DNA Methylation Patterns in the Round Goby Hypothalamus Support an On-The-Spot Decision Scenario for Territorial Behavior
Abstract
:1. Introduction
2. Materials and Methods
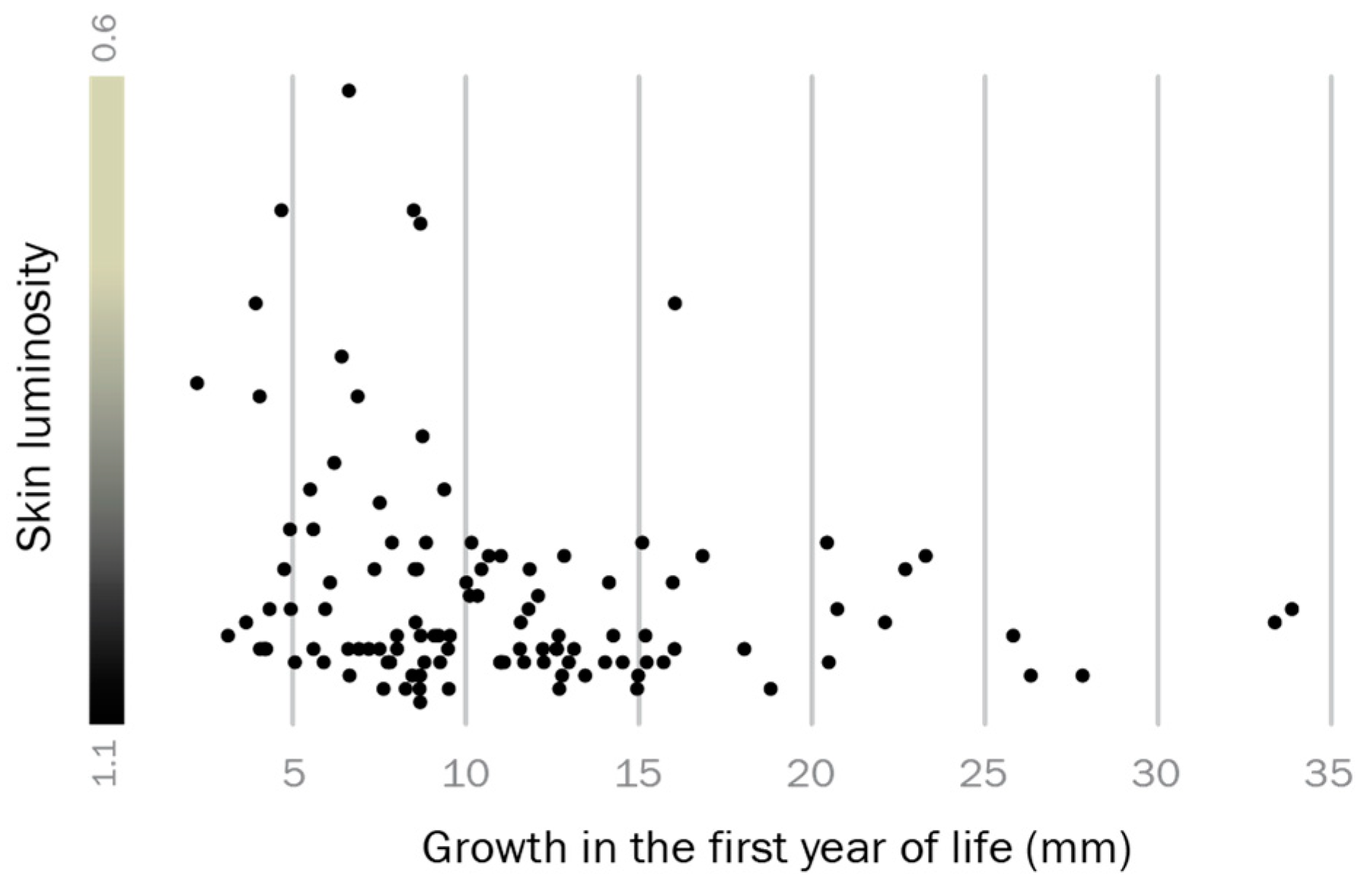
2.1. Growth in First Year and Luminosity
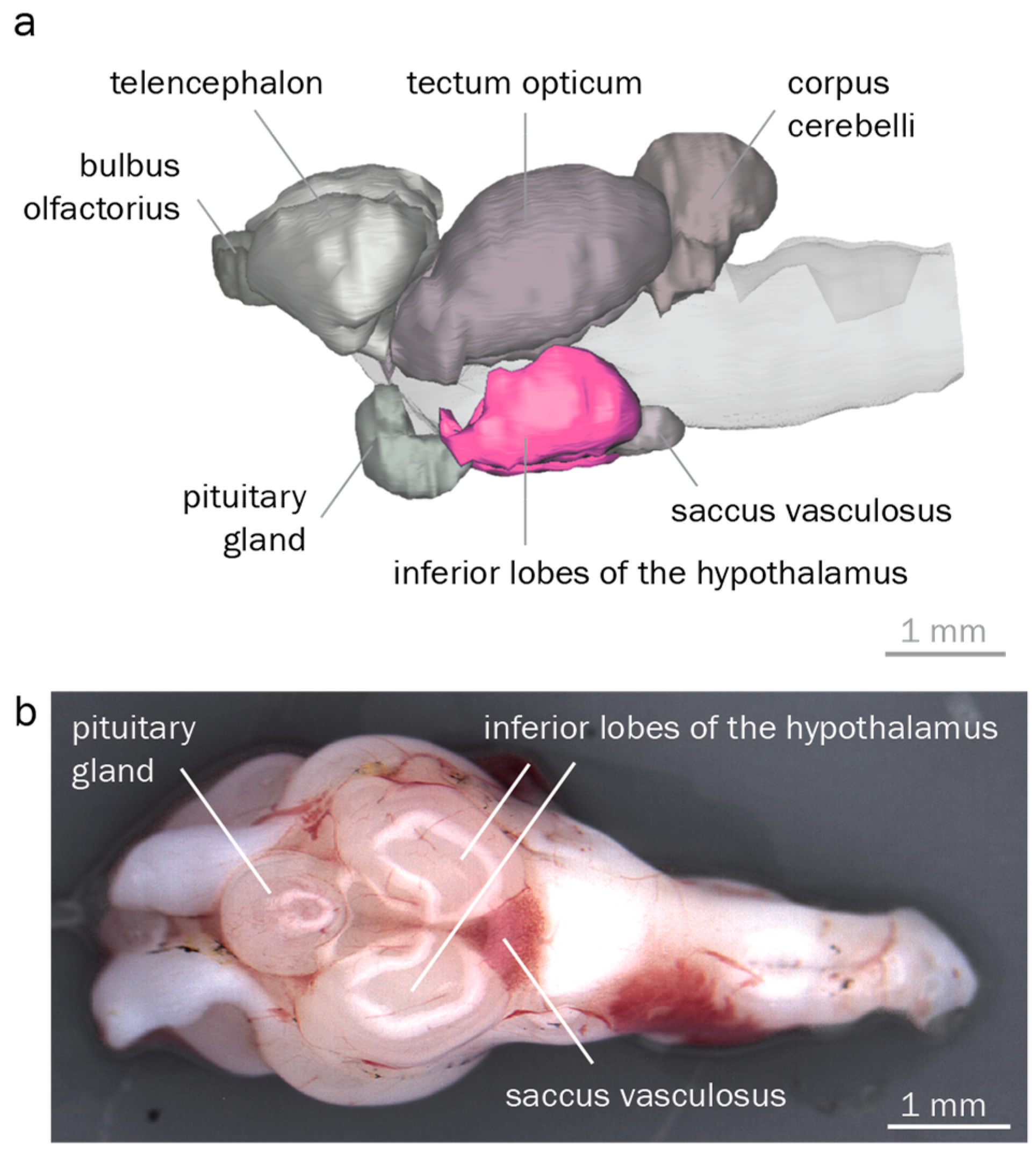
2.2. Brain Morphology
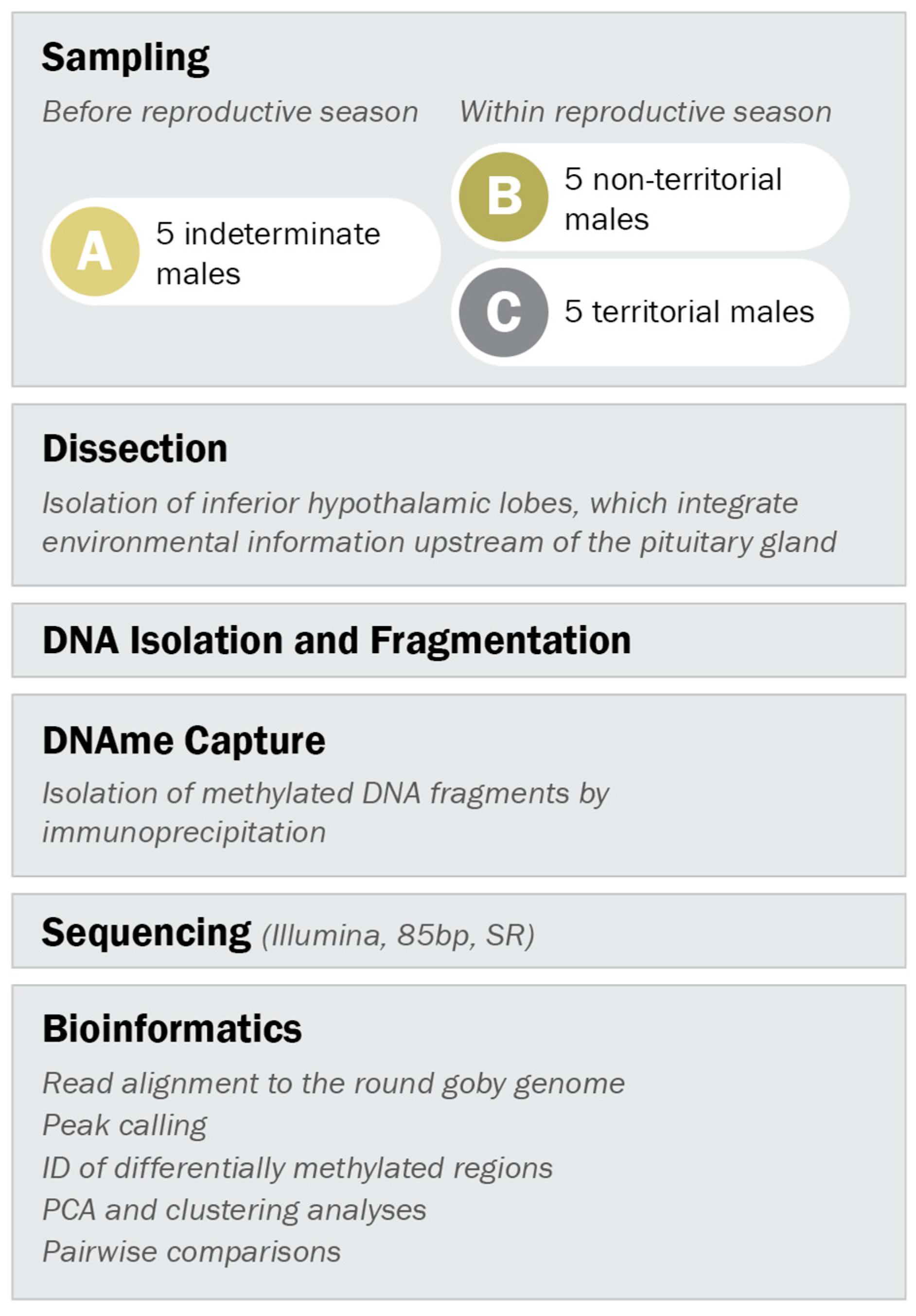
2.3. Sampling of Males for Methylated DNA Analysis
2.4. Brain Dissection
2.5. DNA Isolation
2.6. Enrichment of Methylated DNA
2.7. Library Preparation and Sequencing
2.8. Read Cleaning and Alignment
2.9. Peak Calling
2.10. Princpal Component Analysis and Dendrograms
2.11. Pairwise Comparisons
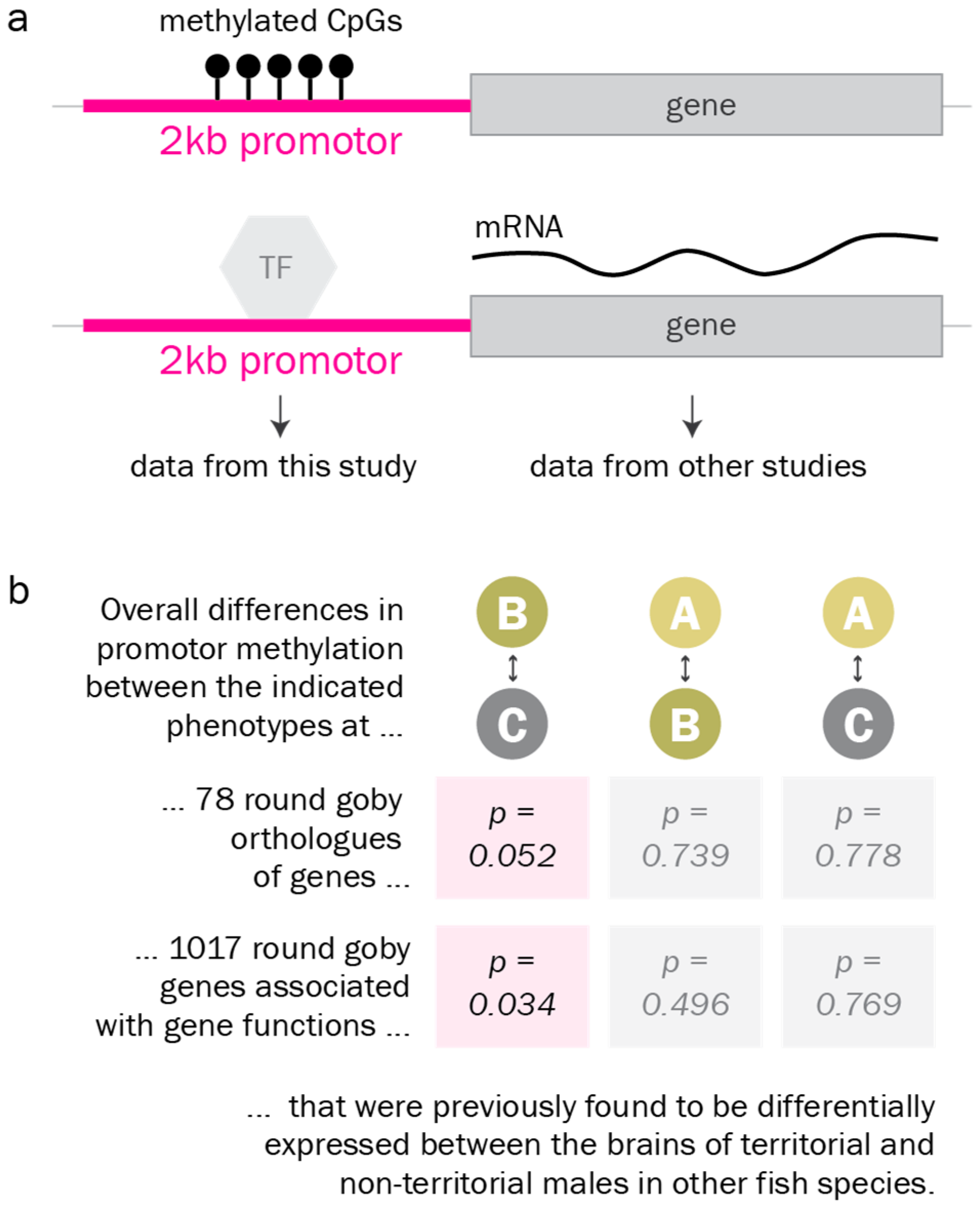
2.12. Analysis of DNA Methylation at Candidate Gene Promotors
3. Results
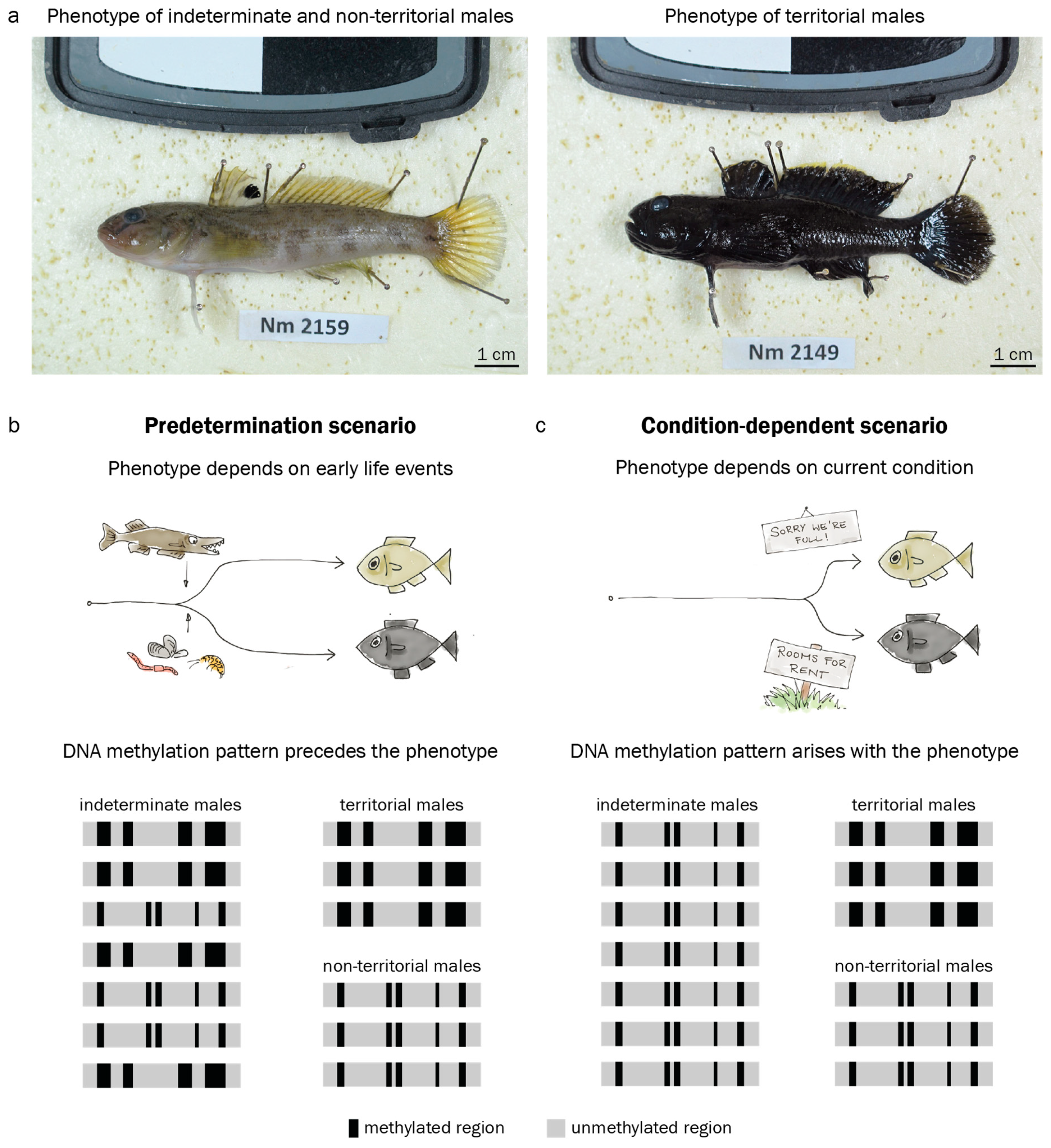
3.1. Adult Male Phenotype Is Related to Early Growth Rate
3.2. The Round Goby Has a Typical Teleost Brain
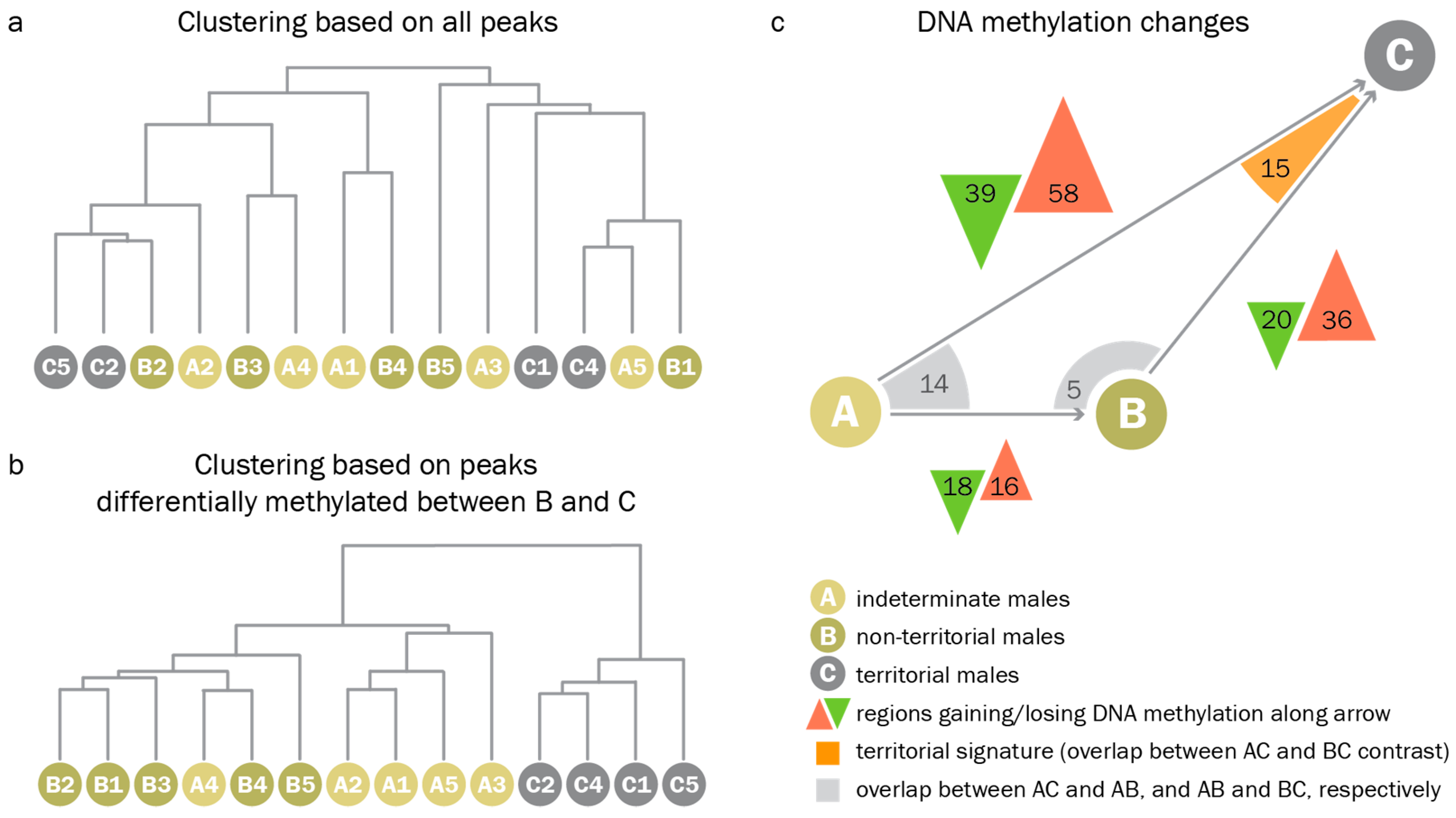
3.3. Territorial DNA Methylation Patterns Arise Concomitantly with the Phenotype
4. Discussion
4.1. The Relationship between Early Growth and Adult Reproductive Phenotype
4.2. Anatomical Identification of Brain Regions Controlling Reproduction
4.3. Assessment of DNA Methylation Across Male Reproductive Phenotypes
4.4. Identification of Differentially Methylated Genes Involved in Reproductive Phenotypes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pechenik, J.A. Larval experience and latent effects—Metamorphosis is not a new beginning. Integr. Comp. Biol. 2006, 46, 323–333. [Google Scholar] [CrossRef] [PubMed]
- Kerr, L.A.; Secor, D.H. Latent effects of early life history on partial migration for an estuarine-dependent fish. Environ. Biol. Fishes 2010, 89, 479–492. [Google Scholar] [CrossRef]
- Shima, J.S.; Swearer, S.E. The legacy of dispersal: Larval experience shapes persistence later in the life of a reef fish. J. Anim. Ecol. 2010, 79, 1308–1314. [Google Scholar] [CrossRef] [PubMed]
- Mirza, R.S.; Chivers, D.P. Predator-recognition training enhances survival of brook trout: Evidence from laboratory and field-enclosure studies. Can. J. Zool. 2000, 78, 2198–2208. [Google Scholar] [CrossRef]
- Mitchell, M.D.; McCormick, M.I.; Ferrari, M.C.O.; Chivers, D.P. Friend or foe? The role of latent inhibition in predator and non-predator labelling by coral reef fishes. Anim. Cogn. 2011, 14, 707–714. [Google Scholar] [CrossRef] [PubMed]
- Cunliffe, V.T. The epigenetic impacts of social stress: How does social adversity become biologically embedded? Epigenomics 2016, 8, 1653–1669. [Google Scholar] [CrossRef] [PubMed]
- Hall, S.E.; Beverly, M.; Russ, C.; Nusbaum, C.; Sengupta, P. A cellular memory of developmental history generates phenotypic diversity in C. elegans. Curr. Biol. 2010, 20, 149–155. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, S.D.; Teles, M.C.; Oliveira, R.F. Neurogenomic mechanisms of social plasticity. J. Exp. Biol. 2015, 218, 140–149. [Google Scholar] [CrossRef] [PubMed]
