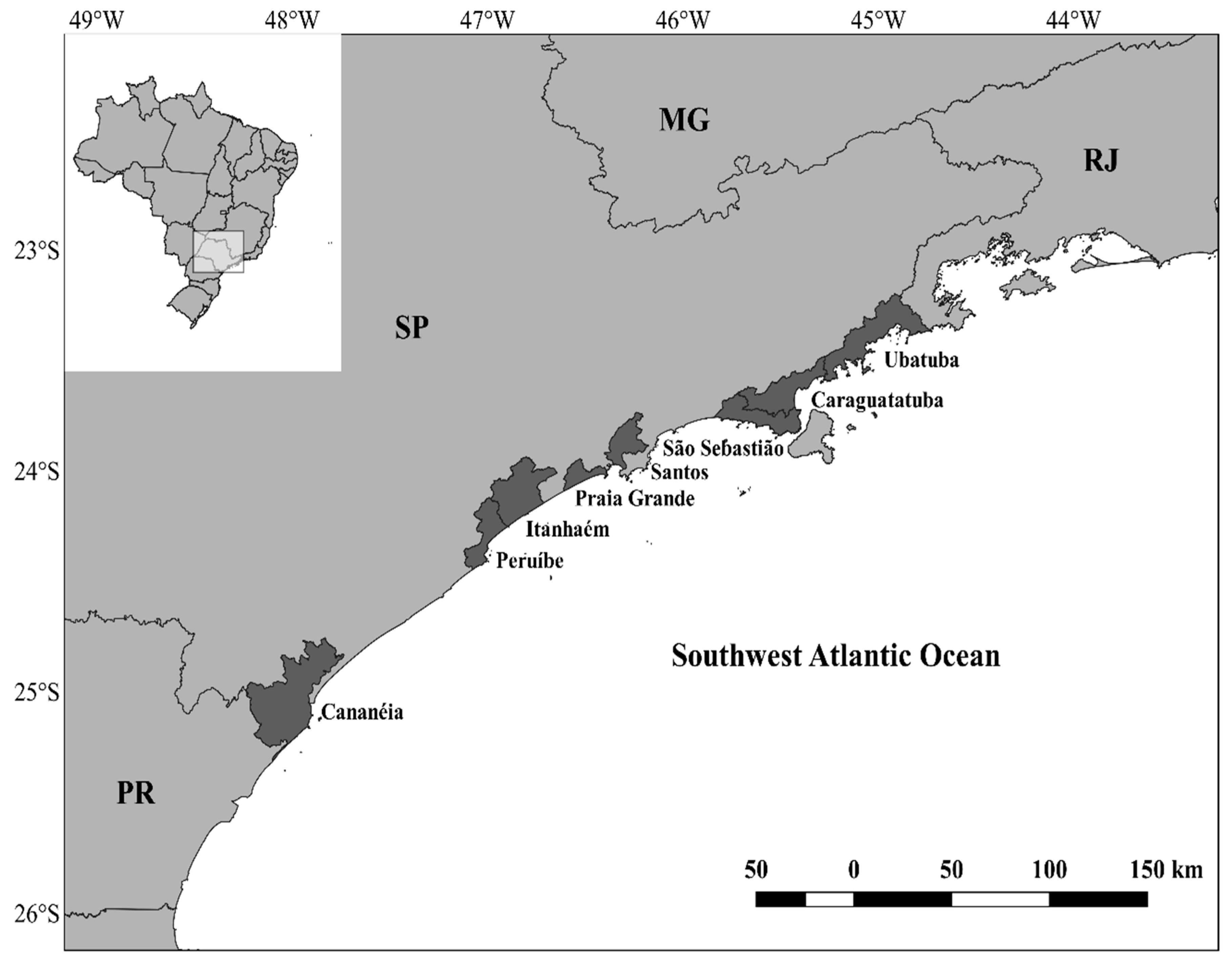
2.1. Sampling
2.2. DNA Extraction, Amplification, and Sequencing
2.3. DNA-Based Species Identification
3. Results
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Species | GenBank Access Number |
|---|---|
| Atlantoraja castelnaui | GU805239 |
| Atlantoraja castelnaui | GU805231 |
| Atlantoraja castelnaui | JX124737 |
| Atlantoraja castelnaui | JQ365232 |
| Hypanus americanus | KJ719465 |
| Hypanus americanus | KT075327 |
| Hypanus americanus | KF461168 |
| Bathytoshia centroura | KT075311 |
| Bathytoshia centroura | KJ719464 |
| Bathytoshia centroura | KF808191 |
| Hypanus guttatus | JX034000 |
| Hypanus guttatus | KJ719467 |
| Dasyatis sp. | JX124776 |
| Gymnura altavela | KC015432 |
| Gymnura altavela | BIM344-13 |
| Gymnura altavela | BIM365-13 |
| Gymnura altavela | KF808207 |
| Myliobatis freminvillei | KT075326 |
| Myliobatis freminvillei | KF930155 |
| Myliobatis goodei | JQ305804 |
| Myliobatis goodei | JQ305809 |
| Myliobatis goodei | JQ305807 |
| Myliobatis goodei | JQ305806 |
| Narcine brasiliensis | JQ365452 |
| Narcine brasiliensis | JX124815 |
| Narcine brasiliensis | JX034007 |
| Narcine brasiliensis | KF930170 |
| Pteroplatytrygon violacea | KT307373 |
| Pteroplatytrygon violacea | KT307370 |
| Pteroplatytrygon violacea | KF930342 |
| Pteroplatytrygon violacea | KF808209 |
| Pseudobatos horkelii | JX034017 |
| Pseudobatos horkelii | HM424136 |
| Pseudobatos horkelii | JX034016 |
| Pseudobatos horkelii | JX124879 |
| Rhinoptera bonasus | JX241049 |
| Rhinoptera bonasus | JX241050 |
| Rhinoptera bonasus | JX241051 |
| Rhinoptera bonasus | KF245598 |
| Rhinoptera brasiliensis | JX124888 |
| Rioraja agassizii | GU805208 |
| Rioraja agassizii | GU805206 |
| Rioraja agassizii | GU805204 |
| Rioraja agassizii | GU805205 |
| Squatina guggenheim | FN431749 |
| Squatina guggenheim | FN431748 |
| Squatina guggenheim | FN431747 |
| Squatina guggenheim | FN431746 |
| Callorhinchus callorynchus | KP719792 |
| Callorhinchus callorynchus | KP719793 |
