Population Genetic and Functional Analysis of a cis-Regulatory Polymorphism in the Drosophila melanogaster Metallothionein A gene
Abstract
1. Introduction
2. Materials and Methods
2.1. Fly Collection and Maintenance
2.2. Detection of the MtnA Deletion in Wild-Caught Flies
2.3. Quantitative Reverse Transcription PCR
2.4. Detection of the MtnA Deletion using Genome Sequence Data
2.5 Detection of Non-African Admixture
2.6. Analysis of MtnA expression and oxidative stress tolerance in DGRP lines
3. Results
3.1. The Deletion is Present at a Stable, High Frequency in a European Population
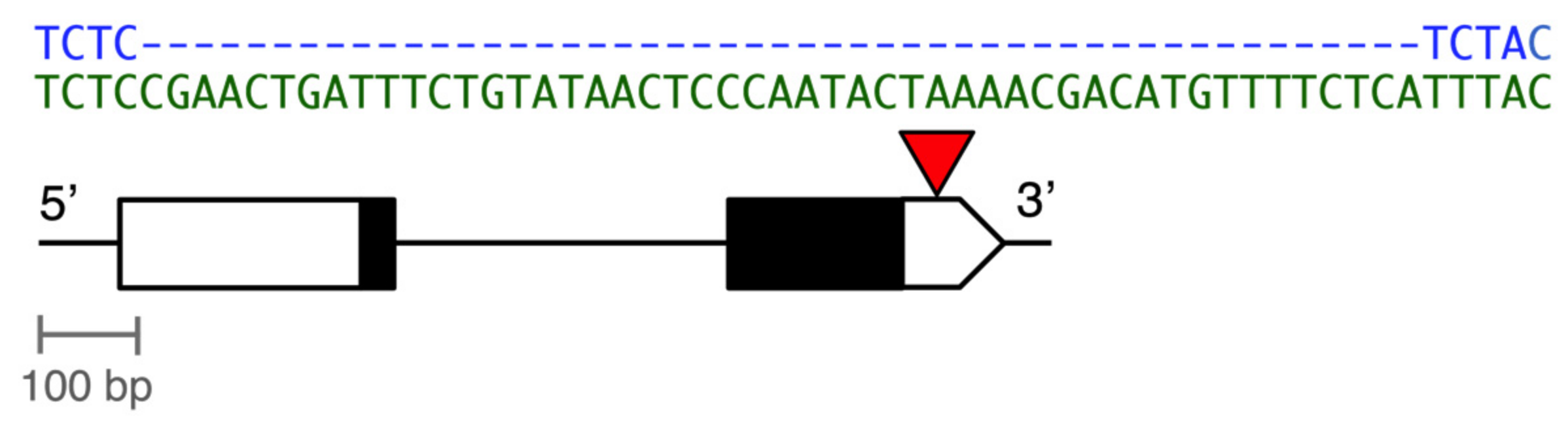
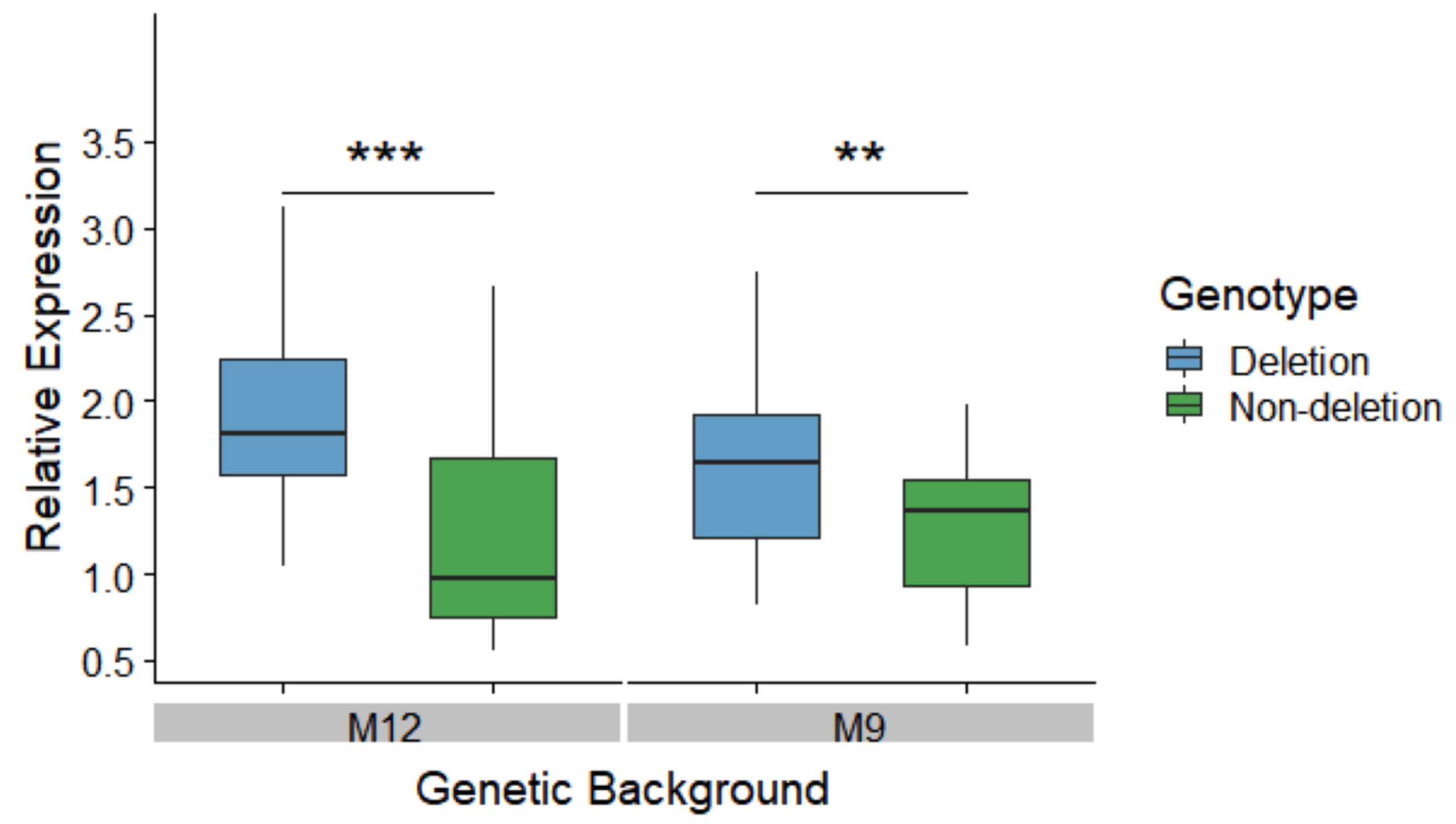
3.2. The Deletion Leads to Higher MtnA Expression in a Common, European Background
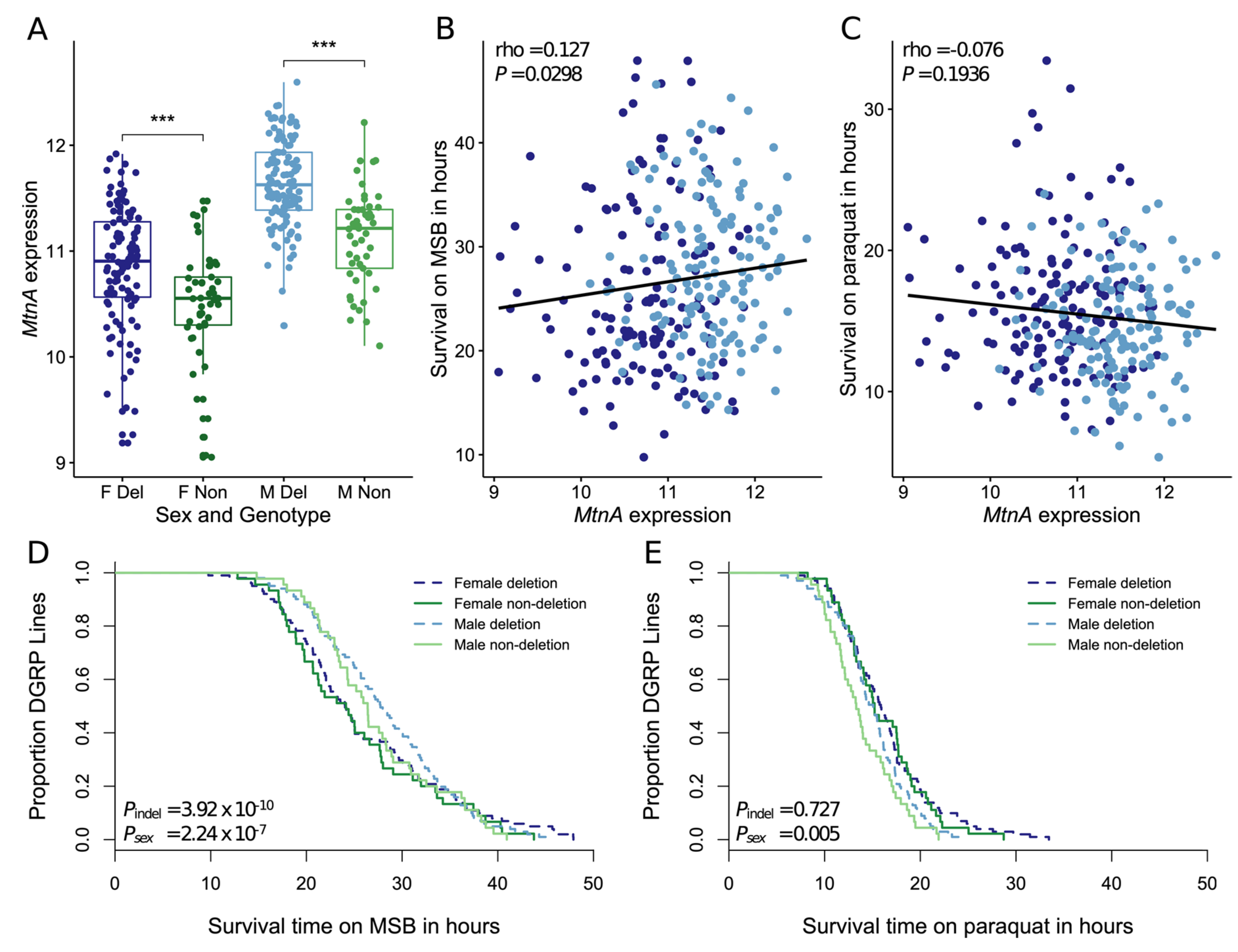
3.3. Association of the Deletion with MtnA Expression and Oxidative Stress Tolerance in a North American Population
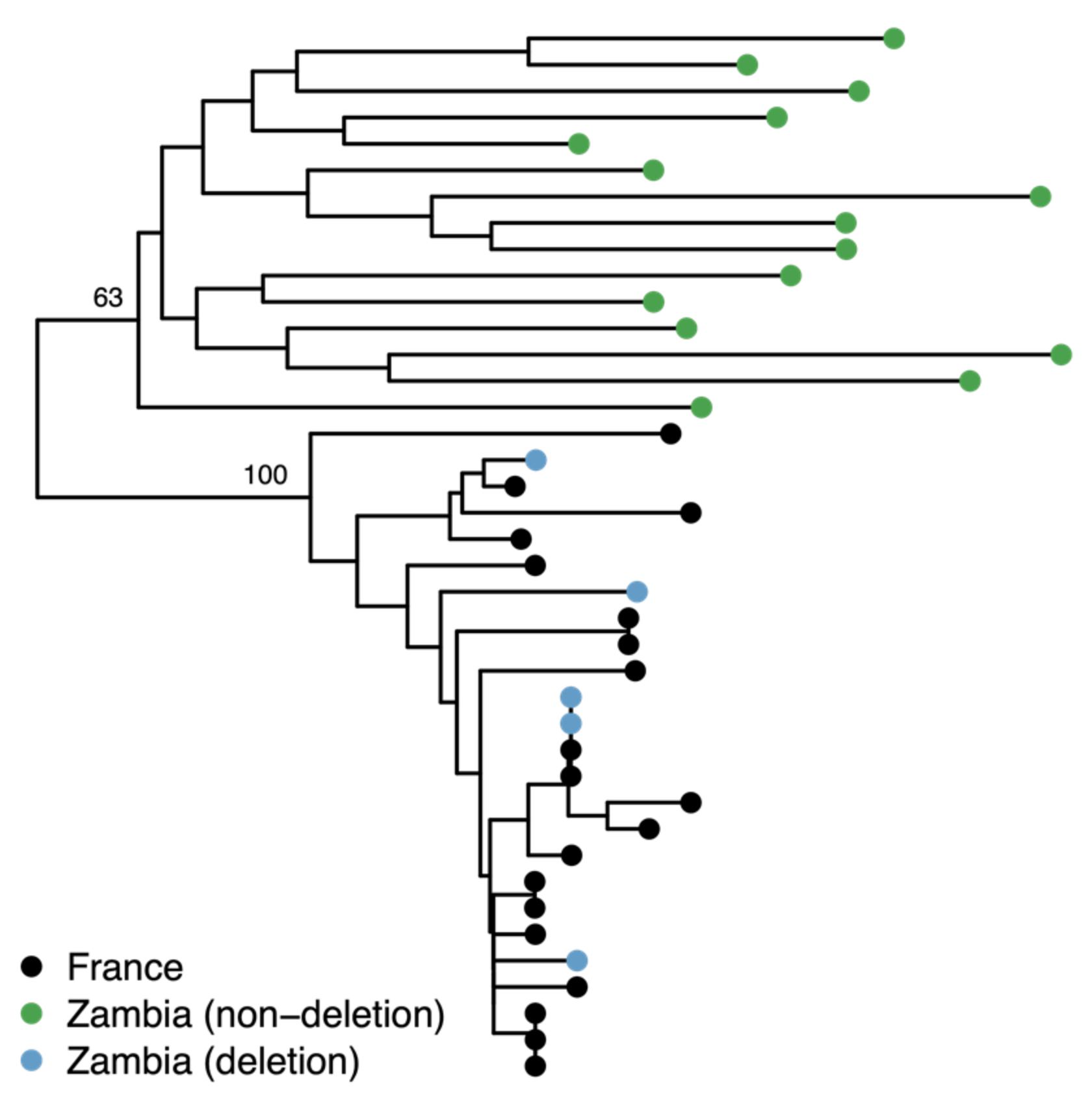
3.4. The Deletion is Rare in Sub-Saharan Africa and is a Result of European Admixture
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Harr, B.; Kauer, M.; Schlötterer, C. Hitchhiking mapping: A population-based fine-mapping strategy for adaptive mutations in Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 2002, 99, 12949–12954. [Google Scholar] [CrossRef] [PubMed]