- Jablonka, E.; Raz, G. Transgenerational epigenetic inheritance: Prevalence, mechanisms, and implications for the study of heredity and evolution. Q. Rev. Biol. 2009, 84, 131–176. [Google Scholar] [CrossRef] [PubMed]
- Szyf, M. Epigenetics, DNA methylation and chromatin modifying drugs. Annu. Rev. Pharmacol. 2009, 49, 243–263. [Google Scholar] [CrossRef] [PubMed]
- Bird, A. DNA methylation patterns and epigenetic memory. Gene Dev. 2002, 16, 6–21. [Google Scholar] [CrossRef] [PubMed]
- Grimm, S.A.; Shimbo, T.; Takaku, M.; Thomas, J.W.; Auerbach, S.; Bennett, B.D.; Bucher, J.R.; Burkholder, A.B.; Day, F.; Du, Y.; et al. DNA methylation in mice is influenced by genetics as well as sex and life experience. Nat. Commun. 2019, 10, 305. [Google Scholar] [CrossRef] [PubMed]
- Weyrich, A.; Benz, S.; Karl, S.; Jeschek, M.; Jewgenow, K.; Fickel, J. Paternal heat exposure causes DNA methylation and gene expression changes of Stat3 in Wild guinea pig sons. Ecol. Evol. 2016, 6, 2657–2666. [Google Scholar] [CrossRef] [PubMed]
- Richards, C.L.; Alonso, C.; Becker, C.; Bossdorf, O.; Bucher, E.; Colome-Tatche, M.; Durka, W.; Engelhardt, J.; Gaspar, B.; Gogol-Doring, A.; et al. Ecological plant epigenetics: Evidence from model and non-model species, and the way forward. Ecol. Lett. 2017, 20, 1576–1590. [Google Scholar] [CrossRef] [PubMed]
- Kucharski, R.; Maleszka, J.; Foret, S.; Maleszka, R. Nutritional control of reproductive status in honeybees via DNA methylation. Science 2008, 319, 1827–1830. [Google Scholar] [CrossRef] [PubMed]
- Smith, Z.D.; Meissner, A. DNA methylation: Roles in mammalian development. Nat. Rev. Genet. 2013, 14, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Potok, M.E.; Nix, D.A.; Parnell, T.J.; Cairns, B.R. Reprogramming the maternal zebrafish genome after fertilization to match the paternal methylation pattern. Cell 2013, 153, 759–772. [Google Scholar] [CrossRef] [PubMed]
- Law, J.A.; Jacobsen, S.E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 2010, 11, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Leenen, F.A.D.; Muller, C.P.; Turner, J.D. DNA methylation: Conducting the orchestra from exposure to phenotype? Clin. Epigenet. 2016, 8, 92. [Google Scholar] [CrossRef] [PubMed]
- Elliott, E.; Ezra-Nevo, G.; Regev, L.; Neufeld-Cohen, A.; Chen, A. Resilience to social stress coincides with functional DNA methylation of the Crf gene in adult mice. Nat. Neurosci. 2010, 13, 1351–1353. [Google Scholar] [CrossRef] [PubMed]
- Knecht, A.L.; Truong, L.; Marvel, S.W.; Reif, D.M.; Garcia, A.; Lu, C.; Simbnich, M.T.; Teeguarden, J.G.; Tanguay, R.L. Transgenerational inheritance of neurobehavioral and physiological deficits from developmental exposure to benzo[a]pyrene in zebrafish. Toxicol. Appl. Pharm. 2017, 329, 148–157. [Google Scholar] [CrossRef] [PubMed]
- Moran, P.; Perez-Figueroa, A. Methylation changes associated with early maturation stages in the Atlantic salmon. BMC Genet. 2011, 12, 86. [Google Scholar] [CrossRef] [PubMed]