| Callorhinchus callorynchus | KP719794 |
| Callorhinchus callorynchus | KP719799 |
| Fishing Gear | 2012 | 2013 | 2014 | 2015 | 2016 | 2017 | 2018 | Total | % |
|---|---|---|---|---|---|---|---|---|---|
| otter-trawl | 135,026.0 | 98,975.8 | 81,566.8 | 82,897.1 | 46,071.4 | 20,935.4 | 28,360.7 | 493,833.0 | 36.34 |
| fish-traps | 1765.74 | 2457.95 | 1891.0 | 804.2 | 970.35 | 1069.9 | 2384.96 | 11,344.1 | 0.83 |
| gillnet | 112,890.0 | 85,540.3 | 70,005.7 | 57,010.3 | 91,245.7 | 76,519.1 | 88,560.2 | 581,771.0 | 42.82 |
| beam-trawl | 117,079.0 | 46,060.0 | 27,992.0 | 28,194.0 | 24,294.0 | 17,703.0 | 10,516.0 | 271,838.0 | 20.01 |
| Total | 366,761.0 | 233,034.0 | 181,456.0 | 168,906.0 | 162,581.0 | 116,227.0 | 129,822.0 | 1,358,786.0 | 100.00 |
| % | 26.99 | 17.15 | 13.35 | 12.43 | 11.97 | 8.55 | 9.55 | 100.00 |
References
- Last, P.; Naylor, G.; Séret, B.; White, W.; de Carvalho, M.; Stehmann, M. Rays of the World, 1st ed.; CSIRO Publishing: Clayton, Australia, 2016; p. 800. [Google Scholar]
- Weigmann, S. Annotated checklist of the living sharks, batoids and chimaeras (Chondrichthyes) of the world, with a focus on biogeographical diversity. J. Fish Biol. 2016, 88, 837–1037. [Google Scholar] [CrossRef] [PubMed]
- Oliver, S.; Braccini, M.; Newman, S.J.; Harvey, E.S. Global patterns in the bycatch of sharks and rays. Mar. Policy 2015, 54, 86–97. [Google Scholar] [CrossRef]
- Dulvy, N.K.; Fowler, S.L.; Musick, J.A.; Cavanagh, R.D.; Kyne, P.M.; Harrison, L.R.; Carlson, J.K.; Davidson, L.N.; Fordham, S.V.; Francis, M.P.; et al. Extinction risk and conservation of the world’s sharks and rays. elife 2014, 3, e00590. [Google Scholar] [CrossRef] [PubMed]
- The IUCN Red List of Threatened Species. Version 2018-2. Available online: http://www.iucnredlist.org (accessed on 25 February 2019).
- FAO Fisheries and Aquaculture Department, Food and Agriculture Organization of the United Nations. Capture Fisheries. Available online: http://www.fao.org/fishery/ (accessed on 25 February 2019).