- Glinka, S.; Ometto, L.; Mousset, S.; Stephan, W.; De Lorenzo, D. Demography and natural selection have shaped genetic variation in Drosophila melanogaster: A multi-locus approach. Genetics 2003, 165, 1269–1278. [Google Scholar] [PubMed]
- Ometto, L.; Glinka, S.; De Lorenzo, D.; Stephan, W. Inferring the effects of demography and selection on Drosophila melanogaster populations from a chromosome-wide scan of DNA variation. Mol. Biol. Evol. 2005, 22, 2119–2130. [Google Scholar] [CrossRef] [PubMed]
- Hutter, S.; Saminadin-Peter, S.S.; Stephan, W.; Parsch, J. Gene expression variation in African and European populations of Drosophila melanogaster. Genome Biol. 2008, 9, R12. [Google Scholar] [CrossRef] [PubMed]
- Müller, L.; Hutter, S.; Stamboliyska, R.; Saminadin-Peter, S.S.; Stephan, W.; Parsch, J. Population transcriptomics of Drosophila melanogaster females. BMC Genom. 2011, 12, 81. [Google Scholar] [CrossRef] [PubMed]
- Catalán, A.; Hutter, S.; Parsch, J. Population and sex differences in Drosophila melanogaster brain gene expression. BMC Genom. 2012, 13, 654. [Google Scholar] [CrossRef] [PubMed]
- Huylmans, A.K.; Parsch, J. Population- and sex-biased gene expression in the excretion organs of Drosophila melanogaster. G3 (Bethesda) 2014, 4, 2307–2315. [Google Scholar] [CrossRef]