- Breder, C.M.; Rosen, D.E. Modes of Reproduction in Fishes; Natural History Press: Garden City, NY, USA, 1966; 941p, p. xv. [Google Scholar]
- Wootton, R.J.; Smith, C. Reproductive Biology of Teleost Fishes; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2015. [Google Scholar]
- Smith, C.; Wootton, R.J. The remarkable reproductive diversity of teleost fishes. Fish Fish. 2016, 17, 1208–1215. [Google Scholar] [CrossRef]
- Cole, K.S. Reproduction and Sexuality in Marine Fishes: Patterns and Processes; University of California Press: Berkeley, CA, USA, 2010; 432p. [Google Scholar]
- Oliveira, R.F.; Ros, A.F.; Goncalves, D.M. Intra-sexual variation in male reproduction in teleost fish: A comparative approach. Horm. Behav. 2005, 48, 430–439. [Google Scholar] [CrossRef] [PubMed]
- Taborsky, M. Sneakers, satellites, and helpers: Parasitic and cooperative behavior in fish reproduction. Adv. Study Behav. 1994, 23, 1–100. [Google Scholar]
- Taborsky, M. Alternative reproductive tactics in fish. In Alternative Reproductive Tactics: An Integrative Approach; Oliveira, R.F., Taborsky, M., Brockmann, H.J., Eds.; Cambridge University Press: Cambridge, UK, 2008. [Google Scholar]
- Taborsky, M. The evolution of bourgeois, parasitic, and cooperative reproductive behaviors in fishes. J. Hered. 2001, 92, 100–110. [Google Scholar] [CrossRef] [PubMed]
- Gross, M.R. Alternative reproductive strategies and tactics: Diversity within sexes. Trends Ecol. Evol. 1996, 11, 92–98. [Google Scholar] [CrossRef]
- Aubin-Horth, N.; Landry, C.R.; Letcher, B.H.; Hofmann, H.A. Alternative life histories shape brain gene expression profiles in males of the same population. Proc. Biol. Sci. 2005, 272, 1655–1662. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, S.D.; Gonçalves, D.; Goesmann, A.; Canário, A.V.M.; Oliveira, R.F. Temporal variation in brain transcriptome is associated with the expression of female mimicry as a sequential male alternative reproductive tactic in fish. Mol. Ecol. 2018, 27, 789–803. [Google Scholar] [CrossRef] [PubMed]
- Maruska, K.P.; Zhang, A.; Neboori, A.; Fernald, R.D. Social Opportunity causes rapid transcriptional changes in the social behaviour network of the brain in an african cichlid fish. J. Neuroendocrinol. 2013, 25, 145–157. [Google Scholar] [CrossRef] [PubMed]
- Renn, S.C.P.; Aubin-Horth, N.; Hofmann, H.A. Fish and chips: Functional genomics of social plasticity in an African cichlid fish. J. Exp. Biol. 2008, 211, 3041–3056. [Google Scholar] [CrossRef] [PubMed]
- Nugent, B.M.; Stiver, K.A.; Alonzo, S.H.; Hofmann, H.A. Neuroendocrine profiles associated with discrete behavioural variation in Symphodus ocellatus, a species with male alternative reproductive tactics. Mol. Ecol. 2016, 25, 5212–5227. [Google Scholar] [CrossRef] [PubMed]
- Stiver, K.A.; Harris, R.M.; Townsend, J.P.; Hofmann, H.A.; Alonzo, S.H. Neural gene expression profiles and androgen levels underlie alternative reproductive tactics in the ocellated wrasse, Symphodus ocellatus. Ethology 2015, 121, 152–167. [Google Scholar] [CrossRef]