- Pauly, D.; Zeller, D. Catch reconstructions reveal that global marine fisheries catches are higher than reported and declining. Nat. Commun. 2016, 7, 10244. [Google Scholar] [CrossRef] [PubMed]
- Hall, M.A. On bycatches. Rev. Fish Biol. Fish. 1996, 6, 319–352. [Google Scholar] [CrossRef]
- Gray, C.A.; Kennelly, S.J. Bycatches of endangered, threatened and protected species in marine fisheries. Rev. Fish Biol. Fish. 2018, 28, 521–541. [Google Scholar] [CrossRef]
- Bellido, J.M.; Santos, M.B.; Pennino, M.G.; Valeiras, X.; Pierce, G.J. Fishery discards and bycatch: Solutions for an ecosystem approach to fisheries management? Hydrobiologia 2011, 670, 317. [Google Scholar] [CrossRef]
- Johnsen, J.P.; Eliasen, S. Solving complex fisheries management problems: What the EU can learn from the Nordic experiences of reduction of discards. Mar. Policy 2011, 35, 130–139. [Google Scholar] [CrossRef]
- Dulvy, N.K.; Baum, J.K.; Clarke, S.; Compagno, L.J.V.; Cortés, E.; Domingo, A.; Fordham, S.; Fowler, S.; Francis, M.P.; Gibson, C.; et al. You can swim but you can’t hide: The global status and conservation of oceanic pelagic sharks and rays. Aquat. Conserv. 2008, 18, 459–482. [Google Scholar] [CrossRef]
- Davies, R.W.; Cripps, S.J.; Nickson, A.; Porter, G. Defining and estimating global marine fisheries bycatch. Mar. Policy 2009, 33, 661–672. [Google Scholar] [CrossRef]
- Stevens, J.D.; Bonfil, R.; Dulvy, N.K.; Walker, P.A. The effects of fishing on sharks, rays, and chimaeras (chondrichthyans), and the implications for marine ecosystems. ICES J. Mar. Sci. 2000, 57, 476–494. [Google Scholar] [CrossRef]
- Zeller, D.; Cashion, T.; Palomares, M.; Pauly, D. Global marine fisheries discards: A synthesis of reconstructed data. Fish Fish. 2018, 19, 30–39. [Google Scholar] [CrossRef]
- Stobutzki, I.C.; Miller, M.J.; Heales, D.S.; Brewer, D.T. Sustainability of elasmobranchs caught as bycatch in a tropical prawn (shrimp) trawl fishery. Fish. Bull. 2002, 100, 800–821. [Google Scholar]
- Bornatowski, H.; Braga, R.R.; Vitule, J.R. Threats to sharks in a developing country: The need for effective simple conservation measures. Nat. Conserv. 2014, 12, 11–18. [Google Scholar] [CrossRef]
- Bornatowski, H.; Braga, R.R.; Vitule, J.R. Shark mislabeling threatens biodiversity. Science 2013, 340, 923. [Google Scholar] [CrossRef]
- Hebert, P.D.; Cywinska, A.; Ball, S.L.; Dewaard, J.R. Biological identifications through DNA barcodes. Proc.-R. Soc. Lond. Biol. Sci. 2003, 270, 313–321. [Google Scholar] [CrossRef] [PubMed]
- Wetterstrand, K.A. DNA Sequencing Costs: Data from the NHGRI Genome Sequencing Program (GSP). Available online: http://www.genome.gov/sequencingcostsdata (accessed on 25 February 2019).
- Dudgeon, C.L.; Blower, D.C.; Broderick, D.; Giles, J.L.; Holmes, B.J.; Kashiwagi, T.; Krück, N.C.; Morgan, J.A.; Tillett, B.J.; Ovenden, J.R. A review of the application of molecular genetics for fisheries management and conservation of sharks and rays. J. Fish Biol. 2012, 80, 1789–1843. [Google Scholar] [CrossRef] [PubMed]
- Shivji, M.; Clarke, S.; Pank, M.; Natanson, L.; Kohler, N.; Stanhope, M. Genetic identification of pelagic shark body parts for conservation and trade monitoring. Conserv. Biol. 2002, 16, 1036–1047. [Google Scholar] [CrossRef]
- Clarke, S.C.; Magnussen, J.E.; Abercrombie, D.L.; McAllister, M.K.; Shivji, M.S. Identification of shark species composition and proportion in the Hong Kong shark fin market based on molecular genetics and trade records. Conserv. Biol. 2006, 20, 201–211. [Google Scholar] [CrossRef]