- Adrion, J.R.; Hahn, M.W.; Cooper, B.S. Revisiting classic clines in Drosophila melanogaster in the age of genomics. Trends Genet. 2015, 31, 434–444. [Google Scholar] [CrossRef]
- Kapun, M.; Fabian, D.K.; Goudet, J.; Flatt, T. Genomic evidence for adaptive inversion clines in Drosophila melanogaster. Mol. Biol. Evol. 2016, 33, 1317–1336. [Google Scholar] [CrossRef]
- Juneja, P.; Quinn, A.; Jiggins, F.M. Latitudinal clines in gene expression and cis-regulatory element variation in Drosophila melanogaster. BMC Genom. 2016, 17, 981. [Google Scholar] [CrossRef]
- Oakeshott, J.G.; Gibson, J.B.; Anderson, P.R.; Knibb, W.R.; Anderson, D.G.; Chambers, G.K. Alcohol dehydrogenase and glycerol-3-phosphate dehydrogenase clines in Drosophila melanogaster on different continents. Evolution 1982, 36, 86–96. [Google Scholar] [CrossRef] [PubMed]
- Verrelli, B.C.; Eanes, W.F. Clinal variation for amino acid polymorphisms at the Pgm locus in Drosophila melanogaster. Genetics 2001, 157, 1649–1663. [Google Scholar] [PubMed]
- Fabian, D.K.; Kapun, M.; Nolte, V.; Kofler, R.; Schmidt, P.S.; Schlötterer, C.; Flatt, T. Genome-wide patterns of latitudinal differentiation among populations of Drosophila melanogaster from North America. Mol. Ecol. 2012, 21, 4748–4769. [Google Scholar] [CrossRef] [PubMed]