- Partridge, C.G.; MacManes, M.D.; Knapp, R.; Neff, B.D. Brain transcriptional profiles of male alternative reproductive tactics and females in Bluegill Sunfish. PLoS ONE 2016, 11, e0167509. [Google Scholar] [CrossRef] [PubMed]
- Schunter, C.; Vollmer, S.V.; Macpherson, E.; Pascual, M. Transcriptome analyses and differential gene expression in a non-model fish species with alternative mating tactics. BMC Genomics 2014, 15, 167. [Google Scholar] [CrossRef] [PubMed]
- Todd, E.V.; Liu, H.; Lamm, M.S.; Thomas, J.T.; Rutherford, K.; Thompson, K.C.; Godwin, J.R.; Gemmell, N.J. Female mimicry by sneaker males has a transcriptomic signature in both the brain and the gonad in a sex-changing fish. Mol. Biol. Evol. 2018, 35, 225–241. [Google Scholar] [CrossRef] [PubMed]
- Bleeker, K.; de Jong, K.; van Kessel, N.; Hinde, C.A.; Nagelkerke, L.A.J. Evidence for ontogenetically and morphologically distinct alternative reproductive tactics in the invasive Round Goby Neogobius melanostomus. PLoS ONE 2017, 12. [Google Scholar] [CrossRef] [PubMed]
- Marentette, J.R.; Fitzpatrick, J.L.; Berger, R.G.; Balshine, S. Multiple male reproductive morphs in the invasive round goby (Apollonia melanostoma). J. Great Lakes Res. 2009, 35, 302–308. [Google Scholar] [CrossRef]
- Mazzoldi, C.; Rasotto, M.B. Alternative male mating tactics in Gobius niger. J. Fish Biol. 2002, 61, 157–172. [Google Scholar] [CrossRef]
- Scaggiante, M.; Grober, M.S.; Lorenzi, V.; Rasotto, M.B. Changes along the male reproductive axis in response to social context in a gonochoristic gobiid, Zosterisessor ophiocephalus (Teleostei, Gobiidae), with alternative mating tactics. Horm. Behav. 2004, 46, 607–617. [Google Scholar] [CrossRef] [PubMed]
- Takegaki, T.; Svensson, O.; Kvarnemo, C. Socially induced tactic change in 2 types of sand goby sneaker males. Behav. Ecol. 2012, 23, 742–750. [Google Scholar] [CrossRef]
- Adrian-Kalchhauser, I. Round Goby Genome Assembly V2, 2017; Unpublished; University of Basel: Basel, Switzerland.
- Shahjahan, M.; Kitahashi, T.; Parhar, I.S. Central pathways integrating metabolism and reproduction in teleosts. Front. Endocrinol. 2014, 5, 36. [Google Scholar] [CrossRef] [PubMed]
- Zohar, Y.; Munoz-Cueto, J.A.; Elizur, A.; Kah, O. Neuroendocrinology of reproduction in teleost fish. Gen. Comp. Endocrinol. 2010, 165, 438–455. [Google Scholar] [CrossRef] [PubMed]
- Gore, A.C. GnRH: The Master Molecule of Reproduction; Springer: New York, NY, USA, 2002. [Google Scholar]
- Greenwood, A.K.; Wark, A.R.; Fernald, R.D.; Hofmann, H.A. Expression of arginine vasotocin in distinct preoptic regions is associated with dominant and subordinate behaviour in an African cichlid fish. Proc. Biol. Sci. 2008, 275, 2393–2402. [Google Scholar] [CrossRef] [PubMed]
- Goncalves, D.; Teles, M.; Alpedrinha, J.; Oliveira, R.F. Brain and gonadal aromatase activity and steroid hormone levels in female and polymorphic males of the peacock blenny Salaria pavo. Horm. Behav. 2008, 54, 717–725. [Google Scholar] [CrossRef] [PubMed]