- Clarke, S.C.; McAllister, M.K.; Milner-Gulland, E.J.; Kirkwood, G.P.; Michielsens, C.G.; Agnew, D.J.; Pikitch, E.K.; Nakano, H.; Shivji, M.S. Global estimates of shark catches using trade records from commercial markets. Ecol. Lett. 2006, 9, 1115–1126. [Google Scholar] [CrossRef]
- Mendonça, F.F.; Hashimoto, D.T.; Porto-Foresti, F.; Oliveira, C.; Gadig, O.B.; Foresti, F. Identification of the shark species Rhizoprionodon lalandii and R. porosus (Elasmobranchii, Carcharhinidae) by multiplex PCR and PCR-RFLP techniques. Mol. Ecol. Resour. 2009, 9, 771–773. [Google Scholar] [CrossRef] [PubMed]
- Bunholi, I.V.; Ferrette, B.L.S.; De Biasi, J.B.; Magalhães, C.O.; Rotundo, M.M.; Oliveira, C.; Foresti, F.; Mendonça, F.F. The fishing and illegal trade of the angelshark: DNA barcoding against misleading identifications. Fish. Res. 2018, 206, 193–197. [Google Scholar] [CrossRef]
- Fields, A.T.; Fischer, G.A.; Shea, S.K.; Zhang, H.; Abercrombie, D.L.; Feldheim, K.A.; Babcock, E.A.; Chapman, D.D. Species composition of the international shark fin trade assessed through a retail-market survey in Hong Kong. Conserv. Biol. 2018, 32, 376–389. [Google Scholar] [CrossRef]
- De-Franco, B.A.; Mendonça, F.F.; Oliveira, C.; Foresti, F. Illegal trade of the guitarfish Rhinobatos horkelii on the coasts of central and southern Brazil: Genetic identification to aid conservation. Aquat. Conserv. 2012, 22, 272–276. [Google Scholar] [CrossRef]
- Flowers, K.I.; Ajemian, M.J.; Bassos-Hull, K.; Feldheim, K.A.; Hueter, R.E.; Papastamatiou, Y.P.; Chapman, D.D. A review of batoid philopatry, with implications for future research and population management. Mar. Ecol. Prog. Ser. 2016, 562, 251–261. [Google Scholar] [CrossRef]
- Domingues, R.R.; Hilsdorf, A.W.; Gadig, O.B. The importance of considering genetic diversity in shark and ray conservation policies. Conserv. Genet. 2018, 19, 501–525. [Google Scholar] [CrossRef]
- Ward, R.D.; Zemlak, T.S.; Innes, B.H.; Last, P.R.; Hebert, P.D. DNA barcoding Australia’s fish species. Proc.-R. Soc. Lond. Biol. Sci. 2005, 360, 1847–1857. [Google Scholar] [CrossRef]
- Van Velzen, R.; Weitschek, E.; Felici, G.; Bakker, F.T. DNA barcoding of recently diverged species: Relative performance of matching methods. PLoS ONE 2012, 7, e30490. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D. BOLD: The Barcode of Life Data System. Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef]
- Sayers, E.W.; Cavanaugh, M.; Clark, K.; Ostell, J.; Pruitt, K.D.; Karsch-Mizrachi, I. GenBank. Nucleic Acids Res. 2018, 47, 94–99. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Anisimova, M.; Gascuel, O. Approximate likelihood-ratio test for branches: A fast, accurate, and powerful alternative. Syst. Biol. 2006, 55, 539–552. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.; Xie, Z.; Salemi, M.; Chen, L.; Wang, Y. An index of substitution saturation and its application. Mol. Phylogenet. Evol. 2003, 26, 1–7. [Google Scholar] [CrossRef]
- Xia, X. DAMBE6: New tools for microbial genomics, phylogenetics, and molecular evolution. J. Hered. 2017, 108, 431–437. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef]
- Towns, J.; Cockerill, T.; Dahan, M.; Foster, I.; Gaither, K.; Grimshaw, A.; Hazlewood, V.; Lathrop, S.; Lifka, D.; Peterson, G.D.; et al. XSEDE: Accelerating scientific discovery. Comput. Sci. Eng. 2014, 16, 62–74. [Google Scholar] [CrossRef]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In 2010 Gateway Computing Environments Workshop (GCE); IEEE: Piscataway, NJ, USA, 2010; pp. 1–8. [Google Scholar]