- Machado, H.E.; Bergland, A.O.; O’Brien, K.R.; Behrman, E.L.; Schmidt, P.S.; Petrov, D.A. Comparative population genomics of latitudinal variation in Drosophila simulans and Drosophila melanogaster. Mol. Ecol. 2016, 25, 723–740. [Google Scholar] [CrossRef] [PubMed]
- Bergland, A.O.; Behrman, E.L.; O’Brien, K.R.; Schmidt, P.S.; Petrov, D.A. Genomic evidence of rapid and stable adaptive oscillations over seasonal time scales in Drosophila. PLoS Genet. 2014, 10, e1004775. [Google Scholar] [CrossRef] [PubMed]
- Laurie-Ahlberg, C.C.; Stam, L.F. Use of P-element-mediated transformation to identify the molecular basis of naturally occurring variants affecting Adh expression in Drosophila melanogaster. Genetics 1987, 115, 129–140. [Google Scholar] [PubMed]
- Chung, H.; Bogwitz, M.R.; McCart, C.; Andrianopoulos, A.; Ffrench-Constant, R.H.; Batterham, P.; Daborn, P.J. Cis-regulatory elements in the Accord retrotransposon result in tissue-specific expression of the Drosophila melanogaster insecticide resistance gene Cyp6g1. Genetics 2007, 175, 1071–1077. [Google Scholar] [CrossRef]