- Abbott, F.S. Endocrine regulation of pigmentation in fish. Am. Zool. 1973, 73, 885–894. [Google Scholar] [CrossRef]
- Leclercq, E.; Taylor, J.F.; Migaud, H. Morphological skin colour changes in teleosts. Fish Fish. 2010, 11, 159–193. [Google Scholar] [CrossRef]
- Bagenal, T.B.; Tesch, F.W. Methods for Assessment of Fish Production in Fresh Waters; Blackwell: Oxford, UK, 1978; pp. 101–136. [Google Scholar]
- Hirsch, P.E.; Adrian-Kalchhauser, I.; Flamig, S.; N’Guyen, A.; Defila, R.; Di Giulio, A.; Burkhardt-Holm, P. A tough egg to crack: Recreational boats as vectors for invasive goby eggs and transdisciplinary management approaches. Ecol. Evol. 2016, 6, 707–715. [Google Scholar] [CrossRef] [PubMed]
- Grul’a, D.; Balazova, M.; Copp, G.H.; Kovac, V. Age and growth of invasive round goby Neogobius melanostomus from middle Danube. Cent. Eur. J. Biol. 2012, 7, 448–459. [Google Scholar]
- Weyrich, A.; Schullermann, T.; Heeger, F.; Jeschek, M.; Mazzoni, C.J.; Chen, W.; Schumann, K.; Fickel, J. Whole genome sequencing and methylome analysis of the wild guinea pig. BMC Genomics 2014, 15, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 2011, 17. [Google Scholar] [CrossRef]
- Schmieder, R.; Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 2011, 27, 863–864. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, U357–U359. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. 1000 genome project data processing subgroup. The sequence alignment/map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- Quinlan, A.R.; Hall, I.M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef] [PubMed]
- Weisenberger, D.J.; Campan, M.; Long, T.I.; Kim, M.; Woods, C.; Fiala, E.; Ehrlich, M.; Laird, P.W. Analysis of repetitive element DNA methylation by MethyLight. Nucleic Acids Res. 2005, 33, 6823–6836. [Google Scholar] [CrossRef] [PubMed]
- Team, R.C. R: A language and environment for statistical computing. In R Foundation for Statistical Computing; Vienna, Austria, 2016. [Google Scholar]
- Wong, R.Y.; McLeod, M.M.; Godwin, J. Limited sex-biased neural gene expression patterns across strains in Zebrafish (Danio rerio). BMC Genomics 2014, 15, 905. [Google Scholar] [CrossRef] [PubMed]
- Martinez, P.; Vinas, A.M.; Sanchez, L.; Diaz, N.; Ribas, L.; Piferrer, F. Genetic architecture of sex determination in fish: Applications to sex ratio control in aquaculture. Front. Genet. 2014, 5, 340. [Google Scholar] [CrossRef] [PubMed]
- Gotz, S.; Garcia-Gomez, J.M.; Terol, J.; Williams, T.D.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talon, M.; Dopazo, J.; Conesa, A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar] [CrossRef] [PubMed]
- Sinnett, D.; Beaulieu, P.; Belanger, H.; Lefebvre, J.F.; Langlois, S.; Theberge, M.C.; Drouin, S.; Zotti, C.; Hudson, T.J.; Labuda, D. Detection and characterization of DNA variants in the promoter regions of hundreds of human disease candidate genes. Genomics 2006, 87, 704–710. [Google Scholar] [CrossRef] [PubMed]