- Suchard, M.A.; Lemey, P.; Baele, G.; Ayres, D.L.; Drummond, A.J.; Rambaut, A. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol. 2018, 4, vey016. [Google Scholar] [CrossRef] [PubMed]
- Drummond, A.J.; Ho, S.Y.; Phillips, M.J.; Rambaut, A. Relaxed phylogenetics and dating with confidence. PLoS Biol. 2006, 4, e88. [Google Scholar] [CrossRef]
- Gernhard, T.; Hartmann, K.; Steel, M. Stochastic properties of generalised Yule models, with biodiversity applications. J. Math. Biol. 2008, 57, 713–735. [Google Scholar] [CrossRef]
- Yule, G.U. A mathematical theory of evolution, based on the conclusions of Dr. J.C. Willis, F.R.S. Proc.-R. Soc. Lond. Biol. Sci. 1925, 213, 21–87. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A.J.; Xie, D.; Baele, G.; Suchard, M.A. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst. Biol. 2018, 67, 901–904. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D. A DNA-based registry for all animal species: The Barcode Index Number (BIN) system. PLoS ONE 2013, 8, e66213. [Google Scholar] [CrossRef]
- Croll, D.A.; Dewar, H.; Dulvy, N.K.; Fernando, D.; Francis, M.P.; Galván-Magaña, F.; Hall, M.; Heinrichs, S.; Marshall, A.; Mccauley, D.; et al. Vulnerabilities and fisheries impacts: The uncertain future of manta and devil rays. Aquat. Conserv. 2016, 26, 562–575. [Google Scholar] [CrossRef]
- Mas, F.; Forselledo, R.; Domingo, A. Mobulid ray by-catch in longline fisheries in the south-western Atlantic Ocean. Mar. Freshw. Res. 2015, 66, 767–777. [Google Scholar] [CrossRef]
- Rohner, C.A.; Pierce, S.J.; Marshall, A.D.; Weeks, S.J.; Bennett, M.B.; Richardson, A.J. Trends in sightings and environmental influences on a coastal aggregation of manta rays and whale sharks. Mar. Ecol. Prog. Ser. 2013, 482, 153–168. [Google Scholar] [CrossRef]
- Bonfil, R.; Munro, G.; Valtysson, H.T.; Wright, M.; Preikshot, D.; Haggan, N.; Pauly, D.; Sumaila, U.R.; Pitcher, T.J. Distant Water Fleets: An Ecological, Economic and Social Assessment; Fisheries Centre, University of British Columbia: Vancouver, BC, Canada, 1998; p. 122. [Google Scholar]
- Chuenpagdee, R.; Rocklin, D.; Bishop, D.; Hynes, M.; Greene, R.; Lorenzi, M.R.; Devillers, R. The global information system on small-scale fisheries (ISSF): A crowdsourced knowledge platform. Mar. Policy 2017, 101, 158–166. [Google Scholar] [CrossRef]
- Dulvy, N.K.; Metcalfe, J.D.; Glanville, J.; Pawson, M.G.; Reynolds, J.D. Fishery stability, local extinctions, and shifts in community structure in skates. Conserv. Biol. 2000, 14, 283–293. [Google Scholar] [CrossRef]
- Brander, K. Disappearance of common skate, Raia batis, from the Irish Sea. Nature 1981, 290, 48–49. [Google Scholar] [CrossRef]
- Casey, J.M.; Myers, R.A. Near extinction of a large, widely distributed fish. Science 1998, 281, 690–692. [Google Scholar] [CrossRef]
- Tamini, L.L.; Chiaramonte, G.E.; Perez, J.E.; Cappozzo, H.L. Batoids in a coastal trawl fishery of Argentina. Fish. Res. 2006, 77, 326–332. [Google Scholar] [CrossRef]
- Žydelis, R.; Wallace, B.P.; Gilman, E.L.; Werner, T.B. Conservation of marine megafauna through minimization of fisheries bycatch. Conserv. Biol. 2009, 23, 608–616. [Google Scholar] [CrossRef]
- Shester, G.G.; Micheli, F. Conservation challenges for small-scale fisheries: Bycatch and habitat impacts of traps and gillnets. Biol. Conserv. 2011, 144, 1673–1681. [Google Scholar] [CrossRef]