- Catalán, A.; Glaser-Schmitt, A.; Argyridou, E.; Duchen, P.; Parsch, J. An indel polymorphism in the MtnA 3′ untranslated region is associated with gene expression variation and local adaptation in Drosophila melanogaster. PLoS Genet. 2016, 12, e1005987. [Google Scholar] [CrossRef]
- Glaser-Schmitt, A.; Parsch, J. Functional characterization of adaptive variation within a cis-regulatory element influencing Drosophila melanogaster growth. PLoS Biol. 2018, 16, e2004538. [Google Scholar] [CrossRef]
- Yang, Y.; Edery, I. Parallel clinal variation in the mid-day siesta of Drosophila melanogaster implicates continent-specific targets of natural selection. PLoS Genet. 2018, 14, e1007612. [Google Scholar] [CrossRef]
- Svetec, N.; Saelao, P.; Cridland, J.M.; Hoffmann, A.A.; Begun, D.J. Functional analysis of a putative target of spatially varying selection in the Menin1 gene of Drosophila melanogaster. G3 (Bethesda) 2019, 9, 73–80. [Google Scholar] [CrossRef] [PubMed]
- Lange, B.W.; Langley, C.H.; Stephan, W. Molecular evolution of Drosophila metallothionein genes. Genetics 1990, 126, 921–932. [Google Scholar] [PubMed]
- Theodore, L.; Ho, A.S.; Maroni, G. Recent evolutionary history of the metallothionein gene Mtn in Drosophila. Genet. Res. 1991, 58, 203–210. [Google Scholar] [CrossRef] [PubMed]