- Krause, E.T.; Honarmand, M.; Wetzel, J.; Naguib, M. Early fasting is long lasting: Differences in early nutritional conditions reappear under stressful conditions in adult female zebra finches. PLoS ONE 2009, 4, e5015. [Google Scholar] [CrossRef] [PubMed]
- Dmitriew, C.M. The evolution of growth trajectories: What limits growth rate? Biol. Rev. 2011, 86, 97–116. [Google Scholar] [CrossRef] [PubMed]
- Helfman, S.; Collette, B.B.; Facey, D.E.; Bowen, B.W. Early life history. In The Diversity of Fishes: Biology, Evolution, and Ecology; John Wiley & Sons: Hoboken, NJ, USA, 2009; p. 129. [Google Scholar]
- Burgerhout, E.; Mommens, M.; Johnsen, H.; Aunsmo, A.; Santi, N.; Andersen, O. Genetic background and embryonic temperature affect DNA methylation and expression of myogenin and muscle development in Atlantic salmon (Salmo salar). PLoS ONE 2017, 12, e0179918. [Google Scholar] [CrossRef] [PubMed]
- Hirsch, P.E.; Eckmann, R.; Oppelt, C.; Behrmann-Godel, J. Phenotypic and genetic divergence within a single whitefish form—Detecting the potential for future divergence. Evol. Appl. 2013, 6, 1119–1132. [Google Scholar] [CrossRef] [PubMed]
- Magnhagen, C. Alternative reproductive-behavior in the common goby, pomatoschistus-microps—An ontogenic gradient. Anim. Behav. 1992, 44, 182–184. [Google Scholar] [CrossRef]
- Maegawa, S.; Hinkal, G.; Kim, H.S.; Shen, L.; Zhang, L.; Zhang, J.; Zhang, N.; Liang, S.; Donehower, L.A.; Issa, J.P. Widespread and tissue specific age-related DNA methylation changes in mice. Genome Res. 2010, 20, 332–340. [Google Scholar] [CrossRef] [PubMed]
- Bauchot, R.; Diagne, M.; Ridet, J.M.; Bauchot, M.L. The Brain of Rhyacichthys-Aspro (Rhyacichthyidae, Gobioidei). Jpn. J. Ichthyol. 1989, 36, 260–266. [Google Scholar] [CrossRef]
- Kassem, M.; Ridet, J.M.; Bauchot, R. Analyse volumetrique des principales subdivisions encephaliques chez les Gobioidei (Teleosteens, Perciformes). J. Hirnforsch. 1989, 30, 59–67. [Google Scholar] [PubMed]
- Vasek, M.; Vsetickova, L.; Roche, K.; Jurajda, P. Diet of two invading gobiid species (Proterorhinus semilunaris and Neogobius melanostomus) during the breeding and hatching season: No field evidence of extensive predation on fish eggs and fry. Limnologica 2014, 46, 31–36. [Google Scholar] [CrossRef]
- Wullimann, M.F.; Rupp, B.; Reichert, H. Neuroanatomy of the Zebrafish Brain: A Topological Atlas; Springer: Basel, Switzerland, 1996. [Google Scholar]
- Meek, J.; Nieuwenhuys, R. Holosteans and teleosts. In The Central Nervous System of Vertebrates; Springer: Berlin/Heidelberg, Germany, 1998. [Google Scholar]
- Zeyl, J.N.; Love, O.P.; Higgs, D.M. Condition-dependent auditory processing in the round goby (Neogobius melanostomus): Links to sex, reproductive condition and female estrogen levels. J. Exp. Biol. 2013, 216, 1075–1084. [Google Scholar] [CrossRef] [PubMed]
- Rollo, A.; Higgs, D. Differential acoustic response specificity and directionality in the round goby, Neogobius melanostomus. Anim. Behav. 2008, 75, 1903–1912. [Google Scholar] [CrossRef]