- Senko, J.; White, E.R.; Heppell, S.S.; Gerber, L.R. Comparing bycatch mitigation strategies for vulnerable marine megafauna. Anim. Conserv. 2014, 17, 5–18. [Google Scholar] [CrossRef]
- Thomas, C.D.; Cameron, A.; Green, R.E.; Bakkenes, M.; Beaumont, L.J.; Collingham, Y.C.; Erasmus, B.F.; De Siqueira, M.F.; Grainger, A.; Hannah, L.; et al. Extinction risk from climate change. Nature 2004, 427, 145–148. [Google Scholar] [CrossRef]
- Myers, N.; Mittermeier, R.A.; Mittermeier, C.G.; Da Fonseca, G.A.; Kent, J. Biodiversity hotspots for conservation priorities. Nature 2000, 403, 853–858. [Google Scholar] [CrossRef]
- Olson, D.M.; Dinerstein, E. The Global 200: Priority ecoregions for global conservation. Ann. Mo. Bot. Gard. 2002, 89, 199–224. [Google Scholar] [CrossRef]
- Lucifora, L.O.; García, V.B.; Worm, B. Global diversity hotspots and conservation priorities for sharks. PLoS ONE 2011, 6, e19356. [Google Scholar] [CrossRef]
- Lessa, R.; Vooren, C.M. Pseudobatos horkelii. IUCN Red List Threatened Species. 2016. Available online: http://dx.doi.org/10.2305/IUCN.UK.2016-3.RLTS.T41064A103933918.en (accessed on 25 February 2019).
- Hozbor, N.; Massa, A.; Vooren, C.M. Atlantoraja castelnaui. IUCN Red List Threatened Species. 2004. Available online: http://dx.doi.org/10.2305/IUCN.UK.2004.RLTS.T44575A10921544.en (accessed on 25 February 2019).
- Vooren, C.M.; Lamónaca, A.F. Rhinoptera brasiliensis. IUCN Red List Threatened Species. 2004. Available online: http://dx.doi.org/10.2305/IUCN.UK.2004.RLTS.T44595A10912274.en (accessed on 25 February 2019).
- Lessa, R.; Charvet-Almeida, P.; Santana, F.M.; Almeida, Z. Isogomphodon oxyrhynchus. IUCN Red List Threatened Species. 2006. Available online: http://dx.doi.org/10.2305/IUCN.UK.2006.RLTS.T60218A12323498.en (accessed on 25 February 2019).
- Lessa, R.; Batista, V.S.; Santana, F.M. Close to extinction? The collapse of the endemic daggernose shark (Isogomphodon oxyrhynchus) off Brazil. Glob. Ecol. Conserv. 2016, 7, 70–81. [Google Scholar] [CrossRef]
- Chiaramonte, G.; Vooren, C.M. Squatina guggenheim. IUCN Red List Threat. Species. 2007. Available online: http://dx.doi.org/10.2305/IUCN.UK.2007.RLTS.T39330A10202558.en (accessed on 25 February 2019).
- Dulvy, N.K.; Simpfendorfer, C.A.; Davidson, L.N.; Fordham, S.V.; Bräutigam, A.; Sant, G.; Welch, D.J. Challenges and priorities in shark and ray conservation. Curr. Biol. 2017, 27, 565–572. [Google Scholar] [CrossRef]
- Butchart, S.H.; Bird, J.P. Data deficient birds on the IUCN Red List: What don’t we know and why does it matter? Biol. Conserv. 2010, 143, 239–247. [Google Scholar] [CrossRef]
- Bland, L.M.; Collen, B.E.; Orme, C.D.; Bielby, J.O. Predicting the conservation status of data-deficient species. Conserv. Biol. 2015, 29, 250–259. [Google Scholar] [CrossRef]
- Cerutti-Pereyra, F.; Meekan, M.G.; Wei, N.W.; O’Shea, O.; Bradshaw, C.J.; Austin, C.M. Identification of rays through DNA barcoding: An application for ecologists. PLoS ONE 2012, 7, e36479. [Google Scholar] [CrossRef]
- Last, P.R.; Stevens, J.D. Sharks and Rays of Australia; CSIRO: Canberra, Australia, 1994; p. 84. [Google Scholar]
- Last, P.R.; Naylor, G.J.; Manjaji-Matsumoto, B.M. A revised classification of the family Dasyatidae (Chondrichthyes: Myliobatiformes) based on new morphological and molecular insights. Zootaxa 2016, 4139, 345–368. [Google Scholar] [CrossRef]
- Holmes, B.H.; Steinke, D.; Ward, R.D. Identification of shark and ray fins using DNA barcoding. Fish. Res. 2009, 95, 280–288. [Google Scholar] [CrossRef]