- Stephan, W.; Rodriguez, V.S.; Zhou, B.; Parsch, J. Molecular evolution of the metallothionein gene Mtn in the melanogaster species group: Results from Drosophila ananassae. Genetics 1994, 138, 135–143. [Google Scholar] [PubMed]
- González, J.; Macpherson, J.M.; Petrov, D.A. A recent adaptive transposable element insertion near highly conserved developmental loci in Drosophila melanogaster. Mol. Biol. Evol. 2009, 26, 1949–1961. [Google Scholar] [CrossRef] [PubMed]
- Guio, L.; Barrón, M.G.; González, J. The transposable element Bari-Jheh mediates oxidative stress response in Drosophila. Mol. Ecol. 2014, 23, 2020–2030. [Google Scholar] [CrossRef] [PubMed]
- Kapun, M.; Barron Aduriz, M.G.; Staubach, F.; Vieira, J.; Obbard, D.; Goubert, C.; Rota Stabelli, R.; Kankare, M.; Haudry, A.; Wiberg, R.A.W.; et al. Genomic analysis of European Drosophila populations reveals longitudinal structure and continent-wide selection (pre-print). bioRxiv 2018, 313759. [Google Scholar] [CrossRef]
- Hu, Y.; Sopko, R.; Foos, M.; Kelley, C.; Flockhart, I.; Ammeux, N.; Wang, X.; Perkins, L.; Perrimon, N.; Mohr, S.E. FlyPrimerBank: An online database for Drosophila melanogaster gene expression analysis and knockdown evaluation of RNAi reagents. G3 (Bethesda) 2013, 3, 1607–1616. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing. Available online: http://www.r-project.org (accessed on 14 January 2019).