- Laframboise, A.J.; Katare, Y.; Scott, A.P.; Zielinski, B.S. The effect of elevated steroids released by reproductive male round gobies, Neogobius melanostomus, on olfactory responses in females. J. Chem. Ecol. 2011, 37, 260–262. [Google Scholar] [CrossRef] [PubMed]
- Marentette, J.R.; Wang, G.; Tong, S.; Sopinka, N.M.; Taves, M.D.; Koops, M.A.; Balshine, S. Laboratory and field evidence of sex-biased movement in the invasive round goby. Behav. Ecol. Sociobiol. 2011, 65, 2239–2249. [Google Scholar] [CrossRef]
- Groen, M.; Sopinka, N.M.; Marentette, J.R.; Reddon, A.R.; Brownscombe, J.W.; Fox, M.G.; Marsh-Rollo, S.E.; Balshine, S. Is there a role for aggression in round goby invasion fronts? Behaviour 2012, 149, 685–703. [Google Scholar]
- Carman, S.M.; Janssen, J.; Jude, D.J.; Berg, M.B. Diel interactions between prey behaviour and feeding in an invasive fish, the round goby, in a North American river. Freshw. Biol. 2006, 51, 742–755. [Google Scholar] [CrossRef]
- McGowan, P.O.; Sasaki, A.; D’Alessio, A.C.; Dymov, S.; Labonte, B.; Szyf, M.; Turecki, G.; Meaney, M.J. Epigenetic regulation of the glucocorticoid receptor in human brain associates with childhood abuse. Nat. Neurosci. 2009, 12, 342–348. [Google Scholar] [CrossRef] [PubMed]
- Fuchikami, M.; Morinobu, S.; Segawa, M.; Okamoto, Y.; Yamawaki, S.; Ozaki, N.; Inoue, T.; Kusumi, I.; Koyama, T.; Tsuchiyama, K.; et al. DNA methylation profiles of the brain-derived neurotrophic factor (BDNF) gene as a potent diagnostic biomarker in major depression. PLoS ONE 2011, 6, e23881. [Google Scholar] [CrossRef] [PubMed]
- Hahn, M.A.; Wu, X.; Li, A.X.; Hahn, T.; Pfeifer, G.P. Relationship between gene body DNA methylation and intragenic H3K9me3 and H3K36me3 chromatin marks. PLoS ONE 2011, 6, e18844. [Google Scholar] [CrossRef] [PubMed]
- Jjingo, D.; Conley, A.B.; Yi, S.V.; Lunyak, V.V.; Jordan, I.K. On the presence and role of human gene-body DNA methylation. Oncotarget 2012, 3, 462–474. [Google Scholar] [CrossRef] [PubMed]
- Mohn, F.; Weber, M.; Rebhan, M.; Roloff, T.C.; Richter, J.; Stadler, M.B.; Bibel, M.; Schubeler, D. Lineage-specific polycomb targets and de novo DNA methylation define restriction and potential of neuronal progenitors. Mol. Cell 2008, 30, 755–766. [Google Scholar] [CrossRef] [PubMed]
- Sheaffer, K.L.; Kim, R.; Aoki, R.; Elliott, E.N.; Schug, J.; Burger, L.; Schubeler, D.; Kaestner, K.H. DNA methylation is required for the control of stem cell differentiation in the small intestine. Gene Dev. 2014, 28, 652–664. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.C.; Wu, X.J.; Jiang, L.; Zhang, Y. Single-Cell RNA-Seq reveals hypothalamic cell diversity. Cell Rep. 2017, 18, 3227–3241. [Google Scholar] [CrossRef] [PubMed]
- Machluf, Y.; Gutnick, A.; Levkowitz, G. Development of the zebrafish hypothalamus. Trends Neuroendocrinol. 2011, 1220, 93–105. [Google Scholar] [CrossRef] [PubMed]