- Zeller, D.; Harper, S.; Zylich, K.; Pauly, D. Synthesis of underreported small-scale fisheries catch in Pacific island waters. Coral Reefs 2015, 34, 25–39. [Google Scholar] [CrossRef]
- Braccini, J.M.; Waltrick, D. Species-specific at-vessel mortality of sharks and rays captured by demersal longlines. Mar. Policy 2019, 99, 94–98. [Google Scholar] [CrossRef]
- Freire, K.M.F.; Aragão, J.A.N.; Araújo, A.R.R.; Ávila-da-Silva, A.O.; Bispo, M.C.S.; Canziani, G.V.; Carneiro, M.H.; Gonçalves, F.D.S.; Keunecke, K.A.; Mendonça, J.T.; et al. Reconstruction of catch statistics for Brazilian marine waters (1950–2010). Fisheries catch reconstructions for Brazil’s mainland and oceanic islands. Fish. Cent. Res. 2015, 23, 3–30. [Google Scholar]
- Hall, S.J.; Mainprize, B.M. Managing by-catch and discards: How much progress are we making and how can we do better? Fish Fish. 2005, 6, 134–155. [Google Scholar] [CrossRef]
- Lewison, R.L.; Soykan, C.U.; Cox, T.; Peckham, H.; Pilcher, N.; LeBoeuf, N.; McDonald, S.; Moore, J.; Safina, C.; Crowder, L.B. Ingredients for addressing the challenges of fisheries bycatch. Bull. Mar. Sci. 2011, 87, 235–250. [Google Scholar] [CrossRef]
- Bradshaw, C.J.; Prowse, T.A.; Drew, M.; Gillanders, B.M.; Donnellan, S.C.; Huveneers, C.; Kuparinen, A. Predicting sustainable shark harvests when stock assessments are lacking. ICES J. Mar. Sci. 2018, 75, 1591–1601. [Google Scholar] [CrossRef]
- Cox, T.M.; Lewison, R.L.; Žydelis, R.; Crowder, L.B.; Safina, C.; Read, A.J. Comparing effectiveness of experimental and implemented bycatch reduction measures: The ideal and the real. Conserv. Biol. 2007, 21, 1155–1164. [Google Scholar] [CrossRef] [PubMed]
- O’keefe, C.E.; Cadrin, S.X.; Stokesbury, K.D. Evaluating effectiveness of time/area closures, quotas/caps, and fleet communications to reduce fisheries bycatch. ICES J. Mar. Sci. 2013, 71, 1286–1297. [Google Scholar] [CrossRef]
- Domingues, R.D.; Amorim, A.F.; Hilsdorf, A.W. Genetic identification of Carcharhinus sharks from the southwest Atlantic Ocean (Chondrichthyes: Carcharhiniformes). J. Appl. Ichthyol. 2013, 29, 738–742. [Google Scholar] [CrossRef]
- Mendonça, F.F.; Hashimoto, D.T.; De-Franco, B.A.; Porto-Foresti, F.; Gadig, O.B.; Oliveira, C.; Foresti, F. Genetic identification of lamniform and carcharhiniform sharks using multiplex-PCR. Conserv. Genet. Resour. 2010, 2, 31–35. [Google Scholar] [CrossRef]
- Palmeira, C.A.; Rodrigues-Filho, L.F.S.; Sales, J.B.S.; Vallinoto, M.; Schneider, H.; Sampaio, I. Commercialization of a critically endangered species (largetooth sawfish, Pristis perotteti) in fish markets of northern Brazil: Authenticity by DNA analysis. Food Control 2013, 34, 249–252. [Google Scholar] [CrossRef]
- Teletchea, F. After 7 years and 1000 citations: Comparative assessment of the DNA barcoding and the DNA taxonomy proposals for taxonomists and non-taxonomists. Mitochondrial DNA 2010, 21, 206–226. [Google Scholar] [CrossRef] [PubMed]
- Wakefield, C.B.; Santana-Garcon, J.; Dorman, S.R.; Blight, S.; Denham, A.; Wakeford, J.; Molony, B.W.; Newman, S.J. Performance of bycatch reduction devices varies for chondrichthyan, reptile, and cetacean mitigation in demersal fish trawls: Assimilating subsurface interactions and unaccounted mortality. ICES J. Mar. Sci. 2017, 74, 343–358. [Google Scholar] [CrossRef]