- Pool, J.E.; Corbett-Detig, R.B.; Sugino, R.P.; Stevens, K.A.; Cardeno, C.M.; Crepeau, M.W.; Duchen, P.; Emerson, J.J.; Saelao, P.; Begun, D.J.; et al. Population genomics of sub-saharan Drosophila melanogaster: African diversity and non-African admixture. PLoS Genet. 2012, 8, e1003080. [Google Scholar] [CrossRef]
- Lack, J.B.; Cardeno, C.M.; Crepeau, M.W.; Taylor, W.; Corbett-Detig, R.B.; Stevens, K.A.; Langley, C.H.; Pool, J.E. The Drosophila genome nexus: A population genomic resource of 623 Drosophila melanogaster genomes, including 197 from a single ancestral range population. Genetics 2015, 199, 1229–1241. [Google Scholar] [CrossRef]
- Lack, J.B.; Lange, J.D.; Tang, A.D.; Corbett-Detig, R.B.; Pool, J.E. A thousand fly genomes: An expanded Drosophila genome nexus. Mol. Biol. Evol. 2016, 33, 3308–3313. [Google Scholar] [CrossRef] [PubMed]
- Sprengelmeyer, Q.D.; Mansourian, S.; Lange, J.D.; Matute, D.R.; Cooper, B.S.; Jirle, E.V.; Stensmyr, M.C.; Pool, J.E. Discovery of Drosophila melanogaster from wild African environments and genomic insights into species history (pre-print). bioRxiv 2018. [Google Scholar] [CrossRef]
- Langley, C.H.; Crepeau, M.; Cardeno, C.; Corbett-Detig, R.; Stevens, K. Circumventing heterozygosity: Sequencing the amplified genome of a single haploid Drosophila melanogaster embryo. Genetics 2011, 188, 239–246. [Google Scholar] [CrossRef] [PubMed]
- Sedlazeck, F.J.; Rescheneder, P.; von Haeseler, A. NextGenMap: Fast and accurate read mapping in highly polymorphic genomes. Bioinformatics 2013, 29, 2790–2791. [Google Scholar] [CrossRef] [PubMed]
- Mackay, T.F.; Richards, S.; Stone, E.A.; Barbadilla, A.; Ayroles, J.F.; Zhu, D.; Casillas, S.; Han, Y.; Magwire, M.M.; Cridland, J.M.; et al. The Drosophila melanogaster genetic reference panel. Nature 2012, 482, 173–178. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Massouras, A.; Inoue, Y.; Peiffer, J.; Ràmia, M.; Tarone, A.M.; Turlapati, L.; Zichner, T.; Zhu, D.; Lyman, R.F.; et al. Natural variation in genome architecture among 205 Drosophila melanogaster Genetic Reference Panel lines. Genome Res. 2014, 24, 1193–1208. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Thurmond, J.; Goodman, J.L.; Strelets, V.B.; Attrill, H.; Gramates, L.S.; Marygold, S.J.; Matthews, B.B.; Millburn, G.; Antonazzo, G.; Trovisco, V.; et al. FlyBase 2.0: The next generation. Nucleic Acids Res. 2019, 47, D759–D765. [Google Scholar] [CrossRef]
- Paradis, E.; Claude, J.; Strimmer, K. APE: Analyses of phylogenetics and evolution in R language. Bioinformatics 2004, 20, 289–290. [Google Scholar] [CrossRef] [PubMed]
- Drosophila Genetic Reference Panel 2. Available online: http://dgrp2.gnets.ncsu.edu (accessed on 14 January 2019).
- Huang, W.; Carbone, M.A.; Magwire, M.M.; Peiffer, J.A.; Lyman, R.F.; Stone, E.A.; Anholt, R.R.; Mackay, T.F. Genetic basis of transcriptome diversity in Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 2015, 112, E6010–E6019. [Google Scholar] [CrossRef] [PubMed]
- Weber, A.L.; Khan, G.F.; Magwire, M.M.; Tabor, C.L.; Mackay, T.F.; Anholt, R.R. Genome-wide association analysis of oxidative stress resistance in Drosophila melanogaster. PLoS ONE 2012, 7, e34745. [Google Scholar] [CrossRef] [PubMed]
- Therneau, T.M.; Grambsch, P.M. Modeling Survival Data: Extending the Cox Model; Springer: New York, NY, USA, 2000. [Google Scholar]
- Therneau, T. A Package for Survival Analysis in R Version 2.38. Available online: https://CRAN.R-project.org/package=survival (accessed on 14 January 2019).