- Kuroyanagi, Y.; Okuyama, T.; Suehiro, Y.; Imada, H.; Shimada, A.; Naruse, K.; Takeda, H.; Kubo, T.; Takeuchi, H. Proliferation zones in adult medaka (Oryzias latipes) brain. Brain Res. 2010, 1323, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Puschina, E.V.; Obukhov, D.K. Processes of proliferation and apoptosis in the brain of the amur sturgeon. Neurophysiology 2011, 43, 271–286. [Google Scholar] [CrossRef]
- Zikopoulos, B.; Kentouri, M.; Dermon, C.R. Cell genesis in the hypothalamus is associated to the sexual phase of a hermaphrodite teleost. Neuroreport 2001, 12, 2477–2481. [Google Scholar] [CrossRef] [PubMed]
- Glasauer, S.M.K.; Neuhauss, S.C.F. Whole-genome duplication in teleost fishes and its evolutionary consequences. Mol. Genet. Genomics 2014, 289, 1045–1060. [Google Scholar] [CrossRef] [PubMed]
- Stone, A.; Zotenko, E.; Locke, W.J.; Korbie, D.; Millar, E.K.A.; Pidsley, R.; Stirzaker, C.; Graham, P.; Trau, M.; Musgrove, E.A.; et al. DNA methylation of oestrogen-regulated enhancers defines endocrine sensitivity in breast cancer. Nat. Commun. 2015, 6, 7758. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Zhao, Z.M. Comparative analysis of CpG islands in four fish genomes. Comp. Funct. Genomics 2008. [Google Scholar] [CrossRef] [PubMed]
- Jiang, N.; Wang, L.; Chen, J.; Wang, L.W.; Leach, L.; Luo, Z.W. Conserved and divergent patterns of DNA methylation in higher vertebrates. Genome Biol. Evol. 2014, 6, 2998–3014. [Google Scholar] [CrossRef] [PubMed]
- McGaughey, D.M.; Abaan, H.O.; Miller, R.M.; Kropp, P.A.; Brody, L.C. Genomics of CpG methylation in developing and developed zebrafish. G3 Genes Genome Genet. 2014, 4, 861–869. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Somerville, V.; Schwaiger, M.; Hirsch, P.E.; Walser, J.-C.; Bussmann, K.; Weyrich, A.; Burkhardt-Holm, P.; Adrian-Kalchhauser, I. DNA Methylation Patterns in the Round Goby Hypothalamus Support an On-The-Spot Decision Scenario for Territorial Behavior. Genes 2019, 10, 219. https://doi.org/10.3390/genes10030219
Somerville V, Schwaiger M, Hirsch PE, Walser J-C, Bussmann K, Weyrich A, Burkhardt-Holm P, Adrian-Kalchhauser I. DNA Methylation Patterns in the Round Goby Hypothalamus Support an On-The-Spot Decision Scenario for Territorial Behavior. Genes. 2019; 10(3):219. https://doi.org/10.3390/genes10030219
Chicago/Turabian StyleSomerville, Vincent, Michaela Schwaiger, Philipp E. Hirsch, Jean-Claude Walser, Karen Bussmann, Alexandra Weyrich, Patricia Burkhardt-Holm, and Irene Adrian-Kalchhauser. 2019. "DNA Methylation Patterns in the Round Goby Hypothalamus Support an On-The-Spot Decision Scenario for Territorial Behavior" Genes 10, no. 3: 219. https://doi.org/10.3390/genes10030219
APA StyleSomerville, V., Schwaiger, M., Hirsch, P. E., Walser, J.-C., Bussmann, K., Weyrich, A., Burkhardt-Holm, P., & Adrian-Kalchhauser, I. (2019). DNA Methylation Patterns in the Round Goby Hypothalamus Support an On-The-Spot Decision Scenario for Territorial Behavior. Genes, 10(3), 219. https://doi.org/10.3390/genes10030219