- Howard, S.; Brill, R.; Hepburn, C.; Rock, J.; Pol, M. Microprocessor-based prototype bycatch reduction device reduces bait consumption by spiny dogfish and sandbar shark. ICES J. Mar. Sci. 2018, 75, 2235–2244. [Google Scholar] [CrossRef]
- Richards, R.J.; Raoult, V.; Powter, D.M.; Gaston, T.F. Permanent magnets reduce bycatch of benthic sharks in an ocean trap fishery. Fish. Res. 2018, 208, 16–21. [Google Scholar] [CrossRef]
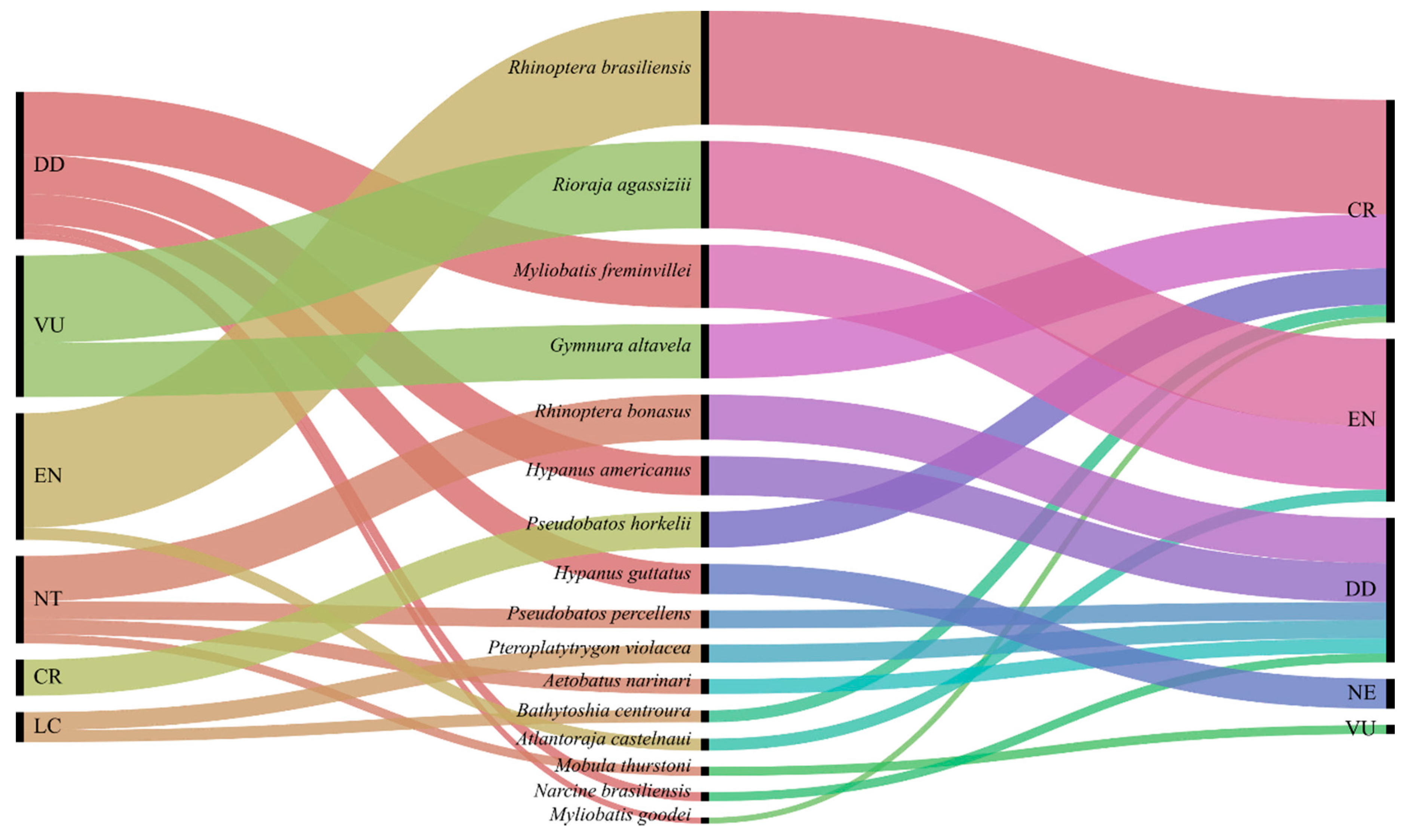
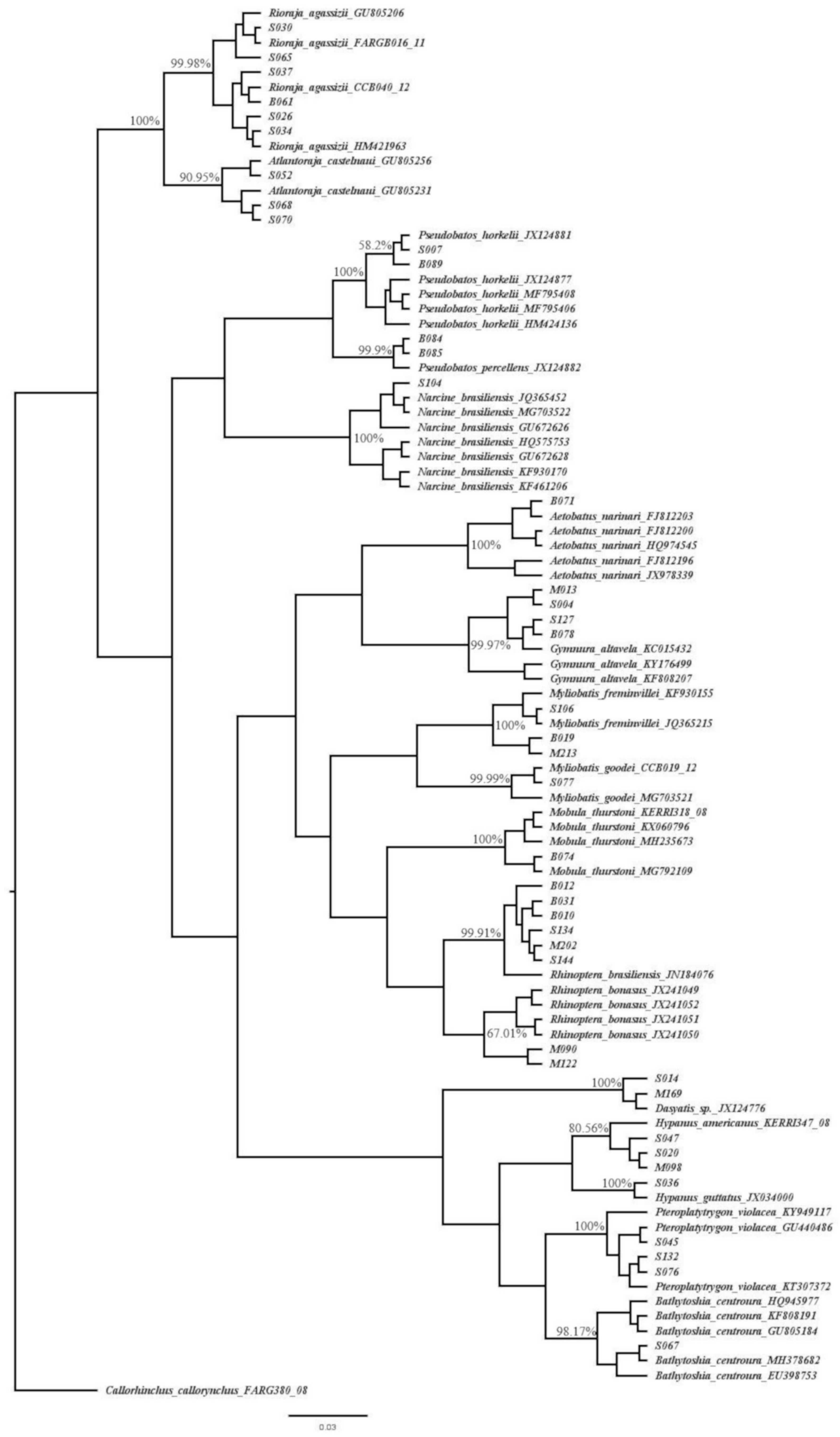
| Order | Family | Common Name | Species | n | % | IUCN | ICMBio |
|---|---|---|---|---|---|---|---|
| Myliobatiformes | Dasyatidae | Roughtail stingray | Bathytoshia centroura | 4 | 1.75% | LC | CR |
| Southern stingray | Hypanus americanus | 13 | 5.70% | DD | DD | ||
| Longnose stingray | Hypanus guttatus | 10 | 4.39% | DD | NE | ||
| Blue pelagic stingray | Pteroplatytrygon violacea | 6 | 2.63% | LC | DD | ||
| Dasyatis sp. | 39 | 17.11% | - | - | |||
| Myliobatidae | Spotted eagle ray | Aetobatus narinari | 5 | 2.19% | NT | DD | |
| Smoothtail mobula | Mobula thurstoni | 3 | 1.32% | NT | VU | ||
| Bullnose ray | Myliobatis freminvillei | 21 | 9.21% | DD | EN | ||
| Southern eagle ray | Myliobatis goodei | 2 | 0.88% | DD | CR | ||
| Rhinopteridae | Cownose ray | Rhinoptera bonasus | 15 | 6.58% | NT | DD | |
| Brazilian cownose ray | Rhinoptera brasiliensis | 38 | 16.67% | EN | CR | ||
| Gymnuridae | Spiny butterfly ray | Gymnura altavela | 18 | 7.89% | VU | CR | |
| Rajiformes | Arhynchobatidae | Spotback skate | Atlantoraja castelnaui | 4 | 1.75% | EN | EN |
| Rio skate | Rioraja agassizii | 29 | 12.72% | VU | EN | ||
| Rhinopristiformes | Rhinobatidae | Brazilian guitarfish | Pseudobatos horkelii | 12 | 5.26% | CR | CR |
| Southern guitarfish | Pseudobatos percellens | 6 | 2.63% | NT | DD | ||
| Torpediniformes | Narcinidae | Brazilian electric ray | Narcine brasiliensis | 3 | 1.32% | DD | DD |
| 228 | 100.00% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ferrette, B.L.d.S.; Domingues, R.R.; Rotundo, M.M.; Miranda, M.P.; Bunholi, I.V.; De Biasi, J.B.; Oliveira, C.; Foresti, F.; Mendonça, F.F. DNA Barcode Reveals the Bycatch of Endangered Batoids Species in the Southwest Atlantic: Implications for Sustainable Fisheries Management and Conservation Efforts. Genes 2019, 10, 304. https://doi.org/10.3390/genes10040304
Ferrette BLdS, Domingues RR, Rotundo MM, Miranda MP, Bunholi IV, De Biasi JB, Oliveira C, Foresti F, Mendonça FF. DNA Barcode Reveals the Bycatch of Endangered Batoids Species in the Southwest Atlantic: Implications for Sustainable Fisheries Management and Conservation Efforts. Genes. 2019; 10(4):304. https://doi.org/10.3390/genes10040304
Chicago/Turabian StyleFerrette, Bruno Lopes da Silva, Rodrigo Rodrigues Domingues, Matheus Marcos Rotundo, Marina Provetti Miranda, Ingrid Vasconcellos Bunholi, Juliana Beltramin De Biasi, Claudio Oliveira, Fausto Foresti, and Fernando Fernandes Mendonça. 2019. "DNA Barcode Reveals the Bycatch of Endangered Batoids Species in the Southwest Atlantic: Implications for Sustainable Fisheries Management and Conservation Efforts" Genes 10, no. 4: 304. https://doi.org/10.3390/genes10040304
APA StyleFerrette, B. L. d. S., Domingues, R. R., Rotundo, M. M., Miranda, M. P., Bunholi, I. V., De Biasi, J. B., Oliveira, C., Foresti, F., & Mendonça, F. F. (2019). DNA Barcode Reveals the Bycatch of Endangered Batoids Species in the Southwest Atlantic: Implications for Sustainable Fisheries Management and Conservation Efforts. Genes, 10(4), 304. https://doi.org/10.3390/genes10040304