- Gnad, F.; Parsch, J. Sebida: A database for the functional and evolutionary analysis of genes with sex-biased expression. Bioinformatics 2006, 22, 2577–2579. [Google Scholar] [CrossRef] [PubMed]
- Jain, K.; Stephan, W. Modes of rapid polygenic adaptation. Mol. Biol. Evol. 2017, 34, 3169–3175. [Google Scholar] [CrossRef] [PubMed]
- Thorpe, G.W.; Fong, C.S.; Alic, N.; Higgins, V.J.; Dawes, I.W. Cells have distinct mechanisms to maintain protection against different reactive oxygen species: Oxidative-stress-response genes. Proc. Natl. Acad. Sci. USA 2004, 101, 6564–6569. [Google Scholar] [CrossRef] [PubMed]
- Krasley, E.; Cooper, K.F.; Mallory, M.J.; Dunbrack, R.; Strich, R. Regulation of the oxidative stress response through Slt2p-dependent destruction of cyclin C in Saccharomyces cerevisiae. Genetics 2006, 172, 1477–1486. [Google Scholar] [CrossRef] [PubMed]
- Girardot, F.; Monnier, V.; Tricoire, H. Genome wide analysis of common and specific stress responses in adult Drosophila melanogaster. BMC Genom. 2004, 5, 74. [Google Scholar] [CrossRef]
- Chen, D.; Wilkinson, C.R.; Watt, S.; Penkett, C.J.; Toone, W.M.; Jones, N.; Bähler, J. Multiple pathways differentially regulate global oxidative stress responses in fission yeast. Mol. Biol. Cell 2008, 19, 308–317. [Google Scholar] [CrossRef]
- Wallace, M.A.; Bailey, S.; Fukuto, J.M.; Valentine, J.S.; Gralla, E.B. Induction of phenotypes resembling CuZn-superoxide dismutase deletion in wild-type yeast cells: An in vivo assay for the role of superoxide in the toxicity of redox-cycling compounds. Chem. Res. Toxicol. 2005, 18, 1279–1286. [Google Scholar] [CrossRef]
- Van Straalen, N.M.; Roelofs, D. An Introduction to Ecological Genomics, 2nd ed.; Oxford University Press: Oxford, UK, 2012. [Google Scholar]
| Collection | Nfemale | Freqfemale (95% CI) | Nmale | Freqmale (95% CI) |
|---|---|---|---|---|
| June 2016 | 224 | 0.906 (0.860–0.941) | 164 | 0.902 (0.846–0.943) |
| Sept 2016 | 192 | 0.927 (0.881–0.960) | 94 | 0.894 (0.813–0.948) |
| June 2017 | 144 | 0.903 (0.842–0.946) | 174 | 0.902 (0.848–0.942) |
| Sept 2017 | 176 | 0.909 (0.857–0.947) | 60 | 0.933 (0.838–0.982) |
| Collection | Del/Del | Del/Non | Non/Non | PHWE |
|---|---|---|---|---|
| June 2016 | 160 | 31 | 3 | 0.731 |
| Sept 2016 | 120 | 22 | 1 | 0.999 |
| June 2017 | 131 | 25 | 3 | 0.653 |
| Sept 2017 | 99 | 18 | 1 | 0.999 |
| All | 510 | 96 | 8 | 0.338 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramnarine, T.J.S.; Glaser-Schmitt, A.; Catalán, A.; Parsch, J. Population Genetic and Functional Analysis of a cis-Regulatory Polymorphism in the Drosophila melanogaster Metallothionein A gene. Genes 2019, 10, 147. https://doi.org/10.3390/genes10020147
Ramnarine TJS, Glaser-Schmitt A, Catalán A, Parsch J. Population Genetic and Functional Analysis of a cis-Regulatory Polymorphism in the Drosophila melanogaster Metallothionein A gene. Genes. 2019; 10(2):147. https://doi.org/10.3390/genes10020147
Chicago/Turabian StyleRamnarine, Timothy J. S., Amanda Glaser-Schmitt, Ana Catalán, and John Parsch. 2019. "Population Genetic and Functional Analysis of a cis-Regulatory Polymorphism in the Drosophila melanogaster Metallothionein A gene" Genes 10, no. 2: 147. https://doi.org/10.3390/genes10020147
APA StyleRamnarine, T. J. S., Glaser-Schmitt, A., Catalán, A., & Parsch, J. (2019). Population Genetic and Functional Analysis of a cis-Regulatory Polymorphism in the Drosophila melanogaster Metallothionein A gene. Genes, 10(2), 147. https://doi.org/10.3390/genes10020147




